Back to Journals » OncoTargets and Therapy » Volume 13
Exosomal miR-548c-5p Regulates Colorectal Cancer Cell Growth and Invasion Through HIF1A/CDC42 Axis
Authors Yan S, Ren X, Yang J , Wang J , Zhang Q, Xu D
Received 27 July 2020
Accepted for publication 15 September 2020
Published 5 October 2020 Volume 2020:13 Pages 9875—9885
DOI https://doi.org/10.2147/OTT.S273008
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Arseniy Yuzhalin
Shushan Yan,1 Xiaoxia Ren,2 Jinghan Yang,3 Jinghua Wang,3 Quan Zhang,4 Donghua Xu3
1Department of Gastrointestinal and Anal Diseases Surgery of Affiliated Hospital, Weifang Medical University, Weifang, Shandong Province, People’s Republic of China; 2Department of Gastrointestinal Surgery, Yantai Shan Hospital, Yantai, Shandong Province, People’s Republic of China; 3Central Laboratory and Department of Rheumatology of the First Affiliated Hospital, Weifang Medical University, Weifang, Shandong Province, People’s Republic of China; 4Department of Cardiology of Affiliated Hospital, Weifang Medical University, Weifang, Shandong Province, People’s Republic of China
Correspondence: Donghua Xu
Central Laboratory and Department of Rheumatology of the First Affiliated Hospital, Weifang Medical University, Weifang 261000, People’s Republic of China
Email [email protected]
Quan Zhang
Department of Cardiology of Affiliated Hospital, Weifang Medical University, Weifang, Shandong Province, People’s Republic of China
Email [email protected]
Background: Mounting evidence has implicated that exosomes-delivered noncoding RNAs are key regulators in carcinogenesis. The effect of miR-548c-5p has been elucidated in some cancers. However, the role of exosomal miR-548c-5p in colorectal cancer (CRC) is not fully understood. We aim to explore the function and mechanism of exosome-delivered miR-548c-5p in CRC. The altering effect of exosome-derived miR-548c-5p on the prognosis of CRC patients is also investigated by estimating overall survival and disease-free survival.
Materials and Methods: The expression of miR-548c-5p in exosomes is determined by real-time PCR. The proliferation and invasion of CRC cells are estimated by MTT, transwell assay and scratch test. The targeted gene of miR-548c-5p is investigated by luciferase reporter assay, real-time PCR, Western blot and chromosome immunoprecipitation (CHIP) assay. CRC cells are transplanted subcutaneously in BALB/c nude mice to estimate their growth in vivo.
Results: MiR-548c-5p derived from CRC cell exosomes inhibits the proliferation and invasion of CRC cells in vitro. Exosomal miR-548c-5p can also prevent from colorectal carcinogenesis in nude mice in vivo. HIF1A is documented to be a target of miR-548c-5p, and HIF1A can targetedly regulate CDC42 in CRC cells. Exosomal miR-548c-5p affects CRC cell growth, migration and invasion via miR-548c-5p/HIF1A/CDC42 axis. In addition, exosomal miR-548c-5p can be a predictive factor for CRC prognosis.
Conclusion: Our study has suggested that exosomal miR-548c-5p can regulate CRC through HIF1A/CDC42 axis, which helps to understand CRC pathogenesis more clearly and identify novel therapeutic strategies for CRC patients.
Keywords: CDC42, colorectal cancer, exosome, HIF1A, microRNA
Introduction
To the best of knowledge, colorectal cancer (CRC) is one of the most common cancers across the world.1 The etiology and pathogenesis of CRC are not clear yet. It has been demonstrated that lifestyle, heredity and colorectal adenoma are closely related to CRC. Besides, familial adenomatous polyposis and hereditary nonpolyposis can elevate the risk of CRC.2,3 Great progress in CRC treatment has been made during the past few years, but the prognosis of patients is not so good particularly among advanced CRC cases. Therefore, identifying biomarkers predicting CRC initiation and progression is essential. Mounting evidence has implicated noncoding RNAs in circulation can serve as useful markers for CRC, including exosomes-encapsulating microRNAs (miRNAs).
Exosomes belong to small extracellular vesicles with important biological activity.4 The role of exosomes in cancer has been widely described, which may serve as promising therapeutic approaches.5,6 Exosomes can mediate communications between cancer cells by delivering active constituents, such as noncoding RNAs, proteins and peptides.7 Among them, miRNA is a kind of noncoding RNA involved in regulating carcinogenesis. During the past few decades, accumulated studies have demonstrated that miRNAs are key regulators in CRC.8–10 Some exosomes-delivering miRNAs have also been identified as ideal makers for CRC diagnosis, treatment and prognosis.11–13 Previously, we have elucidated that miR-548c-5p is dysregulated in CRC.14 However, the function and mechanism of exosomal miR-548c-5p in CRC are unclear yet. This study is aimed to demonstrate the effect of exosomal miR-548c-5p in CRC. In addition, the influence of exosomal miR-548c-5p in CRC outcomes is also estimated.
Materials and Methods
Participants
Table 1 presents the characteristics of 104 CRC patients, who have enrolled in the Affiliated Hospital of Weifang Medical University and received surgical resection. The follow-up time since first surgical resection is up to 3 years since surgical resection. Our study is approved by the Institutional Ethics Committee of the Affiliated Hospital of Weifang Medical University. The patients’ informed consent has been obtained prior to study. This is conducted in accordance with the Declaration of Helsinki.
 |
Table 1 Characteristics of Patients |
Cell Growth and Exosomes Extraction
HCT116 and SW480 CRC cells and normal intestinal epithelial cells Henle-407 are purchased from Genechem Biotechnology Company (Shanghai, China) and cultured in DMEM (GIBCO, USA) adding with 10–20% fetal calf serum (GIBCO, USA). CRC cells are transfected by lentivirus plasmids using polybrene reagent to make CDC42 or HIF1A overexpression (OE) or knocked down (KO) in CRC cells. In addition, CRC cells are treated by miRNA mimics and the controls for tests (Guangzhou, China) using lipofectamine 2000 (Invitrogen, USA). Exosomes are extracted by the instructions of Exosome Isolation and Purification Kit (Umibio, Shanghai, China). In brief, the serum samples are acquired after centrifugation (1000g, 10 min) of fresh peripheral blood. Exosomes in cell culture supernatant and the serum are isolated according to the protocol. The morphology of exosomes is scanned by transmission electron microscope, and their diameter and concentration are estimated by Nanosight instrument. Those purified exosomes are stored at −80°C for subsequent experiments.
Cell Proliferation, Migration and Invasion
MTT kit (Beyotime, Shanghai, China) is adopted to analyze the proliferation of CRC cells growth of cells. The absorption at 490nm is detected according to the protocol. Transwell assay is performed to assess CRC cell invasion by using the transwell chambers with Matrigel (Corning, MA, USA). Briefly, 105 cells (per well) are cultured, respectively, in chambers with or without exosomes (40ug/mL) for 48h, and then fixed by paraformaldehyde (4%) and stained with crystal violet (0.1%). Scratch test is also performed to estimate the migration status of CRC cells when stimulated by exosomes for 48h.
Real-Time Polymerase Chain Reaction (PCR)
MiR-548c-5p derived from serum exosomes is determined by real-time PCR using TaqMan PCR kit (ThermoFisher Scientific, USA). MiR-16-5p is used as an endogenous control. MiR-548c-5p expression in serum exosomes is estimated by 2–ΔΔCT. Primers are as listed below. Human HIF1A, f, 5ʹTCCATGTGACCATGAGGAAA3ʹ; r, 5ʹCCAAGCAGGTCATAGGTGGT3ʹ. Human CDC42, f, 5ʹGCTGGGTGCTAAGAAACTCG3ʹ; r, 5ʹCATAAAGCCACTTCCCTCCA3ʹ. Mouse HIF1A, f, 5ʹGAAATGGCCCAGTGAGAAAA3ʹ; r, 5ʹTATCGAGGCTGTGTCGACTG3ʹ. Mouse GAPDH, f, 5ʹAACTTTGGCATTGTGGAAGG3ʹ; r, 5ʹACACATTGGGGGTAGGAACA3ʹ. Human GAPDH, f, 5ʹCGACCACTTTGTCAAGCTCA3ʹ; r, 5ʹAGGGGAGATTCAGTGTGGTG3ʹ.
Western Blot
CRC cells are treated by mimics and controls. Then, 30μg proteins per well are separated by gel electrophoresis after extraction from CRC cells using RIPA buffer (Beyotime, Shanghai, China). HIF1A (Abcam, Cambridge, UK) and β-actin (CST, Boston, USA) are used for immunoblotting assay based on the instructions.
Luciferase Reporter Assay
The wild and mutant binding sites of HIF1A or CDC42 3ʹ-UTR are cloned into lentivirus plasmids and used to transfect cells. Activity of the luciferase in the system (Promega, WI, USA) is estimated. Experiments are performed for at least three times.
Chromosome Immunoprecipitation (CHIP) Assay
HIF1A overexpressed or siRNA lentivirus plasmids treated CRC cells are treated with formaldehyde (1%) for 10 min at 25°C. Ice-cold PBS containing protease inhibitors is applied to wash cells twice. After centrifugation, cells are resuspended in SDS lysis solution and then sonicated. The lysates are centrifugated, and the supernatant is diluted with ChIP buffer and incubated with antibody and protein G beads according to the protocol. CDC42 expression in the immunoprecipitates is quantified by PCR.
Nude Mice
BALB/c nude mice (4 week old) are used for subcutaneous tumor model construction. CRC cells are harvested and suspended in phosphate-buffered saline, which are injected into mice subcutaneously. Exosomes rich of miR-548c-5p are injected into tumors after tumor formation. Tumor volume is also estimated during tumor formation. Six weeks later, mice are sacrificed by cervical method and the tumor size is evaluated. Animal study is governed by the Animal Ethics Committee of Weifang Medical University and conducted by following the guidelines for the welfare of the laboratory animals.
Statistical Analysis
The 3-year survival of all patients is analyzed by Kaplan–Meier survival curve analysis and Log rank test are performed to assess the overall survival and disease-free survival of CRC patients. Data are analyzed using Graphpad Software (San Diego, CA, USA). The Student t-test or one-way ANOVA is adopted for data calculation. P < 0.05 is statistically significant.
Results
CRC Cells Exosomes Derived miR-548c-5p Prevents from Cell Proliferation, Migration and Invasion
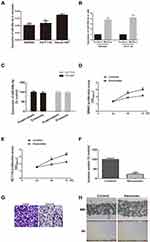
Exosomes from CRC cells have been characterized using electron microscopy, particle-size distribution, and Western blot for exosome markers CD9 and CD63 (Supplementary Figure 1). Previously, we have found miR-548c-5p was reduced in serum exosomes of CRC patients.14 Here, reduced miR-548c-5p is also found in CRC cells (HCT116 and SW480) derived exosomes compared with normal intestinal epithelial cells (Henle-407) derived exosomes (Figure 1A). CRC cells transfected by miR-548c-5p mimics can produce exosomes with high level of miR-548c-5p, and miR-548c-5p primarily exists in exosomes but not free in the supernatant as evidenced by real-time PCR (Figure 1B and C). Subsequently, CRC cells are co-incubated with exosomes from miR-548c-5p mimics treated CRC cells. Exosomal miR-548c-5p prevents from CRC cell proliferation (Figure 1D and E). Besides, the migration and invasion of SW480 cells at 48h is also significantly restrained when they are treated by exosomes delivering high levels of miR-548c-5p, which is demonstrated by scratch test and transwell assay (Figure 1F–H). All these findings have implicated that CRC cells can secret exosomes, and those exosomes are capable of inhibiting CRC cell proliferation, migration and invasion by delivering miR-548c-5p to other CRC cells.
miR-548c-5p Targetedly Regulates HIF1A in CRC Cells
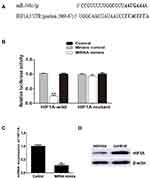
In order to elucidate the mechanism of miR-548c-5p in CRC, we screen the potential target of miR-548c-5p in the miRBase database. It is implicated that HIF1A, also known as HIF1α, is a potential-targeted gene of miR-548c-5p (Figure 2A). HIF1A has been strongly suggested to be involved in the pathogenesis of carcinogenesis. As a result, we hypothesize that miR-548c-5p might regulate CRC via regulating the expression and function of HIF1A. Luciferase reporter assay has further demonstrated that miR-548c-5p targetedly regulated HIF1A (Figure 2B). Besides, the mRNA and protein of HIF1A in SW480 cells are significantly decreased when cells are administrated into miR-548c-5p mimics (Figure 2C and D). Accordingly, we conclude that miR-548c-5p can regulate HIF1A posttranscriptionally.
miR-548c-5p/HIF1A/CDC42 Axis in CRC Cells
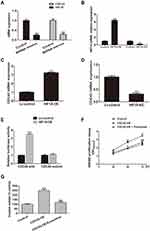
The expression of CDC42 and HIF1A is significantly reduced in miR-548c-5p mimics treated CRC cells as evidenced by real-time PCR (Figure 3A). We hypothesize that the decreased expression of CDC42 in miR-548c-5p mimics treated CRC cells is attributed to downregulation of HIF1A. In order to elucidate this, we have performed the following experiments. HIF1A is up- and down-regulated by lentivirus transfection of SW480 cells (Figure 3B). As a transcriptional factor, HIF1A can bind to the promoter of CDC42 and enhance its expression suggested by CHIP assay, while the expression of CDC42 is decreased when HIF1A is knocked down (Figure 3C and D). Besides, as demonstrated by luciferase reporter assay, the overexpressed HIF1A promotes the transcription of CDC42 demonstrated by enhanced luciferase activity (Figure 3E). After screening in databases of Target Scan and miRBase, we have found miR-548c-5p could not recognize the 3ʹUTR of CDC42. Therefore, CDC42 can be targetedly regulated by transcriptional factor HIF1A but not miR-548c-5p directly. To investigate the function of CDC42 in CRC cells, SW480 cells are overexpressed with CDC42 and administrated into exosomes rich of miR-548c-5p. It is demonstrated that CDC42 overexpression enhances the proliferation and invasion of SW480 cells, whereas exosomal miR-548c-5p can restrain its effects probably by downregulating HIF1A in CRC cells (Figure 3F and G). Accordingly, exosomal miR-548c-5p can regulate CRC cell growth and invasion through miR-548c-5p/HIF1A/CDC42 axis. All these findings have suggested that exosome-encapsulated miR-548c-5p participates in CRC tumorigenesis.
Exosome-Delivered miR-548c-5p Inhibits Tumor Growth in Nude Mice
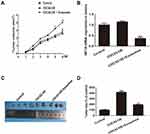
CDC42 overexpressed SW480 cells are subcutaneously planted in nude mice. As presented in Figure 4A, exosome-delivered miR-548c-5p injected into tumors can slow down the growth of CDC42-overexpressed CRC cells via downregulating HIF1A, which is demonstrated by real-time PCR (Figure 4B). In addition, the tumor size in nude mice is smaller in exosomes-encapsulated miR-548c-5p treated cells compared with that from CDC42 overexpressed SW480 cells (Figure 4C and D). Accordingly, exosome-delivered miR-548c-5p prevents from the tumorigenesis of CRC by targeting HIF1A/CDC42.
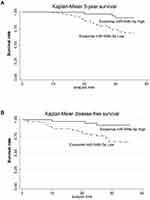
miR-548c-5p in Exosomes Predicts CRC Prognosis
Exosomal miR-548c-5p in serum is determined in our study. We have found that CRC patients with low miR-548c-5p expression in serum exosomes had poor outcomes (Figure 5). The survival of patients with higher miR-548c-5p in exosomes is longer than those with lower expression miR-548c-5p in exosomes (Figure 5A, P for Log rank test = 0.008). Similarly, CRC patients with lower miR-548c-5p in exosomes have poor disease-free survival (Figure 5B, P for Log rank test = 0.007). Table 2 shows univariate and multivariate cox regression results, which has implicated that reduced miR-548c-5p in exosomes is associated with poor prognosis of CRC patients independent of late TNM stage, low tumor grade and liver metastasis. Accordingly, exosomal miR-548c-5p is a predictive factor for CRC prognosis.
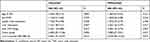 |
Table 2 Cox Regression Analysis |
Discussion
This study shows that miR-548c-5p in exosomes is dysregulated in CRC and inhibits the growth of cancer cells. Besides, HIF1A is documented to be a target of miR-548c-5p, and transcriptional factor HIF1A can regulate CDC42 in CRC cells. Moreover, exosomal miR-548c-5p regulates CRC oncogenesis via miR-548c-5p/HIF1A/CDC42 axis. Additionally, exosomal miR-548c-5p is a promising prognostic factor for CRC. As a result, exosome-encapsulated miR-548c-5p is a key biomarker for CRC by influencing its oncogenesis and prognosis.
Exosomes are common membrane-bound nanovesicles, which encapsulate proteins, lipids, carbohydrates, nucleic acids and many other contents with biological activity.15–17 As extracellular vesicles (EVs), exosomes participate in cell-to-cell communications by delivering diverse biomolecules between cells. Therefore, exosomes can be explored as promising drug carriers for diverse diseases, particularly cancer. Accumulating studies have strongly suggested a number of exosomes-encapsulated miRNAs play vital roles in cancer in the past few decades, such as breast cancer, liver cancer and CRC.18–20 Previously, we have demonstrated the specific profile of dysregulated miRNAs in exosomes in CRC, among which miR-548c-5p is one of the most significantly reduced miRNAs.14 Certain differentially expressed miRNAs in the profile have specific function in CRC, including miR-638, miR-6869-5p and miR-6803-5p.9,11,21,22 The role of miR-548c-5p in cancer has drawn wide attention.23–25 The study by Peng et al has suggested exosomal miR-548c-5p is correlated with cancer prognosis.26 However, the function and mechanism of exosomal miR-548c-5p in CRC pathogenesis are largely unknown. In this study, we have demonstrated miR-548c-5p is reduced in HCT116 and SW480 CRC cells derived exosomes compared with those from Henle-407 normal intestinal epithelial cells. MiR-548c-5p derived from CRC cell exosomes can restrain the proliferation, migration and invasion of other CRC cells. Besides, miR-548c-5p is capable of downregulating the expression of HIF1A posttranscriptionally. As a transcriptional factor, HIF1A can targetedly regulate CDC42 in CRC cells. Currently published studies have suggested the key effect of HIF1A in cancer, including CRC.27,28 HIF1A can regulate the tumorigenesis and tumor progression of CRC through interacting with certain miRNAs.29,30 In this study, we have demonstrated HIF1A can be regulated by miR-548c-5p in CRC cells. Accordingly, HIF1A and its linkage with miRNA provide new therapeutic targets for CRC.
CDC42 is an essential GTPase protein for cancer metastasis, which belongs to the Rho family member.31,32 CDC42 participates in regulating cell-to-cell adhesion, cell proliferation and cell cycle.32 It has been well established that CDC42 is upregulated in a number of malignancies and serves as a promising target for cancer treatment, such as colon cancer and hepatocellular carcinoma.33–35 CDC42 regulates oncogenesis and cancer metastasis by influencing cancer cell proliferation, migration and invasion.31,36,37 There are several miRNAs demonstrated to directly regulate CDC42 as targeted gene in CRC, such as miR-384 and miR-330.38,39 However, there is no data demonstrating the association between HIF1A and CDC42. Our findings firstly show the evidence that CDC42 can be targetedly regulated by transcriptional factor HIF1A but not miR-548c-5p directly. Accordingly, exosomal miR-548c-5p is involved in oncogenesis of CRC through miR-548c-5p/HIF1A/CDC42 axis. MiR-548c-5p targetedly regulates HIF1A, while HIF1A can directly influence CDC42 expression in CRC cells. The miR-548c-5p/HIF1A/CDC42 axis may serve as prospective strategies for the diagnosis and therapy of CRC.
In conclusion, our study has shown that exosomal miR-548c-5p is involved in colorectal carcinogenesis via regulating HIF1A/CDC42. Besides, exosomes-encapsulated miR-548c-5p can function as an independent factor of CRC prognosis. As a result, exosomal miR-548c-5p is a promising target for CRC treatment, and miR-548c-5p/HIF1A/CDC42 axis provides new insight into CRC pathogenesis and treatment.
Data Sharing Statement
All data were available upon requesting from the corresponding author.
Funding
The study is funded by grants from Natural Science Foundation for Young Scholars, Shandong, China (ZR2019QH012 and ZR2015HL011), and China National Natural Science Foundation (81601408, 81870237, and 31570941), and Weifang Science and Technology Development Program, China (2019YX020 and 2019GX031), and Shandong Medical and Health Science and Technology Development Program (2016WS0687), and Outstanding Youth Innovation Plan of Shandong Province (2019).
Disclosure
The authors report no conflicts of interest for this work.
References
1. Siegel RL, Miller KD, Fedewa SA, et al. Colorectal cancer statistics, 2017. CA Cancer J Clin. 2017;67(3):177–193. doi:10.3322/caac.21395
2. Valle L, de Voer RM, Goldberg Y, et al. Update on genetic predisposition to colorectal cancer and polyposis. Mol Aspects Med. 2019;69:10–26. doi:10.1016/j.mam.2019.03.001
3. Kanth P, Grimmett J, Champine M, et al. Hereditary colorectal polyposis and cancer syndromes: a primer on diagnosis and management. Am J Gastroenterol. 2017;112(10):1509–1525. doi:10.1038/ajg.2017.212
4. Koniusz S, Andrzejewska A, Muraca M, et al. Extracellular vesicles in physiology, pathology, and therapy of the immune and central nervous system, with focus on extracellular vesicles derived from mesenchymal stem cells as therapeutic tools. Front Cell Neurosci. 2016;10:109.
5. Bastos N, Ruivo CF, da Silva S, et al. Exosomes in cancer: use them or target them? Semin Cell Dev Biol. 2018;78:13–21. doi:10.1016/j.semcdb.2017.08.009
6. Al-Sowayan BS, Al-Shareeda AT, Al-Hujaily EM. Exosomes, cancer’s little army. Stem Cell Investig. 2018;9:9. doi:10.21037/sci.2019.03.02
7. Bae S, Brumbaugh J, Bonavida B. Exosomes derived from cancerous and non-cancerous cells regulate the anti-tumor response in the tumor microenvironment. Genes Cancer. 2018;9(3–4):87–100. doi:10.18632/genesandcancer.172
8. Shirafkan N, Mansoori B, Mohammadi A, et al. MicroRNAs as novel biomarkers for colorectal cancer: new outlooks. Biomed Pharmacother. 2018;97:1319–1330. doi:10.1016/j.biopha.2017.11.046
9. Yan S, Cheng M, Duan Q, et al. MiR-6803-5p promotes cancer cell proliferation and invasion via PTPRO/NF- κ B axis in colorectal cancer. Mediators Inflamm. 2019;2019:8128501. doi:10.1155/2019/8128501
10. Fadaka AO, Pretorius A, Klein A. MicroRNA assisted gene regulation in colorectal cancer. Int J Mol Sci. 2019;20(19):19. doi:10.3390/ijms20194899
11. Yan S, Liu G, Jin C, et al. MicroRNA-6869-5p acts as a tumor suppressor via targeting TLR4/NF-kappaB signaling pathway in colorectal cancer. J Cell Physiol. 2018;233(9):6660–6668. doi:10.1002/jcp.26316
12. Vautrot V, Chanteloup G, Elmallah M, et al. Exosomal miRNA: small molecules, big impact in colorectal cancer. J Oncol. 2019;2019:8585276. doi:10.1155/2019/8585276
13. Nedaeinia R, Manian M, Jazayeri MH, et al. Circulating exosomes and exosomal microRNAs as biomarkers in gastrointestinal cancer. Cancer Gene Ther. 2017;24(2):48–56.
14. Yan S, Han B, Gao S, et al. Exosome-encapsulated microRNAs as circulating biomarkers for colorectal cancer. Oncotarget. 2017;8(36):60149–60158. doi:10.18632/oncotarget.18557
15. He C, Zheng S, Luo Y, et al. Exosome theranostics: biology and translational medicine. Theranostics. 2018;8(1):237–255. doi:10.7150/thno.21945
16. Walker S, Busatto S, Pham A, et al. Extracellular vesicle-based drug delivery systems for cancer treatment. Theranostics. 2019;9(26):8001–8017. doi:10.7150/thno.37097
17. Clayton SM, Archard J, Wagner J, et al. Immunoregulatory potential of exosomes derived from cancer stem cells. Stem Cells Dev. 2020;29(6):327–335.
18. Zhao L, Gu C, Gan Y, et al. Exosome-mediated siRNA delivery to suppress postoperative breast cancer metastasis. J Control Release. 2020;318:1–15. doi:10.1016/j.jconrel.2019.12.005
19. Redd Bowman KE, Lu P, Vander Mause ER, et al. Advances in delivery vectors for gene therapy in liver cancer. Ther Deliv. 2020;11(1):833–850.
20. Ruiz-Lopez L, Blancas I, Garrido JM, et al. The role of exosomes on colorectal cancer: a review. J Gastroenterol Hepatol. 2018;33(4):792–799. doi:10.1111/jgh.14049
21. Yan S, Dang G, Zhang X, et al. Downregulation of circulating exosomal miR-638 predicts poor prognosis in colon cancer patients. Oncotarget. 2017;8(42):72220–72226. doi:10.18632/oncotarget.19689
22. Yan S, Jiang Y, Liang C, et al. Exosomal miR-6803-5p as potential diagnostic and prognostic marker in colorectal cancer. J Cell Biochem. 2018;119(5):4113–4119. doi:10.1002/jcb.26609
23. Block I, Burton M, Sorensen KP, et al. Association of miR-548c-5p, miR-7-5p, miR-210-3p, miR-128-3p with recurrence in systemically untreated breast cancer. Oncotarget. 2018;9(10):9030–9042. doi:10.18632/oncotarget.24088
24. Fang L, Zhang H-B, Li H, et al. miR-548c-5p inhibits proliferation and migration and promotes apoptosis in CD90+ HepG2 cells. Radiol Oncol. 2012;46(3):233–241. doi:10.2478/v10019-012-0025-z
25. Ge J, Li J, Na S, et al. miR-548c-5p inhibits colorectal cancer cell proliferation by targeting PGK1. J Cell Physiol. 2019;234(10):18872–18878. doi:10.1002/jcp.28525
26. Peng ZY, Gu RH, Yan B. Downregulation of exosome-encapsulated miR-548c-5p is associated with poor prognosis in colorectal cancer. J Cell Biochem. 2019;120(2):1457–1463.
27. Lu Y, Li Y, Wang Z, et al. Downregulation of RGMA by HIF-1A/miR-210-3p axis promotes cell proliferation in oral squamous cell carcinoma. Biomed Pharmacother. 2019;112:108608. doi:10.1016/j.biopha.2019.108608
28. Martinez C-A, Kerr B, Jin C, et al. Obstructive sleep apnea activates hif-1 in a hypoxia dose-dependent manner in HCT116 colorectal carcinoma cells. Int J Mol Sci. 2019;20(2):2. doi:10.3390/ijms20020445
29. Jin F, Yang R, Wei Y, et al. HIF-1α-induced miR-23a223C27a223C24 cluster promotes colorectal cancer progression via reprogramming metabolism. Cancer Lett. 2019;440–441:211–222. doi:10.1016/j.canlet.2018.10.025
30. Xu Z, Zhu C, Chen C, et al. CCL19 suppresses angiogenesis through promoting miR-206 and inhibiting Met/ERK/Elk-1/HIF-1α/VEGF-A pathway in colorectal cancer. Cell Death Dis. 2018;9(10):974. doi:10.1038/s41419-018-1010-2
31. Maldonado MDM, Dharmawardhane S. Targeting rac and Cdc42 GTPases in cancer. Cancer Res. 2018;78(12):3101–3111.
32. Stengel K, Zheng Y. Cdc42 in oncogenic transformation, invasion, and tumorigenesis. Cell Signal. 2011;23(9):1415–1423. doi:10.1016/j.cellsig.2011.04.001
33. Zhu G-F, Xu Y-W, Li J, et al. Mir20a/106a-WTX axis regulates RhoGDIa/CDC42 signaling and colon cancer progression. Nat Commun. 2019;10(1):112. doi:10.1038/s41467-018-07998-x
34. Zhang Q, Chen Y, Liu K. miR-185 inhibits cell migration and invasion of hepatocellular carcinoma through CDC42. Oncol Lett. 2018;16(3):3101–3107.
35. Arias-Romero LE, Chernoff J. Targeting Cdc42 in cancer. Expert Opin Ther Targets. 2013;17(11):1263–1273. doi:10.1517/14728222.2013.828037
36. Xiao XH, Lv LC, Duan J, et al. Regulating Cdc42 and its signaling pathways in cancer: small molecules and MicroRNA as new treatment candidates. Molecules. 2018;23:4.
37. Aguilar BJ, Zhou H, Lu Q. Cdc42 signaling pathway inhibition as a therapeutic target in ras- related cancers. Curr Med Chem. 2017;24(32):3485–3507. doi:10.2174/0929867324666170602082956
38. Wang Y-X, Chen Y-R, Liu -S-S, et al. MiR-384 inhibits human colorectal cancer metastasis by targeting KRAS and CDC42. Oncotarget. 2016;7(51):84826–84838. doi:10.18632/oncotarget.12704
39. Li Y, Zhu X, Xu W, et al. miR-330 regulates the proliferation of colorectal cancer cells by targeting Cdc42. Biochem Biophys Res Commun. 2013;431(3):560–565. doi:10.1016/j.bbrc.2013.01.016
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.