Back to Journals » Infection and Drug Resistance » Volume 12
Escherichia coli O25b-ST131 and O16-ST131 causing urinary tract infection in women in Changsha, China: molecular epidemiology and clinical characteristics
Authors Zhong YM, Liu WE, Meng Q, Li Y
Received 17 April 2019
Accepted for publication 24 July 2019
Published 30 August 2019 Volume 2019:12 Pages 2693—2702
DOI https://doi.org/10.2147/IDR.S212658
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Joachim Wink
Yi-Ming Zhong,1 Wen-En Liu,1 Qian Meng,2 Yuan Li2
1Department of Clinical Laboratory, Xiangya Hospital, Central South University, Changsha, People’s Republic of China; 2Faculty of Laboratory Medicine, Xiangya School of Medicine, Central South University, Changsha, People’s Republic of China
Correspondence: Wen-En Liu
Department of Clinical Laboratory, Xiangya Hospital, Central South University, Changsha, Hunan Province 410008, People’s Republic of China
Tel +86 7 318 432 7437
Fax +86 7 318 432 7332
Email [email protected]
Objective: This study aimed to investigate the prevalence of Escherichia coli ST131 and molecularly characterize the O25b-ST131 and O16-ST131 subgroups among urinary tract infection (UTI) E. coli isolates from women in central China. We also assessed the clinical characteristics and outcomes of infections caused by E. coli ST131.
Methods: Between January 2014 and December 2015, a total of 216 consecutive, non-repetitive E. coli isolates were recovered from UTI urine samples from women in Changsha, China. All isolates were analyzed for phylogenetic groups, antimicrobial resistance and virulence genotypes. ST131 clonal groups were identified using PCR and characterized using O serotyping, CTX-M genotypes, fimH, gyrA, and parC alleles, fluoroquinolone resistance genes and pulsed-field gel electrophoresis (PFGE). Clinical data were obtained from medical records.
Results: Overall, 41 (19.0%) of 216 E. coli isolates were identified to contain ST131 strains, among which 27 were O25b-ST131 strains and 14 were O16-ST131 strains. The clinical characteristics and outcomes of the ST131 group did not differ significantly from those of the non-ST131 group, except for the presence of urinary stones (43.9% vs 27.4%, P=0.039). Ciprofloxacin resistance was found to be significantly higher in O25b-ST131 isolates than O16-ST131 isolates (96.3% vs 14.3%, P<0.001). The majority of O25b-ST131 isolates belonged to fimH30 (92.6%), followed by fimH41 (3.7%) and fimH27 (3.7%). O25b-H30 and O25b-H41 isolates were resistant to ciprofloxacin, and possessed gyrA1AB/parC1aAB combination. All of the O16-S131 isolates were found to belong to fimH41, and of which, two of the ciprofloxacin-resistant strains harbored gyrA1AB/parC3A combination. Three PFGE clusters, consisting of 38 (92.7%) isolates, with more than 70% similarity were identified.
Conclusion: The O25b and O16 sub-lineages have emerged as an important group of E. coli ST131 in UTI isolates from women in China. UTI patients with a history of urinary stones may need to be particularly vigilant against ST131 infection.
Keywords: E. coli, ST131, fimH-based subclones, fluoroquinolone resistance, urinary tract infection
Introduction
Urinary tract infection (UTI) is one of the most common infectious diseases. UTIs are much more common in women.1,2 It is estimated that almost half of all women will experience at least one UTI in their lifetime.1,2 Recurrent UTIs occur in more than 25% of affected women.3 Escherichia coli is the most common pathogen causing UTIs in humans and belongs to the large group of extraintestinal pathogenic E. coli (ExPEC).4–6 ExPEC is the leading cause of community-onset UTIs and nosocomial UTIs.6
In the last 10 years, the emergence and rapid global spread of E. coli sequence type 131 (ST131) with high virulence potential presents a severe threat to public health.7 ST131 is considered to be the predominant ExPEC strain throughout the world.7 Regarding the spread of ST131 among E. coli isolates causing UTIs, most studies have focused on extended-spectrum beta-lactamase (ESBL)-producing or fluoroquinolone-resistant (FQ-R) isolates.7 Fewer studies have evaluated the prevalence of ST131 among unselected UTI E. coli isolates, with dissimilarities between different subjects and geographic areas.8–10 ST131 in UTI E. coli isolates has been sporadically reported in two recent studies in mainland China,11,12 but this has not yet been investigated systematically. In addition, although the majority of ST131 isolates belong to the O25b:H4 serotype, a small subset of ST131 isolates with the O16:H5 serotype have recently been identified in several countries.13–16 In China, we first reported O16-ST131 in a previous study, and found that this type was the predominant subset among ST131 fecal E. coli isolates.17 However, two recent studies found that O16-ST131 accounted for 33.7% of ST131 among clinical E. coli isolates in mainland China,11 and 27.0% of ST131 among urinary E. coli isolates in Hong Kong.18 Considering the huge population of China (1.4 billion), more data are urgently needed. Furthermore, women, as a high-risk population, have not yet received the attention they deserve in most studies.8–12
Therefore, the aim of the current study was to determine the prevalence of ST131 and molecularly characterize the O25b-ST131 and O16-ST131 subgroups among unselected UTI E. coli isolates from women in central China. We also assessed the clinical characteristics and outcomes of infections caused by E. coli ST131.
Materials and methods
Bacterial isolates and clinical data collection
This study was conducted at Xiangya Hospital of Central South University, a 3500-bed tertiary care center located in Changsha, central China, with 7500–10,000 daily patient visits. A total of 216 consecutive, non-repetitive E. coli clinical isolates from female patients with UTIs, one from each patient, were collected from January 2014 to December 2015. Isolates were recovered from non-duplicate urine samples with significant E. coli bacteriuria (colony count >105 cfu/mL) and pyuria.
The demographic and clinical information of the patients was retrieved retrospectively, and included age, location, history of underlying diseases, use of urinary catheters or central venous catheters, received radiotherapy or chemotherapy within 1 month, corticosteroid or immunosuppressant use within 1 month, recent surgical procedures, treatment with antibiotics and outcomes.
Antimicrobial susceptibility testing
Susceptibility testing was performed using the Vitek 2 system (bioMérieux, Marcy-l’Étoile, France) and interpreted in accordance with Clinical and Laboratory Standards Institute (CLSI) criteria.19 ESBL producers were confirmed using the double-disc synergy test.19 Isolates with intermediate susceptibility were considered to be resistant. The resistance score was the number of antimicrobial agents to which the isolate was non-susceptible.17 Multidrug resistance was defined as resistance to three or more antimicrobial categories.17,20
Phylogenetic group, virulence genotyping, and ST131 detection
Major E. coli phylogenetic groups (A, B1, B2, C, D and E) were determined using PCR.21 The presence of 31 virulence genes was assessed through multiplex PCR and the virulence score was calculated as described previously.22,23 The ST131 lineages and O serotyping were identified as previously described.24
Molecular characterization of ST131
Detection of blaCTX-M was performed in ST131 isolates through multiplex PCR and sequencing.25,26 fimH, gyrA and parC subtyping was performed as previously described.27–29 The screening of plasmid-mediated quinolone resistance (PMQR) determinants [qnrA, qnrB, qnrC, qnrS, aac(6ʹ)-Ib-cr and qepA] was carried out using a multiplex PCR method.30
Pulsed-field gel electrophoresis (PFGE)
PFGE was performed on all ST131 isolates in accordance with a standardized protocol.31 Banding patterns were analyzed using BioNumerics software (Applied Maths, Sint-Martens-Latem, Belgium) and the Dice similarity coefficient. A dendrogram was constructed in accordance with the unweighted pair group method with arithmetic averages (UPGMA). Isolates were considered to belong to the same PFGE pattern if their Dice similarity index was ≥85%.
Statistical analysis
Categorical variables (clinical information, antimicrobial resistance patterns and virulence traits) were compared between different groups using a χ2 or Fisher’s exact test, as appropriate. Continuous variables (age, resistance and virulence score) were compared using Student’s t-test or the Mann-Whitney U test. The criterion for statistical significance was P<0.05. All statistical analyses were performed using SPSS Statistics for Windows, version 20.0 (SPSS Inc., Chicago, IL, USA).
Results
Distribution of phylogenetic groups, ST131 and O types
Phylogenetic analysis revealed that 100 (46.3%) of the 216 E. coli strains belonged to phylogroup B2, 36 (16.7%) to phylogroup D, 29 (13.4%) to phylogroup F, 28 (13.0%) to phylogroup A, 10 (4.6%) to phylogroup B1, 9 (4.2%) to phylogroup C, and 4 (1.8%) to phylogroup E.
Overall, 41 (19.0%) of the 216 E. coli strains were identified as ST131, and of these, 27 (65.9%) were from serogroup O25b and 14 (34.1%) from serogroup O16.
Epidemiological and clinical associations of ST131 status and O types
The epidemiological and clinical data of the 216 female patients with ST131 and non-ST131 E. coli isolates causing UTIs are demonstrated in Table 1. The clinical characteristics of the ST131 group did not differ significantly from the findings for the non-ST131 group, except for the presence of urinary stones (43.9% vs 27.4%, P=0.039). In addition, the clinical presentation and outcomes of patients infected with O16-ST131 were similar to those of patients infected with O25b-ST131 isolates.
 |
Table 1 Epidemiological and clinical data of 216 female patients with ST131 and non-ST131 E. coli isolates causing urinary tract infection |
Antimicrobial susceptibility associated with ST131 status and O types
ST131 isolates showed higher resistance proportions than non-ST131 isolates, except for resistance to nitrofurantoin. This difference was significant for ampicillin, ceftriaxone and ampicillin/sulbactam (Table 2). ST131 accounted for 23.9% of ESBL-producing isolates and 23.0% of the FQ-R isolates, but only 11.0% of non-ESBL-producing isolates and 13.8% of the FQ-susceptible (FQ-S) isolates (Table 3). However, there was no significant difference in the resistance scores of ST131 and non-ST131 isolates.
 |
Table 2 Prevalence of antimicrobial drug resistance in relation to ST131 status among 216 UTI E. coli isolates from women |
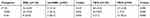 |
Table 3 Distribution by resistance group of ST131 among 216 E. coli isolates and O25b/O16 subgroups among 41 ST131 isolates |
As indicated in Table 2, the O25b-ST131 isolates had a significantly higher prevalence of resistance to cefazolin and ciprofloxacin compared with the O16-ST131 isolates. O25b subgroups accounted for 75.0% and 92.9% of ST131 isolates within ESBL and the FQ-R groups, but only 33.3% and 7.7% of those within non-ESBL-producing and FQ-S groups, respectively. In contrast, O16 subgroups accounted for 66.7% and 92.3% of ST131 isolates within non-ESBL-producing and FQ-S groups, but only 25.0% and 7.1% of those within ESBL and FQ-R groups, respectively (Table 3). Resistance scores were significantly higher for the O25b-ST131 isolates than the O16-ST131 isolates.
Virulence traits associated with ST131 status and O types
The positive rate of virulence genes and virulence scores in ST131 isolates were higher than those in non-ST131 isolates. This difference was significant for malX, bmaE, iutA, hlyA, kpsM II, traT, afa/draBC, cnf1, iha, ompT, sat, afa FM955459, usp, iucD, fyuA, F10 papA, and fimH (Table 4). However, the virulence genotypes and virulence scores of the O16-ST131 isolates resembled those of the O25b-ST131 isolates.
 |
Table 4 Prevalence of virulence traits in relation to ST131 status among 216 UTI E. coli isolates from women |
ST131 characterization
ESBL-producing E. coli ST131 prevalence was 78.0% (32/41). A total of 93.8% (30/32) of the isolates harbored blaCTX-M. Of the 30 isolates, 15 (50.0%) produced CTX-M-14, 6 (20.0%) CTX-M-55, 4 (13.3%) CTX-M-15, 4 (13.3%) CTX-M-27, and 1 (3.3%) CTX-M-24.
As indicated in Figure S1, the majority of the O25b-ST131 isolates belonged to fimH30 (25/27, 92.6%), followed by fimH41 (n=1) and fimH27 (n=1). O25b-H30 and O25b-H41 isolates were resistant to ciprofloxacin, and possessed a set of four amino acid replacement mutations (gyrA1AB Ser-83-Leu, Asp-87-Asn and parC1aAB Ser-80-Ile, Glu-84-Val) in the quinolone resistance-determining regions (QRDRs). However, all of the O16-ST131 isolates were found to belong to fimH41, and of which, two of the ciprofloxacin-resistant strains harbored three replacement mutations (gyrA1AB Ser-83-Leu, Asp-87-Asn and parC3A Ser-80-Ile). The aac(6ʹ)-Ib-cr gene was found in two ST131 isolates. None of the other types of PMQR determinant were detected.
The PFGE results showed that the 41 ST131 isolates comprised 28 PFGE patterns (defined at ≥85% similarity), the majority of which were grouped into three large clusters with a 70% similarity. These three PFGE clusters accounted for 38 (92.7%, 38/41) isolates (Figure S1).
Discussion
The overall rate of the ST131 clone in UTI-associated E. coli isolates from women was 19.0%, which is similar to that reported (18.5%) in male and female patients in Hong Kong in 2015 and slightly higher than that observed (15.4%) in the general population in mainland China in 2017.11,18 The high prevalence of this worldwide pandemic clone highlights the necessity to monitor the spread of the highly successful multidrug-resistant ST131 clonal group throughout China. The positivity rate of ESBLs in the ST131 strains was 78.0%, which is higher than that observed (35.1%) in the UK.8 The ST131 isolates were more resistant to ampicillin, ceftriaxone and ampicillin/sulbactam, but more susceptible to nitrofurantoin than the non-ST131 isolates. These results suggest that nitrofurantoin may be effective in treating UTIs caused by E. coli ST131.
Similar to previous studies,7,11 we found that O25b-ST131 was the predominant subclone among ST131 isolates. O16-ST131 accounted for a high proportion (34.1%) of the ST131 isolates in the current study, whereas this type accounted for 1% in Australia, 4.3% in Spain, 8% in France and 12% in Japan.7,13–16 The first report of O16-ST131 was from China in 2015 and occurred in fecal samples from healthy individuals,17 and it has now been identified across China. The distribution of serogroups we observed in the current study is very close to that reported recently from consecutive E. coli isolates from various clinical samples in Fuzhou, China.11 This emphasizes the importance of studying the O16 subclone further in this high-incidence country.
Some studies have found that older age was associated with ST131 infection.18,23,32 Although the differences did not reach statistical significance, patients with ST131 isolates were older than those with non-ST131 isolates in the present study. We found for the first time that the presence of urinary stones was significantly associated with patients infected with ST131 isolates, indicating that UTI patients with a history of urinary stones may need to be particularly vigilant against ST131 infections. As has been found in other studies,23,32,33 other clinical characteristics and mortality were similar between the different groups.
Some studies have reported that the ST131 isolates significantly surpassed the non-ST131 isolates in terms of the presence of antimicrobial resistance and/or virulence genes.23,34 The present study demonstrated similar results. Johnson et al considered that the resistance-plus-virulence combination associated with ST131 gave it a fitness advantage in the pathogenic niche.14,34 However, several recent studies using mouse models yielded discordant results as to whether ST131 is more virulent than other E. coli strains.14,35,36 We also found that patients with the ST131 isolates with more virulence factors and antimicrobial resistance did not have worse outcomes than patients with non-ST131 isolates, as previous studies reported.23,32 Therefore, further studies are needed to assess the correlation between pathogenic properties and the carriage of virulence genes as well as that between clinical implications and the carriage of virulence genes in ST131 strains.
We found that ST131 isolates mainly carried fimH30, in uropathogenic E. coli isolates in the current study. This result is in disagreement with our previous study in which H41 was the predominant fimH allele among ST131 isolates from intestinal commensal E. coli.17 Paul et al reported that the tip FimH adhesin of type 1 fimbriae, encoded by the fimH gene, was under strong positive selection for functional changes.37 In uropathogenic E. coli isolates, the monomannose-specific adherence of the NA114 variant (encoded by the fimH30 allele) was higher than the adherence of the SE15 FimH variant (encoded by the fimH41 allele).37 Additionally, increased monomannose-specific uroepithelial adhesion is considered to be commonly associated with point FimH mutations in uropathogenic strains of E. coli.38 In the current study, we reported for the first time that an O25b-ST131 isolate contained the fimH27 allele. This may be because fimH is under selection and probably gained point mutations. Further studies are required to verify this speculation and elucidate the evolution of the fimH allele in ST131 isolates.
Consistent with Johnson et al,24 we found that most of the O16-ST131 isolates were susceptible to ciprofloxacin. In contrast, 96.3% of the O25b-ST131 isolates were resistant to ciprofloxacin, suggesting that the O16-ST131 isolates have recently emerged among the fluoroquinolone-susceptible ST131 isolates. The main mechanism of fluoroquinolone resistance in ST131 isolates is amino acid substitutions within the QRDRs of GyrA and ParC, the fluoroquinolone targets.7 Our results also confirm this. The aac(6ʹ)-Ib-cr gene had been sporadically detected in ST131 isolates, which is similar to the results of previous studies.7,11 Additionally, the fluoroquinolone-susceptible isolates in the O16-ST131 subgroup were commonly found to have one or two mutations in the QRDRs. With long-term use and exposure to quinolones, these fluoroquinolone-susceptible isolates may be at risk of acquiring drug resistance.39
A limitation was that certain subgroups, particularly within non-ESBL-producing and FQ-S O25b/O16 sub-lineages, were small, increasing the probability of type II errors.40 Therefore, further multicenter and broad scale studies with larger sample sizes are needed to molecularly characterize these clonal groups in the future.
Conclusion
In summary, to the best of our knowledge, this is the first report to systematically characterize O25b-ST131 and O16-ST131 clonal groups among clinical UTI isolates from women in China. These results are particularly valuable as they are specific to women, a population at high risk of UTIs. Despite the lower ciprofloxacin resistance of the O16-ST131 isolates, these isolates did not exhibit significant differences in virulence genotypes or virulence scores compared with classic O25b-ST131 isolates. The results reveal for the first time that UTI patients with a history of urinary stones may need to be particularly vigilant against ST131 infection.
Ethical approval
This study was approved by the Ethics Committee of the Xiangya Hospital of Central South University, and the requirement for informed consent from patients was waived because the study was retrospective and used a database that ensured confidentiality.
Acknowledgments
We thank Prof. Fangyou Yu, Xin Xia , Jiayu Jiang and all the staff in the Microbiology Department of Xiangya Hospital for their kind help. This work was supported by National Natural Science Foundation of China (81672066).
Disclosure
The authors report no conflicts of interest in this work.
References
1. Foxman B. Epidemiology of urinary tract infections: incidence, morbidity, and economic costs. Am J Med. 2002;113 Suppl 1A:5S–13S. doi:10.1016/S0002-9343(02)01054-9
2. Foxman B, Barlow R, D’Arcy H, et al. Urinary tract infection: self-reported incidence and associated costs. Ann Epidemiol. 2000;10(8):509–515. doi:10.1016/s1047-2797(00)00072-7
3. Foxman B. Recurring urinary tract infection: incidence and risk factors. Am J Public Health. 1990;80(3):331–333. doi:10.2105/ajph.80.3.331
4. Yang Q, Zhang H, Wang Y, et al. Antimicrobial susceptibilities of aerobic and facultative gram-negative bacilli isolated from Chinese patients with urinary tract infections between 2010 and 2014. BMC Infect Dis. 2017;17(1):192. doi:10.1186/s12879-017-2296-x
5. Zhang H, Kong H, Yu Y, et al. Carbapenem susceptibilities of Gram-negative pathogens in intra-abdominal and urinary tract infections: updated report of SMART 2015 in China. BMC Infect Dis. 2018;18(1):493. doi:10.1186/s12879-018-3405-1
6. Pitout JD. Extraintestinal pathogenic Escherichia coli: a combination of virulence with antibiotic resistance. Front Microbiol. 2012;3:9. doi:10.3389/fmicb.2012.00009
7. Nicolas-Chanoine MH, Bertrand X, Madec JY. Escherichia coli ST131, an intriguing clonal group. Clin Microbiol Rev. 2014;27(3):543–574. doi:10.1128/CMR.00125-13
8. Gibreel TM, Dodgson AR, Cheesbrough J, et al. Population structure, virulence potential and antibiotic susceptibility of uropathogenic Escherichia coli from Northwest England. J Antimicrob Chemother. 2012;67(2):346–356. doi:10.1093/jac/dkr451
9. Brisse S, Diancourt L, Laouánan C, et al. Phylogenetic distribution of CTX-M- and non-extended-spectrum-β-lactamase-producing Escherichia coli isolates: group B2 isolates, except clone ST131, rarely produce CTX-M enzymes. J Clin Microbiol. 2012;50(9):2974–2981. doi:10.1128/jcm.00919-12
10. Banerjee R, Strahilevitz J, Johnson JR, et al. Predictors and molecular epidemiology of community-onset extended-spectrum β-lactamase-producing Escherichia coli infection in a Midwestern community. Infect Control Hosp Epidemiol. 2013;34(9):947–953. doi:10.1086/671725
11. Li B, Lu Y, Lan F, He Q, Li C, Cao Y. Prevalence and characteristics of ST131 clone among unselected clinical Escherichia coli in a Chinese university hospital. Antimicrob Resist Infect Control. 2017;6:118. doi:10.1186/s13756-017-0274-0
12. Qin X, Hu F, Wu S, et al. Comparison of adhesin genes and antimicrobial susceptibilities between uropathogenic and intestinal commensal Escherichia coli strains. PLoS One. 2013;8(4):e61169. doi:10.1371/journal.pone.0061169
13. Platell JL, Cobbold RN, Johnson JR, et al. Commonality among fluoroquinolone-resistant sequence type ST131 extraintestinal Escherichia coli isolates from humans and companion animals in Australia. Antimicrob Agents Chemother. 2011;55(8):3782–3787. doi:10.1128/AAC.00306-11
14. Banerjee R, Johnson JR. A new clone sweeps clean: the enigmatic emergence of Escherichia coli sequence type 131. Antimicrob Agents Chemother. 2014;58(9):4997–5004. doi:10.1128/AAC.02824-14
15. Dahbi G, Mora A, López C, et al. Emergence of new variants of ST131 clonal group among extraintestinal pathogenic Escherichia coli producing extended-spectrum beta-lactamases. Int J Antimicrob Agents. 2013;42(4):347–351. doi:10.1016/j.ijantimicag.2013.06.017
16. Blanc V, Leflon-Guibout V, Blanco J, et al. Prevalence of day-care centre children (France) with faecal CTX-M-producing Escherichia coli comprising O25b: H4and O16: H5 ST131 strains. J Antimicrob Chemother. 2014;69(5):1231–1237. doi:10.1093/jac/dkt519
17. Zhong YM, Liu WE, Liang XH, et al. Emergence and spread of O16-ST131 and O25b-ST131 clones among faecal CTX-M-producing Escherichia coli in healthy individuals in Hunan Province, China. J Antimicrob Chemother. 2015;70(8):2223–2227. doi:10.1093/jac/dkv114
18. Ho PL, Chu YP, Lo WU, et al. High prevalence of Escherichia coli sequence type 131 among antimicrobial-resistant E. coli isolates from geriatric patients. J Med Microbiol. 2015;64(Pt 3):243–247. doi:10.1099/jmm.0.000012
19. Clinical and Laboratory Standards Institute. Performance Standards for Antimicrobial Susceptibility Testing: Twenty-sixth Informational Supplement M100-S26. Wayne, PA: Clinical and Laboratory Standards Institute; 2016.
20. Jia H, Li W, Hou T, et al. The attributable direct medical cost of healthcare associated infection caused by multidrug resistance organisms in 68 hospitals of China. Biomed Res Int. 2019;2019:7634528. doi:10.1155/2019/7634528
21. Clermont O, Christenson JK, Denamur E, et al. The Clermont Escherichia coli phylo-typing method revisited: improvement of specificity and detection of new phylo-groups. Environ Microbiol Rep. 2013;5(1):58–65. doi:10.1111/1758-2229.12019
22. Johnson JR, Stell AL. Extended virulence genotypes of Escherichia coli strains from patients with urosepsis in relation to phylogeny and host compromise. J Infect Dis. 2000;181(1):261–272. doi:10.1086/315217
23. Blanco J, Mora A, Mamani R, et al. Four main virotypes among extended-spectrum-beta-lactamase-producing isolates of Escherichia coli O25b: H4-B2-ST131:bacterial, epidemiological, and clinical characteristics. J Clin Microbiol. 2013;51(10):3358–3367. doi:10.1128/JCM.01555-13
24. Johnson JR, Clermont O, Johnston B, et al. Rapid and specific detection, molecular epidemiology, and experimental virulence of the O16 subgroup within Escherichia coli sequence type 131. J Clin Microbiol. 2014;52(5):1358–1365. doi:10.1128/JCM.03502-13
25. Woodford N, Fagan EJ, Ellington MJ. Multiplex PCR for rapid detection of genes encoding CTX-M extended-spectrum (beta)-lactamases. J Antimicrob Chemother. 2006;57(1):154–155. doi:10.1093/jac/dki412
26. Grobner S, Linke D, Schutz W, et al. Emergence of carbapenem-non-susceptible extended-spectrum beta-lactamase- producing Klebsiella pneumoniae isolates at the university hospital of Tubingen, Germany. J Med Microbiol. 2009;58(Pt 7):912–922. doi:10.1099/jmm.0.005850-0
27. Weissman SJ, Johnson JR, Tchesnokova V, et al. High-resolution two-locus clonal typing of extraintestinal pathogenic Escherichia coli. Appl Environ Microbiol. 2012;78(5):1353–1360. doi:10.1128/AEM.06663-11
28. Leflon-Guibout V, Jurand C, Bonacorsi S, et al. Emergence and spread of three clonally related virulent isolates of CTX-M-15-producing Escherichia coli with variable resistance to aminoglycosides and tetracycline in a French geriatric hospital. Antimicrob Agents Chemother. 2004;48(10):3736–3742. doi:10.1128/AAC.48.10.3736-3742.2004
29. Johnson JR, Tchesnokova V, Johnston B, et al. Abrupt emergence of a single dominant multidrug-resistant strain of Escherichia coli. J Infect Dis. 2013;207(6):919–928. doi:10.1093/infdis/jis933
30. Kim HB, Park CH, Kim CJ, et al. Prevalence of plasmid-mediated quinolone resistance determinants over a 9-year period. Antimicrob Agents Chemother. 2009;53(2):639–645. doi:10.1128/AAC.01051-08
31. Ribot EM, Fair MA, Gautom R, et al. Standardization of pulsed-field gel electrophoresis protocols for the subtyping of Escherichia coli O157: H7, Salmonella, and Shigella for PulseNet. Foodborne Pathog Dis. 2006;3(1):59–67. doi:10.1089/fpd.2006.3.59
32. Banerjee R, Johnston B, Lohse C, et al. Escherichia coli sequence type 131 is a dominant, antimicrobial-resistant clonal group associated with healthcare and elderly hosts. Infect Control Hosp Epidemiol. 2013;34(4):361–369. doi:10.1086/669865
33. Kim YA, Lee K, Chung JE. Risk factors and molecular features of sequence type (ST) 131 extended-spectrum-β-lactamase-producing Escherichia coli in community-onset female genital tract infections. BMC Infect Dis. 2018;18(1):250. doi:10.1186/s12879-018-3168-8
34. Johnson JR, Johnston B, Clabots C, et al. Escherichia coli sequence type ST131 as the major cause of serious multidrug-resistant E. coli infections in the United States. Clin Infect Dis. 2010;51(3):286–294. doi:10.1086/653932
35. Vimont S, Boyd A, Bleibtreu A, et al. The CTX-M-15-producing Escherichia coli clone O25b: H4-ST131 has high intestine colonization and urinary tract infection abilities. PLoS One. 2012;7(9):e46547. doi:10.1371/journal.pone.0046547
36. Johnson JR, Porter SB, Zhanel G, et al. Virulence of Escherichia coli clinical isolates in a murine sepsis model in relation to sequence type ST131 status, fluoroquinolone resistance, and virulence genotype. Infect Immun. 2012;80(4):1554–1562. doi:10.1128/IAI.06388-11
37. Paul S, Linardopoulou EV, Billig M, et al. Role of homologous recombination in adaptive diversification of extraintestinal Escherichia coli. J Bacteriol. 2013;195(2):231–242. doi:10.1128/JB.01524-12
38. Weissman SJ, Beskhlebnaya V, Chesnokova V, et al. Differential stability and trade-off effects of pathoadaptive mutations in the Escherichia coli fimH adhesin. Infect Immun. 2007;75(7):3548–3555. doi:10.1128/iai.01963-06
39. Matsumura Y, Yamamoto M, Nagao M, et al. Association of fluoroquinolone resistance, virulence genes, and IncF plasmids with extended-spectrum-beta-lactamase-producing Escherichia coli sequence type 131 (ST131) and ST405 clonal groups. Antimicrob Agents Chemother. 2013;57(10):4736–4742. doi:10.1128/AAC.00641-13
40. Kudinha T, Johnson JR, Andrew SD, et al. Escherichia coli sequence type 131 as a prominent cause of antibiotic resistance among urinary Escherichia coli isolates from reproductive-age women. J Clin Microbiol. 2013;51(10):3270–3276. doi:10.1128/JCM.01315-13
Supplementary material
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.

