Back to Journals » OncoTargets and Therapy » Volume 13
ERα, A Key Target for Cancer Therapy: A Review
Received 29 October 2019
Accepted for publication 20 February 2020
Published 11 March 2020 Volume 2020:13 Pages 2183—2191
DOI https://doi.org/10.2147/OTT.S236532
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Prof. Dr. Takuya Aoki
Yanfang Liu,* Hong Ma,* Jing Yao
Cancer Center, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Jing Yao
Cancer Center, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Jiefang Road 1277, Wuhan, Hubei Province 430022, People’s Republic of China
Tel +86-189 8627 1157
Email [email protected]
Abstract: Estrogen receptor α (ERα) is closely associated with both hormone-dependent and hormone-independent tumors, and it is also essential for the development of these cancers. The functions of ERα are bi-faceted; it can contribute to cancer progression as well as cancer inhibition. Therefore, understanding ERα is vital for the treatment of those cancers that are closely associated with its expression. Here, we will elaborate on ERα based on its structure, localization, activation, modification, and mutation. Also, we will look at co-activators of ERα, elucidate the signaling pathway activated by ERα, and identify cancers related to its activation. A comprehensive understanding of ERα could help us to find new ways to treat cancers.
Keywords: ERα, estrogen receptors, estradiol, signaling pathway, cancer
Introduction
Estrogen receptors (ERs) consist of nuclear ERs, extra-nuclear ERs, and G protein-coupled ERs (GPERs).1 Nuclear ERs, including estrogen receptor α (ERα) and estrogen receptor β (ERβ), are located in the nucleus and are encoded by ESR1 and ERS2, respectively.2 Once activated, nuclear ERs transcriptionally regulate the expression of targeted genes.3 Extra-nuclear ERs include cytosolic ERα and ERβ, both of which are located in the plasma membrane.4 GPERs are expressed both in the plasma membrane and cytoplasm,5 and are structurally different from ERα and ERβ.6 ERs show similar main structures; however, their sequential homology is as low as 47%.2 The different functions of ERs depend on structural differences. ERs can be activated when cells are exposed to estrogen.7–9 Emerging evidence shows that the activation of ERs is highly associated with cancer formation and metastasis, 10–12 extracellular matrix (ECM) remodeling2,13 and drug resistance.14–17
Here, we focus on providing a comprehensive understanding of ERα. We hope this will help doctors to find more effective ways to treat ERα-related cancers.
The Structure of ERα
ERα was the first ER to be discovered and cloned.9 The gene ESR1 that encodes ERα is located on chromosome 6.18 As shown in Figure 1, the ERα protein consists of 595 amino acids with a molecular weight of approximately 66.2kD.18 The ERα protein contains six domains (A-F), three of which are functionally significant.19 The three functional domains are the N-terminal A/B domain (NTD), the C domain (which includes the DNA-binding domain, DBD), and the E domain (the ligand-binding domain, LBD).19 NTD has a low degree of conservation and contains AF-1, which has the function of transcriptional activation and is also the main reason for ERα’s endocrine-sensitivity.20 AF-1 is critical to the transactivation function and shows the highest variability among ERs.2 DBD in the C domain is highly conserved and exerts its function by binding to the estrogen-responsive element (ERE), which subsequently regulates the expression of target genes.21 The D domain shows 30% homology among ERs and links the C and E domains.22,23 LBD (also called AF-2) or the E domain, showing 55% homology with other ERs, is mainly involved in protein and estradiol (E2) binding.22 LBD combines with estrogen to form a homodimer that regulates gene suppression and activation and contributes to transcriptional activation.22,23 Studies have also shown that LBD is responsible for nuclear localization.22,24 The F domain, which is not conserved and shows only 18% homology, is regarded as an extension of the E domain.22 Although the structure of ERα has been studied extensively, the function of the F domain has not been clarified. Understanding the structure of ERα is essential for the treatment of ERα-over-expressing cancers.
Localization and Activation of ERα
ERα is widely expressed in human tissues, including breast, prostate, uterus, liver, and bone.25 As stated above, there are two types of ERα, nuclear and extra-nuclear. Proteins are generally synthesized in the ribosome and then relocated under the guidance of a signal peptide.26 In the nuclear ERα, the LBD region contains nuclear localization signals that guide the estrogen-ERα homodimer transfer from the cytoplasm to the nucleus.24,27 Once ERα has been relocated to the nucleus, its DBD then links with an ERE on the DNA.4,9,28 Through this process, nuclear ERα is activated.4,9,28
Activated nuclear ERα regulates the expression of target genes by activating transcription factors downstream.29 The E domain is fundamental to membrane translocation of ERα.30 Studies have shown that membrane ERα acts as a kind of G protein-coupled receptor, activates G proteins, and stimulates G protein-induced signal transduction.31,32 Therefore, the interaction between E2 and membrane ERα activates various signaling pathways and signaling molecules, subsequently triggering downstream gene transcription and affecting cancer progression.33–39 It is, for that reason, understandable that different locations of ERα exert distinct functions in multiple ways.
Post-Translational Modification and Function of ERα
Proteins exert their functions, including phosphorylation and dephosphorylation, lipidation or palmitoylation, methylation, acetylation, and SUMOylation, after post-translational modifications.40–42 Common post-translational modifications of For ERα include phosphorylation, palmitoylation, and ubiquitination.43–47 Studies have revealed that frequent phosphorylation sites of ERα are Ser118, Ser167, and Ser305.43,44 The phosphorylation of these three sites leads to cancer progression, tumor metastasis, and endocrine therapy resistance.43,44 Interestingly, the phosphorylation of Ser305 activates the phosphorylation of Ser118, which subsequently promotes cancer development.48
The palmitoylation site of ERα is Cys-447, and studies have demonstrated that the palmitoylation of ERα is essential for locating ERα in the plasma membrane.4,49 By binding to E2, the palmitoylation of ERα activates downstream signaling pathways.45 The ubiquitination of ERα is the primary way of degrading ERα. However, emerging evidence shows that the function of the ubiquitination of ERα is complicated.46,47,50 ERα ubiquitination promotes tumorigenesis in hepatocellular carcinoma,46 resulting in the slow growth of cancer cells in breast cancer.47,50 In conclusion, the function of ERα is dependent on post-translational modifications.
Mutation of ERα
ER-positive (ER+) breast cancer has a good prognosis, mainly owing to endocrine therapy,51,52 which has shown great success.52 However, endocrine resistance is partially responsible for patient relapse,53–55 and the mutation of ERα plays a significant part in endocrine resistance.56 Modification of ERα frequently results in changes in the activity of ERα and variations in protein expression and function, which lead to the proliferation of cancer cells.56,57
ERα mutations are commonly observed in ER+ breast cancer. Two ESR1 mutations, Y537S and D538G, are most easily identified.56,58 Investigations have demonstrated that ESR1 mutations result in cancer cell resistance to tamoxifen (TAM) in breast cancer patients.56,58 Y537S mutants reportedly are not dependent on estrogen, but D538G mutants are.56 Both mutants have been shown to be associated with endocrine resistance,56 and neither change the ability of ERs to bind to transcription factors.59–61 We may, therefore, conclude that the mutation of ERα is critical for cancer development and drug resistance.
Co-Activators of ERα
ERα regulates the expression of its target genes through the participation of its co-activators.62 In the presence of E2, co-activators combine with ERα and subsequently activate transcription factors, which contribute to the transcription of target genes (Figure 2).62 Many co-regulators have been found; however, their mechanism of action is not always clear Co-activators act as co-regulators, exerting their effect through various mechanisms. Specifically, SRC-1 and SRC-2 are functionally similar and contribute to the activation of ERα.63–65 Previous research revealed that SRC-1 and SRC-2 could lead to resistance to TAM in ER+ breast cancer patients,66 while another investigation demonstrated that SRC-3 is overexpressed in breast cancer and acts as a selective activator of ERα.66 In vivo experiments showed that SWI2/SNF2 protein enhanced gene transcription by interacting with the AF-2 domain,67 and PBP contributed to mammary epithelial differentiation in breast cancer.68 AIB1 interacts with ERs and resulting enhancement of estrogen-related gene transcription, which leads to development of breast and ovarian cancer.64 There are other co-activators whose functions are unclear.65 In all, many co-activators have been discovered that work together with ERα to co-regulate the expression of target genes. More co-activators will undoubtedly be studied in the future, which should be very helpful in understanding the mechanisms by which ERα regulates its target genes.
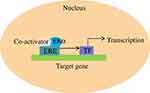 |
Figure 2 ERα’s contribution to the transcription of target genes with the help of co-activators. |
ERα and Signaling Pathways
Studies have shown that the activation of ERα leads to the activation of downstream signaling pathways.69,70 In endometrial carcinoma, estrogen contributes to carcinogenesis by activating ERα, which subsequently activates the downstream signaling pathways of phosphatidylinositol 3-kinase (PI3K)/AKT and mitogen-activated protein kinase/extracellular signal-regulated kinase (MAPK/ERK) (Figure 3). 69,71 In ER+ breast cancer, estrogen activates the PI3K/AKT/mTOR signaling pathway by associating with extra-nuclear ERα, which results in drug resistance and epithelial-to-mesenchymal transition (EMT) (Figure 3).70,72 Targeting ERα reportedly causes changes in the expression of components of the PI3K/AKT- protein kinase Cα signaling pathway, resulting in cell apoptosis.73 Also, the activation of ERα results in increased expression of the PI3K/AKT/NF-κB signaling pathway, leading to tumor invasion and metastasis in breast cancer.74
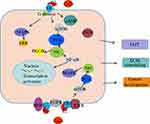 |
Figure 3 The signaling pathways in which ERα is involved. |
As discussed above, membrane ERα is linked to G proteins, transmitting signals from the outside to the inside of the cell.75 Downstream signaling pathways, including adenosine monophosphate (cAMP) signaling,33 PI3K/AKT, and endothelial nitric oxide synthase, are activated after receiving signals.76,77 As a result, cAMP levels increase, and the mobilization of Ca2+ is rapidly enhanced in the presence of estrogen; this contributes to the activation of estrogen signaling by activating the C-terminal of ERα (Figure 3).78,79 Emerging evidence shows that the membrane ERα activated by E2 interacts with signaling molecules, including PI3K, MAPK, AKT, p21ras, and PKC, contributing to the cascade amplification reaction of signaling molecules.2,80 Reportedly, the activation of ERα leads to the activation of human epidermal growth factor receptor 2 and epidermal growth factor receptor (EGFR), resulting in the upregulation of the mTOR/PI3K/AKT/MAPK signaling pathway.81 In breast cancer, ERα activation contributes to cancer progression by binding to IGF-IR, which subsequently activates the IGF pathway (Figure 3).82,83
Overall, ERα is extremely important in cancer progression. Understanding the mechanisms involving ERα is key to treating cancers.
ERα and Cancer
ERα is critical to the development of ER+ breast cancer,84 which accounts for approximately 70% of all breast cancers.7,85 Overexpression of ERα frequently sensitizes tumors to endocrine therapy.84 When exposed to E2, ERα activation stimulates downstream signaling pathways,86 and leads to EMT and ECM remodeling (Figure 3).87,88 In ER+ breast cancer, estrogen contributes to cancer progression by activating the PI3K/AKT signaling pathway.89,90 In the ER+ breast cancer cell line MCF-7, calcium mediates the activation of estrogen signaling.78 Overall in all, ERα plays a significant part in the progression of ER+ breast cancer.
ERα is widely expressed in cells and has a critical role in both hormone-dependent and hormone-in dependent cancers. In hormone-related cancers, such as breast, endometrial and ovarian cancers, ERα expression contributes to disease progression mostly by regulating the PI3K/AKT signaling pathway.69,73 Emerging evidence shows that ERα is also crucial to the progression of prostate cancer.91 Overexpression of ERα in prostate cancer is strongly associated with adverse survival outcomes.91 ERα acts as an oncogene and contributes to the development of prostate cancer by inducing EMT and the activation of matrix metalloproteinases.92,93 However, ERα also has a key role in inhibiting tumor development, maintaining the luminal phenotype, and restoring the sensitivity of breast cancer to hormone therapy.94 In hormone-independent cancers, such as colorectal cancer, ERα expression was shown to inhibit tumors in women.95 In non-small-cell lung cancer, ERα expression contributed to sensitivity to pemetrexed and carboplatin.96 However, high ERα expression is also significantly related to poor survival outcomes in colorectal cancer patients.97 Therefore, we can conclude that the regulation of ERα is complicated, and its role is bi-faceted.
Conclusions and Perspectives
Study have shown that changes in expression of ERα, ERβ, and GPERs greatly affect cell proliferation and cancer development.98 As discussed above, the functions of ERs are bi-faceted. ERβ also exerts its functions through various mechanisms. In triple-negative breast cancer cells, ERβ suppresses tumor progression by interacting with androgen receptors.99 ERβ also contributes to beneficial gut microbiota diversity, which suppresses colorectal cancer development.100 However, in prostate cancer cell line PC-3, ERβ exerts its oncogenic effect by activating β-catenin and regulating the PI3K/AKT signaling pathway.101 Therefore, the effects of ERβ in cancer cells are complicated.
The functions of GPERs are also multi-faceted. In hormone-dependent cancers, such as breast cancer and endometrial cancer, GPER expression leads to tumor progression. Specifically, analysis of data from a subset of breast cancer patients showed that GPER-1 expression was positively correlated with overexpression of EGFR.102 In TAM-resistant breast cancer cells, GPER-1/EGFR receptor signaling contributes to the development of TAM resistance,103 indicating that either GPER-1 exerts its function by regulating EGFR or there is a mutual regulation between the two. In breast cancer MDA-MB-231 cells, down-regulation of GPER induces inhibition of cell proliferation and tumor metastasis.104 In endometrial cancer, GPER-1 promotes cell growth by binding to autocrine motility factor.105 GPER also contributes to insulin-driven endometrial cancer cell proliferation by regulating the PI3K/AKT signaling pathway.106 Overall, GPER expression contributes to the development of hormone-dependent cancers. However, in hormone-independent cancers, such as colorectal cancer, the relationship between GPER expression and tumor progression is more complicated. In ERβ-negative colorectal cancer cells, GPER-induced hypoxic condition leads to tumor development.107 However, another study reported that GPER −1 inhibits the activation of NF-κB by the canonical IKKα/IκBα pathway. In vivo experiments confirmed that GPER-1 suppresses progression of colorectal cancer.108 Overall, GPER has complicated functions in cancers.
As important ERs, ERα, ERβ, and GPER do not function independently from each other. Cross-regulation among ERs has an important role in physiological activities and biological behaviors. In zebrafish, ERα is a core factor, interacting with ERβ and GPER to regulate vitellogenesis.109 In vivo experiments showed that ERβ and GPER-1 co-regulate the effects of E2 on arginine-vasopressin immunoreactivity.110 In human renal tubular epithelial cells, E2 leads to cell proliferation via ERα and GPER-1.111 In vitro experiments showed that ERβ suppressed the transcriptional and oncogenic effects of ERα.112,113 The functions of ERα and ERβ are antagonistic; therefore, their ratio is important in the development of diseases. An ERβ/ERα ratio lower than 0.85 was associated with and could potentially be used to predict endoscopic activity in Crohn’s disease.114 In conclusion, the expression changes of different ERs are associated with abnormal regulation and disorders.
ERα is localized in the nucleus and the plasma membrane; however, the membrane-localized receptors mediate faster signal transduction via the MAPK/ERK, PI3K/AKT, and p38/MAPK signaling pathways.115,116 In this review, we emphasize that ERα expression is closely linked to cancer development.33 The activation of ERα by estrogen leads to tumor progression and metastasis, which subsequently promotes the transduction of downstream signaling pathways.82,83 Currently, ERα antagonists such as TAM are widely used in clinical settings with great success.117 Nevertheless, endocrine resistance remains partially responsible for patient relapse.53–55 TAM is structurally similar to estrogen and competitively combines with ERs, subsequently blocking the entry of estrogen into tumor cells and inhibiting the development of cancers.118 However, resistance to TAM has multiple mechanisms, including ER mutation, loss of ER expression, overexpression of ER co-activators, activation of the EGFR or PI3K/AKT signaling pathway, epigenomic and post-translational modifications in ER, and enhanced mitochondrial metabolism of TAM.56,119–124 Endocrine therapy resistance is a challenge, and successfully solving this problem would greatly benefit cancer patients. This review provides a comprehensive understanding of ERα, which we hope will help in the search for new ways to treat ERα-related cancers.
Acknowledgments
This review was supported by the Joint Fund of Wuhan Union Hospital (No.02.03.2019-95). In addition, we thank Ye Wang and Xinyang Du for valuable suggestions during the process of manuscript revision.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Butler MJ, Hildebrandt RP, Eckel LA. Selective activation of estrogen receptors, ERalpha and GPER-1, rapidly decreases food intake in female rats. Horm Behav. 2018;103:54–61. doi:10.1016/j.yhbeh.2018.05.018
2. Piperigkou Z, Karamanos NK. Estrogen receptor-mediated targeting of the extracellular matrix network in cancer. Semin Cancer Biol. 2019. doi:10.1016/j.semcancer.2019.07.006
3. Nilsson S, Makela S, Treuter E, et al. Mechanisms of estrogen action. Physiol Rev. 2001;81(4):1535–1565. doi:10.1152/physrev.2001.81.4.1535
4. Pedram A, Razandi M, Sainson RC, Kim JK, Hughes CC, Levin ER. A conserved mechanism for steroid receptor translocation to the plasma membrane. J Biol Chem. 2007;282(31):22278–22288. doi:10.1074/jbc.M611877200
5. Tominna R, Chokr S, Feri M, Chuon T, Sinchak K. Plasma membrane G protein-coupled estrogen receptor 1 (GPER) mediates rapid estradiol facilitation of sexual receptivity through the orphanin-FQ-ORL-1 system in estradiol primed female rats. Horm Behav. 2019;112:89–99. doi:10.1016/j.yhbeh.2019.04.004
6. Carmeci C, Thompson DA, Ring HZ, Francke U, Weigel RJ. Identification of a gene (GPR30) with homology to the G-protein-coupled receptor superfamily associated with estrogen receptor expression in breast cancer. Genomics. 1997;45(3):607–617. doi:10.1006/geno.1997.4972
7. Leone S, Busonero C, Acconcia F. A high throughput method to study the physiology of E2: eRalphasignaling in breast cancer cells. J Cell Physiol. 2018;233(5):3713–3722. doi:10.1002/jcp.26251
8. He WH, Jin MM, Liu AP, et al. Estradiol promotes trophoblast viability and invasion by activating SGK1. Biomed Pharmacother. 2019;117:109092. doi:10.1016/j.biopha.2019.109092
9. Dahlman-Wright K, Cavailles V, Fuqua SA, et al. International union of pharmacology. LXIV. Estrogen receptors. Pharmacol Rev. 2006;58(4):773–781. doi:10.1124/pr.58.4.8
10. Li Q, Gao H, Yang H, Wei W, Jiang Y. Estradiol promotes the progression of ER+ breast cancer through methylation-mediated RSK4 inactivation. Onco Targets Ther. 2019;12:5907–5916. doi:10.2147/OTT.S208988
11. Jallow F, O’Leary KA, Rugowski DE, Guerrero JF, Ponik SM, Schuler LA. Dynamic interactions between the extracellular matrix and estrogen activity in progression of ER+ breast cancer. Oncogene. 2019;38(43):6913–6925.
12. Che Q, Xiao X, Jun X, et al. 17beta-estradiol promotes endometrial cancer proliferation and invasion through IL-6 pathway. Endocr Connect. 2019.
13. Menendez-Menendez J, Hermida-Prado F, Granda-Diaz R, et al. Deciphering the molecular basis of melatonin protective effects on breast cells treated with doxorubicin: TWIST1 a transcription factor involved in EMT and metastasis, a novel target of melatonin. Cancers (Basel). 2019;11:7. doi:10.3390/cancers11071011
14. Wu Y, Zhang Z, Cenciarini ME, et al. Tamoxifen resistance in breast cancer is regulated by the EZH2-ERalpha-GREB1 transcriptional axis. Cancer Res. 2018;78(3):671–684. doi:10.1158/0008-5472.CAN-17-1327
15. Selmin OI, Donovan MG, Skovan B, Paine-Murieta GD, Romagnolo DF. Arsenicinduced BRCA1 CpG promoter methylation is associated with the downregulation of ERalpha and resistance to tamoxifen in MCF7 breast cancer cells and mouse mammary tumor xenografts. Int J Oncol. 2019;54(3):869–878. doi:10.3892/ijo.2019.4687
16. Button B, Croessmann S, Chu D, et al. The estrogen receptor-alpha S118P variant does not affect breast cancer incidence or response to endocrine therapies. Breast Cancer Res Treat. 2019;174(2):401–412. doi:10.1007/s10549-018-05087-7
17. Arpino G, Wiechmann L, Osborne CK, Schiff R. Crosstalk between the estrogen receptor and the HER tyrosine kinase receptor family: molecular mechanism and clinical implications for endocrine therapy resistance. Endocr Rev. 2008;29(2):217–233. doi:10.1210/er.2006-0045
18. Jameera Begam A, Jubie S, Nanjan MJ. Estrogen receptor agonists/antagonists in breast cancer therapy: a critical review. Bioorg Chem. 2017;71:257–274. doi:10.1016/j.bioorg.2017.02.011
19. Ogawa S, Inoue S, Watanabe T, et al. The complete primary structure of human estrogen receptor beta (hER beta) and its heterodimerization with ER alpha in vivo and in vitro. Biochem Biophys Res Commun. 1998;243(1):122–126. doi:10.1006/bbrc.1997.7893
20. Kuiper GG, Enmark E, Pelto-Huikko M, Nilsson S, Gustafsson JA. Cloning of a novel receptor expressed in rat prostate and ovary. Proc Natl Acad Sci U S A. 1996;93(12):5925–5930. doi:10.1073/pnas.93.12.5925
21. Kumar V, Chambon P. The estrogen receptor binds tightly to its responsive element as a ligand-induced homodimer. Cell. 1988;55(1):145–156. doi:10.1016/0092-8674(88)90017-7
22. Ruff M, Gangloff M, Wurtz JM, Moras D. Estrogen receptor transcription and transactivation: structure-function relationship in DNA- and ligand-binding domains of estrogen receptors. Breast Cancer Res. 2000;2(5):353–359. doi:10.1186/bcr80
23. Huang W, Peng Y, Kiselar J, et al. Multidomain architecture of estrogen receptor reveals interfacial cross-talk between its DNA-binding and ligand-binding domains. Nat Commun. 2018;9(1):3520. doi:10.1038/s41467-018-06034-2
24. Beato M, Sanchez-Pacheco A. Interaction of steroid hormone receptors with the transcription initiation complex. Endocr Rev. 1996;17(6):587–609. doi:10.1210/edrv-17-6-587
25. Mokarram P, Alizadeh J, Razban V, Barazeh M, Solomon C, Kavousipour S. Interconnection of estrogen/testosterone metabolism and mevalonate pathway in breast and prostate cancers. Curr Mol Pharmacol. 2017;10(2):86–114. doi:10.2174/1874467209666160112125631
26. Frydman J. Folding of newly translated proteins in vivo: the role of molecular chaperones. Annu Rev Biochem. 2001;70:603–647. doi:10.1146/annurev.biochem.70.1.603
27. Evans RM. The steroid and thyroid hormone receptor superfamily. Science. 1988;240(4854):889–895. doi:10.1126/science.3283939
28. Vanacker JM, Pettersson K, Gustafsson JA, Laudet V. Transcriptional targets shared by estrogen receptor- related receptors (ERRs) and estrogen receptor (ER) alpha, but not by ERbeta. EMBO J. 1999;18(15):4270–4279. doi:10.1093/emboj/18.15.4270
29. Kos M, Reid G, Denger S, Gannon F. Minireview: genomic organization of the human ERalpha gene promoter region. Mol Endocrinol. 2001;15(12):2057–2063. doi:10.1210/mend.15.12.0731
30. Razandi M, Alton G, Pedram A, Ghonshani S, Webb P, Levin ER. Identification of a structural determinant necessary for the localization and function of estrogen receptor alpha at the plasma membrane. Mol Cell Biol. 2003;23(5):1633–1646. doi:10.1128/MCB.23.5.1633-1646.2003
31. Razandi M, Pedram A, Greene GL, Levin ER. Cell membrane and nuclear estrogen receptors (ERs) originate from a single transcript: studies of ERalpha and ERbeta expressed in Chinese hamster ovary cells. Mol Endocrinol. 1999;13(2):307–319. doi:10.1210/mend.13.2.0239
32. Wyckoff MH, Chambliss KL, Mineo C, et al. Plasma membrane estrogen receptors are coupled to endothelial nitric-oxide synthase through Galpha(i). J Biol Chem. 2001;276(29):27071–27076. doi:10.1074/jbc.M100312200
33. Aronica SM, Kraus WL, Katzenellenbogen BS. Estrogen action via the cAMP signaling pathway: stimulation of adenylate cyclase and cAMP-regulated gene transcription. Proc Natl Acad Sci U S A. 1994;91(18):8517–8521. doi:10.1073/pnas.91.18.8517
34. Castoria G, Barone MV, Di Domenico M, et al. Non-transcriptional action of oestradiol and progestin triggers DNA synthesis. EMBO J. 1999;18(9):2500–2510. doi:10.1093/emboj/18.9.2500
35. Cooper LF, Tiffee JC, Griffin JP, Hamano H, Guo Z. Estrogen-induced resistance to osteoblast apoptosis is associated with increased hsp27 expression. J Cell Physiol. 2000;185(3):401–407. doi:10.1002/(ISSN)1097-4652
36. Le Mellay V, Grosse B, Lieberherr M. Phospholipase C beta and membrane action of calcitriol and estradiol. J Biol Chem. 1997;272(18):11902–11907. doi:10.1074/jbc.272.18.11902
37. Levin ER. Cellular functions of the plasma membrane estrogen receptor. Trends Endocrinol Metab. 1999;10(9):374–377. doi:10.1016/S1043-2760(99)00192-7
38. Razandi M, Pedram A, Levin ER. Plasma membrane estrogen receptors signal to antiapoptosis in breast cancer. Mol Endocrinol. 2000;14(9):1434–1447. doi:10.1210/mend.14.9.0526
39. Wise PM, Dubal DB, Wilson ME, Rau SW, Bottner M. Minireview: neuroprotective effects of estrogen-new insights into mechanisms of action. Endocrinology. 2001;142(3):969–973. doi:10.1210/endo.142.3.8033
40. Xu FQ, Xue HW. The ubiquitin-proteasome system in plant responses to environments. Plant Cell Environ. 2019.
41. Ficarro SB, McCleland ML, Stukenberg PT, et al. Phosphoproteome analysis by mass spectrometry and its application to Saccharomyces cerevisiae. Nat Biotechnol. 2002;20(3):301–305. doi:10.1038/nbt0302-301
42. Davis BG. Biochemistry. Mimicking posttranslational modifications of proteins. Science. 2004;303(5657):480–482. doi:10.1126/science.1093449
43. Anbalagan M, Rowan BG. Estrogen receptor alpha phosphorylation and its functional impact in human breast cancer. Mol Cell Endocrinol. 2015;418(Pt 3):264–272. doi:10.1016/j.mce.2015.01.016
44. Park J, Lee Y. Hypoxia induced phosphorylation of estrogen receptor at serine 118 in the absence of ligand. J Steroid Biochem Mol Biol. 2017;174:146–152. doi:10.1016/j.jsbmb.2017.08.013
45. Adlanmerini M, Solinhac R, Abot A, et al. Mutation of the palmitoylation site of estrogen receptor alpha in vivo reveals tissue-specific roles for membrane versus nuclear actions. Proc Natl Acad Sci U S A. 2014;111(2):E283–E290. doi:10.1073/pnas.1322057111
46. Wu H, Yao S, Zhang S, et al. Elevated expression of Erbin destabilizes ERalpha protein and promotes tumorigenesis in hepatocellular carcinoma. J Hepatol. 2017;66(6):1193–1204. doi:10.1016/j.jhep.2017.01.030
47. Yang H, Yu N, Xu J, et al. SMURF1 facilitates estrogen receptor a signaling in breast cancer cells. J Exp Clin Cancer Res. 2018;37(1):24. doi:10.1186/s13046-018-0672-z
48. Oladimeji P, Skerl R, Rusch C, Diakonova M. Synergistic activation of ERalpha by estrogen and prolactin in breast cancer cells requires tyrosyl phosphorylation of PAK1. Cancer Res. 2016;76(9):2600–2611. doi:10.1158/0008-5472.CAN-15-1758
49. Acconcia F, Ascenzi P, Bocedi A, et al. Palmitoylation-dependent estrogen receptor alpha membrane localization: regulation by 17beta-estradiol. Mol Biol Cell. 2005;16(1):231–237. doi:10.1091/mbc.e04-07-0547
50. Xue M, Zhang K, Mu K, et al. Regulation of estrogen signaling and breast cancer proliferation by an ubiquitin ligase TRIM56. Oncogenesis. 2019;8(5):30. doi:10.1038/s41389-019-0139-x
51. Turner NC, Neven P, Loibl S, Andre F. Advances in the treatment of advanced oestrogen-receptor-positive breast cancer. Lancet. 2017;389(10087):2403–2414. doi:10.1016/S0140-6736(16)32419-9
52. Musgrove EA, Sutherland RL. Biological determinants of endocrine resistance in breast cancer. Nat Rev Cancer. 2009;9(9):631–643. doi:10.1038/nrc2713
53. Early Breast Cancer Trialists’ Collaborative Group. Effects of chemotherapy and hormonal therapy for early breast cancer on recurrence and 15-year survival: an overview of the randomised trials. Lancet. 2005;365(9472):1687–1717. doi:10.1016/S0140-6736(05)66544-0
54. Bosch A, Li Z, Bergamaschi A, et al. PI3K inhibition results in enhanced estrogen receptor function and dependence in hormone receptor-positive breast cancer. Sci Transl Med. 2015;7(283):283ra251. doi:10.1126/scitranslmed.aaa4442
55. Toska E, Osmanbeyoglu HU, Castel P, et al. PI3K pathway regulates ER-dependent transcription in breast cancer through the epigenetic regulator KMT2D. Science. 2017;355(6331):1324–1330. doi:10.1126/science.aah6893
56. Fiorillo M, Sanchez-Alvarez R, Sotgia F, Lisanti MP. The ER-alpha mutation Y537S confers Tamoxifen-resistance via enhanced mitochondrial metabolism, glycolysis and Rho-GDI/PTEN signaling: implicating TIGAR in somatic resistance to endocrine therapy. Aging (Albany NY). 2018;10(12):4000–4023. doi:10.18632/aging.v10i12
57. Jia S, Miedel MT, Ngo M, et al. Clinically observed estrogen receptor alpha mutations within the ligand-binding domain confer distinguishable phenotypes. Oncology. 2018;94(3):176–189. doi:10.1159/000485510
58. Bahreini A, Li Z, Wang P, et al. Mutation site and context dependent effects of ESR1 mutation in genome-edited breast cancer cell models. Breast Cancer Res. 2017;19(1):60. doi:10.1186/s13058-017-0851-4
59. Weis KE, Ekena K, Thomas JA, Lazennec G, Katzenellenbogen BS. Constitutively active human estrogen receptors containing amino acid substitutions for tyrosine 537 in the receptor protein. Mol Endocrinol. 1996;10(11):1388–1398. doi:10.1210/mend.10.11.8923465
60. Zhang QX, Borg A, Wolf DM, Oesterreich S, Fuqua SA. An estrogen receptor mutant with strong hormone-independent activity from a metastatic breast cancer. Cancer Res. 1997;57(7):1244–1249.
61. Fanning SW, Mayne CG, Dharmarajan V, et al. Estrogen receptor alpha somatic mutations Y537S and D538G confer breast cancer endocrine resistance by stabilizing the activating function-2 binding conformation. Elife. 2016;5.
62. McKenna NJ, Lanz RB, O’Malley BW. Nuclear receptor coregulators: cellular and molecular biology. Endocr Rev. 1999;20(3):321–344. doi:10.1210/edrv.20.3.0366
63. Smith CL, Nawaz Z, O’Malley BW. Coactivator and corepressor regulation of the agonist/antagonist activity of the mixed antiestrogen, 4-hydroxytamoxifen. Mol Endocrinol. 1997;11(6):657–666. doi:10.1210/mend.11.6.0009
64. Anzick SL, Kononen J, Walker RL, et al. AIB1, a steroid receptor coactivator amplified in breast and ovarian cancer. Science. 1997;277(5328):965–968. doi:10.1126/science.277.5328.965
65. Heery DM, Kalkhoven E, Hoare S, Parker MG. A signature motif in transcriptional co-activators mediates binding to nuclear receptors. Nature. 1997;387(6634):733–736. doi:10.1038/42750
66. Spencer TE, Jenster G, Burcin MM, et al. Steroid receptor coactivator-1 is a histone acetyltransferase. Nature. 1997;389(6647):194–198. doi:10.1038/38304
67. Ichinose H, Garnier JM, Chambon P, Losson R. Ligand-dependent interaction between the estrogen receptor and the human homologues of SWI2/SNF2. Gene. 1997;188(1):95–100. doi:10.1016/S0378-1119(96)00785-8
68. Zhu Y, Qi C, Jain S, et al. Amplification and overexpression of peroxisome proliferator-activated receptor binding protein (PBP/PPARBP) gene in breast cancer. Proc Natl Acad Sci U S A. 1999;96(19):10848–10853. doi:10.1073/pnas.96.19.10848
69. Tian W, Teng F, Gao J, et al. Estrogen and insulin synergistically promote endometrial cancer progression via crosstalk between their receptor signaling pathways. Cancer Biol Med. 2019;16(1):55–70. doi:10.20892/j.issn.2095-3941.2018.0157
70. Cuesta R, Gritsenko MA, Petyuk VA, et al. Phosphoproteome analysis reveals estrogen-ER pathway as a modulator of mTOR activity via DEPTOR. Mol Cell Proteomics. 2019;18(8):1607–1618. doi:10.1074/mcp.RA119.001506
71. Chen R, Zhang M, Liu W, et al. Estrogen affects the negative feedback loop of PTENP1-miR200c to inhibit PTEN expression in the development of endometrioid endometrial carcinoma. Cell Death Dis. 2018;10(1):4. doi:10.1038/s41419-018-1207-4
72. Salmeron-Hernandez A, Noriega-Reyes MY, Jordan A, Baranda-Avila N, Langley E. BCAS2 enhances carcinogenic effects of estrogen receptor alpha in breast cancer cells. Int J Mol Sci. 2019;20:4. doi:10.3390/ijms20040966
73. Arun A, Ansari MI, Popli P, et al. New piperidine derivative DTPEP acts as dual-acting anti-breast cancer agent by targeting ERalpha and downregulating PI3K/Akt-PKCalpha leading to caspase-dependent apoptosis. Cell Prolif. 2018;51(6):e12501. doi:10.1111/cpr.12501
74. Han R, Gu S, Zhang Y, et al. Estrogen promotes progression of hormone-dependent breast cancer through CCL2-CCR2 axis by upregulation of Twist via PI3K/AKT/NF-kappaB signaling. Sci Rep. 2018;8(1):9575. doi:10.1038/s41598-018-27810-6
75. Dohlman HG. Thematic minireview series: cell biology of G protein signaling. J Biol Chem. 2015;290(11):6679–6680. doi:10.1074/jbc.R114.631093
76. Chambliss KL, Wu Q, Oltmann S, et al. Non-nuclear estrogen receptor alpha signaling promotes cardiovascular protection but not uterine or breast cancer growth in mice. J Clin Invest. 2010;120(7):2319–2330. doi:10.1172/JCI38291
77. Banerjee S, Chambliss KL, Mineo C, Shaul PW. Recent insights into non-nuclear actions of estrogen receptor alpha. Steroids. 2014;81:64–69. doi:10.1016/j.steroids.2013.11.002
78. Divekar SD, Storchan GB, Sperle K, et al. The role of calcium in the activation of estrogen receptor-alpha. Cancer Res. 2011;71(5):1658–1668. doi:10.1158/0008-5472.CAN-10-1899
79. Tesarik J, Mendoza C. Nongenomic effects of 17 beta-estradiol on maturing human oocytes: relationship to oocyte developmental potential. J Clin Endocrinol Metab. 1995;80(4):1438–1443. doi:10.1210/jcem.80.4.7714121
80. Levin ER, Pietras RJ. Estrogen receptors outside the nucleus in breast cancer. Breast Cancer Res Treat. 2008;108(3):351–361. doi:10.1007/s10549-007-9618-4
81. Marquez DC, Chen HW, Curran EM, Welshons WV, Pietras RJ. Estrogen receptors in membrane lipid rafts and signal transduction in breast cancer. Mol Cell Endocrinol. 2006;246(1–2):91–100. doi:10.1016/j.mce.2005.11.020
82. Becker MA, Ibrahim YH, Cui X, Lee AV, Yee D. The IGF pathway regulates ERalpha through a S6K1-dependent mechanism in breast cancer cells. Mol Endocrinol. 2011;25(3):516–528. doi:10.1210/me.2010-0373
83. Yu Z, Gao W, Jiang E, et al. Interaction between IGF-IR and ER induced by E2 and IGF-I. PLoS One. 2013;8(5):e62642. doi:10.1371/journal.pone.0062642
84. Tang Y, Wang Y, Kiani MF, Wang B. Classification, treatment strategy, and associated drug resistance in breast cancer. Clin Breast Cancer. 2016;16(5):335–343. doi:10.1016/j.clbc.2016.05.012
85. Nicolini A, Ferrari P, Duffy MJ. Prognostic and predictive biomarkers in breast cancer: past, present and future. Semin Cancer Biol. 2018;52(Pt 1):56–73. doi:10.1016/j.semcancer.2017.08.010
86. Su X, Xu X, Li G, Lin B, Cao J, Teng L. ER-alpha36: a novel biomarker and potential therapeutic target in breast cancer. Onco Targets Ther. 2014;7:1525–1533. doi:10.2147/OTT.S65345
87. Mishra SK, Talukder AH, Gururaj AE, et al. Upstream determinants of estrogen receptor-alpha regulation of metastatic tumor antigen 3 pathway. J Biol Chem. 2004;279(31):32709–32715. doi:10.1074/jbc.M402942200
88. Bouris P, Skandalis SS, Piperigkou Z, et al. Estrogen receptor alpha mediates epithelial to mesenchymal transition, expression of specific matrix effectors and functional properties of breast cancer cells. Matrix Biol. 2015;43:42–60. doi:10.1016/j.matbio.2015.02.008
89. Zhu C, Wang S, Wang B, et al. 17beta-Estradiol up-regulates Nrf2 via PI3K/AKT and estrogen receptor signaling pathways to suppress light-induced degeneration in rat retina. Neuroscience. 2015;304:328–339. doi:10.1016/j.neuroscience.2015.07.057
90. Leung HW, Wang Z, Yue GG, et al. Cyclopeptide RA-V inhibits cell adhesion and invasion in both estrogen receptor positive and negative breast cancer cells via PI3K/AKT and NF-kappaB signaling pathways. Biochim Biophys Acta. 2015;1853(8):1827–1840. doi:10.1016/j.bbamcr.2015.04.020
91. Royuela M, de Miguel MP, Bethencourt FR, et al. Estrogen receptors alpha and beta in the normal, hyperplastic and carcinomatous human prostate. J Endocrinol. 2001;168(3):447–454. doi:10.1677/joe.0.1680447
92. Mishra S, Tai Q, Gu X, et al. Estrogen and estrogen receptor alpha promotes malignancy and osteoblastic tumorigenesis in prostate cancer. Oncotarget. 2015;6(42):44388–44402. doi:10.18632/oncotarget.v6i42
93. Slavin S, Yeh CR, Da J, et al. Estrogen receptor alpha in cancer-associated fibroblasts suppresses prostate cancer invasion via modulation of thrombospondin 2 and matrix metalloproteinase 3. Carcinogenesis. 2014;35(6):1301–1309. doi:10.1093/carcin/bgt488
94. Dou XW, Liang YK, Lin HY, et al. Notch3 maintains luminal phenotype and suppresses tumorigenesis and metastasis of breast cancer via trans-activating estrogen receptor-alpha. Theranostics. 2017;7(16):4041–4056. doi:10.7150/thno.19989
95. Hasson RM, Briggs A, Carothers AM, et al. Estrogen receptor alpha or beta loss in the colon of Min/+ mice promotes crypt expansion and impairs TGFbeta and HNF3beta signaling. Carcinogenesis. 2014;35(1):96–102. doi:10.1093/carcin/bgt323
96. Lund-Iversen M, Scott H, Strom EH, Theiss N, Brustugun OT, Gronberg BH. Expression of estrogen receptor-alpha and survival in advanced-stage non-small cell lung cancer. Anticancer Res. 2018;38(4):2261–2269. doi:10.21873/anticanres.12470
97. Liang R, Lin Y, Yuan CL, et al. High expression of estrogen-related receptor alpha is significantly associated with poor prognosis in patients with colorectal cancer. Oncol Lett. 2018;15(4):5933–5939. doi:10.3892/ol.2018.8011
98. Ignatov T, Treeck O, Kalinski T, Ortmann O, Ignatov A. GPER-1 expression is associated with a decreased response rate to primary tamoxifen therapy of breast cancer patients. Arch Gynecol Obstet. 2020;301:565–571. doi:10.1007/s00404-019-05384-6
99. Anestis A, Sarantis P, Theocharis S, et al. Estrogen receptor beta increases sensitivity to enzalutamide in androgen receptor-positive triple-negative breast cancer. J Cancer Res Clin Oncol. 2019;145(5):1221–1233. doi:10.1007/s00432-019-02872-9
100. Ibrahim A, Hugerth LW, Hases L, et al. Colitis-induced colorectal cancer and intestinal epithelial estrogen receptor beta impact gut microbiota diversity. Int J Cancer. 2019;144(12):3086–3098. doi:10.1002/ijc.v144.12
101. Lombardi AP, Pisolato R, Vicente CM, Lazari MF, Lucas TF, Porto CS. Estrogen receptor beta (ERbeta) mediates expression of beta-catenin and proliferation in prostate cancer cell line PC-3. Mol Cell Endocrinol. 2016;430:12–24. doi:10.1016/j.mce.2016.04.012
102. Ignatov A, Ignatov T, Weissenborn C, et al. G-protein-coupled estrogen receptor GPR30 and tamoxifen resistance in breast cancer. Breast Cancer Res Treat. 2011;128(2):457–466. doi:10.1007/s10549-011-1584-1
103. Ignatov A, Ignatov T, Roessner A, Costa SD, Kalinski T. Role of GPR30 in the mechanisms of tamoxifen resistance in breast cancer MCF-7 cells. Breast Cancer Res Treat. 2010;123(1):87–96. doi:10.1007/s10549-009-0624-6
104. Yang K, Yao Y. Mechanism of GPER promoting proliferation, migration and invasion of triple-negative breast cancer cells through CAF. Am J Transl Res. 2019;11(9):5858–5868.
105. Li Y, Jia Y, Bian Y, et al. Autocrine motility factor promotes endometrial cancer progression by targeting GPER-1. Cell Commun Signal. 2019;17(1):22. doi:10.1186/s12964-019-0336-4
106. Xie BY, Lv QY, Ning CC, et al. TET1-GPER-PI3K/AKT pathway is involved in insulin-driven endometrial cancer cell proliferation. Biochem Biophys Res Commun. 2017;482(4):857–862. doi:10.1016/j.bbrc.2016.11.124
107. Bustos V, Nolan AM, Nijhuis A, et al. GPER mediates differential effects of estrogen on colon cancer cell proliferation and migration under normoxic and hypoxic conditions. Oncotarget. 2017;8(48):84258–84275. doi:10.18632/oncotarget.20653
108. Liu Q, Chen Z, Jiang G, et al. Epigenetic down regulation of G protein-coupled estrogen receptor (GPER) functions as a tumor suppressor in colorectal cancer. Mol Cancer. 2017;16(1):87. doi:10.1186/s12943-017-0654-3
109. Chen Y, Tang H, He J, et al. Interaction of nuclear ERs and GPER in vitellogenesis in zebrafish. J Steroid Biochem Mol Biol. 2019;189:10–18. doi:10.1016/j.jsbmb.2019.01.013
110. Lagunas N, Marraudino M, de Amorim M, et al. Estrogen receptor beta and G protein-coupled estrogen receptor 1 are involved in the acute estrogenic regulation of arginine-vasopressin immunoreactive levels in the supraoptic and paraventricular hypothalamic nuclei of female rats. Brain Res. 2019;1712:93–100. doi:10.1016/j.brainres.2019.02.002
111. Sanchez DS, Fischer Sigel LK, Azurmendi PJ, et al. Estradiol stimulates cell proliferation via classic estrogen receptor-alpha and G protein-coupled estrogen receptor-1 in human renal tubular epithelial cell primary cultures. Biochem Biophys Res Commun. 2019;512(2):170–175. doi:10.1016/j.bbrc.2019.03.056
112. Jonsson P, Katchy A, Williams C. Support of a bi-faceted role of estrogen receptor beta (ERbeta) in ERalpha-positive breast cancer cells. Endocr Relat Cancer. 2014;21(2):143–160. doi:10.1530/ERC-13-0444
113. Lu W, Katzenellenbogen BS. Estrogen Receptor-β modulation of the ERα-p53 loop regulating gene expression, proliferation, and apoptosis in breast cancer. Horm Cancer. 2017;8(4):230–242. doi:10.1007/s12672-017-0298-1
114. Linares PM, Algaba A, Urzainqui A, et al. Ratio of circulating estrogen receptors beta and alpha (ERβ/ERα) indicates endoscopic activity in patients with Crohn’s disease. Dig Dis Sci. 2017;62(10):2744–2754. doi:10.1007/s10620-017-4717-5
115. Sosa L, Gutierrez S, Petiti JP, et al. 17beta-Estradiol modulates the prolactin secretion induced by TRH through membrane estrogen receptors via PI3K/Akt in female rat anterior pituitary cell culture. Am J Physiol Endocrinol Metab. 2012;302(10):E1189–E1197. doi:10.1152/ajpendo.00408.2011
116. Acconcia F, Totta P, Ogawa S, et al. Survival versus apoptotic 17beta-estradiol effect: role of ER alpha and ER beta activated non-genomic signaling. J Cell Physiol. 2005;203(1):193–201. doi:10.1002/jcp.20219
117. Reinbolt RE, Mangini N, Hill JL, et al. Endocrine therapy in breast cancer: the neoadjuvant, adjuvant, and metastatic approach. Semin Oncol Nurs. 2015;31(2):146–155. doi:10.1016/j.soncn.2015.02.002
118. Liu X, Miao W, Huang M, Li L, Dai X, Wang Y. Elevated Hexokinase II expression confers acquired resistance to 4-hydroxytamoxifen in breast cancer cells. Mol Cell Proteomics. 2019;18(11):2273–2284. doi:10.1074/mcp.RA119.001576
119. Khan SA, Rogers MA, Khurana KK, Meguid MM, Numann PJ. Estrogen receptor expression in benign breast epithelium and breast cancer risk. J Natl Cancer Inst. 1998;90(1):37–42. doi:10.1093/jnci/90.1.37
120. Widschwendter M, Siegmund KD, Muller HM, et al. Association of breast cancer DNA methylation profiles with hormone receptor status and response to tamoxifen. Cancer Res. 2004;64(11):3807–3813. doi:10.1158/0008-5472.CAN-03-3852
121. Haque MM, Desai KV. Pathways to endocrine therapy resistance in breast cancer. Front Endocrinol (Lausanne). 2019;10:573. doi:10.3389/fendo.2019.00573
122. Weiner M, Skoog L, Fornander T, Nordenskjold B, Sgroi DC, Stal O. Oestrogen receptor co-activator AIB1 is a marker of tamoxifen benefit in postmenopausal breast cancer. Ann Oncol. 2013;24(8):1994–1999. doi:10.1093/annonc/mdt159
123. Nicholson RI, Hutcheson IR, Jones HE, et al. Growth factor signalling in endocrine and anti-growth factor resistant breast cancer. Rev Endocr Metab Disord. 2007;8(3):241–253. doi:10.1007/s11154-007-9033-5
124. Hasson SP, Rubinek T, Ryvo L, Wolf I. Endocrine resistance in breast cancer: focus on the phosphatidylinositol 3-kinase/akt/mammalian target of rapamycin signaling pathway. Breast Care (Basel). 2013;8(4):248–255. doi:10.1159/000354757
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.

