Back to Journals » Infection and Drug Resistance » Volume 11
Epidemiology and molecular characterization of multidrug-resistant Escherichia coli isolates harboring blaCTX-M group 1 extended-spectrum β-lactamases causing bacteremia and urinary tract infection in Manhiça, Mozambique
Authors Guiral E , Pons MJ, Vubil D , Marí-Almirall M , Sigaúque B, Soto SM, Alonso PL, Ruiz J , Vila J, Mandomando I
Received 9 October 2017
Accepted for publication 20 February 2018
Published 3 July 2018 Volume 2018:11 Pages 927—936
DOI https://doi.org/10.2147/IDR.S153601
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Sahil Khanna
Elisabet Guiral,1 Maria Jesús Pons,1 Delfino Vubil,2 Marta Marí-Almirall,1 Betuel Sigaúque,2,3 Sara Maria Soto,1 Pedro Luís Alonso,1,2 Joaquim Ruiz,1 Jordi Vila,1,4 Inácio Mandomando2,3
1Barcelona Institute for Global Health (ISGlobal), Hospital Clínic-Universitat de Barcelona, Barcelona, Spain; 2Centro de Investigação em Saúde de Manhiça (CISM), Maputo, Mozambique; 3Instituto Nacional de Saúde (INS), Ministério da Saúde, Maputo, Mozambique; 4Microbiology Department, Hospital Clínic, School of Medicine, University of Barcelona, Barcelona, Spain
Background: The emergence and spread of extended-spectrum β-lactamases (ESBLs), especially CTX-M, is an important public health problem with serious implications for low-income countries where second-line treatment is often unavailable. Knowledge of the local prevalence of ESBL is critical to define appropriate empirical therapeutic strategies for multidrug-resistant (MDR) organisms. This study aimed to assess and characterize the presence of ESBL and especially CTX-M-producing Escherichia coli MDR isolates from patients with urinary tract infections (UTIs) and bacteremia in a rural hospital in Mozambique.
Materials and methods: One hundred and fifty-one E. coli isolates from bacteremia and UTI in children were screened for CTX-M, TEM, SHV and OXA β-lactamases by polymerase chain reaction and sequencing. Isolates carrying CTX-M group 1 β-lactamases were further studied. The resistance to other antibiotic families was determined by phenotypic and genotypic methods, the location of the blaCTX-M gene and the epidemiology of the isolates were studied, and extensive plasmid characterization was performed.
Results: Approximately 11% (17/151) of E. coli isolates causing bacteremia and UTI were ESBL producers. CTX-M-15 was the most frequently detected ESBL, accounting for 75% of the total isolates characterized. The blaCTX-M gene is located in different plasmids belonging to different incompatibility groups and can be found in non-epidemiologically related isolates, indicating the high capacity of this resistance determinant to spread widely.
Conclusion: Our data suggest the presence of a co-selection of third-generation cephalosporin-resistant determinants in the study area despite limited access to these antibiotics. This highlights the importance of continuous surveillance of antimicrobial resistance of both genetic elements of resistance and resistant isolates in order to monitor the emergence and trends of ESBL-producing isolates to promote adequate therapeutic strategies for the management of MDR bacterial infections.
Keywords: CTX-M-15, multidrug-resistance, Enterobacteriaceae, resistance determinant location
Introduction
Infections caused by members of the Enterobacteriaceae family are among the major causes of hospital admission and associated morbidity and mortality in children, particularly in Africa.1,2 Infections caused by these microorganisms in low- and middle-income countries (LMIC) have been successfully treated with the inexpensive antibiotics available. Nevertheless, with the widespread development of multidrug-resistant (MDR) strains, the usefulness of the early effective antibiotics has greatly decreased,3,4 leading to the introduction of broad-spectrum antibiotics such as fluoroquinolones or third-generation cephalosporins (cefotaxime, ceftriaxone, or ceftazidime). Unfortunately, these agents are often unaffordable in most LMIC, especially in remote rural areas.
On the other hand, since their first description in 1983, extended-spectrum β-lactamases (ESBLs) produced by enteric pathogens have spread worldwide.5
The emergence and spread of ESBLs, especially those included in the CTX-M group, is an important public health problem.6 In fact, it has been considered that ESBL-carrying Enterobacteriaceae cause >1700 deaths yearly in the USA alone,7 and these pathogens have had a tremendous impact on the treatment of severe or MDR-associated infections, particularly in LMIC where second-line antibiotics are often unaffordable or unavailable. In addition, few new antibiotics against Gram-negative bacteria have been marketed in the last decades8 which may favor the emergence of new resistances, further challenging the management of infectious diseases in this setting. This may play a role in the high morbidity and mortality observed in these countries, particularly in children <5 years of age.
Although different types of ESBLs have been reported among the Enterobacteriaceae family, CTX-M-15, a community-acquired ESBL that was originally described in India in the 1990s, is one of the most frequent type I CTX-M disseminated worldwide.9 The genes encoding ESBL enzymes are usually located in plasmids but can also be found in the chromosomal DNA as described elsewhere.10 It has been reported that the blaCTX-M-15 gene is usually found downstream from the insertion sequence ISEcp1 that may be involved in their dissemination and expression.11 Plasmid-mediated ESBL genes are of special interest due to their capability of getting transferred between strains or even species, favoring their dissemination among the bacterial population and from region to region. Moreover, these plasmids usually carry other antibiotic resistance determinants, resulting not only in the spread of ESBL but also in the dissemination of other resistance genes.12 The selection of one resistance gene due to environmental pressure harbored in the same genetic element as another resistance gene or genes is known as the co-selection of resistance genes phenomenon. Since ESBL-producing microorganisms are also often resistant to other commonly available antibiotics, including fluoroquinolones, especially in most LMIC,13 knowledge of their prevalence and characterization is important for defining local empirical stewardship programs for infections caused by MDR organisms.
In Africa, ESBLs have increasingly been reported.14,15 In Mozambique, the prevalence of these pathogens is extremely high, although the data available are limited to only a few studies.16 Herein, we report the prevalence of Escherichia coli harboring the ESBL gene blaCTX-M group I as well as its molecular characterization and epidemiology among isolates recovered from blood cultures and urine in a rural hospital in Southern Mozambique.
Materials and methods
Study population and clinical isolates
The study was conducted by the Centro de Investigação em Saúde de Manhiça (CISM) at the Manhiça District Hospital, a rural referral hospital of the Manhiça district, located 80 km north of Maputo, in Southern Mozambique. Invasive bacterial disease surveillance has been conducted in the pediatric population in this area since 1997. The full description and characteristics of the study area are detailed elsewhere.17 As described previously in standard clinical protocols, blood cultures are systematically collected upon admission of all children up to 14 years of age with an axillary temperature ≥37.5°C or meeting criteria of severe infection.2 We analyzed E. coli isolates recovered from children with community-acquired bacteremia between August 2004 and December 2009. Urine samples were also collected during the same study period from patients (adults and children) visited at the outpatient department or admitted to the hospital with clinical suspicion of urinary tract infection (UTI). All the isolates included in the present study were recovered from different patients.
Bacterial culture and identification
Blood culture tubes were incubated in an automated system (BACTEC® 9050; Becton Dickinson, Franklin Lakes, NJ, USA). Positive blood cultures were subcultured in solid media after Gram staining as appropriate. Urine samples were microscopically screened after centrifugation, and those with pathologic sediment (presence of leucocytes or bacteria) were cultured in MacConkey and blood agar media. Pathogens were identified according to conventional microbiology protocols. Among the Enterobacteriaceae isolates identified, ceftriaxone susceptibility was tested by disk diffusion in Mueller–Hinton agar (Oxoid®, Basingstoke, Hampshire, UK) according to the Clinical and Laboratory Standard Institute (CLSI) 2013 guidelines.18 The selected ceftriaxone non-susceptible E. coli isolates were screened for the ESBL enzyme CTX-M group 1 and the positive isolates were included in the study. The isolates were confirmed by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) prior to further analysis.19
Antimicrobial susceptibility testing
The susceptibility phenotype for ampicillin, chloramphenicol, ceftriaxone, gentamicin, tetracycline, trimethoprim-sulfamethoxazole, rifampicin, and amikacin was determined by a conventional disk diffusion method and for nalidixic acid and ciprofloxacin by minimum inhibitory concentration (MIC). The interpretative category of resistance for disk diffusion and MIC were done according to the CLSI 2013 guidelines.18 Antimicrobial susceptibility testing of the isolates was also performed by Siemens MicroScan panels NEG MIC TYPE 37. The E. coli American Type Culture Collection 25922 strain was used as the quality control. Multidrug resistance was defined as resistance to 3 or more unrelated antibiotic families.20 Resistance genes to quinolones and rifampicin were also studied by polymerase chain reaction (PCR) and sequencing methods using primers for aac (6’)-Ib-cr, qnrA, qnrB, qnrC, qnrD, qnrS, qepA, gyrA, parC, arr2,3, arr4, arr5, arr6, arr7, arr3, and rpoB already described.21–29
ESBL phenotype detection
All ceftriaxone non-susceptible E. coli isolates were phenotypically screened for the presence of ESBL. Phenotypic confirmation of ESBL expression was carried out using the ESBL disk synergy test with disks containing cefotaxime, amoxicillin with clavulanate, and ceftazidime on Mueller–Hinton agar (Oxoid) as described elsewhere.30 E. coli isolates with an ESBL phenotype were tested by PCR for the presence of genes encoding β-lactamases and were further characterized as follows.
β-lactamases analysis
The presence of blaCTX-M was detected by PCR using universal primers, while the blaCTX-M groups 1, 2, 8, 9, and blaCTX-M-15 were determined using specific CTX-M group primers and subsequently sequenced.31 Moreover, the presence of blaSHV, blaTEM, blaOXA-1-like, blaOXA-2-like, and blaOXA-5-like genes was also determined by PCR and sequencing as described elsewhere.32 The presence of the insertion sequence ISEcp1 upstream from the blaCTX-M genes was determined by PCR and sequencing as previously described.11 Sequencing of the genes was performed by the Macrogen® DNA Sequencing Service (Macrogen, Amsterdam, the Netherlands) using sets of consecutive primers specific for each gene type.
Class I integron analysis
The presence of class I integrons was analyzed in all the E. coli isolates. A PCR was carried out with primers 3′CS and 5′CS as described by Lévesque et al33 and the amplicons obtained were sequenced by Beckman Coulter Sequencing Genomics® sequencing facilities (Takeley, UK).
Typing
Pulsed-field gel electrophoresis (PFGE) was performed with the XbaI restriction enzyme (New England Biolabs, Beverly, MA, USA) as described previously.34 PFGE profiles were analyzed with InfoQuest FP software version 4.5 (Bio-Rad Laboratories Inc., Hercules, CA, USA). In order to establish the epidemiological relationship among the isolates from the electrophoretic patterns, the Dice coefficient was used and clustering was based on the unweighted pair group method with arithmetic mean with a 1% tolerance in band position differences. The isolates were considered to belong to the same epidemiological group when the PFGE-XbaI profiles showed ≥80% of homology, adapting the criteria described by Tenover et al.35 Multi-locus sequence typing (MLST) was carried out by amplification and sequencing of the 7 E. coli housekeeping genes as described previously.36 The database available at http://mlst.warwick.ac.uk/mlst/dbs/Ecoli/ was used for assigning sequence types (STs) and clonal complexes (CCs). Classification of isolates into E. coli phylogenetic groups was done using a previously described triplex PCR-based protocol.37
Plasmid transferability analysis
Conjugation assays were carried out with all the isolates in order to determine if the blaCTX-M group 1 gene was located in a conjugative plasmid adapting the protocol described elsewhere.38 The E. coli K7 759 lac- kanamycin-resistant isolate was used as the recipient strain. Both parental and recipient strains were cultured over-night (ON) with Luria Bertani (LB) broth (Laboratorios Conda, Barcelona, Spain). The parental isolates were grown in LB medium supplemented with 32 µg/mL of cefotaxime in order to force resistance determinant replication. An aliquot of 500 µL of a parental isolate subcultured for 2 hours was mixed with the same volume of the recipient strain and cultured ON at 37°C. Transconjugant strains were finally grown in MacConkey agar plates supplemented with 256 µg/mL of kanamycin and 32 µg/mL of cefotaxime. Repetitive extragenic palindromic PCRs and a blaCTX-M group 1 PCR were performed to ensure the correct selection of the transconjugants.39
The blaCTX-M gene location
Considering the previously described large size of the plasmids carrying blaCTX-M group 1 ESBL genes,40 a S1 nuclease (Promega, Madison, WI, USA) digestion followed by PFGE analysis were performed in the 12 E. coli isolates and the transconjugants obtained as described elsewhere.41 To determine the plasmid or chromosomal location of the ESBL-encoding gene, a Southern blot of the PFGE gel followed by hybridization with a blaCTX-M group 1 probe was carried out.
PCR-based Replicon Typing (PBRT)
Plasmids from both parental and transconjugant isolates were assigned to incompatibility groups depending on the presence of specific replicon sequences identified by PCR using the primers designed by Carattoli et al in 2005 but employing the adapted amplification protocols for commensal and pathogenic E. coli isolates described by Johnson et al.42
Ethical clearance
The strains characterized here were isolated from the ongoing invasive bacterial surveillance system that included several research protocols reviewed and approved by the Mozambican National Bioethics Committee for Health (IR00002657) and by Institutional Review Boards of Hospital Clinic of Barcelona, Spain; the US Centers for Disease Control and Prevention; and the School of Medicine, University of Maryland. Written informed consent was obtained from parents or caretakers of the eligible children.
Results
Study population and clinical isolates
During the study period, a total of 15,057 blood cultures were collected and 1325 (8.8%) were found to be positive for any pathogen evaluable. Of these, 27.7% were identified as belonging to the Enterobacteriaceae family, with E. coli being the second most frequent after non-typhoidal Salmonella, accounting for 29% (106/368). Among these, 8 out of 12 E. coli isolates non-susceptible to ceftriaxone and positive for the ESBL disk synergy test (11.3% of total E. coli isolates) were found to carry a blaCTX-M group I gene.
Of the 298 urine samples cultured for bacterial isolation, 35% (n=103) were positive for pathogenic bacteria. Among these, 81 (78.6%) corresponded to Enterobacteriaceae with E. coli being the most prevalent species with 45 isolates of which 5 (11.1%) were non-susceptible to ceftriaxone and positive for the ESBL double-disk synergy test. Four were found to carry a blaCTX-M group I gene.
A total of 12 E. coli isolates from bacteremia and UTI carrying a blaCTX-M group I gene were selected for further characterization.
β-lactamases analysis
Gene amplification sequencing revealed that 92% (n=11) of the isolates harbored blaCTX-M-15, while the remaining strain presented the ESBL gene blaCTX-M-37. The non-ESBL resistance genes also detected, blaTEM-1 and blaOXA-1, were found in 100% and 58.3% of the isolates, respectively. Another ESBL-encoding gene detected was blaSHV-12, which was found in 2 of the isolates also presenting blaCTX-M-15. In all cases, the insertion sequence ISEcp1 was found upstream from the blaCTX-M group 1 gene. The overall results are summarized in Table 1.
Antimicrobial susceptibility testing
All the isolates were MDR, presenting not only resistance to third-generation β-lactams (ceftriaxone) but also to other classes of antimicrobial agents. All the isolates were resistant to rifampicin, gentamicin, chloramphenicol, and trimethoprim-sulfamethoxazole, while 66.7% were resistant to quinolones (Table 1). Regarding the resistance genotype of the E. coli isolates to quinolones, among the isolates with a MIC=1 µg/mL of ciprofloxacin, 4 showed the presence of the qnrB gene without mutations in the gyrA and parC genes; 1 isolate showed only a mutation in amino acid codon Ser83 of gyrA and 1 did not show any of the resistance determinants studied. The isolate with a MIC=64 µg/mL of ciprofloxacin showed 3 mutations (2 in gyrA and 1 in parC) and the other isolate with a MIC >256 µg/mL has the same mutations plus the presence of the qnrB gene (Table 2).
The resistance genotype to rifampicin was not well elucidated as arr genes were not detected and any significant mutations in rpoB gene were observed in any isolate.
All the isolates were susceptible to fosfomycin, nitrofurantoin, and carbapenems.
Class 1 integron analysis
Seven isolates were found to carry class 1 integrons. Two isolates harbored an integron of ~1000 bp carrying the resistance gene aadA1, conferring resistance to streptomycin and spectinomycin. One isolate had a 2000 bp class 1 integron carrying 2 resistance genes: dfrA12 and aadA2 that confer resistance to trimethoprim and streptomycin-spectinomycin, respectively. Four isolates presented 2 integrons of ~800 and 1000 bp containing the dfrA16 and aadA1 genes, respectively, conferring the same resistances as those mentioned earlier (Table 1).
Molecular typing
According to the PFGE analysis constructed from the electrophoresis patterns of the XbaI restriction and considering the same profile of ≥80% of similarity, there were 8 different epidemiological groups among the 12 isolates studied. The analysis showed 4 and 2 other isolates to be in the same epidemiological group, thereby being epidemiologically related isolates. This association, however, involved grouping isolates harboring different blaCTX-M group 1 genes. The MLST analysis also showed the same number of ST groups as epidemiologically unrelated isolates (singletons). This data correlates 100% with the epidemiological grouping established by the PFGE analysis and with the genetic characterization of non-β-lactam resistance genes and the antimicrobial susceptibility profiles. Only 2 out of the 8 STs described belonged to the same CC (ST10). Four E. coli phylogenetic groups were represented in the collection of isolates (A, B1, B2, and D), with none having a statistically significant prevalence taking into account the epidemiological associations. All the isolates causing UTI belonged to phylogenetic group A (Table 3).
  | Table 3 Typing and epidemiological relationship between the Escherichia coli isolates Abbreviations: PFGE, pulsed-field gel electrophoresis; ST, sequence type. |
Plasmid characterization
Transconjugants were obtained from 10 parental isolates as shown in Table 4. S1 endonuclease digestion allowed visualizing the plasmid profile of each isolate. The parental isolates carried 1, 2, or even 3 plasmids each, ranging from <48.5 kb to ~380 kb (Figure 1). Hybridization with the blaCTX-M group 1 probe allowed the localization of the plasmid carrying the antimicrobial resistance determinant in each isolate, which appeared to be of 3 different plasmid sizes among the 8 epidemiological groups (Figure 2). However, epidemiologically related isolates carried plasmids with different sizes harboring the resistance determinant. The conjugative plasmid in isolate E8 carrying blaCTX-M-37 was the smallest (<48.5 Kb), whereas in the epidemiologically related isolates carrying blaCTX-M-15, the conjugative plasmid was the largest (~290 Kb). The transconjugant E11T showed a hybridization signal in a larger-sized plasmid from its donor isolate. Isolates in which conjugation was not possible (E2 and E4) only showed a hybridization signal in the chromosome.
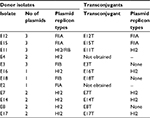  | Table 4 Plasmid transferability assay and characterization |
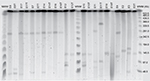  | Figure 1 S1 endonuclease pulsed-field gel electrophoresis (PFGE). Abbreviation: MWM, molecular weight marker. |
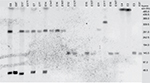  | Figure 2 Hybridization of S1 endonuclease pulsed-field gel electrophoresis (PFGE) with blaCTX-M group 1 probe. |
The plasmid incompatibility groups amplified in the PBRT analysis of the parental isolates were IncFIIA, IncFIA, IncHI2, and IncFIB, but only the incompatibility groups IncFIIA and IncHI2 were found in the plasmids carrying the blaCTX-M group 1 gene (such as those amplified in the transconjugants; Table 4).
Discussion
This is one of the few studies on the prevalence of CTX-M group 1 ESBLs in E. coli causing both bacteremia and UTIs among children in Mozambique. The prevalence of ESBLs reported here is a matter of concern as MDR pathogens causing infectious diseases are common in this area, limiting the therapeutic options for treating severe infections often associated with a poor outcome. The rates of ESBLs and other antibiotic resistances observed in this study may be associated with the high prevalences of other infectious diseases, such as tuberculosis, respiratory infections, malaria, and human immunodeficiency virus, which requires the frequent use of antibacterial agents.43–45 Despite the high prevalence of ESBLs reported in this study, it is lower compared with the prevalence of those causing UTIs in children admitted to the malnutrition and pediatric wards described in the central region of the country (Beira City).46
Regarding the resistance mechanisms to non-β-lactamic antibiotic families found among the E. coli isolates studied, the resistance determinants to quinolones correspond to those described so far,47 whereas the lack of detection of resistance determinants to rifampicin suggests other mechanisms of resistance, such as the effect of efflux pumps, as described elsewhere.29
The most prevalent resistance mechanism to third-generation cephalosporins found in the collection of MDR isolates studied is the blaCTX-M gene belonging to a sublineage or group I (accounting for 70.6% of the ESBL-carrying isolates), which is consistent with other reports5 showing isolates harboring a blaCTX-M group I gene with almost 92% being blaCTX-M-15. Within the CTX-M-1 group, blaCTX-M-15 is the most frequently described resistance gene in isolates causing both community-acquired bacteremia and UTI.6,16 In fact, blaCTX-M-15 is currently the most common variant detected worldwide in clinically important Gram-negative bacteria together with blaCTX-M-14.9 According to the current data and the widely reported blaCTX-M-15 dissemination, this is not the first description of this gene in the area since it has previously been described in ESBL-carrying Klebsiella pneumoniae isolates.16
Regarding the other blaCTX-M group I gene detected, this is the first description of blaCTX-M-37 in Mozambique. This infrequently detected CTX-M was first described in an Enterobacter cloacae isolated in Mongolia in 2002, and to our knowledge, it has only been previously reported in 1 other African country, the neighboring South Africa, as well as in the chromosome of an isolate of Kluivera cryocrescences from Argentina (GenBank access No: FN813246.1).48
Whereas the mechanism of resistance to third-generation cephalosporins in the collection of isolates of our study was the same, the isolates showed low relatedness at an epidemiological level, being distributed in 4 phylogenetic groups and 8 epidemiological groups. Although the range of phylogenetic groups represented is wide, it is important to highlight that all the isolates from UTIs were phylogenetic group A. This phylogenetic group has been associated with MDR strains causing UTI.49 Based on PFGE and MLST results, 4 isolates within the same epidemiological group belonged to ST ST216. However, these isolates showed some divergent evolution concerning the resistant determinants, as one harbored CTX-M-37 (isolate E8) instead of CTX-M-15 and another (isolate E17) also showed SHV-12. Furthermore, isolates E12 and E15 belonged to the same epidemiological group although they were isolated at a different period. The non-related isolates belonged to different STs and even different CCs, indicating that it was not a clonal dissemination.
Regarding the plasmid analysis to determine the location of blaCTX-M group 1, the collection presented a wide range of different plasmid incompatibility groups. Furthermore, 2 epidemiologically unrelated isolates harbored the resistance determinant in the chromosome and, therefore, no transconjugants were obtained. The transconjugant E11 showed a hybridization signal in a larger plasmid from its donor isolate, but its size corresponded to the recombination of the 2 plasmids harbored by the parental isolate, a common phenomenon in conjugation assays. Moreover, together with E16, which belongs to ST ST10, this isolate harbored the same plasmid (in terms of size and incompatibility group) as the isolates belonging to the clone ST216, suggesting a potential dissemination of the same plasmid among E. coli strains belonging to different STs.
The location of the blaCTX-M gene upstream from the insertion sequence ISEcp1 in all the isolates and in a conjugative plasmid in most of the isolates implies a high potential of dissemination of this ESBL, suggesting that this is not a result of the dissemination of particular clones but rather is due to the spread of multiple specific clones and/or mobile genetic elements. However, it is interesting that these strains presented such resistance to third-generation cephalosporins, as these antimicrobial agents are little used in Mozambique.
With regard to the resistance phenotype, 100% of the isolates were resistant to gentamicin, chloramphenicol, and trimethoprim-sulfamethoxazole. The first 2 antibiotics are used in the empirical treatment of bacteremia (chloramphenicol or penicillin plus gentamicin) whereas ceftriaxone is reserved for MDR cases,2 which would not be effective in the isolates studied. The transconjugant isolates showed not only β-lactam resistance but also the same resistance profile to other antibiotic families, which may have an impact on the clinical management of the patients in this setting. There are 3 main non-exclusive explanations for the finding of CTX-M group 1 ESBL resistance genes in these isolates: 1) the use of third-generation cephalosporins as second-line treatment may play a role in the emergence and subsequent dissemination of β-lactamase resistance; 2) the resistance to β-lactam antibiotics is the result of a co-selection from another family of antibiotic resistance mechanisms located in the same genetic mobile element; 3) globalization may influence the dissemination of ESBLs-carrying isolates in the community similar to what has been shown with New Dehli metallo-β-lactamase.50 The historical movement of the population between Manhiça and neighboring South Africa,17 as well as the increasingly more frequent presence of international travelers to the Manhiça District may support the international dissemination of these strains, as third-generation cephalosporins are not commonly used in this community.
Conclusion
As observed in this study, continuously increasing resistance among Gram-negative bacteria associated with the emergence and spread of MDR isolates, including ESBL producers, is an important public health problem with serious implications in low-income countries. Taking into account that the availability of effective antibiotics is a challenge worldwide and is of special concern in LMIC due to limited resources, clinical microbiology services in Mozambique – and in all the Sub-Saharan African countries – need to be reinforced in order to perform coordinated antimicrobial resistance surveillance and establish national policies to control this public health problem.
Acknowledgments
The authors thank all the clinical and laboratory staff from the CISM for their contribution in different stages of the study, specifically for Dinis Jantilal, Oscar Fraile, Tacilta Nhampossa, and Pedro Aide. Special thanks to the bacteriology laboratory technicians for their excellent support in performing antimicrobial susceptibility testing and ESBL phenotyping. The authors are also grateful to Alessandra Carattoli for providing the control strains for the PBRT and to Donna Pringle for language correction.
This study received funding from Planes Nacionales de I+D+i 2008–2011/2013–2016 and the Instituto de Salud Carlos III, Subdirección General de Redes y Centros de Investigación Cooperativa, Ministerio de Economía y Competitividad, Spanish Network for Research in Infectious Diseases (REIPI RD12/0015/0013 and REIPI RD16/0016/0010) and was co-financed by European Development Regional Fund “A way to achieve Europe” and operative program Intelligent Growth 2014–2020. The CISM received core funding from the Agencia Española de Cooperación Internacional y Desarrollo (AECID). This work was also supported by grant 2009 SGR 1256 from the Agència de Gestió d’Ajuts Universitaris i de Recerca of the Generalitat de Catalunya. JR had a fellowship from program I3 of the Instituto de Salud Carlos III (ISCIII) (grant number: CES11/012).
ISGlobal is a member of the Centres de Recerca de Catalunya (CERCA) Programme, Generalitat de Catalunya.
Disclosure
The authors report no conflicts of interest in this work.
References
Berkley JA, Lowe BS, Mwangi I, et al. Bacteremia among children admitted to a rural hospital in Kenya. N Engl J Med. 2005;352(1):39–47. | ||
Sigaúque B, Roca A, Mandomando I, et al. Community-acquired bacteremia among children admitted to a rural hospital in Mozambique. Pediatr Infect Dis J. 2009;28(2):108–113. | ||
Mandomando I, Sigaúque B, Morais L, et al. Antimicrobial drug resistance trends of bacteremia isolates in a rural hospital in southern Mozambique. Am J Trop Med Hyg. 2010;83(1):152–157. | ||
Okeke IN, Laxminarayan R, Bhutta ZA, et al. Antimicrobial resistance in developing countries. Part I: recent trends and current status. Lancet Infect Dis. 2005;5(8):481–493. | ||
Rossolini GM, D’Andrea MM, Mugnaioli C. The spread of CTX-M-type extended-spectrum β-lactamases. Clin Microbiol Infect. 2008;14(Suppl 1):33–41. | ||
Peirano G, Pitout JD. Molecular epidemiology of Escherichia coli producing CTX-M beta-lactamases: the worldwide emergence of clone ST131 O25:H4. Int J Antimicrob Agents. 2010;35(4):316–321. | ||
Centers for Disease Control and Prevention. US Department of Health and Human Services. Antibiotic Resistance Threats in the United States, 2013. Atlanta, GE: 2013. | ||
Fair RJ, Tor Y. Perspectives in medicinal chemistry antibiotics and bacterial resistance in the 21th century. Perspect Medicin Chem. 2014;6:25–64. | ||
Coque TM, Novais Â, Carattoli A, et al. Dissemination of clonally related Escherichia coli strains expressing extended-spectrum beta-lactamase CTX-M-15. Emerg Infect Dis. 2008;14(2):195–200. | ||
Rodríguez I, Thomas K, Van Essen A, et al; SAFEFOODERA-ESBL consortium. Chromosomal location of blaCTX-M genes in clinical isolates of Escherichia coli from Germany, The Netherlands and the UK. Int J Antimicrob Agents. 2014;43(6):553–557. | ||
Chouchani C, El Salabi A, Marrakchi R, Ferchichi L, Walsh TR. Characterization of IncA/C conjugative plasmid harboring blaTEM-52 and blaCTX-M-15 extended-spectrum β-lactamases in clinical isolates of Escherichia coli in Tunisia. Eur J Clin Microbiol Infect Dis. 2012;31(6):1081–1087. | ||
Carattoli A. Plasmids and the spread of resistance. Int J Med Microbiol. 2013;303(6–7):298–304. | ||
Tumbarello M, Sali M, Trecarichi EM, et al. Bloodstream infections caused by extended-spectrum-beta-lactamase-producing Escherichia coli: risk factors for inadequate initial antimicrobial therapy. Antimicrob Agents Chemother. 2008;52(9):3244–3252. | ||
Gray KJ, Wilson LK, Phiri A, Corkill JE, French N, Hart CA. Identification and characterization of ceftriaxone resistance and extended-spectrum beta-lactamases in Malawian bacteraemic Enterobacteriaceae. J Antimicrob Chemother. 2006;57(4):661–665. | ||
Blomberg B, Jureen R, Manji KP, et al. High rate of fatal cases of pediatric septicemia caused by gram-negative bacteria with extended-spectrum beta-lactamases in Dar es Salaam, Tanzania. J Clin Microbiol. 2005;43(2):745–749. | ||
Pons MJ, Vubil D, Guiral E, et al. Characterisation of extended-spectrum β-lactamases among Klebsiella pneumoniae isolates causing bacteraemia and urinary tract infection in Mozambique. J Glob Antimicrob Resist. 2015;3(1):19–25. | ||
Sacoor C, Nhacolo A, Nhalungo D, et al. Profile: Manhiça health research centre (Manhiça HDSS). Int J Epidemiol. 2013;42(5):1309–1318. | ||
Clinical and Laboratory Standards Institute. Performance Standards for Antimicrobial Susceptibility Testing. Twenty-Third Informational Supplement. M100-S23. Wayne, PE: Clinical and Laboratory Standards Institute; 2013. | ||
Sogawa K, Watanabe M, Sato K, et al. Use of the MALDI BioTyper system with MALDI-TOF mass spectrometry for rapid identification of microorganisms. Anal Bioanal Chem. 2011;400(7):1905–1911. | ||
Magiorakos AP, Srinivasan A, Carey RB, et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect. 2012;18(3):268–281. | ||
Park CH, Robicsek A, Jacoby GA, Sahm D, Hooper DC. Prevalence in the United States of aac(6’)-Ib-cr encoding a ciprofloxacin-modifying enzyme. Antimicrob Agents Chemother. 2006;50(11):3953–3955. | ||
Cattoir V, Poirel L, Rotimi V, Soussy CJ, Nordmann P. Multiplex PCR for detection of plasmid-mediated quinolone resistance qnr genes in ESBL-producing enterobacterial isolates. J Antimicrob Chemother. 2007;60(2):394–397. | ||
Kim HB, Park CH, Kim CJ, Kim E-C, Jacoby GA, Hooper DC. Prevalence of plasmid-mediated quinolone resistance determinants over a 9-year period. Antimicrob Agents Chemother. 2009;53(2):639–645. | ||
Zhao J, Chen Z, Chen S, et al. Prevalence and dissemination of oqxAB in Escherichia coli isolates from animals, farmworkers, and the environment. Antimicrob Agents Chemother. 2010;54(10):4219–4224. | ||
Heisig P. Genetic evidence for a role of parC mutations in development of high-level fluoroquinolone resistance in Escherichia coli. Antimicrob Agents Chemother. 1996;40(4):879–885. | ||
Heisig P, Kratz B, Halle E, et al. Identification of DNA gyrase a mutations in ciprofloxacin-resistant isolates of Salmonella typhimurium from men and cattle in Germany. Microb Drug Resist. 1995;1(3):211–218. | ||
Hopkins KL, Mushtaq S, Richardson JF, et al. In vitro activity of rifaximin against clinical isolates of Escherichia coli and other enteropathogenic bacteria isolated from travellers returning to the UK. Int J Antimicrob Agents. 2014;43(5):431–437. | ||
Lee K, Yum JH, Yong D, et al. Novel acquired metallo-β-lactamase gene, blaSIM-1, in a class 1 integron from Acinetobacter baumannii clinical isolates from Korea. Antimicrob Agents Chemother. 2005;49(11):4485–4491. | ||
Pons MJ, Mensa L, Gascón J, Ruiz J. Fitness and molecular mechanisms of resistance to rifaximin in in vitro selected Escherichia coli mutants. Microb Drug Resist. 2012;18(4):376–379. | ||
Drieux L, Brossier F, Sougakoff W, Jarlier V. Phenotypic detection of extended-spectrum β-lactamase production in Enterobacteriaceae: review and bench guide. Clin Microbiol Infect. 2008;14(Suppl 1):90–103. | ||
Woodford N, Fagan EJ, Ellington MJ. Multiplex PCR for rapid detection of genes encoding CTX-M extended-spectrum β-lactamases. J Antimicrob Chemother. 2006;57(1):154–155. | ||
Medina AM, Rivera FP, Pons MJ, et al. Comparative analysis of antimicrobial resistance in enterotoxigenic Escherichia coli isolates from two paediatric cohort studies in Lima, Peru. Trans R Soc Trop Med Hyg. 2015;109(8):493–502. | ||
Lévesque C, Piché L, Larose C, Roy PH. PCR mapping of integrons reveals several novel combinations of resistance genes. Antimicrob Agents Chemother. 1995;39(1):185–191. | ||
Ribot EM, Fair MA, Gautom R, et al. Standardization of pulsed-field gel electrophoresis protocols for the subtyping of Escherichia coli O157:H7, Salmonella, and Shigella for PulseNet. Foodborne Pathog Dis. 2006;3(1):59–67. | ||
Tenover FC, Arbeit RD, Goering RV, et al. Interpreting chromosomal DNA restriction patterns produced by pulsed-field gel electrophoresis: criteria for bacterial strain typing. J Clin Microbiol. 1995;33(9):2233–2239. | ||
Wirth T, Falush D, Lan R, et al. Sex and virulence in Escherichia coli : an evolutionary perspective. Mol Microbiol. 2006;60(5):1136–1151. | ||
Clermont O, Bonacorsi S, Bingen E. Rapid and simple determination of the Escherichia coli phylogenetic group. Appl Environ Microbiol. 2000;66(10):4555–4558. | ||
Wachino J, Kurokawa H, Suzuki S, et al. Horizontal transfer of blaCMY-bearing plasmids among clinical Escherichia coli and Klebsiella pneumoniae isolates and emergence of cefepime-hydrolyzing CMY-19. Antimicrob Agents Chemother. 2006;50(2):534–541. | ||
Eckert C, Gautier V, Saladin-Allard M, et al. Dissemination of CTX-M-type β-lactamases among clinical isolates of Enterobacteriaceae in Paris, France. Antimicrob Agents Chemother. 2004;48(4):1249–1255. | ||
Wang J, Stephan R, Karczmarczyk M, Yan Q, Hächler H, Fanning S. Molecular characterization of blaESBL-harboring conjugative plasmids identified in multi-drug resistant Escherichia coli isolated from food-producing animals and healthy humans. Front Microbiol. 2013;4:188. | ||
Barton BM, Harding GP, Zuccarelli AJ. A general method for detecting and sizing large plasmids. Anal Biochem. 1995;226(2):235–240. | ||
Johnson TJ, Wannemuehler YM, Johnson SJ, et al. Plasmid replicon typing of commensal and pathogenic Escherichia coli isolates. Appl Environ Microbiol. 2007;73(6):1976–1983. | ||
González R, Munguambe K, Aponte JJ, et al. High HIV prevalence in a southern semi-rural area of Mozambique: a community-based survey. HIV Med. 2012;13(10):581–588. | ||
García-Basteiro A, Miranda Ribeiro R, Brew J, et al. Tuberculosis on the rise in southern Mozambique (1997–2012). Eur Respir J. 2017;49(3):1601683. | ||
Maltha J, Guiraud I, Kaboré B, et al. Frequency of severe malaria and invasive bacterial infections among children admitted to a rural hospital in Burkina Faso. PLoS One. 2014;9(2):e89103. | ||
Van der Meeren BT, Chhaganlal KD, Pfeiffer A, et al. Extremely high prevalence of multi-resistance among uropathogens from hospitalised children in Beira, Mozambique. S Afr Med J. 2013;103(6):382–386. | ||
Hopkins KL, Davies RH, Threlfall EJ. Mechanisms of quinolone resistance in Escherichia coli and Salmonella: recent developments. Int J Antimicrob Agents. 2005;25(5):358–373. | ||
Lee K, Yong D, Jeong SH, et al. Genetic and biochemical characterisation of CTX-M-37 extended-spectrum β-lactamase from an Enterobacter cloacae clinical isolate from Mongolia. J Glob Antimicrob Resist. 2017;10:3–7. | ||
Ejrnæs K. Bacterial characteristics of importance for recurrent urinary tract infections caused by Escherichia coli. Dan Med Bull. 2011;58(4):B4187. | ||
Lee CR, Lee JH, Park KS, Kim YB, Jeong BC, Lee SH. Global dissemination of carbapenemase-producing Klebsiella pneumoniae: epidemiology, genetic context, treatment options, and detection methods. Front Microbiol. 2016;7:895. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.


