Back to Journals » International Journal of Nanomedicine » Volume 10 » Issue 1
Effect of the protein corona on nanoparticles for modulating cytotoxicity and immunotoxicity
Authors Lee YK, Choi E, Webster T , Kim S, Khang D
Received 20 August 2014
Accepted for publication 10 November 2014
Published 18 December 2014 Volume 2015:10(1) Pages 97—113
DOI https://doi.org/10.2147/IJN.S72998
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 5
Editor who approved publication: Professor Carlos Rinaldi
Yeon Kyung Lee,1,* Eun-Ju Choi,2,* Thomas J Webster,3 Sang-Hyun Kim,4 Dongwoo Khang1
1Department of Molecular Medicine, School of Medicine, Gachon University, Incheon, South Korea; 2Division of Sport Science, College of Science and Technology, Konkuk University, Chungju, South Korea; 3Department of Chemical Engineering and Program in Bioengineering, Northeastern University, Boston, MA, USA; 4Department of Pharmacology, School of Medicine, Kyungpook National University, Daegu, South Korea
*These authors contributed equally to this work
Abstract: Although the cytotoxicity of nanoparticles (NPs) is greatly influenced by their interactions with blood proteins, toxic effects resulting from blood interactions are often ignored in the development and use of nanostructured biomaterials for in vivo applications. Protein coronas created during the initial reaction with NPs can determine the subsequent immunological cascade, and protein coronas formed on NPs can either stimulate or mitigate the immune response. Along these lines, the understanding of NP-protein corona formation in terms of physiochemical surface properties of the NPs and NP interactions with the immune system components in blood is an essential step for evaluating NP toxicity for in vivo therapeutics. This article reviews the most recent developments in NP-based protein coronas through the modification of NP surface properties and discusses the associated immune responses.
Keywords: nanostructured biomaterials, blood response, cytotoxicity, immunotoxicity, protein corona
Introduction
The development and rational design of nanoparticles (NPs) has created a new paradigm for overcoming current limitations and concerns in medicine.1–3 Compared with micron-sized particles, NPs can retain unique material characteristics, such as size, solubility, shape, surface charge, and chemistry, that can be tailored to increase the efficacy of NP-based therapeutics for various biomedical applications.1,2,4 In drug delivery systems, for example, conventional (micron-sized) particles are quickly cleared by the immune system and can only enter phagocytic cells, whereas NP-based drugs can be delivered to all organs and, thus, all cells.5,6
In this regard, NPs have been the subject of increasing concern from the medical community because of their immunological toxicity. Once NPs enter various organs and interact with blood, they come into immediate contact with various plasma constituents. These NP-bound blood proteins can influence the immune system by mediating subsequent immune cell responses.7–10 Protein corona or biological macromolecule adsorption is influenced by physiochemical material properties and can affect the in vivo toxicity of the material. For example, a study on polymer-based NPs designed for intravenous administration has demonstrated that NPs can stimulate and/or suppress immune cell responses, depending on the formation of protein coronas formed in blood.11 Furthermore, in vivo organ distribution and the clearance rate of nanomaterial-derived drug carriers in blood are significantly influenced by the formation of plasma proteins.7,11,12 As such, evaluating and understanding the toxicity of nanostructure-based materials coated with protein coronas is a critical step toward manipulating their subsequent immune response and cytotoxic effects.
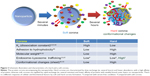
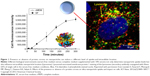
Surprisingly, it was determined that the same NPs can induce different biological outcomes, depending on the control presence or absence of a protein corona. For example, silica NPs in serum conditions showed stronger accumulation at lysosomes. However, silica NPs without serum proteins exposed to cells showed a higher degree of attachment to the cell membrane and greater internalization (both lysosomes and cytosols; Figure 1).13 In addition, carboxylated polystyrene NPs under serum-free conditions exhibit a higher adhesion to the cell membrane than adhesion of the NP surface to the cell membrane under serum conditions.14
|
Figure 1 Presence or absence of protein corona on nanoparticles can induce a different level of uptake and intracellular location. |
The mechanism of selective uptake of NPs with or without serum proteins was identified by a recent study that analyzed this mechanism by a two-step process. The NPs were initially adhered to the cell membrane at 4°C and internalized by increasing temperature (37°C). High surface energy of the bare NPs can cause unspecific interactions and adsorb strongly to the cell membranes by the process of quite reactive, and chemically lowering energy. It was interpreted that the formation of a corona surrounding the NP lowered the energy by unspecific interactions and, thus, leads to less attachment of NPs at the cell membrane in the presence of biomolecules.14
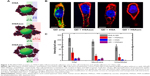
Selective cellular uptake by engineered biochemical structures of proteins also was studied. In this study, pure human serum albumin (HSA), modified HSA by succinic anhydride (HSAsuc), and modified HSA by succinic anhydride by ethylenediamine (HSAam) were coated on dihydrolipoic acid-coated quantum dots (DHLA-QDs) to investigate the effect of biochemical structures on cellular uptake and membrane binding (Figure 2). The HSAsuc-coated DHLA-QDs showed enhanced binding to the cell membrane compared with native HSA (Figure 2B).15 Uncoated DHLA-QDs recorded the highest uptake amount and membrane localization by cells compared with HSA, HSAsuc, and HSAam (Figure 2C).15 The results obtained highlighted that both the presence of a protein corona and modified chemical structures markedly suppress the membrane binding and uptake level of DHLA-QDs. Specifically, suggested articles indicate that engineered protein structures on NPs not only affect cellular uptake levels but also determine the intracellular location of NPs. For this reason, the biological effect of NPs cannot be solely understood by considering the intrinsic properties of the NP; we must also consider protein dynamics on NPs.14–16
|
Figure 2 Modified HSA proteins on DHLA-QDs influenced both adhesion rate to the cell membrane and uptake efficiency for HeLa cells. |
Various biomolecular coronas, antibodies, proteins, and peptides on NPs were specifically designed for targeting in nanomedicine.2 Specifically, the number of publications of protein corona has increased about 65% during the last 5 years (2009–2013). The majority of previous studies focused on understanding protein corona composition affected by NP properties, such as size, charge, and surface modification.17–21 Moreover, several studies have attempted to address the toxicity associated with various routes of NP administration.22,23 As such, few guidelines have been established and even fewer studies have been conducted to understand how protein coronas on nanomaterials are triggering NP toxicity. This may be attributed to a lack of comprehensive evaluation of the NP-protein corona that might dictate the immunological response of cells. For this reason, the objective of this article is to introduce the most recent advances in NP-protein corona formation with various components of the immune system and their subsequent cytotoxicity in blood systems.
General composition types of protein corona on NPs
“Soft” and “Hard” proteins
The process of corona formation is determined by the competition of countless proteins to adsorb at the approaching NP surface. Protein coronas are categorized as either “soft” or “hard” (Figure 3). Specifically, the soft corona represents loosely bound proteins on a NP surface over short time scales (ie, seconds to minutes) or weak interactions compared with the hard corona. In contrast, the “hard” corona represents tightly bound proteins with high affinity on the NP surface and for longer periods (ie, hours; Figure 4).24,25 The core NP (and other multiparticle assemblies) surrounded by a hard corona demonstrated slowly exchanging proteins, and these complexes represent “what the cell sees” (Figures 3 and 4)9,25 and are more significant in determining immunologic response than the bare material properties of the particle itself. In addition, the plasma-derived hard corona (ie, the final long-term corona) may be important when NPs are accumulated in various organs.9
 
|
Figure 4 NP-protein complexes based on sulfonated polystyrene and silica NPs. |
Cedervall et al determined that the kinetic and equilibrium binding properties of NP-protein coronas depend not only on the NP surface and size but also on plasma protein identity, such as with HSA, fibrinogen, and lipoproteins.24 For example, HSA and fibrinogen exhibited higher rates of association and dissociation than apolipoprotein A-I and other plasma proteins.24 In addition, it has been demonstrated that HSA and fibrinogen adsorption dominate on hydrophobic particles more than on hydrophilic particles for short times.24
However, initially attached proteins were subsequently replaced by lower-abundance proteins with higher affinity (ie, slower kinetics), such as lipoproteins, and especially apolipoprotein A-I.24 Thus, greater proteins with low affinity made up the protein corona during the initial period (soft corona) but were replaced with proteins of lower abundance and higher affinity at longer time scales (hard corona).
As a consequence, the hard corona may possess a greater role than the soft corona in determining the physiological response, and because of the long residence time, the hard corona on NP experiences further biological process (ie, endocytosis).9,21 As such, if adsorbed hard coronas with specific physiochemical nanomaterial properties do not invoke immune cell responses, they could be advantageous for reducing NP toxicity.
Protein adsorption on various NPs
The adsorptions of blood proteins on NP have been analyzed in recent studies.26–32 It was found that proteins (eg, albumin, apolipoprotein, immunoglobulins [Igs], complement, and fibrinogen) adsorbed on various NP (polymeric NPs, iron oxide NPs, gold NPs [Au-NPs], liposomes, QDs, and carbon nanotubes [CNTs]) surfaces exhibited structural changes in the bloodstream (ie, bioactivity).33–35 For example, Roach et al studied the binding events of bovine serum albumin (BSA) and bovine fibrinogen (BFG) on silica nanospheres that exhibited both hydrophilic and hydrophobic surface curvature and performed secondary structure analysis (ie, infrared spectroscopy on surface-bound proteins).35 Compared with BSA, fibrinogen tended to undergo greater conformational changes in its secondary structure when proteins were adsorbed on NPs with high surface curvature (ie, small dimension of radius).35 In addition, surface properties of NPs, such as charge, size, and the effect of particle coating, affected the compatibility of NP-protein corona complexes with the immune system.36
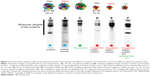
Lundqvist et al categorized different hard corona formations on NPs by different chemistries (ie, untreated polystyrene NP, carboxyl-modified, and amine-modified) with different sizes (50 and 100 nm) (Figure 5).36 All separated and identified proteins were categorized according to their functions, such as Igs, lipoproteins, complements, acute-phase proteins, and coagulation factors. Interestingly, both size and surface charge played a significant role in determining the NP-coronas, even for identical NP materials. For example, the 100-nm carboxyl-modified particles showed high Igs adsorption, whereas the 50-nm carboxyl- and amine-modified particles exhibited relatively low Igs adsorption (Figure 5). In this regard, specific blood protein adsorption and changes in protein conformation at certain dimensions of NPs can influence the subsequent immunotoxicity of cells.36–38
|
Figure 5 Excised sodium dodecyl sulfate-polyacrylamide gel electrophoresis gels and different formations of hard protein coronas on different polystyrene particle types. |
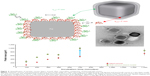
Recently, protein coronas adsorbed on polyvinylpyrrolidone-stabilized silver nanocubes in fetal bovine serum (FBS)-containing media were also quantified, using localized surface plasmon resonances.32 The soft corona formation was identified at the cube edges/corners versus facets at short incubation times.32 The soft corona also contained more proteins than the hard corona at all points (eightfold difference in 10% serum condition; Figure 6).32 This study suggests that polymer-coated NP coating may influence the kinetics of corona formation or protein distribution on NPs in biological media.
 
|
Figure 6 Quantification of protein corona layers around silver nanocubes; comparison of hard and soft corona. |
Charge-dependent interactions of NPs with biological media and associated cellular uptake were investigated. Colloidal Au-NPs were modified with amphiphilic polymer NPs to obtain identical physical properties, with the exception of the signs of surface charge (negative or positive) and, thus, removed other influential factors, such as different size and colloidal stability.39 This study clarified that the number of adsorbed HSA molecules per NP was not influenced by their surface charge. However, the cellular uptake rate of NPs was lower for negative NPs than for positive NPs, both in serum-free and serum-containing media.39 Furthermore, cytotoxicity assays (ie, Alamar Blue assay) exhibit a reduced cytotoxicity for negatively charged NPs compared with positively charged NPs by reduced uptake of negatively charged NPs.39
As a consequence, internalization and cytotoxicity of NPs are significantly influenced by the surface charge of NPs.39 This study speculated that a different hydrodynamic radius was induced by changed conformational structures on negative and positive NPs.
It was also demonstrated that rapid corona formation can influence hemolysis, thrombocyte activation, NP uptake, and endothelial cell death. Interestingly, 300 different protein coronas on NP rapidly interacted (within 30 seconds). As time advanced, the composition percentage was sustained (although the amount of bounded proteins changed). This study highlighted that very rapidly established complex protein coronas at the NP–biological interface can modulate early pathobiological effects, which has a great implications in nanomedicine and nanotoxicology.37 As such, this study addressed the concept that early exposure time to plasma in biological fluids can be another important factor for determining cytotoxicity.
Mitigated toxicity on various NPs by the protein corona
Blood protein interaction with carbonic NPs
CNTs are broadly used in various biomaterials applications because of their unique mechanical, electrical, optical, and biological properties.40,41 The first large-scale characterization of the protein corona using mass spectrometry-based proteomics demonstrated that more than 750 proteins were bound to CNTs.42
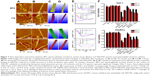
The competitive binding of blood proteins on single-wall CNTs (SWCNTs) influenced cellular pathways and resulted in reduced cytotoxicity that depended on the presence of protein adsorption. Ge et al explored interactions between SWCNTs and human blood proteins and examined the cytoxicity of SWCNTs for two types of human cell lines (human acute monocytic leukemia cell line [THP-1] and human umbilical vein endothelial cells) by coating the SWCNTs with blood proteins (Figure 7).43 Atomic force microscopy (AFM) data demonstrated that the adsorption of transferrin and BSA quickly reaches thermodynamic equilibrium in 10 minutes, whereas BFG and gamma globulin adsorbed onto the SWCNT surface over longer periods (approximately 5 hours; Figure 7A–E).43 This study suggested that the highly competitive binding of blood proteins on the SWCNT surface can affect subsequent cellular responses in a time-dependent manner. Specifically, viability of cells using a cell counting kit-8 assay in the presence of BFG-, BSA-, transferrin-, and gamma globulin-coated SWCNTs represented less cytotoxicity than uncoated SWCNTs (Figure 7F).43 In particular, BFG-coated SWCNTs showed no toxicity and reduced toxicity might be related that BFG proteins have structures with more protein layers (five layers) than fibrinogen, BSA, and transferrin (two or three protein layers) and, thus, can protect cells from direct exposure to bare SWCNT surfaces. Further analysis demonstrated that the π–π stacking interactions between SWCNTs and aromatic residues (Trp, Phe, and Tyr) notably affected the binding capabilities. This study suggested that the binding of various protein types onto the SWCNT surface can elicit different cytotoxic cellular responses and can provide an important guideline for elucidating structures of protein corona on NPs that are nontoxic to cells.43
 
|
Figure 7 Time-dependent interactions between BFG, gamma globulin, transferrin, bovine serum albumin, and SWCNTs and the associated viability of human cell lines. |
In the bloodstream, the occurrence of blood toxicity via the formation of protein corona on a CNT is another significant issue. In this regard, the role of protein corona composition of CNTs and their interaction with human blood platelets was studied.38
Specifically, protein corona interaction on carboxylated, multiwalled CNTs (CNT-COOH) with common human serum proteins, such as HSA, fibrinogen, immunoglobulin G (IgG), and histone H1 (H1), revealed that bare CNT-COOH and H1 corona induced platelet aggregation and increased lactate dehydrogenase (one of cytotoxicity markers), whereas HSA, fibrinogen, and IgG corona on CNT-COOH attenuated subsequent cytotoxicity.38
The cytotoxicity and serum protein interaction of multiwalled CNTs with three different grades of carbon blacks were also investigated.44 The kinetics of various carbon NPs (CNPs) and serum proteins in the culture medium indicated that the adsorption of serum proteins (ie, FBS) on CNPs reached maximum values within 5 minutes.44
AFM studies showed that CNPs were enveloped in serum proteins (NP-protein coronas) after simple mixing of CNPs and FBS proteins. It was identified that cellular uptake of CNPs was greater in serum-free medium than in medium containing serum (Figure 6A and B). Serum protein adsorption on CNPs attenuated the inherent cytotoxicity of CNPs and resulted in decreased cytotoxicity by increasing the amount of serum proteins adsorbed on the CNPs.44 A possible mechanism governing this behavior is that the presence of serum proteins in the medium significantly reduced the intracellular uptake of CNPs. As such, serum proteins adsorbed on CNPs can inhibit their toxicity by shielding impurities of metal catalyzers and suppressing the competitive adsorption of other proteins in the medium.44,45
Graphene is a material composed of either a single layer or several layers of sp2-bonded carbons that has unique and highly attractive electrical, mechanical, and thermal properties.41,46,47
Despite these unique properties, the use of graphene and its derivatives (eg, graphene oxide [GO]) has raised considerable concerns about human and environmental health. Recently, the immunotoxicity of hard corona composition on GO was analyzed.47 Long-term exposure to GO in plasma conditions (sufficient time to generate hard corona on NP) decreased reactive oxygen species production and reduced the cytotoxicity of GO.47
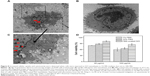
In addition, Hu et al reported a protein corona-mediated reduction of cytotoxicity of GO.16 In their study, the authors performed a systematic investigation of the cellular toxicity of GO nanosheets and identified that GO interactions with FBS or a protein component in cell culture medium resulted in a decrease of cytotoxicity (Figure 8).16 Specifically, at low concentrations of FBS (ie, 1%), cytotoxicity of human lung cancer cells (A549) was sensitive to the presence of GO and showed concentration-dependent cytotoxicity.16
|
Figure 8 Increased cellular uptake and cytotoxicity were observed when cells were exposed to GO nanosheets in 1% FBS medium, but not in 10% FBS. |
However, the cytotoxicity of GO was greatly mitigated in 10% FBS media and showed no concentration-dependent toxicity. Specifically, this study addressed the stimulated cytotoxicity of A549 via the direct interactions with bare GO nanosheets at the cell membrane that, ultimately, resulted in physical damage to the membrane.16
Blood protein interaction with polymeric NPs and other inorganic NPs
Polymeric NPs [eg, poly(sodium acrylate), poly(ethylene glycol), chitosan, etc] have been widely used as coating materials of NPs to avoid activation of the immune system, to reduce cytotoxicity, and to prolong drug circulation time in the blood system. Lemarchand et al investigated the NP–plasma protein interactions with dextran-grafted poly(e-caprolactone).48 The dextran modification on the NP surface significantly inhibited the complement system on human THP-1 and J774.A1 murine macrophage-like cell lines (phagocytosis). Specifically, dextran-coated poly(e-caprolactone) showed increased plasma protein adsorption compared with uncoated poly(e-caprolactone), with the exception of Ig adsorption.48
The adsorption of proteins to inorganic NPs (eg, Au, Ag, FeO4, cobalt oxide, and CeO2) in serum-containing medium also has been investigated. Casals et al determined that the production of reactive oxygen species was decreased in THP-1 cells when cobalt oxide NPs were incubated with serum for 48 hours.49 Silver NPs (Ag-NPs) recorded the highest cytotoxicity to human cells compared with other metallic NPs, such as MoO3 and Fe3O4.50,51 To improve biocompatibility of Ag-NPs or Au-NPs, various polymers have been coated on the NP surfaces. One study examined the cytotoxicity of spherical Ag-NPs (diameter of 50±20 nm) that were stabilized with either polyvinylpyrrolidone or citrate and dispersed in cell culture media (ie, pure media and media containing 10% BSA or 10% fetal calf serum).52 In this study, the release of Ag ions from polyvinylpyrrolidone-stabilized Ag-NPs and their biological effect on human mesenchymal stem cells (hMSCs) were examined. The formation of Ag-protein coronas with BSA and fetal calf serum reduced the cytotoxicity of hMSCs by inhibiting the release of free Ag ions from the silver.
Citrate-coated Au-NPs, in different sizes, were examined with two types of culture media and cancer cell lines.53 This study showed that Au-NP-protein coronas formed in Roswell Park Memorial Institute medium exerted greater cytotoxic effects compared with Dulbecco modified Eagle’s medium. This was attributed to the greater abundance and stability of Au-NP-protein coronas formed in Dulbecco’s medium than in the Roswell medium. These findings indicated that different compositions of cell culture media can determine the formation of NP-protein corona and, ultimately, the effect on cellular toxicity.
Activated immune response by protein coronas
General recognition of NP-corona by immune cells
When NPs are injected into blood systems, they are considered foreign objects by a series of defense and recognition mechanisms. Interactions with plasma proteins and plasma factors can affect clearance and systemic toxicity of the NP delivery system. Recently, several studies focused on understanding the effect of the NP-protein corona on leukocytes and macrophages.23,54 Synthetic hydroxyapatite particles with a fine needle shape or hydroxyapatite aggregate can activate the NLR (nucleotide-binding oligomerization domain receptors) family, pyrin domain-containing 3 inflammasome in lipopolysaccharide-induced macrophages, and can induce secretion of proinflammatory cytokines such as interleukin 1β and interleukin 18 in blood.55,56 It has been shown that the mineral hydroxyapatite NPs exhibit significant dose-dependent cytotoxicity via human macrophages.55,56
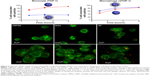
The protein corona can significantly influence NP–cell interactions through different internalization and pathway activations.10 Selective cellular uptake of disulfide-stabilized poly-(methacrylic acid) nanoporous polymer particles with presence and without presence of protein corona conditions was recently reported.10 Interestingly, depending on the corona formation on NP, the uptake levels of monocytes and macrophages (differentiated monocyte) were opposites. Specifically, BSA adsorption on poly-(methacrylic acid) nanoporous polymer particles underwent conformational changes and showed decreased internalization efficiency by THP-1 compared with pure NPs (Figure 9).10 However, the BSA on nanoporous polymer particles triggered significant internalization and proinflammatory cytokine secretion (as a result of phagocytosis activity) by differentiated macrophage cells (Figure 9).10 This study highlighted selective cell recognition for protein corona formation on NPs.
|
Figure 9 Opposite cellular uptake by monocyte (THP-1) and macrophage (dTHP-1) was observed in the presence and absence of protein corona. |
The recognition of foreign NPs could be opsonin-dependent, opsonin-independent, or a combination of both. An opsonin is any protein (ie, IgG or complement proteins) that promotes phagocytosis by marking an antigen for an immune response.57 In addition, molecules that activate the complement system are considered opsonins.57 In a series of in vitro experiments, positive correlations between complement protein (C3) adsorption and uptake/clearance were observed for liposomes and polylactic acid and poly (methyl methacrylate) NPs interacting with immune cells.58,59 Along these lines, the physiochemical surface properties of NPs can mediate the protein corona formation in which NP surface properties can enhance the probability of immune cell activation.
Complement activation by NP-protein coronas
Although protein corona formation can mitigate nanotoxicity, the immune cell response can be activated, depending on the type of corona formation. The complement system is a vital component of the immune system and acts as a first line of defense against foreign materials. Complement can be activated via three general pathways (ie, classical, lectin, and alternative) after surface binding of key complement recognition proteins.60 To recognize and eliminate foreign materials, the function of the immune system can facilitate NP uptake by phagocytic cells via cell-killing mechanisms.60,61 When NPs are exposed to blood, various NP-protein coronas can activate complement mechanisms and can cause inflammation and damage to the host.60 To attenuate complement activation, the surface functionality and availability of reactive functional groups of NPs need to be regulated.29,62 Hulander et al showed that complement activation can be attenuated by immobilizing Au-NPs on a smooth gold substrate (with an average size of 58 nm) by creating nanostructures.63 In this study, immune complement was activated through the classical pathway when the complement protein (ie, C1) bound to IgG molecules adsorbed on a nanosurface.64 Specifically, the authors observed that complement activation was significantly attenuated (nearly a 50% reduction) by the nanostructured hydrophilic gold surfaces, even after preadsorption with human IgG. As such, the activation of the complement system was suppressed when in the presence of surface-bound hydrophilic Au-NPs, whereas it was highly activated when the nanostructured surface was hydrophobized. From these findings, it can be concluded that surface nanotopography with different surface energies can influence complex blood protein adsorption mechanisms and, to this extent, can influence complement pathways in the blood system.63
Salvador-Morales et al demonstrated a method to control the levels of complement activation of poly(d,l-lactide-co-glycolide)–lipid poly(ethylene glycol) NPs by controlling surface chemical modification with methoxyl, carboxyl, and amine groups on NPs.65 Human serum and plasma protein-binding studies identified the binding of two key complement proteins, factor H (fH) and C3b β chain on the hybrid NPs played a key role in complement activation via the alternative pathway. In particular, fH can prevent the formation of C3 convertase by binding to C3b and can enhance decay activity of the convertase by dissociating Bb from the C3 and C5 convertase complexes. As a consequence, the suggested process can inhibit alternative complement pathways.66 In this study, hybrid polymeric structures of NPs with methoxyl surface groups bind with complement deactivating proteins (fH) and, thus, induce the lowest complement activation.65 Thus, attached fH on NPs can down-regulate signal pathways in the complement system via the alternative pathway by controlling surface chemical modification.65
Fibrinogen adsorption on NPs
Exposure of foreign NPs to blood can result in the adsorption of several protein layers (in this case, immune-activating proteins) that often triggers activation of the immune system.36 When foreign NPs come in contact with blood, plasma proteins adsorb onto the NP surfaces within a second and form a NP-protein corona. Although forming a NP-protein corona reduces cytotoxicity and deactivates the immune system, the adsorption of certain types of plasma proteins can induce adverse effects on the immune system, such as complement activation.
For example, specific plasma proteins can affect platelet adhesion and initiate thrombogenesis.67 Fibrinogen, fibronectin, vitronectin, and von Willebrand factors are major mediators of platelet aggregation by binding with platelet receptors, such as GP IIb/IIIa.64 Among these proteins, fibrinogen is a 340 kDa soluble plasma glycoprotein that is converted by thrombin into fibrin during the formation of a blood clot.67,68 A recent study has shown that fibrinogen in plasma can adsorb to NP surfaces at significantly higher quantities than other adhesion proteins.67 In addition, the correlation between fibrinogen adsorption and platelet was examined.68 In a separate study, Park and Khang also demonstrated that the chemical functional intensity of carboxyl groups on SWCNTs, which corresponds to the water dispersity or hydrophilicity of CNTs, can induce conformational changes in the fibrinogen domains.45 Alleviating the density of carboxyl groups of SWCNT can alter the secondary structure of fibrinogen and, thus, alter bioactivity. Different patterns of heat denaturation (change of three-dimensional structures of proteins) were observed on fibrinogen with NPs compared with free fibrinogen when increasing the hydrophilicity and concentration of SWCNTs.
Importantly, fibrinogen can trigger hemostasis (stops bleeds from a damaged blood vessel) and leukocyte activation via a well-known regulator of the inflammatory response.69
The integrin receptor Mac-1 (CD11b/CD18, αMβ2, and CR3) is the most common complex member of the entire integrin family. Mac-1 is dominantly expressed on activated leukocytes, primarily neutrophils and monocytes, and can mediate critical adhesive reactions during the inflammatory response.69 In particular, Mac-1 can promote the strong adhesion of neutrophils to endothelial cells and activate diapedesis, a process in which neutrophils migrate through the interstitial matrix (ie, a type of extracellular matrix found in interstitial connective tissue).70 The Mac-1 integrin is also involved in other neutrophil responses, such as phagocytosis, homotypic aggregation, degranulation, and adhesion to microorganisms.70
Ligands of Mac-1 can be stimulated by many extracellular matrix proteins (eg, fibronectin, collagens, thrombospondin, Cyr61, etc), blood proteins (eg, fibrinogen, iC3b, kininogen, factor H, factor X, t-PA, and so on), and proteases (eg, elastase, myeloperoxidase, plasminogen, etc).
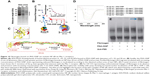
Recently, inflammatory responses to fibrinogen-adsorbed NPs by Mac-1 activation have been reported. Deng et al showed that negatively charged poly(acrylic acid)-conjugated Au-NPs can bind and induce the unfolding of fibrinogen by promoting interactions with Mac-1 (Figure 10).71 Activation of this receptor can stimulate the NF-κB signaling pathway and lead to the release of inflammatory cytokines. In this study, poly(acrylic acid)–Au-NP induced conformational changes in the fibrinogen structure significantly involved with binding to Mac-1-receptor-positive THP-1 cells, but not to Mac-1-receptor-negative HL-60 cells. In addition, fibrinogen/poly(acrylic acid)–Au-NP complexes increased levels of nuclear factor-κB in THP-1 cells (Figure 10E). As such, this study confirmed that NP surface density is another critical factor in fibrinogen binding. The study also demonstrated that negatively charged NPs induced the unfolding of fibrinogen, and thus, fibrinogen-bound NPs triggered subsequent proinflammatory responses.
|
Figure 10 Fibrinogen bound on PAA-GNP can activate NF-κB via Mac-1 receptors. |
Conclusion
In spite of the improved efficacy of many NP-based drug delivery systems, unexpected interactions with different blood components can limit their use in various biomedical applications. Protein adsorption to NPs (ie, the NP-protein corona) is critical to understanding how immune cells interact with NPs and how the NP-protein corona can regulate immunotoxicity. Although NP-protein coronas generally reduce cytotoxicity and immunotoxicity, immunotoxicity can be mitigated or activated depending on the type of NP and adsorbed plasma protein (Table 1). At this time, there is a limited understanding of the connection between the physicochemical properties of NPs and their corresponding effect on the physiological system. As such, it still remains unclear how to optimally synthesize and chemically modify NPs for in vivo application. It is clear, however, that understanding NP-protein corona formation and routes of administration can support the ability to control immune responses and cytotoxicity. The formation and immunological response to NP-protein coronas is significantly influenced by the physiochemical surface properties of the NPs (ie, physical surface architecture and chemical functionality), and thus, future works should address the effective physiochemical properties of NP for determining protein corona and associated toxicological evaluation by analyzing protein distribution and examining in vitro and in vivo responses. With an improved understanding of NP-protein corona interactions in the immune system, the adverse aspects of NPs will be anticipated in advance and inhibited through rational design.
Acknowledgments
This research was supported by the National Research Foundation of Korea funded by the Ministry of Science (2012R1A1A2041157 and 2014R1A2A1A11052615), Korea Health technology R&D Project through the KHIDI, funded by the Ministry of Health & Welfare (HI14C1802) and Korea Food and Drug Administration (KFDA).
Disclosure
The authors report no conflicts of interest in this work.
References
De M, Ghosh P, Rotello V. Applications of nanoparticles in biology. Adv Mater. 2008;20:4225–4241. | ||
Ferrari M. Cancer nanotechnology: opportunities and challenges. Nat Rev Cancer. 2005;5(3):161–171. | ||
Stephan MT, Moon JJ, Um SH, Bershteyn A, Irvine DJ. Therapeutic cell engineering with surface-conjugated synthetic nanoparticles. Nat Med. 2010;16(9):1035–1041. | ||
Zhang L, Gu FX, Chan JM, Wang AZ, Langer RS, Farokhzad OC. Nanoparticles in medicine: therapeutic applications and developments. Clin Pharmacol Ther. 2008;83(5):761–769. | ||
Mause SF, Weber C. Microparticles: protagonists of a novel communication network for intercellular information exchange. Circ Res. 2010;107(9):1047–1057. | ||
Rak J. Microparticles in cancer. Semin Thromb Hemost. 2010;36(8):888–906. | ||
Gref R, Minamitake Y, Peracchia MT, Trubetskoy V, Torchilin V, Langer R. Biodegradable long-circulating polymeric nanospheres. Science. 1994;263(5153):1600–1603. | ||
Göppert TM, Müller RH. Polysorbate-stabilized solid lipid nanoparticles as colloidal carriers for intravenous targeting of drugs to the brain: comparison of plasma protein adsorption patterns. J Drug Target. 2005;13(3):179–187. | ||
Walczyk D, Bombelli FB, Monopoli MP, Lynch I, Dawson KA. What the cell “sees” in bionanoscience. J Am Chem Soc. 2010;132(16):5761–5768. | ||
Yan Y, Gause KT, Kamphuis MM, et al. Differential roles of the protein corona in the cellular uptake of nanoporous polymer particles by monocyte and macrophage cell lines. ACS Nano. 2013;23:7(12):10960–10970. | ||
Leu D, Manthey B, Kreuter J, Speiser P, DeLuca PP. Distribution and elimination of coated polymethyl [2–14C]methacrylate nanoparticles after intravenous injection in rats. J Pharm Sci. 1984;73(10):1433–1437. | ||
Bala I, Hariharan S, Kumar MN. PLGA nanoparticles in drug delivery: the state of the art. Crit Rev Ther Drug Carrier Syst. 2004;21(5):387–422. | ||
Lesniak A, Fenaroli F, Monopoli MP, Åberg C, Dawson KA, Salvati A. Effects of the presence or absence of a protein corona on silica nanoparticle uptake and impact on cells. ACS Nano. 2012;6(7):5845–5857. | ||
Lesniak A, Salvati A, Santos-Martinez MJ, Radomski MW, Dawson KA, Åberg C. Nanoparticle adhesion to the cell membrane and its effect on nanoparticle uptake efficiency. J Am Chem Soc. 2013;135(4):1438–1444. | ||
Treuel L, Brandholt S, Maffre P, Wiegele S, Shang L, Nienhaus GU. Impact of protein modification on the protein corona on nanoparticles and nanoparticle-cell interactions. ACS Nano. 2014;8(1):503–513. | ||
Hu W, Peng C, Lv M, et al. Protein corona-mediated mitigation of cytotoxicity of graphene oxide. ACS Nano. 2011;5(5):3693–3700. | ||
Yang ST, Liu Y, Wang YW, Cao A. Biosafety and bioapplication of nanomaterials by designing protein-nanoparticle interactions. Small. 2013;9(9–10):1635–1653. | ||
Walkey CD, Chan WC. Understanding and controlling the interaction of nanomaterials with proteins in a physiological environment. Chem Soc Rev. 2012;41(7):2780–2799. | ||
Fenoglio I, Fubini B, Ghibaudi EM, Turci F. Multiple aspects of the interaction of biomacromolecules with inorganic surfaces. Adv Drug Deliv Rev. 2011;63(13):1186–1209. | ||
Monopoli MP, Aberg C, Salvati A, Dawson KA. Biomolecular coronas provide the biological identity of nanosized materials. Nat Nanotechnol. 2012;7(12):779–786. | ||
Nel AE, Mädler L, Velegol D, et al. Understanding biophysicochemical interactions at the nano-bio interface. Nat Mater. 2009;8(7):543–557. | ||
Wang X, Reece SP, Brown JM. Immunotoxicological impact of engineered nanomaterial exposure: mechanisms of immune cell modulation. Toxicol Mech Methods. 2013;23(3):168–177. | ||
Karmali PP, Simberg D. Interactions of nanoparticles with plasma proteins: implication on clearance and toxicity of drug delivery systems. Expert Opin Drug Deliv. 2011;8(3):343–357. | ||
Cedervall T, Lynch I, Lindman S, et al. Understanding the nanoparticle-protein corona using methods to quantify exchange rates and affinities of proteins for nanoparticles. Proc Natl Acad Sci U S A. 2007;104(7):2050–2055. | ||
Monopoli MP, Walczyk D, Campbell A, et al. Physical-chemical aspects of protein corona: relevance to in vitro and in vivo biological impacts of nanoparticles. J Am Chem Soc. 2011;133(8):2525–2534. | ||
Peracchia MT, Harnisch S, Pinto-Alphandary H, et al. Visualization of in vitro protein-rejecting properties of PEGylated stealth polycyanoacrylate nanoparticles. Biomaterials. 1999;20(14):1269–1275. | ||
Gref R, Lück M, Quellec P, et al. ‘Stealth’ corona-core nanoparticles surface modified by polyethylene glycol (PEG): influences of the corona (PEG chain length and surface density) and of the core composition on phagocytic uptake and plasma protein adsorption. Colloids Surf B Biointerfaces. 2000;18(3–4):301–313. | ||
Gessner A, Waicz R, Lieske A, Paulke B, Mäder K, Müller RH. Nanoparticles with decreasing surface hydrophobicities: influence on plasma protein adsorption. Int J Pharm. 2000;196(2):245–249. | ||
Gessner A, Lieske A, Paulke BR, Müller RH. Functional groups on polystyrene model nanoparticles: influence on protein adsorption. J Biomed Mater Res A. 2003;65(3):319–326. | ||
Lacerda SH, Park JJ, Meuse C, et al. Interaction of gold nanoparticles with common human blood proteins. ACS Nano. 2010;4(1):365–379. | ||
Casals E, Pfaller T, Duschl A, Oostingh GJ, Puntes V. Time evolution of the nanoparticle protein corona. ACS Nano. 2010;4(7):3623–3632. | ||
Miclăuş T, Bochenkov VE, Ogaki R, Howard KA, Sutherland DS. Spatial mapping and quantification of soft and hard protein coronas at silver nanocubes. Nano Lett. 2014;14(4):2086–2093. | ||
Medintz IL, Konnert JH, Clapp AR, et al. A fluorescence resonance energy transfer-derived structure of a quantum dot-protein bioconjugate nanoassembly. Proc Natl Acad Sci U S A. 2004;101(26):9612–9617. | ||
Aubin-Tam ME, Hamad-Schifferli K. Gold nanoparticle-cytochrome C complexes: the effect of nanoparticle ligand charge on protein structure. Langmuir. 2005;21(26):12080–12084. | ||
Roach P, Farrar D, Perry CC. Surface tailoring for controlled protein adsorption: effect of topography at the nanometer scale and chemistry. J Am Chem Soc. 2006;128(12):3939–3945. | ||
Lundqvist M, Stigler J, Elia G, Lynch I, Cedervall T, Dawson KA. Nanoparticle size and surface properties determine the protein corona with possible implications for biological impacts. Proc Natl Acad Sci U S A. 2008;105(38):14265–14270. | ||
Tenzer S, Docter D, Kuharev J, et al. Rapid formation of plasma protein corona critically affects nanoparticle pathophysiology. Nat Nanotechnol. 2013;8(10):772–781. | ||
De Paoli SH, Diduch LL, Tegegn TZ, et al. The effect of protein corona composition on the interaction of carbon nanotubes with human blood platelets. Biomaterials. 2014;35(24):6182–6194. | ||
Hühn D, Kantner K, Geidel C, et al. Polymer-coated nanoparticles interacting with proteins and cells: focusing on the sign of the net charge. ACS Nano. 2013;7(4):3253–3263. | ||
Lacerda L, Bianco A, Prato M, Kostarelos K. Carbon nanotubes as nanomedicines: from toxicology to pharmacology. Adv Drug Deliv Rev. 2006;58(14):1460–1470. | ||
Liu Z, Robinson JT, Tabakman SM, Yang K, Dai H. Carbon materials for drug delivery and cancer therapy. Mater Today. 2011;14(7–8):316–323. | ||
Cai X, Ramalingam R, Wong HS, et al. Characterization of carbon nanotube protein corona by using quantitative proteomics. Nanomedicine (Lond). 2013;9(5):583–593. | ||
Ge C, Du J, Zhao L, et al. Binding of blood proteins to carbon nanotubes reduces cytotoxicity. Proc Natl Acad Sci U S A. 2011;108(41):16968–16973. | ||
Zhu Y, Li W, Li Q, et al. Effects of serum proteins on intracellular uptake and cytotoxicity of carbon nanoparticles. Carbon. 2009;47(5):1351–1358. | ||
Park SJ, Khang D. Conformational changes of fibrinogen in dispersed carbon nanotubes. Int J Nanomedicine. 2012;7:4325–4333. | ||
Mao HY, Laurent S, Chen W, et al. Graphene: promises, facts, opportunities, and challenges in nanomedicine. Chem Rev. 2013;113(5):3407–3424. | ||
Mao H, Chen W, Laurent S, et al. Hard corona composition and cellular toxicities of the graphene sheets. Colloids Surf B Biointerfaces. 2013;109:212–218. | ||
Lemarchand C, Gref R, Passirani C, et al. Influence of polysaccharide coating on the interactions of nanoparticles with biological systems. Biomaterials. 2006;27(1):108–118. | ||
Casals E, Pfaller T, Duschl A, Oostingh GJ, Puntes VF. Hardening of the nanoparticle-protein corona in metal (Au, Ag) and oxide (Fe3O4, CoO, and CeO2) nanoparticles. Small. 2011;7(24):3479–3486. | ||
Braydich-Stolle L, Hussain S, Schlager JJ, Hofmann MC. In vitro cytotoxicity of nanoparticles in mammalian germline stem cells. Toxicol Sci. 2005;88(2):412–419. | ||
AshaRani PV, Low Kah Mun G, Hande MP, Valiyaveettil S. Cytotoxicity and genotoxicity of silver nanoparticles in human cells. ACS Nano. 2009;3(2):279–290. | ||
Kittler S, Greulich C, Diendorf J, Köller M, Epple M. Toxicity of Silver Nanoparticles Increases during Storage Because of Slow Dissolution under Release of Silver Ions. Chem Mater. 2010;22(16):4548–4554. | ||
Maiorano G, Sabella S, Sorce B, Brunetti V, Malvindi MA, Cingolani R, Pompa PP. Effects of cell culture media on the dynamic formation of protein-nanoparticle complexes and influence on the cellular response. ACS Nano. 2010;4(12):7481–7491. | ||
Lynch I, Salvati A, Dawson KA. Protein-nanoparticle interactions: What does the cell see? Nat Nanotechnol. 2009;4(9):546–547. | ||
Müller KH, Motskin M, Philpott AJ, et al. The effect of particle agglomeration on the formation of a surface-connected compartment induced by hydroxyapatite nanoparticles in human monocyte-derived macrophages. Biomaterials. 2014;35(3):1074–1088. | ||
Motskin M, Müller KH, Genoud C, Monteith AG, Skepper JN. The sequestration of hydroxyapatite nanoparticles by human monocyte-macrophages in a compartment that allows free diffusion with the extracellular environment. Biomaterials. 2011;32(35):9470–9482. | ||
Salmon JE, Kapur S, Kimberly RP. Opsonin-independent ligation of Fc gamma receptors. The 3G8-bearing receptors on neutrophils mediate the phagocytosis of concanavalin A-treated erythrocytes and nonopsonized Escherichia coli. J Exp Med. 1987;166(6):1798–1813. | ||
Harashima H, Sakata K, Funato K, Kiwada H. Enhanced hepatic uptake of liposomes through complement activation depending on the size of liposomes. Pharm Res. 1994;11(3):402–406. | ||
Leroux JC, De Jaeghere F, Anner B, Doelker E, Gurny R. An investigation on the role of plasma and serum opsonins on the internalization of biodegradable poly(d,l-lactic acid) nanoparticles by human monocytes. Life Sci. 1995;57(7):695–703. | ||
Sim RB, Tsiftsoglou SA. Proteases of the complement system. Biochem Soc Trans. 2004;32(Pt 1):21–27. | ||
Rybak-Smith MJ, Sim RB; Rybak-Smith MJ. Complement activation by carbon nanotubes. Adv Drug Deliv Rev. 2011;63(12):1031–1041. | ||
Bertholon I, Vauthier C, Labarre D. Complement activation by core-shell poly(isobutylcyanoacrylate)-polysaccharide nanoparticles: influences of surface morphology, length, and type of polysaccharide. Pharm Res. 2006;23(6):1313–1323. | ||
Hulander M, Lundgren A, Berglin M, Ohrlander M, Lausmaa J, Elwing H. Immune complement activation is attenuated by surface nanotopography. Int J Nanomedicine. 2011;6:2653–2666. | ||
Gorbet MB, Sefton MV. Biomaterial-associated thrombosis: roles of coagulation factors, complement, platelets and leukocytes. Biomaterials. 2004;25(26):5681–5703. | ||
Salvador-Morales C, Flahaut E, Sim E, Sloan J, Green ML, Sim RB, Salvador-Morales C. Complement activation and protein adsorption by carbon nanotubes. Mol Immunol. 2006;43(3):193–201. | ||
Kouser L, Abdul-Aziz M, Nayak A, Stover CM, Sim RB, Kishore U. Properdin and factor h: opposing players on the alternative complement pathway “see-saw”. Front Immunol. 2013;4:93. | ||
Horbett T. Chapter 13 Principles underlying the role of adsorbed plasma proteins in blood interactions with foreign materials. Cardiovasc Pathol. 1993;2(3):137–148. | ||
Chinn JA, Horbett TA, Ratner BD. Baboon fibrinogen adsorption and platelet adhesion to polymeric materials. Thromb Haemost. 1991;65(5):608–617. | ||
Lishko VK, Podolnikova NP, Yakubenko VP, et al. Multiple binding sites in fibrinogen for integrin alphaMbeta2 (Mac-1). J Biol Chem. 2004 22;279(43):44897–44906. | ||
Ding ZM, Babensee JE, Simon SI, et al. Relative contribution of LFA-1 and Mac-1 to neutrophil adhesion and migration. J Immunol. 1999;163(9):5029–5038. | ||
Deng ZJ, Liang M, Monteiro M, Toth I, Minchin RF. Nanoparticle-induced unfolding of fibrinogen promotes Mac-1 receptor activation and inflammation. Nat Nanotechnol. 2011;6(1):39–44. |
 © 2014 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms.php
and incorporate the Creative Commons Attribution
- Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2014 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms.php
and incorporate the Creative Commons Attribution
- Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.