Back to Journals » Journal of Inflammation Research » Volume 7
Effect of plant extracts on H2O2-induced inflammatory gene expression in macrophages
Authors Pomari E, Stefanon B, Colitti M
Received 28 January 2014
Accepted for publication 6 March 2014
Published 24 June 2014 Volume 2014:7 Pages 103—112
DOI https://doi.org/10.2147/JIR.S61471
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Elena Pomari, Bruno Stefanon, Monica Colitti
Department of Agricultural and Environmental Sciences, University of Udine, Udine, Italy
Background: Arctium lappa (AL), Camellia sinensis (CS), Echinacea angustifolia, Eleutherococcus senticosus, Panax ginseng (PG), and Vaccinium myrtillus (VM) are plants traditionally used in many herbal formulations for the treatment of various conditions. Although they are well known and already studied for their anti-inflammatory properties, their effects on H2O2-stimulated macrophages are a novel area of study.
Materials and methods: Cell viability was tested after treatment with increasing doses of H2O2 and/or plant extracts at different times of incubation to identify the optimal experimental conditions. The messenger (m)RNA expression of TNFα, COX2, IL1β, NFκB1, NFκB2, NOS2, NFE2L2, and PPARγ was analyzed in macrophages under H2O2 stimulation. The same genes were also quantified after plant extract treatment on cells pre-stimulated with H2O2.
Results: A noncytotoxic dose (200 µM) of H2O2 induced active mRNA expression of COX2, IL1β, NFE2L2, NFκB1, NFκB2, NOS2, and TNFα, while PPARγ was depressed. The expression of all genes tested was significantly (P<0.001) regulated by plant extracts after pre-stimulation with H2O2. COX2 was downregulated by AL, PG, and VM. All extracts depressed IL1β expression, but upregulated NFE2L2. NFκB1, NFκB2, and TNFα were downregulated by AL, CS, PG, and VM. NOS2 was inhibited by CS, PG, and VM. PPARγ was decreased only after treatment with E. angustifolia and E. senticosus.
Conclusion: The results of the present study indicate that the stimulation of H2O2 on RAW267.4 cells induced the transcription of proinflammatory mediators, showing that this could be an applicable system by which to activate macrophages. Plant extracts from AL, CS, PG, and VM possess in vitro anti-inflammatory activity on H2O2-stimulated macrophages by modulating key inflammation mediators. Further in vitro and in vivo investigation into molecular mechanisms modulated by herbal extracts should be undertaken to shed light on the development of novel modulating therapeutic strategies.
Keywords: inflammation, qRT-PCR, RAW264.7 cells, nutraceuticals
Introduction
The use of herbal/botanical products worldwide enjoys increasing popularity.1,2 It is known that the great variety of plant secondary metabolites are excellent sources for components such as polyphenols, which exhibit antioxidant, anticarcinogenic, antimutagenic, and anti-inflammatory effects.3–5 The understanding of the cellular and molecular mechanisms involved in the inflammatory process has increased in recent decades, and this has permitted the discovery of many promising targets for the development of new bioactive compounds to treat chronic inflammatory diseases.6
Arctium lappa (AL), commonly known as burdock, has anti-inflammatory,7 free radical scavenging,8 antioxidant,9 and liver10 and skin11 protective effects. On lipopolysaccharide (LPS)-stimulated macrophages, arctigenin exhibited potent inhibitory effects on the production of nitric oxide (NO) and the release of tumor necrosis factor α (TNFα) and interleukin 6 (IL6), but not cyclooxygenase 2 (COX2) expression or COX2 activity.12
Camellia sinensis (CS), as green tea, is a species of plant whose leaves and leaf buds are used to produce the popular tea beverage. It is widely known for its anti-inflammatory effect, and there is growing interest in its cardiovascular health benefits.13 Green tea and its major ingredient, epigallocatechin gallate, effectively inhibited the release of high mobility group box 1 (HMGB1) in LPS-induced macrophage cultures14 by promoting its aggregation and autophagic degradation in these cells.15
Echinacea spp. plant preparations are claimed to have several pharmacological effects, such as immune enhancement, anti-inflammation, and prevention of common cold.16 Numerous reports have demonstrated that Echinacea spp. constituents have inflammation-related bioactivities such as stimulation of cytokine production17 or inhibition of COX1 or COX2 activity in vitro.18 Hou et al,19 in murine macrophages, demonstrated that alkamides significantly inhibited COX2 activity and the LPS-induced expression of COX2, inducible NO synthase (NOS2), and specific cytokines or chemokines.
Eleutherococcus senticosus (ES) is used as an immunological modulator in Oriental medicine.20 It has been used to treat stress-induced physiological changes and various allergic disorders.21–23 However, it is still unclear how it inhibits allergic responses and how effective it is in experimental models. On activated murine macrophage RAW264.7 cells, ES downregulated NOS2 expression, showing pro-inflammatory action by blocking activation of JNK and Akt signaling pathways.20
The beneficial effects of Panax ginseng (PG), have been attributed to the biological activities of its constituents, the ginsenosides. Ginseng extracts and ginsenosides have been reported to have anti-inflammatory properties.24 Ginseng can modulate the generation of inflammatory mediators (such as NO, synthesized by the activity of NOSs25) and the phagocytic activity of macrophages.26
Vaccinium myrtillus (VM) was one of the most frequently used antidiabetic remedies before the discovery of insulin.27 However, Hou et al28 demonstrated that delphinidin, one of the five anthocyanidins present in bilberry, suppressed COX2 by blocking MAPK-mediated pathways with the attendant activation of nuclear factor κB (NFκB), activator protein 1 (AP1), and CCAAT-enhancer-binding protein delta (C/EBPδ) in LPS-activated murine macrophage RAW264.7 cells. Moreover, gene expression profiling of inflammatory cell model RAW264.7 treated with bilberry extract was provided, showing that bilberry extracts affected genes related to inflammation and defense.29
A broad range of roles of secondary plant metabolites has been identified for several inflammatory cells and for a large number of inflammatory mediators in important pathologies not previously known to be linked to inflammation, such as obesity, atherosclerosis, and cancer.30,31 However, evidence exists that nutraceuticals can modulate various inflammatory mediators (eg, cytokines), the production and/or action of second messengers (cGMP, cAMP, protein kinases), the expression of transcription factors (AP1, NFκB), and the expression of key proinflammatory molecules such as NOS2, COX2, interleukin 1β (IL1β) and TNFα, neuropeptides (substance P), and proteases (matrix proteases).6 Moreover, whereas the NFκB plays an important role as transcription factor in coordinating the expression of proinflammatory genes, the peroxisome proliferator-activated receptor γ (PPARγ) plays an important role in controlling various diseases based on its anti-inflammatory properties.32,33 In this regard, expression of PPARγ is not restricted to adipocytes, but is also found in immune cells, such as B and T lymphocytes, monocytes, macrophages, dendritic cells, and granulocytes.34 All this evidence suggests a role of PPARγ in the control of the inflammatory response with potential therapeutic applications in inflammation-related diseases.35
It has been shown that nuclear factor E2-related factor 2 (NFE2L2) is responsible for both constitutive and inducible expression of antioxidant response element-regulated genes,36 indicating an important protective role in regulating antioxidant and anti-inflammatory cellular responses.
In this regard, the aim of this research was to investigate the molecular activity of AL, CS, Echinacea angustifolia (EA), ES, PG, and VM extracts on cell model RAW264.7, in order to investigate their effect in modulating messenger (m)RNA expression of genes involved in the inflammatory process: TNFα, COX2, IL1β, NFκB1, NFκB2, NOS2, NFE2L2, and PPARγ. RAW264.7 cells were pre-stimulated with H2O2 to mimic inflammatory conditions and then treated with plant extracts.
Materials and methods
Plant extracts
All the extracts were obtained from commercial companies, standardized for the content of bioactive compounds (Table 1). AL was provided by Bayer S.p.A. (Milan, Italy), and CS, EA, ES, PG, and VM were provided by ACEF S.p.a. (Piacenza, Italy). The extracts (50 mg) were dissolved in 1 mL of 10% dimethyl sulfoxide (DMSO), passed through 0.22 μm pore-sized membrane filter (EMD Millipore, Billerica, MA, USA), and kept in the dark at −20°C until further analysis.
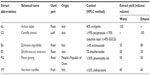  | Table 1 Abbreviation, botanical name, used part, and yield of extract of each screened plant |
Cell culture
RAW264.7 murine macrophages (American Type Culture Collection [ATCC], Manassas, VA, USA) were cultured in Dulbecco’s Modified Eagle’s Medium supplemented with 10% heat-inactivated fetal bovine serum, 2 mM L-glutamine, 100 U/mL penicillin, and 100 μg/mL streptomycin. All reagents were purchased from Euroclone (Milan, Italy). Cells were maintained in humidified air with 5% CO2 at 37°C. For all experiments, the cells were grown to approximately 70%–90% confluence and culture medium with 2% heat-inactivated fetal bovine serum was used.37
H2O2 cell treatment
A preliminary assay was performed by treating macrophages with different final concentrations of H2O2 (50, 100, and 200 μM) diluted in phosphate-buffered saline (PBS) and further diluted in culture medium. The control cells (CTRL) were incubated for 24 hours with the same final amount of <0.1% DMSO in the culture medium.
This assay was used to identify the concentration of H2O2 that would be used to induce an inflammatory condition in the subsequent experiments. Cell viability was measured after 30 and 60 minutes of incubation with H2O2 (50, 100, and 200 μM).
Plant extracts cell treatment
To test the effect of plant extracts on macrophages, cells were washed with PBS and treated with different doses of plant extract (1, 10, 100, and 200 μg/mL) in a final concentration of <0.1% DMSO in the culture medium for 24 hours. In addition, a combined effect of each plant extract and H2O2 was assayed by incubating cells for 60 minutes with H2O2 (200 μM) and, after washing with PBS, with the specific dose of each plant extract in a final concentration of <0.1% DMSO in the culture medium for 24 hours. The CTRL were incubated for 24 hours with the same final amount of <0.1% DMSO in the culture medium.
Cell viability assay
To ascertain the cytotoxic effect of H2O2 and plant extracts, viability of RAW264.7 cells was measured by the 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) colorimetric assay.11,38 For the assay, cells were seeded in a 96-well plate at a density of 2×104 cells/well and left to grow for 24 hours. Per well, 10 μL of MTT reagent was added and the mixture was further incubated for 3 hours. Next, the mixture in each well was removed, and formed formazan crystals were dissolved in 100 μL of DMSO. Optical density measurement of the mixture was performed in a microplate reader (Tecan GENios, Männedorf, Switzerland) at 550 nm. Cell viability was expressed as percentage of each treatment versus the CTRL.
Quantitative reverse transcriptase polymerase chain reaction (qRT-PCR)
To determine the modulation of H2O2 on mRNA expression of IL1β, NFE2L2, NFκB1, NFκB2, PPARγ, NOS2, COX2, and TNFα, qRT-PCR was preliminarily performed on macrophages treated with 50, 100, and 200 μM H2O2 for 30 and 60 minutes.
In addition, macrophages were treated with 10 μg/mL of each extract for 24 hours after a pre-stimulation using 200 μM H2O2 for 60 minutes. To determine whether plant extracts had an effect on H2O2 pre-stimulated cells, the mRNA expression of the same genes was analyzed.
After treatment, cells were washed with cold PBS before total RNA extraction using the RNeasy Plus Mini Kit (Qiagen NV, Venlo, the Netherlands), according to the manufacturer’s instructions. Then, complementary (c)DNA was synthesized from the same purified RNA samples according to ImProm-II™ Reverse Transcription System (Promega Corporation, Fitchburg, WI, USA) and using random and oligo(dT)15 primers. Briefly, 1 μg of the purified total RNA was denatured by incubating at 70°C for 5 minutes and then placed at 25°C for 5 minutes, 42°C for 60 minutes, and 70°C for 15 minutes.
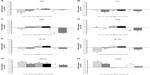
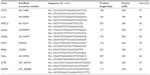
Primer3 Input software (Whitehead Institute for Biomedical Research, Cambridge, MA, USA)39 was used to design the primer sequences encoding for β actin (ACTB), glyceraldehyde-3-phosphate dehydrogenase (GAPDH), IL1β, NFE2L2, NFκB1, NFκB2, PPARγ, NOS2, COX2, and TNFα (Table 2). The relative qRT-PCR was performed in triplicate with the Platinum® SYBR® Green qPCR SuperMix-UDG with ROX (Life Technologies, Carlsbad, CA, USA) on CFX96 Real-Time PCR Detection System (Bio-Rad Laboratories Inc., Hercules, CA, USA). Reaction mixes were prepared with 200 nM of each primer pair (Table 2) and the cDNA (10 ng) added last. Differences between samples and controls were calculated using the 2−ΔΔCt method,40,41 where 2−ΔΔCt represents the difference of a given target gene in H2O2-treated cells versus the CTRL (Table 3) and each sample treated with plant extract versus H2O2-treated cells (Figure 1). The expression of target genes was normalized using ACTB and GAPDH, and ΔCts were calculated by the difference between Ct of the gene target and the geometric mean of the two housekeeping genes. The n-fold expression of a given target gene was calculated as log2(2−ΔΔCt).
Statistical analysis
Cell viability data related to H2O2 treatments and computed as % mean ± standard error (SE), were analyzed using the analysis of variance (ANOVA) model to assess the significance of doses (50, 100, and 200 μM H2O2) and time of treatments (30 minutes and 60 minutes). The effect of plant extracts on cell viability was analyzed using the ANOVA model. The differences in cell viability were analyzed among doses (1, 10, 100, and 200 μg/mL) of each extract. Means were compared adjusting the P-values with Bonferroni correction for multiple comparisons (Statistical Package for the Social Sciences [SPSS] Version 16.0; SPSS Inc., Chicago, USA).
Relative gene expression data, after H2O2 treatments, were analyzed using the general linear model to assess the significance of doses (50, 100, and 200 μM H2O2) and time of treatments (30 minutes and 60 minutes). Expression data are reported as a mean of log2(n-fold) ± SE. Means were compared adjusting P-values with Bonferroni correction for multiple comparisons (SPSS Version 16.0; SPSS Inc.).
The effect of plant extracts on gene expressions, computed as log2(n-fold), were checked by the general linear model. The differences within each gene were analyzed among different extracts. Data were reported as mean of log2(n-fold) ± SE. For mean comparisons, the P-values were adjusted with Bonferroni correction for multiple comparisons (SPSS Version 16.0; SPSS Inc.).
Results
Cell viability
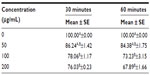
To analyze the effect of dose and time of H2O2 exposure, a preliminary cell viability assay was performed by treating macrophages with increasing doses of H2O2 from 1 to 200 μM at two times of incubation (30 minutes and 60 minutes). Cell viability (% versus CTRL) after exposure for 30 minutes and 60 minutes slowly decreased, remaining at over 60% (P<0.001) up to 200 μM (67.89%±1.66%) (Table 4). The dose of 200 μM of H2O2 was chosen to evoke inflammation for 60 minutes without affecting the cell viability (see also the next paragraph).
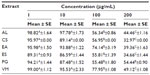
In addition, cells were treated for 24 hours with plant extracts (Table 5). A concentration of 10 μg/mL of each AL, CS, EA, ES, PG, and VM significantly (P<0.001) reduced cell viability in comparison to the CTRL, remaining, nevertheless, over 80%. On the contrary, concentrations of 100 and 200 μg/mL led to a strong decrease (P<0.001) in viability. Based on this evidence, 10 μg/mL of each plant extract was chosen for the further experiments.
H2O2 upregulates mRNA expression of inflammatory genes in macrophages
As shown in Table 3, the H2O2 treatment induced the upregulation of all genes, except PPARγ, at both incubation times. In particular, the mRNA expression of NFκB1, NFκB2, COX2, and NFE2L2 significantly increased (P<0.001) after 60 minutes of incubation. TNFα expression was significantly induced at P<0.05. The mRNA expression of COX2 was also significantly (P<0.001) upregulated in a concentration-dependent manner. The mRNA expression of TNFα, NFκB1, NFκB2, and NFE2L2 showed a similar trend, at P<0.01. The interaction between time and dose of incubation was significant in the expression of COX2 (P<0.001) and NFκB2, NOS2, and NFE2L2 (P<0.05). The relative expression of IL1β and PPARγ did not present any significant variation between doses, times of incubation, or in the interaction between time and dose. Taken together, these results show that 60 minutes of exposure and a concentration of 200 μM were significantly more effective than the other times or concentrations in stimulating the mRNA expression, and therefore were chosen for the further experiments.
Plant extracts modulate the expression of cytokines in pretreated H2O2 macrophages
As shown in Figure 1, all of the treatments significantly (P<0.001) modulated the mRNA expression of each gene, but with some specific effect. AL showed an increase of NOS2, NFE2L2, and PPARγ expression, while depressing the expression of TNFα, IL1β, COX2, NFκB1, and NFκB2. CS stimulated COX2, NFE2L2, and PPARγ, and negatively modulated TNFα, IL1β, NOS2, NFκB1, and NFκB2. EA and ES upregulated all genes except IL1β and PPARγ. PG and VM caused a decreased expression of all genes apart from NFE2L2 and PPARγ, which were stimulated.
Discussion
Epidemiological studies have provided convincing evidence that natural dietary compounds possess many biological activities against various chronic inflammatory diseases.42,43 In the present study, we evaluated the effect of AL, CS, EA, ES, PG, and VM extracts on the expression of relevant genes involved in inflammatory response. The analyses were performed on macrophages, key cells of immunity mechanisms.44 In particular, these cells are essential in the shift of the immune response from the so-called type 1 response with cytotoxic effector cells toward the type 2 response that is characterized by antibody production. Previous studies have been commonly conducted on LPS-induced macrophages, while, in this paper, macrophages were stimulated by direct addition of H2O2. In fact, H2O2 acts as a secondary messenger in diverse stimuli, which are also not infective, that activate NFκB.45 In this study, the data demonstrated that 200 μM of H2O2 affects cell viability similarly to lower doses. Moreover, the choice of 200 μM H2O2 is a good compromise between the doses that activate inflammatory mediators, such as NFκB, and those naturally produced in the inflammatory process.46 In fact, the addition of 200 μM H2O2 for 60 minutes on RAW264.7 cells significantly stimulated the transcription of TNFα, COX2, NFκB1, NFκB2, NOS2, and NFE2L2. Interestingly, no differences were found between times of exposure and doses in IL1β and PPARγ expression. H2O2 clearly stimulated the expression of NFκB-dependent genes in RAW264.7 cells, likely induced by proinflammatory cytokine TNFα, as already observed in MCF-7 cells.47,48 Moreover, it is known that several proteins can be oxidized and thus modified by H2O2. These redox-regulated proteins include transcription factors such as p53, Jun, Fos, and the p50 subunit of NFκB (NFκB1). The oxidation of these proteins can either prevent (p53, Jun, Fos) or stimulate (p50) the transcriptional activity of these genes.49
The plant extract treatments on pre-stimulated macrophages showed an anti-inflammatory action, downregulating inflammatory genes that typically can activate NFκB1 and NFκB2 pathways, as previously observed.12,50
Flavonoids, which specifically affect the modulation of inflammatory gene expression and the function of enzymes critically involved in inflammation,51,52 are also present in AL,53 CS,54 PG,55 and VM.56
Zhao et al,12 using LPS to stimulate RAW264.7 cells, demonstrated a block in NO production after addition of the active compound arctigenin. Moreover, the major polyphenol in CS extracts, epigallocatechin gallate, has been recently found to suppress NOS2 mRNA and protein expression in macrophages57 through the inhibition of the activation of NFκB.58 In the present study, NOS2 was upregulated by AL, and by up to twofold by EA and ES, in comparison to the control. CS downregulated NOS2 expression related to the control, possibly through inhibition of NFκB1 and NFκB2, but without any significant difference compared to other compounds. In general, controversial data could be due to the different extraction solvent, stimulant (LPS or H2O2), and different active compounds present in the extract.
Anthocyanins, from bilberry (VM), constitute the major bioactive components responsible for the observed downregulating effects on genes involved in inflammatory response, as already observed in a microarray study.29 Among the tested extracts, VM showed the highest anti-inflammatory activity, as also reported in LPS-stimulated human monocytic cell line by downregulating all the proinflammatory genes, in particular COX2 and NOS2.59 It has previously been shown that, in murine RAW264.7 cells, anthocyanidins inhibited LPS-induced COX2 expression by blocking MAPK pathways with the attendant activation of NFκB, C/EBPδ, and AP1.28
EA and ES seem to act as immunostimulating agents, downregulating IL1β only. It is known that Echinacea spp. plant extract has been used for immunomodulation,60 but the evidence supporting its therapeutic potential is still controversial. In vitro studies have shown that the commercial preparations of Echinacea spp. extract stimulate cytokine production by human peripheral blood macrophages17 and enhance NK cell function and antibody-dependent cytotoxicity of peripheral blood mononuclear cells.61,62 In the present study, EA caused an upregulation of NFκBs, TNFα, and COX2. An enhanced release of cytokines (such as TNFα and interferon-γ) in response to Echinacea spp. components was also detectable in rat spleen macrophage.60 The existing results suggest that the Echinacea spp. preparations are potentially effective in stimulating an in vitro and in vivo nonspecific immune response.63,64
Similarly to EA, ES downregulated IL1β only. Previous reports indicated that the isolated compounds from ES inhibit the NOS2 as well as COX2 expression.65,66 However, it has also been shown that 80% ethanol extract from ES attenuated LPS-induced NOS2 expression but not COX2 expression.20 Dissimilar results may be due to efficacy according to concentration between whole extract and its derived compounds. On the other hand, EA and ES are the only extracts that downregulated the expression of PPARγ, therefore promoting the expression of inflammatory mediators. The PPARγ, a member of the nuclear hormone receptor superfamily, is essential for adipocyte differentiation and glucose homeostasis. PPARγ has been found to regulate macrophage activation in response to mitogens and inflammation,67 and it was recognized as a negative regulator of the inflammatory activation of both macrophage and T-cells.68 Christensen et al69 demonstrated that, on 3T3-L1 adipocytes, various fatty acids and alkylamides from the flowers of Echinacea purpurea at high concentrations activated PPARγ. An involvement of PPARγ in the inhibition of IL2 secretion by T-cells in response to EA alkylamides has also been observed.70 Interestingly, only EA and ES did not stimulate the expression of PPARγ, and this was in agreement with the attendant overexpression of the proinflammatory genes.
EA and ES, as well as AL, CS, PG, and VM, upregulated NFE2L2, a transcriptional regulator of antioxidant enzymes with an important role in attenuating oxidative stress associated with diseases.71 CS has been shown to activate NFE2L2 in HL-1 murine cardiomyocytes and to induce phase II enzymes through the antioxidant response element and concomitant protection against H2O2.72 Furthermore, echinacoside, one of the major active compounds in EA,73 resulted in strong activation of NFE2L2 in HaCaT cells, providing a potential skin protection.74 Interestingly, in the present study, phytochemical treatments modulated the NFE2L2 expression, which was increased by the oxidant treatment.
Conclusion
Although the important role that H2O2 plays as an intracellular messenger is not well understood, it is known that local H2O2 concentration in extracellular fluids during inflammatory processes are sufficient to activate the NFκB pathway, thereby allowing induction of the synthesis of cytokines and other inflammatory mediators. The stimulation of H2O2 on RAW264.7 cells induced the transcription of proinflammatory mediators, showing that this could be an applicable system by which to activate macrophages. The extracts from AL, CS, PG, and VM decreased the expression of proinflammatory genes on H2O2-induced RAW264.7 cells in vitro, while EA and ES, on the contrary, promoted these inflammatory mediators.
The promising anti-inflammatory molecular mechanisms of plant extracts in macrophages of murine RAW264.7 cells require validation in future in vivo dietary intervention studies.
Acknowledgments
This work was supported by Progetto Nutriheart POR FESR 2007–2013 Friuli Venezia Giulia, Italy.
Author contributions
Elena Pomari performed the experiments concerning cell viability, isolation of RNA, and qRT-PCR. Moreover, she assisted with interpretation of the results and helped draft the manuscript. Bruno Stefanon and Monica Colitti assisted with interpretation of the results, supervised the analyses, and helped to draft the manuscript. All authors read and approved the final manuscript.
Disclosure
The authors report no conflicts of interest in this work.
References
Tapsell LC, Hemphill I, Cobiac L, et al. Health benefits of herbs and spices: the past, the present, the future. Med J Aust. 2006;185:S4–S24. | |
Iriti M, Vitalini S, Fico G, Faoro F. Neuroprotective herbs and foods from different traditional medicines and diets. Molecules. 2010;15:3517–3555. | |
Farinacci M, Colitti M, Sgorlon S, Stefanon B. Immunomodulatory activity of plant residues on ovine neutrophils. Vet Immunol Immunopathol. 2008;126:54–63. | |
Pandey KB, Rizvi SI. Plant polyphenols as dietary antioxidants in human health and disease. Oxid Med Cell Longev. 2009;2:270–278. | |
Colitti M, Gaspardo B, Della Pria A, Scaini C, Stefanon B. Transcriptome modification of white blood cells after dietary administration of curcumin and non-steroidal anti-inflammatory drug in osteoarthritic affected dogs. Vet Immunol Immunopathol. 2012;147:136–146. | |
Gautam R, Jachak SM. Recent developments in anti-inflammatory natural products. Med Res Rev. 2009;29:767–820. | |
Sohn EH, Jang SA, Joo H, et al. Anti-allergic and anti-inflammatory effects of butanol extract from Arctium Lappa L. Clin Mol Allergy. 2011;9:4. | |
Lin CC, Lu JM, Yang JJ, Chuang SC, Ujiie T. Anti-inflammatory and radical scavenge effects of Arctium lappa. Am J Chin Med. 1996;24:127–137. | |
Chan YS, Cheng LN, Wu JH, et al. A review of the pharmacological effects of Arctium lappa (burdock). Inflammopharmacology. 2011;19:245–254. | |
Lin S, Lin C, Lin C, et al. Hepatoprotective effects of Arctium lappa Linne on liver injuries induced by chronic ethanol consumption and potentiated by carbon tetrachloride. J Biomed Sci. 2002;9:401–409. | |
Pomari E, Stefanon B, Colitti M. Effect of Arctium lappa (burdock) extract on canine dermal fibroblasts. Vet Immunol Immunopathol. 2013;156:159–166. | |
Zhao F, Wang L, Liu K. In vitro anti-inflammatory effects of arctigenin, a lignan from Arctium lappa L., through inhibition on iNOS pathway. J Ethnopharmacol. 2009;122:457–462. | |
Dell’Agli M, Di Lorenzo C, Badea M, et al. Plant food supplements with anti-inflammatory properties: a systematic review (I). Crit Rev Food Sci Nutr. 2013;53:403–413. | |
Chen X, Li W, Wang H. More tea for septic patients? – Green tea may reduce endotoxin-induced release of high mobility group box 1 and other pro-inflammatory cytokines. Med Hypotheses. 2006;66:660–663. | |
Li W, Zhu S, Li J, et al. EGCG stimulates autophagy and reduces cytoplasmic HMGB1 levels in endotoxin-stimulated macrophages. Biochem Pharmacol. 2011;81:1152–1163. | |
Barrett B. Medicinal properties of Echinacea: a critical review. Phytomed. 2003;10:66–86. | |
Burger RA, Torres AR, Warren RP, Caldwell VD, Hughes BG. Echinacea-induced cytokine production by human macrophages. Int J Immunopharmacol. 1997;19:371–379. | |
Hinz B, Woelkart K, Bauer R. Alkamide from Echinacea inhibit cyclooxygenase-2 activity in human neuroglioma cells. Biochem Biophys Res Commun. 2007;360:441–446. | |
Hou CC, Chen CH, Yang NS, et al. Comparative metabolomics approach coupled with cell- and gene-based assays for species classification and anti-inflammatory bioactivity validation of Echinacea plants. J Nutr Biochem. 2010;21:1045–1059. | |
Jung CH, Jung H, Shin YC, et al. Eleutherococcus senticosus extract attenuates LPS-induced iNOS expression through the inhibition of Akt and JNK pathways in murine macrophage. J Ethnopharmacol. 2007;113:183–187. | |
Gaffney BT, Hügel HM, Rich PA. The effects of Eleutherococcus senticosus and Panax ginseng on steroidal hormone indices of stress and lymphocyte subset numbers in endurance athletes. Life Sci. 2001;70:431–442. | |
Kimura Y, Sumiyoshi M. Effects of various Eleutherococcus senticosus cortex on swimming time, natural killer activity and corticosterone level in forced swimming stressed mice. J Ethnopharmacol. 2004;95:447–453. | |
Zhang N, Van Crombruggen K, Holtappels G, Bachert C. A herbal composition of Scutellaria baicalensis and Eleutherococcus senticosus shows potent anti-inflammatory effects in an ex vivo human mucosal tissue model. Evid Based Complement Alternat Med. 2012;2012:673145. | |
Xie W, Du L. Diabetes is an inflammatory disease: evidence from traditional Chinese medicines. Diabetes Obes Metab. 2011;13:289–301. | |
Redington AE. Modulation of nitric oxide pathways: therapeutic potential in asthma and chronic obstructive pulmonary disease. Eur J Pharmacol. 2006;533:263–276. | |
Shin JY, Song JY, Yun YS, Yang HO, Rhee DK, Pyo S. Immunostimulating effects of acidic polysaccharides extract of Panax ginseng on macrophage function. Immunopharmacol Immunotoxicol. 2002;24:469–482. | |
Helmstädter A, Schuster N. Vaccinium myrtillus as an antidiabetic medicinal plant – research through the ages. Pharmazie. 2010;65:315–321. | |
Hou DX, Yanagita Y, Uto T, Masuzaki S, Fujii M. Anthocyanidins inhibit cyclooxygenase-2 expression in LPS-evoked macrophages: structure activity relationship and molecular mechanisms involved. Biochem Pharmacol. 2005;70:417–425. | |
Chen J, Uto T, Tanigawa S, Kumamoto T, Fujii M, Hou DX. Expression profiling of genes targeted by bilberry (Vaccinium myrtillus) in macrophages through DNA microarray. Nutr Cancer. 2008;60 Suppl 1:43–50. | |
Burke KE. Photoaging: the role of oxidative stress. G Ital Dermatol Venereol. 2010;145:445–459. | |
Drummond GR, Selemidis S, Griendling KK, Sobey CG. Combating oxidative stress in vascular disease: NADPH oxidases as therapeutic targets. Nat Rev Drug Discov. 2011;10:453–471. | |
Spiegelman BM. PPARgamma in monocytes: less pain, any gain? Cell. 1998;93:153–155. | |
Gurnell M. PPARgamma and metabolism: insights from the study of human genetic variants. Clin Endocrinol (Oxf). 2003;59:267–277. | |
Yang J, Zhou Y, Guan Y. PPARγ as a therapeutic target in diabetic nephropathy and other renal diseases. Curr Opin Nephrol Hypertens. 2012;21:97–105. | |
Belvisi MG, Hele DJ, Birrell MA. Peroxisome proliferator-activated receptor gamma agonists as therapy for chronic airway inflammation. Eur J Pharmacol. 2006;533:101–109. | |
Pedruzzi LM, Stockler-Pinto MB, Leite M Jr, Mafra D. Nrf2-keap1 system versus NF-κB: the good and the evil in chronic kidney disease? Biochimie. 2012;94:2461–2466. | |
Cho M, Hunt TK, Hussain MZ. Hydrogen peroxide stimulates macrophage vascular endothelial growth factor release. Am J Physiol Heart Circ Physiol. 2001;280:H2357–H2363. | |
Pruett SB, Loftis AY. Characteristics of MTT as an indicator of viability and respiratory burst activity of human neutrophils. Int Arch Allergy Appl Immunol. 1990;92:189–192. | |
Rozen S, Skaletsky H. Primer3 on the WWW for general users and for biologist programmers. Methods Mol Biol. 2000;132:365–386. | |
Bustin SA, Benes V, Garson JA, et al. The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem. 2009;55:611–622. | |
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods. 2001;25:402–408. | |
Bellik Y, Boukraâ L, Alzahrani HA, et al. Molecular mechanism underlying anti-inflammatory and anti-allergic activities of phytochemicals: an update. Molecules. 2012;18:322–353. | |
Lai YS, Hsu WH, Huang JJ, Wu SC. Antioxidant and anti-inflammatory effects of pigeon pea (Cajanus cajan L.) extracts on hydrogen peroxide- and lipopolysaccharide-treated RAW264.7 macrophages. Food Funct. 2012;3:1294–1301. | |
Al-Mulla F, Leibovich SJ, Francis IM, Bitar MS. Impaired TGF-β signaling and a defect in resolution of inflammation contribute to delayed wound healing in a female rat model of type 2 diabetes. Mol Biosyst. 2011;7:3006–3020. | |
Schreck R, Rieber P, Baeuerle PA. Reactive oxygen intermediates as apparently widely used messengers in the activation of the NF-kappa B transcription factor and HIV-1. EMBO J. 1991;10:2247–2258. | |
Liu X, Zweier JL. A real-time electrochemical technique for measurement of cellular hydrogen peroxide generation and consumption: evaluation in human polymorphonuclear leukocytes. Free Radic Biol Med. 2001;31:894–901. | |
de Oliveira-Marques V, Cyrne L, Marinho HS, Antunes F. A quantitative study of NF-kappaB activation by H2O2: relevance in inflammation and synergy with TNF-alpha. J Immunol. 2007;178:3893–3902. | |
Solt LA, Madge LA, Orange JS, May MJ. Interleukin-1-induced NF-kappaB activation is NEMO-dependent but does not require IKKbeta. J Biol Chem. 2007;282:8724–8733. | |
Reth M. Hydrogen peroxide as second messenger in lymphocyte activation. Nat Immunol. 2002;3:1129–1134. | |
Wang BS, Yen GC, Chang LW, Wen-Jye Yen, Pin-Der Duh. Protective effects of burdock (Arctium lappa Linne) on oxidation of low-density lipoprotein and oxidative stress in RAW264.7 macrophages. Food Chem. 2007;101:729–738. | |
Middleton E, Kandaswami C, Theoharides TC. The effects of plant flavonoids on mammalian cells: implications for inflammation, heart disease, and cancer. Pharmacol Rev. 2000;52:673–751. | |
Kim HP, Son KH, Chang HW, Kang SS. Anti-inflammatory plant flavonoids and cellular action mechanisms. J Pharmacol Sci. 2004;96:229–245. | |
Ferracane R, Graziani G, Gallo M, Fogliano V, Ritieni A. Metabolic profile of the bioactive compounds of burdock (Arctium lappa) seeds, roots and leaves. J Pharm Biomed Anal. 2010;51:399–404. | |
van Dam RM, Naidoo N, Landberg R. Dietary flavonoids and the development of type 2 diabetes and cardiovascular diseases: review of recent findings. Curr Opin Lipidol. 2013;24:25–33. | |
Qian ZM, Lu J, Gao QP, Li SP. Rapid method for simultaneous determination of flavonoid, saponins and polyacetylenes in folium ginseng and radix ginseng by pressurized liquid extraction and high-performance liquid chromatography coupled with diode array detection and mass spectrometry. J Chromatogr A. 2009;1216:3825–3830. | |
Triebel S, Trieu HL, Richling E. Modulation of inflammatory gene expression by a bilberry (Vaccinium myrtillus L.) extract and single anthocyanins considering their limited stability under cell culture conditions. J Agric Food Chem. 2012;60:8902–8910. | |
Zhong Y, Chiou YS, Pan MH, Shahidi F. Anti-inflammatory activity of lipophilic epigallocatechin gallate (EGCG) derivatives in LPS-stimulated murine macrophages. Food Chem. 2012;134:742–748. | |
Surh YJ, Chun KS, Cha HH, et al. Molecular mechanism underlying chemopreventive activities of anti-inflammatory phytochemicals: down-regulation of COX-2 and iNOS through suppression of NF-kappa B activation. Mutat Res. 2001;480–481:243–268. | |
Karlsen A, Paur I, Bøhn SK, et al. Bilberry juice modulates plasma concentration of NF-kappaB related inflammatory markers in subjects at increased risk of CVD. Eur J Nutr. 2010;49:345–355. | |
Goel V, Chang C, Slama J, et al. Echinacea stimulates macrophage function in the lung and spleen of normal rats. J Nutr Biochem. 2002;13:487. | |
Roesler J, Emmendörffer A, Steinmüller C, Luettig B, Wagner H, Lohmann-Matthes ML. Application of purified polysaccharides from cell cultures of the plant Echinacea purpurea to test subjects mediates activation of the phagocyte system. Int J Immunopharmacol. 1991;13:931–941. | |
See DM, Broumand N, Sahl L, Tilles JG. In vitro effects of echinacea and ginseng on natural killer and antibody-dependent cell cytotoxicity in healthy subjects and chronic fatigue syndrome or acquired immunodeficiency syndrome patients. Immunopharmacol. 1997;35:229–235. | |
Stimpel H, Proksch A, Wagner H, Lohmann-Matthes ML. Macrophage activation and induction of macrophage cytotoxicity by purified polysaccharide fractions from the plant Echinacea purpurea. Infect Immun. 1984;46:845–849. | |
Leuttig B, Steinmuller C, Gifford GE, Wagner H, Lohmann-Matthes ML. Macrophage activation by the polysaccharide arabinogalactan isolated from plant cell cultures of Echinacea purpurea. J Natl Cancer Inst. 1989;81:669–675. | |
Jung HJ, Park HJ, Kim RG, et al. In vivo anti-inflammatory and antinociceptive effects of liriodendrin isolated from the stem bark of Acanthopanax senticosus. Planta Med. 2003;69:610–616. | |
Tokiwa T, Yamazaki T, Sakurai S. Anti-inflammatory effect of eleuthreoside E from Acanthopanax senticosus. Foods and Food Ingredients Journal Japan. 2006;211:137–145. | |
Ricote M, Li AC, Willson TM, Kelly CJ, Glass CK. The peroxisome proliferator activated receptor-gamma is a negative regulator of macrophage activation. Nature. 1998;391:79–82. | |
Wang P, Anderson PO, Chen S, Paulsson KM, Sjögren HO, Li S. Inhibition of the transcription factors AP-1 and NF-kappaB in CD4 T cells by peroxisome proliferator-activated receptor gamma ligands. Int Immunopharmacol. 2001;1:803–812. | |
Christensen KB, Petersen RK, Petersen S, Kristiansen K, Christensen LP. Activation of PPARgamma by metabolites from the flowers of purple coneflower (Echinacea purpurea). J Nat Prod. 2009;72:933–937. | |
Spelman K, Iiams-Hauser K, Cech NB, Taylor EW, Smirnoff N, Wenner CA. Role for PPARgamma in IL-2 inhibition in T cells by Echinacea-derived undeca-2E-ene-8,10-diynoic acid isobutylamide. Int Immunopharmacol. 2009;9:1260–1264. | |
Xu X, Luo P, Wang Y, Cui Y, Miao L. Nuclear factor (erythroid-derived 2)-like 2 (NFE2L2) is a novel therapeutic target for diabetic complications. J Int Med Res. 2013;41:13–19. | |
Reuland DJ, Khademi S, Castle CJ, et al. Upregulation of phase II enzymes through phytochemical activation of Nrf2 protects cardiomyocytes against oxidant stress. Free Radic Biol Med. 2013;56:102–111. | |
Jia C, Shi H, Jin W, et al. Metabolism of echinacoside, a good antioxidant, in rats: isolation and identification of its biliary metabolites. Drug Metab Dispos. 2009;37:431–438. | |
Sgarbossa A, Dal Bosco M, Pressi G, Cuzzocrea S, Dal Toso R, Menegazzi M. Phenylpropanoid glycosides from plant cell cultures induce heme oxygenase 1 gene expression in a human keratinocyte cell line by affecting the balance of NRF2 and BACH1 transcription factors. Chem Biol Interact. 2012;199:87–95. |
 © 2014 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2014 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.