Back to Journals » OncoTargets and Therapy » Volume 13
DUB3 Facilitates Growth and Inhibits Apoptosis Through Enhancing Expression of EZH2 in Oral Squamous Cell Carcinoma
Authors Luo F, Zhou Z, Cai J, Du W
Received 10 September 2019
Accepted for publication 15 January 2020
Published 18 February 2020 Volume 2020:13 Pages 1447—1460
DOI https://doi.org/10.2147/OTT.S230577
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Carlos E Vigil
Fei Luo, Zunyan Zhou, Jun Cai, Wei Du
Department of Oncology, First People’s Hospital of Jinzhou, Jinzhou 434000, People’s Republic of China
Correspondence: Wei Du
Department of Oncology, First People’s Hospital of Jinzhou, Tumor Hospital, No. 40, Jinlong Road, Jinzhou, Hubei Province 434000, People’s Republic of China
Tel +86-18972161688
Email [email protected]
Background: Here, we probed the action mechanism of ubiquitin-specific processing proteases 17 (DUB3) in the evolution of oral squamous cell carcinoma (OSCC).
Methods: The expression of genes were calculated by qRT-PCR, and proteins were assessed by Western blot and immunohistochemistry. The cells viability and proliferation were checked by MTT and EdU assay, respectively. Flow cytometry was implemented to detect the cell cycle and apoptosis. The activity of EZH2 gene promoter was measured by luciferase reporter assay. Co-immunoprecipitation assay was used to ensure the ubiquitination of bromodomain-containing protein 4 (BRD4). The cell apoptosis of tumor tissues was assessed by TUNEL assay.
Results: DUB3 was overexpressed in OSCC tissues and cell lines, and negatively correlated with patient’s survival time. DUB3 downregulation could effectively curb OSCC cells viability and proliferation, promote cell apoptosis and the expression of cleaved-caspase-3, cleaved PARP and p21, while inhibit cyclin D1. Besides, DUB3 production was positivity correlated with enhancer of zeste homolog-2 (EZH2) and BRD4. BRD4 downregulation could repress DUB3-induced EZH2 production, and MG132 reversed DUB3 decreasing-mediated BRD4 downregulation. Downregulation of DUB3 promoted BRD4 ubiquitination. DUB3 promoted OSCC cells proliferation, while suppressing apoptosis via facilitating EZH2 production. At last, in vivo experiment indicated that the downregulation of DUB3 significantly inhibited the growth of xenograft tumor.
Conclusion: In summary, we found that DUB3 enhanced OSCC cells proliferation and xenograft tumor growth, while inhibited their apoptosis via promoting BRD4-mediated upregulation of EZH2. Our study indicated that DUB3 may be an effective anti-cancer target for OSCC therapy.
Keywords: oral squamous cell carcinoma, ubiquitin-specific processing proteases 17, enhancer of zeste homolog-2, bromodomain-containing protein 4
Introduction
Oral squamous cell carcinoma (OSCC) is the most universal and worse categorization of oral cancer, and its recurrent rate and metastatic remain high.1 The metastasis of OSCC cancer cells is associated with 90% cancer death.2 Tobacco use, radiation exposure, alcohol consumption, viral infections, betel quid chewing, and immunoincompetence are the risk factors associated with OSCC.3 Despite the advance in therapeutic strategies, such as chemotherapy and surgery, the prognosis of OSCC is still poor, and it has no significant improvement in the overall survival rate of OSCC patients.4,5 Hence, it is important to verify novel candidate molecules and more efficacious therapeutic approaches for OSCC.
Ubiquitin-proteasome system is an essential pathway for protein degradation, consists by abnormal and short-lived protein, and located in the nucleus and cytoplasm.6 The balance between ubiquitination and deubquitination is necessary for protein degradation.7 The ubiquitination process completed by a series of proteins containing Ub-activating (E1), Ub-conjugating (E2) and Ub-ligating (E3) enzymes, and the deubquitination process regulated by deubiquitinating enzymes (DUBs).8,9 DUB3, also named as ubiquitin-specific processing proteases 17 (USP17), belongs to DUB/USP family and has been announced to induce in response to cytokine stimulation.10 It has been demonstrated that DUB3 involves in regulating a series of cellular reactions including cell cycle progression, proliferation, and metastasis.11,12 Recently, high expressed DUB3 has been measured in non-small cell lung cancer, and it has been also indicated that DUB3 suppression could inhibit tumorigenesis and invasion of cancer cells13 Moreover, Wu et al indicated that DUB3 is over-expressed in breast cancer, and decreasing of DUB3 could suppress the migration, invasion, and metastasis of tumor cells via promoting Snail1 degradation.14 Increasing evidence suggested that DUB3 could contribute to tumor progression. However, the effects of DUB3 on OSCC progression are unknown.
Epigenetic modifications, such as histone modification, have been proved to affect almost every component of gene regulation. Histone acts as a crucial role in gene activation.15 Bromodomain-containing protein 4 (BRD4) belongs to the bromodomain and extra-terminal (BET) family, which regulates gene expression via targeting to the gene promoter.16 It has been indicated that BRD4, a nuclear protein, binds to acetylated lysine residues on histone via its bromodomain.17 In 1996, Denis et al firstly demonstrated that the expression and function of BET protein is associated with human cancer.18 Recently, several lines of evidence have suggested that BRD4 regulates tumor progression. For instance, BRD4 has been proved to regulate the dissemination of breast cancer via targeting the Jagged1/Notch1 signaling pathway.19 BRD4 could contribute to the progression of gastric cancer via regulating c-MYC in transcriptional and epigenetic.20 Furthermore, Hu et al proved that inhibitor of BRD4 could attenuate the growth and metastasis of colorectal cancer.21 These studies confirmed that BRD4 maybe play an oncogene.
Enhancer of zeste homolog-2 (EZH2), a histone methyltransferase, belongs to the polycomb repressor complex 2 catalytic subunit.22 It has been demonstrated that EZH2 could mediate gene silencing via regulating the posttranslational modifications of histone.23 EZH2 has been shown to be overexpressed in many cancer, and widely recognized to participate in the initiation and progression of cancer as an oncogene. For example, Xu et al demonstrated that EZH2 could advance the proliferation of gastric cancer cells through downregulation of p21, and decreasing of EZH2 could inhibit the growth of tumor.24 Moreover, curcumol has been indicated to inhibit the proliferation and promote the apoptosis of bladder cancer cell lines via suppression of the production of EZH2.25 Currently, EZH2 has emerged as a potential target of natural products with anti-cancer effect, such as curcumin and berberine.26 Furthermore, a recent study demonstrated that EZH2 could promote the metastasis of OSCC cell lines via facilitating the expression of ROS1.27
Here, we researched the expression level and the potential role of DUB3 in OSCC progression. Our study indicated that DUB3 furthered the proliferation, while suppressed the apoptosis of OSCC cells via accelerating the expression of EZH2 through DUB4. Moreover, we also proved that decreasing in DUB3 could inhibit the growth of tumor in the mouse model. Our data demonstrated that DUB3 may be a novel biomarker for OSCC therapy.
Materials and Methods
Patient Samples
Fifty cases of OSCC and matched peritumoral tissues were collected from patients accepted surgical resection at our hospital, and stored at −80°C until to follow experiments. No patients had accepted any surgery, and all examples were confirmed by experienced physician. Written informed consent from all patients were obtained. The study was approved by the Ethics Committee of the First People’s Hospital of Jinzhou (IRB015047). This study was conducted in accordance with the Declaration of Helsinki.
Cell Culture and Transfection
The human oral epithelial cell HOEC was purchased from BNCC (BeNa Culture Collection, China), and OSCC cell lines CAL27, H157, and HSC-2 were purchased from American Type Culture Collection (ATCC; Manassas, VA, USA). All cells were maintained in RPMI-1640 medium (Gibco, Waltham, MA, USA) contained with 10% fetal bovine serum (FBS; Life Technologies, Carlsbad, CA), 100 μg/mL streptomycin (Gibco, Carlsbad, CA), and 100 U/mL penicillin (Gibco, Carlsbad, CA). All cells were cultured in a humidified incubator with 5% CO2 at 37°C. shRNA-DUB3 (shDUB3), shRNA negative control (shNC), pcDNA3.1-DUB3, pcDNA3.1, shRNA-BRD4 (shBRD4), and pcDNA3.1-EZH2 were transfected into OSCC cell lines using the Lipofectamine 2000 (Invitrogen, USA) according to the direction.
qRT-PCR Assay
TRIzol reagent (Invitrogen, USA) was needed in separating total RNA from OSCC tissues and cells. Then, the MiRcute First-strand cDNA Synthesis Kit (Tiangen Biotech, Beijing, China) was used to reversely transcribe RNA into cDNA. Subsequently, PCR analysis was determined by an SYBR Premix Ex TaqTM Kit (Takara, Japan). All experiments were performed according to the instructions. GAPDH was considered as the internal control for gene expression. The primer sequences were as following, DUB3: F: 5′-CAGTGAATTCGTGGGAATGGAGGAC GACTCACTCTAC-3′ and R: 5′-AGTCATCGATCTGGCACACAAGCATAGCCCT C-3′. EZH2: F: 5′-CCCTGACCTCTGTCTTACTTGTGGA-3′ and R: 5′-ACGTCAG ATGGTGCCAGCAATA-3′. BRD4: F: 5′-GTGGGAGGAAAGAAACAGGGACA-3′ and R: 5′-AGGAGGAGGATTCGGCTGAGG-3′. GAPDH: F: 5′-TCAAGAAGGT GGTGAAGCAG-3′ and R: 5′-CGTCAAAGGTGGAGGAGTG-3′.
Western Blot Assay
Total protein was collected from OSCC tissues or cell lines using the RIPA lysis buffer (KeyGen, China) according to the instruction. The concentration of proteins was calculated using a BCA Protein Assay Kit (Beyotime, China). Protein (20 μg) was added into 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE), and then relocated into polyvinylidene fluoride (PVDF) membranes. The protein was blocked with 5% skim milk at room temperature followed by incubating with the primary antibodies against DUB3, cyclinD1, p21, cleaved caspase-3, caspase-3, cleaved PARP, un-cleaver PARP, EZH2, BRD4 and β-actin (Cell Signaling Technology, Boston, MA) overnight at 4°C. Then, the membranes were incubated with HRP-linked secondary antibodies (Cell Signaling Technology, Boston, MA) for 1 h at room temperature. Finally, an ECL Kit was needed for the protein bands visualizing, and the grayscale value was ensured using the Image J software (NIH, Bethesda, MD, USA). β-actin was considered as the internal control for proteins.
Immunohistochemistry Assay
The human or mouse OSCC tissues and control tissues were cut into slices, and fixed with ice code 4% paraformaldehyde for 30 mins. Then, the slices were incubated with 0.2% Triton X-100 for 6 mins, goat serum for 1 hr, primary antibodies, including Ki67, DUB3 and EZH2 overnight at 4°C, secondary antibodies for 1 hr at room temperature, and DAB reagent (Venata Medical Systems, Basal, Switzerland), successively. Finally, the Olympus microscope and Image J software were used to collect and analyze the images, respectively.
MTT Assay
MTT assay was completed to distinguish the viability of OSCC cells. 24, 48, 72, and 96 hrs later for transfection, MTT solution was added into 96-well microplate, and then the plate was placed in the dark for 4 hrs at 37°C. At last, the generated formation of optical density (OD) was calculated at 490 nm.
EdU Assay
OSCC cells proliferation was measured by EdU assay. Forty-eight hours later for transfection, the cells were maintained with EdU solution for 2 hrs in 96-well microplate. Then, treatment OSCC cells with 4% paraformaldehyde for 30 mins followed by staining with Cell-LightTM EdU Apollo®488 In vitro Imaging Kit (RioBio, China) according to the synopsis. At last, the images were collected using a microscope.
Flow Cytometry Assay
Flow cytometry analysis was accomplished to explore the cell cycle and apoptosis of OSCC cells. Forty-eight hours later for transfection, the cells were fixed with absolute ethanol overnight at 4°C, and then treated with RNase. Next, the cells were incubated with propidium iodide (PI, Sigma-Aldrich, St. Louis, MO, USA). At last, the cell cycle was measured by flow cytometry. Apoptosis of OSCC cells were examined employing an Annexin-VFITC/PI Apoptosis Detection Kit (Sigma-Aldrich, St. Louis, MO, USA) according to the manual.
Luciferase Reporter Assay
The activity of the EZH2 promoter was measured using luciferase reporter assay. Next, PCR reaction was needed for obtaining the sequence of EZH2 gene promoter, which was cloned into pGL3-basic plasmid. pGL3-basic-EZH2 promoter or pGL3-basic were transfected into OSCC cell lines treated with pcDNA3.1, pcDNA3.1-DUB3, shNC, shDUB3, or shBRD4. Twenty-four hour later, a Dual Luciferase Reporter Assay System Kit (Promega, Madison, WI) was used to test the luciferase activity for each group.
Co-Immunoprecipitation Assay
Ubiquitinated BRD4 was measured by co-immunoprecipitation assay. shNC, Flag-Ubiquitin (Ub) or shDUB3 were transfected into HSC-2 cells for 24 hrs; then, each group was treated with MG132 for 8 hrs. Next, the cell lysate was obtained, and incubated with primary antibodies (DUB3 and DRB4) overnight at 4°C followed by protein G magnetic beads for 5 hrs at 4°C. Subsequently, Western blot assay was accomplished to analyze the ubiquitination of BRD4.
In vivo Treatment
BALB/C nude mice (18–20 g) were obtained from Guangdong Medical Laboratory Animal Center (Foshan, China). Two cell lines HSC-2-shNC and HSC-2-shDUB3-2# were digested with 0.25% trypsin and adjusted to the concentration of 1.0×107, and then divided into 0.1 mL for each group. Subsequently, HSC-2-shNC or HSC-2-shDUB3 were injected into subcutaneously between the abdominal ribs following mixed with matrigel matrix (Beijing Xia Si Biotechnology Co., Ltd., Beijing, China). The tumor size and weight were measured for every 10 days. All animal experiments have been approved by the Animal Ethics Committee of First People’s Hospital of Jinzhou, and were performed in accordance with the guidelines of the National Institutes of Health for the Care and Use of Laboratory Animals.
TUNEL Assay
Apoptosis of OSCC tumor tissues were measured using a TUNEL kit (Roche, Welwyn Garden, UK) according to the manufacture’s introduction. In brief, the slices were subjected to deparaffinization followed by rehydration, and then incubated with TUNEL reaction solution for 2 h at 37°C. Next, the slices were maintained with 3% H2O2, and then incubated with decoloration, DAB, hematoxylin, and alcohol solution. Finally, microscope and Image J software were used to obtain and analyze the images, respectively.
Statistical Analysis
SPSS 17.0 software (SPSS, Inc.) was used to analyze the data. The comparing of two experimental groups was fulfilled using the Student’s t-test. All data were present as mean ± standard deviation (SD). Moreover, p < 0.01 and p < 0.05 were known to indicate a significant difference. All experiments were repeated at least three times.
Results
DUB3 Was Overexpressed in Both OSCC Tissues and Cell Lines
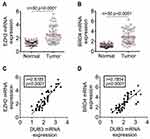
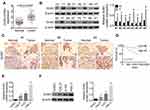
It has been demonstrated that DUB3 is overexpressed in many cancer, involving in non-small cell lung cancer.13 In this present study, the tumor and normal tissues were obtained from 50 confirmed OSCC patients. As displayed in Figure 1A, the production of DUB3 mRNA was markedly heightened in OSCC tumor (p < 0.0001). Meanwhile, Western blot also showed that DUB3 protein was markedly heightened in the tumor group compared with the normal group (Figure 1B). Furthermore, DUB3 production also was detected using immunohistochemistry analysis. Consisted with the result of Western blot, DUB3 was highly expressed in OSCC tumor (Figure 1C). The expression level of DUB3 was categorized as low or high according to the median level of DUB3 in the OSCC tumors. Then, we further explored the links between DUB3 and survival rate, and found that low expressed DUB3 was associated with long survival time of OSCC patients, and the difference was statistically significant (Figure 1D). Besides, we found that the production of DUB3 was not related to gender, age, T stage, differentiation, and bone invasion, while related to N stage, overall stage, perineural invasion, and tumor depth (Table 1). Next, we also detected the production of DUB3 in OSCC cell lines. The human oral epithelial cell HOEC and OSCC cell lines CAL27, H157, and HSC-2 were purchased and maintained with RPMI-1640. Increased DUB3 mRNA and protein were measured by qRT-PCR (Figure 1E) and Western blot (Figure 1F), respectively, in OSCC cell lines. Together, these data recommended that DUB3 was overexpressed in OSCC tumors and cell lines, and associated with OSCC patient’s survival time.
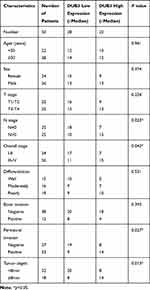 |
Table 1 The Clinicopathological Characteristics Related to the Expression of DUB3 in Samples of Oral Cavity Squamous Cell Carcinomas |
Downregulation of DUB3 Attenuated OSCC Cells Proliferation
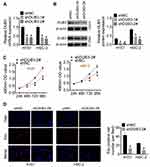
To examine the effects of DUB3 on the proliferation of OSCC cells, next experiments were done. The expression level of DUB3 in H157 and HSC-2 cells was significantly higher than in HOEC and CAL27 (Figure 1E and F); hence, H157 and HSC-2 cells were used to follow experiments. Two shRNA sequences were designed and used to downregulate the production of DUB3. shDUB3-1#, shDUB3-2#, and shNC were transfected into OSCC cells, respectively. Forty-eight hours later, the expression level of DUB3 was detected. Our results indicated that the production of DUB3 gene (Figure 2A) and protein (Figure 2B) were obviously downregulated by shDUB3-1# and shDUB3-2#, and the inhibitory effect of shDUB3-2# was higher than shDUB3-1# (p < 0.05). Thereby, shDUB3-2# was used to the next study. Next, the OD value was, respectively, measured at 24, 48, 72, and 96 hrs later for transfection of shNC or shDUB3-2#, and the MTT assay indicated that the cell viability was obviously suppressed by shDUB3-2# both in H157 and HSC-2 (Figure 2C). Besides, the proliferation of OSCC cell lines were detected by EdU assay. As shown in Figure 2D, the positive cell number of EdU was notably downregulated by shDUB3-2#. MTT and EdU analysis showed that decreasing in DUB3 using the shRNA could significantly attenuate OSCC cells viability and proliferation.
Downregulation of DUB3 Promoted OSCC Cells Apoptosis
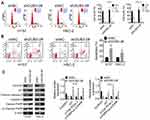
Next, the effects of DUB3 downregulation on OSCC cells apoptosis were explored. Treatment OSCC cell lines with shNC or shDUB3-2#, and 48 h later, the cell cycle and apoptosis rate of OSCC cells were examined. Flow cytometry experiment indicated that the cell number in G0/G1 stage was increased, while S and G2/M stages cell numbers were decreased in the shDUB3-2# group compared to the group of shNC (Figure 3A). Moreover, apoptosis of OSCC cells also was detected by flow cytometry, and the result proved that the downregulation of DUB3 using shDUB3-2# could induce OSCC cells apoptosis (Figure 3B). Besides, we further investigated the expression levels of cyclinD1, p21, cleaved caspase-3, caspase-3, un-cleaved PARP and cleaved PARP using Western blot assay. Our data indicated that the expression level of cell cycle-related protein cyclinD1 was downregulated, while p21 was upregulated, and the production of apoptosis-related proteins cleaved caspase-3 and cleaved PARP also were upregulated in the shDUB3-2# group compared to shNC group (Figure 3C). These data indicated that decreasing of DUB3 could markedly promote OSCC cells apoptosis.
DUB3 Promoted the Expression of EZH2 via Inhibiting BRD4 Degradation
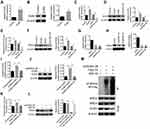
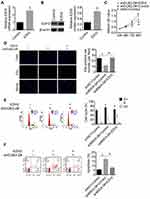
We further explored the action mechanism of DUB3 downregulation suppressed OSCC cells proliferation and promoted apoptosis. The total DUB3 gene sequence was cloned into pcDNA3.1 plasmid for constructing the overexpression system of DUB3. Empty plasmid (pcDNA3.1) was designed as the control of pcDNA3.1-DUB3. Here, we found that the expression level of DUB3 was significantly upregulated by pcDNA3.1-DUB3 treatment, as shown in the results of qRT-PCR (Figure 4A) and Western blot (Figure 4B). Then, to investigate the effects of DUB3 on its target protein EZH2, pcDNA3.1, pcDNA3.1-DUB3, shNC and shDUB3-2# were transfected into HSC-2 cells, respectively. qRT-PCR (Figure 4C) and Western blot (Figure 4D) indicated that increasing of DUB3 facilitated EZH2 expression, oppositely, decreasing of DUB3 repressed the expression of EZH2. Besides, overexpression of DUB3 could increase the activity of EZH2 gene promoter, and suppression of DUB3 led to the decrease of EZH2 gene promoter activity (Figure 4E). Furthermore, we also found that the upregulation of DUB3 promoted BRD4 production, and downregulation of DUB3 inhibited the expression of BRD4 (Figure 4F). Subsequently, two BRD4 shRNA sequences were designed and transfected into HSC-2 cells, shNC was served as the control of shBRD4. The inhibitory efficacy was measured, and the result of qRT-PCR proved that shBRD4-1# and shBRD4-2# could markedly downregulate the expression level of BRD4 (Figure 4G). Western blot assay also showed that expression of BRD4 was repressed by shBRD4-1# and shBRD4-2#, compared to the shNC group (Figure 4H). To ensure whether BRD4 mediates the effects of DUB3 on EZH2, following experiments were done. Treatment HSC-2 cells with only pcDNA3.1-DUB3 or together with shBRD4, and 48 hrs later, DUB3 production was detected. As shown in Figure 4I and J, the upregulation of DUB3 could notably promote the expression of EZH2, while decreasing of BRD4 could inhibit DUB3-mediated upregulation of EZH2. In addition, the downregulation of BRD4 using shBRD4-2# significantly repressed the increasing of EZH2 gene promoter activity induced by DUB3 was proved by luciferase reporter assay (Figure 4K). Subsequently, treatment HSC-2 cells with the proteasome inhibitor MG132 for 4 hrs following shNC or shDUB3 transfection for 24 hrs. Western blot indicated that MG132 treatment dose does not affect the expression of BRD4 compared to the control + shNC group, while significantly inhibited the decrease in DUB3-induced BRD4 suppression (Figure 4L). Downregulation of DUB3 could promote the degradation of BRD4 via the ubiquitination pathway. Next, Flag-Ub with or without shDUB3-2# was transfected into HSC-2 cells treated with MG132. Co-immunoprecipitation assay demonstrated that no difference in BRD4 expression between each group, and the ubiquitination level of BRD4 is most in Flag-Ub + shDUB3-2# group (Figure 4M). Downregulation of DUB3 could promote BRD3 ubiquitination. Overall, these data suggested that DUB3 could boost EZH2 production via inhibiting ubiquitin-mediated BRD4 degradation.
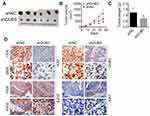
Moreover, we also detected the expressions of EZH2 and BRD4 in OSCC clinical samples and analyzed the correlation of DUB3 expression with EZH2 expression or BRD4 expression. Our data indicated that EZH2 (Figure 5A) and BRD4 (Figure 5B) were increased in OSCC tissues. Importantly, the expressions of EZH2 (Figure 5C) and BRD4 (Figure 5D) were positively associated with DUB3 expression in OSCC tissues.
DUB3 Promoted Proliferation and Inhibited Apoptosis of OSCC Cells via Facilitating EZH2 Production
We next examined whether EZH2 mediates the regulatory effects of DUB3 on HSC-2 cell proliferation and apoptosis. EZH2 overexpression plasmid (pcDNA3.1-EZH2) was constructed, and our results revealed that the expression level of EZH2 was notably heightened in pcDNA3.1-EZH2 (EZH2) group compared with control (empty plasmid) group (Figure 6A and B). Then, pcDNA3.1-EZH2 was transfected into HSC-2 cells with shNC or shDUB3-2#. MTT assay indicated that DUB3 decreasing could meaningfully attenuate OSCC cells viability, while increasing of EZH2 could inhibit the DUB3 decreasing-induced cell viability downregulation (Figure 6C). Besides, our data proved that the downregulation of DUB3 could significantly decrease EdU positive cell number, and overexpression of EZH2 could reverse the inhibitory effects of DUB3 downregulation on HSC-2 cells proliferation (Figure 6D). We also detected the cell cycle and apoptosis of HSC-2 using flow cytometry assay. Our results expounded that the downregulation of DUB3 markedly increased the G0/G1 stage cell number, while decreased the cell number in S and G2/M stage. However, the effect of DUB3 silencing on cell cycle was reversed by EZH2 (Figure 6E). Moreover, we also found that downregulation of DUB3 could significantly facilitate HSC-2 cell apoptosis; however, EZH2 overexpression could attenuate DUB3 downregulation-induced apoptosis of HSC-2 cells (Figure 6F). Above all, we can know that DUB3 could promote HSC-2 cell proliferation and suppress its apoptosis via upregulation of EZH2.
Decreasing of DUB3 Suppressed the Growth of Xenograft Tumors
Finally, we established a xenograft mouse model to probe the biological function of DUB3 downregulation in vivo. The tumor size was measured for every 10 days following HSC-2 cells transfected with shNC or shDUB3-2# injection. Our data demonstrated that the downregulation of DUB3 could significantly suppress the growth of tumor compared with the shNC group (Figure 7A and B). At 35 days later, the tumor was separated, and then weighted. As shown in Figure 7C, DUB3 downregulation could markedly decrease the tumor weight compared with shNC group. Subsequently, expression of Ki67, DUB3, and EZH2 were measured by immunohistochemistry, and the TUNEL assay was performed to detect the cell apoptosis. Our data have shown that the production of Ki67, DUB3, and EZH2 were downregulated, while the cell apoptosis in tumor tissues was promoted in the shDUB3-2# group compared to shNC group (Figure 7D). Together, these results proved that suppression of DUB3 could inhibit xenograft tumor growth.
Discussion
Ubiquitination, a crucial mechanism for regulating protein degradation and cell cycle progression, is a reversible post-translational modification. The balance between ubiquitination and deubiquitination is important for maintaining the cellular homeostasis.28 DUB3, a deubiquitination enzyme, has been proved to be demanded for cell cycle regulating and G1-S progression.29 In addition, several lines of evidence have implicated that DUB3 is overexpressed in cancer tissues and cell lines, such as glioma.30 However, how about the expression level of DUB3 in OSCC tissues and cells remain unclear. It has been revealed that the depletion of DUB3 in non-small cell lung cancer cell could induce cell apoptosis and attenuate cell proliferation.31 This research pointed out that DUB3 participates in the regulating of cell proliferation and apoptosis. Moreover, Burrows et al reported that DUB3 co-localizes in endoplasmic reticulum with RCE1 to attenuate the activity of RCE1, and modulates the processing and activation of Ras, and then regulates the cell proliferation.32 In this present study, our data revealed that DUB3 was upregulated in OSCC tissues and cells. Meanwhile, low expressed DUB3 was associated with long survival time of OSCC patients. Decreasing of DUB3 could suppress the proliferation and facilitate apoptosis of OSCC cell lines. Besides, we also found that the downregulation of DUB3 in the mouse model could inhibit the growth of xenograft tumors induced by HSC-2 cells.
It is a series of dynamic and plastic processes of transcript epigenetic regulation, which is coordinated by complicated machinery including methylation, acetylation, and ubiquitination. EZH2, a histone methyltransferase, is one of the essential catalytic enzymes for histone lysine 27 methylation and is associated with an aberrant transcript in the cancer cell.33 It has been demonstrated that EZH2 could facilitate the occurrence of tumor via targeting to regulate the expression of a series of tumor suppressor genes. Moreover, EZH2 has been reported to regulate numerous molecular processes associated with tumorigenesis including activation of NF-κB signaling and silencing of microRNA (miRNA).34 For example, Song et al proved that EZH2 could directly bind with miR-214 to repress its expression, and lncRNA LINC01535 could promote the growth, invasion, and migration of cervical cancer cells, and contribute to the progression of cervical cancer.35 EZH2 has been served as a transcription factor of EPHA7 and has been proved to suppress the transcription of EPHA2 via enhancing EPHA7 gene promoter methylation. Di et al reported that lncRNA SNHN14 could promote the metastasis of colorectal cancer by upregulation the expression of EZH3 though repressing the production of EPHA7.36 Furthermore, a recent research has reported that EZH2 is indispensable for maintaining epithelial cell barrier integrity under the condition of inflammatory, and decreasing the level of EZH2 could block the protective role of the NF-κB signaling pathway in cell survival.37 Increasing evidence has suggested that EZH2 is a potential target in cancer therapy. In this present study, we certified that DUB3 could regulate the proliferation and apoptosis of OSCC cell lines via boosting the production of EZH2.
It is demonstrated that BRD4 is an epigenetic and transcriptional regulator, and plays a pivotal role during cancer development. For instance, Shen et al informed that decreasing of BRD4 using its inhibitor SF2523 could restrain the progression of human prostate cancer.38 Tan et al revealed that BRD4 is increased in human prostate cancer cells. Decreasing of BRD4 could inhibit cancer cells proliferation, and lead to cell cycle arrest and apoptosis by facilitating FOXO1 production.39 Besides, it has been verified that BRD4 binds to acetylated lysine-310 to regulate the transcriptional activity of NF-κB in cancer cells, as well as contribute to cancer progression.40 Importantly, a recent study reported that DUB3 could bind to and promote BRD4 deubiquitination and stabilization, and protect prostate cancer cells against bromodomain and extra-terminal domain inhibitor via promoting BRD4 deubiquitination.41 The ubiquitin-proteasome system is a selective cellular mechanism of protein degradation and has been recognized that exists an interact with apoptosis. For instance, it has been demonstrated that Rhus coriaria could facilitate protein ubiquitination and proteasomal degradation, and then induce autophagy and apoptosis in colon cancer cells.42 Here, our data indicated that downregulation of DUB3 suppressed BRD4 production, and facilitated the ubiquitination of it. DUB3 could inhibit BRD4 degradation via deubiquitination. Moreover, Wu et al have verified that BRD4 is a key upstream regulator of EZH2 enhancer.43 In this present study, we found that DUB3 could upregulate EZH2 expression via suppression of the degradation of BRD4.
Conclusion
In summary, our data demonstrated that DUB3is highly expressed in OSCC, and DUB3 promoted the progression of OSCC via promoting BRD4-mediated EZH2 production. Our research extended the knowledge about the molecular mechanism underlying OSCC progression.
Data Sharing Statement
All data generated and/or analyzed during this study are included in this published article.
Author Contributions
All authors contributed to data analysis, drafting and revising the article, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Disclosure
The authors declare that they have no competing interests in this work.
References
1. Yang MD, Chang WS, Tsai CW, et al. Inhibitory effects of AVEMAR on proliferation and metastasis of oral cancer cells. Nutr. Cancer. 2016;68(3):473–480. doi:10.1080/01635581.2016.1153668
2. Pan L, Yang H, Xu C, et al. ZNF750 inhibited the malignant progression of oral squamous cell carcinoma by regulating tumor vascular microenvironment. Biomed Pharmacother. 2018;105:566. doi:10.1016/j.biopha.2018.06.001
3. Jemal A, Bray F, Center MM, et al. Global cancer statistics. CA Cancer J Clin. 2011;61(2):69–90. doi:10.3322/caac.v61:2
4. Sieviläinen M, Almahmoudi R, Al‐Samadi A, et al. The prognostic value of immune checkpoints in oral squamous cell carcinoma. Oral Dis. 2018;25(6):1435–45.
5. Warnakulasuriya S. Living with oral cancer: epidemiology with particular reference to prevalence and life-style changes that influence survival. Oral Oncol. 2010;46(6):407–410. doi:10.1016/j.oraloncology.2010.02.015
6. Komander D, Rape M. The ubiquitin code. Annu Rev Biochem. 2012;81:203–229. doi:10.1146/annurev-biochem-060310-170328
7. Chen X, Wu J, Chen Y, et al. Ubiquitin-specific protease 14 regulates cell proliferation and apoptosis in oral squamous cell carcinoma. Int J Biochem Cell Biol. 2016;79:350–359. doi:10.1016/j.biocel.2016.08.038
8. Maspero E, Polo S, In vitro ubiquitination: self-ubiquitination, chain formation, and substrate ubiquitination assays. 2016.
9. Liu F, Fu Q, Li Y, et al. USP21 modulates Goosecoid function through deubiquitination. Biosci Rep. 2019;39(7):BSR20182148. doi:10.1042/BSR20182148
10. Mj MG, Rascle A, Humbert M, et al. DUB-3, a cytokine-inducible deubiquitinating enzyme that blocks proliferation. J Bio Chem. 2004;279(14):13993–14000. doi:10.1074/jbc.M311291200
11. Hu B, Deng T, Ma H, et al. Deubiquitinase DUB3 regulates cell cycle progression via stabilizing cyclin A for proliferation of non-small cell lung cancer cells. Cells. 2019;8(4):297. doi:10.3390/cells8040297
12. Zhang Q, Zhang Z, Du H, et al. DUB3 deubiquitinates and stabilizes NRF2 in chemotherapy resistance of colorectal cancer. Cell Death Differ. 2019;26(11):2300–13.
13. Zhang S, Yuan J, Zheng R. Suppression of ubiquitin-specific peptidase 17 (USP17) inhibits tumorigenesis and invasion in non-small cell lung cancer cells. Oncol Res Featuring Preclinical Clin Cancer Ther. 2016;24(4):263–269. doi:10.3727/096504016X14666990347392
14. Wu Y, Wang Y, Lin Y, et al. Dub3 inhibition suppresses breast cancer invasion and metastasis by promoting Snail1 degradation. Nat Commun. 2017;8:14228. doi:10.1038/ncomms14228
15. Grunstein M. Histone acetylation in chromatin structure and transcription. Nature. 1997;389(6649):349–352. doi:10.1038/38664
16. Khoueiry P, Gahlawat AW, Petretich M, et al. BRD4 bimodal binding at promoters and drug-induced displacement at Pol II pause sites associates with I-BET sensitivity. Epigenetics Chromatin. 2019;12(1):39. doi:10.1186/s13072-019-0286-5
17. Chen R, Yik JH, Lew QJ, et al. Brd4 and HEXIM1: multiple roles in P-TEFb regulation and cancer. Biomed Res Int. 2014;2014.
18. Denis GV, Green MR. A novel, mitogen-activated nuclear kinase is related to a Drosophila developmental regulator. Genes Dev. 1996;10(3):261–271. doi:10.1101/gad.10.3.261
19. Andrieu G, Tran AH, Strissel KJ, et al. BRD4 regulates breast cancer dissemination through Jagged1/Notch1 signaling. Cancer Res. 2016;76(22):6555–6567. doi:10.1158/0008-5472.CAN-16-0559
20. Ba M, Long H, Yan Z, et al. BRD4 promotes gastric cancer progression through the transcriptional and epigenetic regulation of c‐MYC. J Cell Biochem. 2018;119(1):973–982. doi:10.1002/jcb.26264
21. Hu Y, Zhou J, Ye F, et al. BRD4 inhibitor inhibits colorectal cancer growth and metastasis. Int J Mol Sci. 2015;16(1):1928–1948. doi:10.3390/ijms16011928
22. Lou X, Zhu H, Ning L, et al. EZH2 regulates intestinal inflammation and necroptosis through the JNK signaling pathway in intestinal epithelial cells. Dig Dis Sci. 2019;64:1–10.
23. Madani GK, Rad A, Molavi M, et al. Predicting the correlation of EZH2 and cancer stem cell markers in esophageal squamous cell carcinoma. J Gastrointest Cancer. 2018;49(4):437–441. doi:10.1007/s12029-017-9985-y
24. Xu J, Wang Z, Lu W, et al. EZH2 promotes gastric cancer cells proliferation by repressing p21 expression. Pathol Res Pract. 2019;215:152374. doi:10.1016/j.prp.2019.03.003
25. Zhou L, Wei E, Zhou B, et al. Anti-proliferative benefit of curcumol on human bladder cancer cells via inactivating EZH2 effector. Biomed Pharmacother. 2018;104:798–805. doi:10.1016/j.biopha.2018.05.101
26. Shahabipour F, Caraglia M, Majeed M, et al. Naturally occurring anti-cancer agents targeting EZH2. Cancer Lett. 2017;400:325–335. doi:10.1016/j.canlet.2017.03.020
27. Shih C, Chang Y, Huang W, et al. EZH2-mediated upregulation of ROS1 oncogene promotes oral cancer metastasis. Oncogene. 2017;36(47):6542. doi:10.1038/onc.2017.262
28. Tan T, Zimmermann M, Reichert AS. Controlling quality and amount of mitochondria by mitophagy: insights into the role of ubiquitination and deubiquitination. Biol Chem. 2016;397(7):637–647. doi:10.1515/hsz-2016-0125
29. McFarlane C, Kelvin AA, de la Vega M, et al. The deubiquitinating enzyme USP17 is highly expressed in tumor biopsies, is cell cycle regulated, and is required for G1-S progression. Cancer Res. 2010;70(8):3329–3339. doi:10.1158/0008-5472.CAN-09-4152
30. Yildirim N, Kocal GC, Isik Z, et al. Ubiquitin–proteasome axis, especially ubiquitin-specific protease-17 (USP17) gene family, is a potential target for epithelial–mesenchymal transition in high-grade serous ovarian cancer. Reprod Sci. 2019;26(6):794–805. doi:10.1177/1933719118799189
31. Song C, Liu W, Li J. USP17 is upregulated in osteosarcoma and promotes cell proliferation, metastasis, and epithelial–mesenchymal transition through stabilizing SMAD4. Tumor Bio. 2017;39(7):1010428317717138. doi:10.1177/1010428317717138
32. Burrows JF, Kelvin AA, McFarlane C, et al. USP17 regulates Ras activation and cell proliferation by blocking RCE1 activity. J Bio Chem. 2009;284(14):9587–9595. doi:10.1074/jbc.M807216200
33. Wang J, Cheng P, Pavlyukov MS, et al. Targeting NEK2 attenuates glioblastoma growth and radioresistance by destabilizing histone methyltransferase EZH2. J Clin Invest. 2017;127(8):3075–3089. doi:10.1172/JCI89092
34. Yamagishi M, Uchimaru K. Targeting EZH2 in cancer therapy. Curr Opin Oncol. 2017;29(5):375–381. doi:10.1097/CCO.0000000000000390
35. Song H, Liu Y, Jin X, et al. Long non‐coding RNA LINC01535 promotes cervical cancer progression via targeting the miR‐214/EZH2 feedback loop. J Cell Mol Med. 2019;23:6098–6111. doi:10.1111/jcmm.v23.9
36. Di W, Weinan X, Xin L, et al. Long noncoding RNA SNHG14 facilitates colorectal cancer metastasis through targeting EZH2-regulated EPHA7. Cell Death Dis. 2019;10(7):514. doi:10.1038/s41419-019-1707-x
37. Liu Y, Peng J, Sun T, et al. Epithelial EZH2 serves as an epigenetic determinant in experimental colitis by inhibiting TNFα-mediated inflammation and apoptosis. Proc National Acad Sci. 2017;114(19):E3796–E3805. doi:10.1073/pnas.1700909114
38. Shen G, Jiang M, Pu J. Dual inhibition of BRD4 and PI3K by SF2523 suppresses human prostate cancer cell growth in vitro and in vivo. Biochem Biophys Res Commun. 2018;495(1):567–573. doi:10.1016/j.bbrc.2017.11.062
39. Tan Y, Wang L, Du Y, et al. Inhibition of BRD4 suppresses tumor growth in prostate cancer via the enhancement of FOXO1 expression. Int J Oncol. 2018;53(6):2503–2517. doi:10.3892/ijo.2018.4577
40. Zou Z, Huang B, Wu X, et al. Brd4 maintains constitutively active NF-κB in cancer cells by binding to acetylated RelA. Oncogene. 2014;33(18):2395. doi:10.1038/onc.2013.179
41. Jin X, Yan Y, Wang D, et al. DUB3 promotes BET inhibitor resistance and cancer progression by deubiquitinating BRD4. Mol Cell. 2018;71(4):592–605. e4. doi:10.1016/j.molcel.2018.06.036
42. Athamneh K, El Hasasna H, Al Samri H, et al. Rhus coriaria increases protein ubiquitination, proteasomal degradation and triggers non-canonical Beclin-1-independent autophagy and apoptotic cell death in colon cancer cells. Sci Rep. 2017;7(1):11633. doi:10.1038/s41598-017-11202-3
43. Wu X, Liu D, Tao D, et al. BRD4 regulates EZH2 transcription through upregulation of C-MYC and represents a novel therapeutic target in bladder cancer. Mol Cancer Ther. 2016;15(5):1029–1042. doi:10.1158/1535-7163.MCT-15-0750
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.