Back to Journals » Infection and Drug Resistance » Volume 13
Detection of Resistance to Fluoroquinolones and Second-Line Injectable Drugs Among Mycobacterium tuberculosis by a Reverse Dot Blot Hybridization Assay
Authors Li G, Guo Q, Liu H , Wan L, Jiang Y, Li M, Zhao LL, Zhao X, Liu Z, Wan K
Received 1 July 2020
Accepted for publication 7 October 2020
Published 11 November 2020 Volume 2020:13 Pages 4091—4104
DOI https://doi.org/10.2147/IDR.S270209
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Suresh Antony
Guilian Li,1,* Qian Guo,1,2,* Haican Liu,1 Li Wan,1 Yi Jiang,1 Machao Li,1 Li-li Zhao,1 Xiuqin Zhao,1 Zhiguang Liu,1 Kanglin Wan1
1State Key Laboratory for Infectious Disease Prevention and Control, Collaborative Innovation Center for Diagnosis and Treatment of Infectious Diseases, National Institute for Communicable Disease Control and Prevention, Chinese Center for Disease Control and Prevention, Beijing 102206, People’s Republic of China; 2Department of Molecular Biology, Shanghai Jiao Tong University Affiliated Sixth People’s Hospital, Shanghai 200233, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Kanglin Wan Email [email protected]
Background: Reliable and timely determination of second-line drug resistance is essential for early initiation effective anti-tubercular treatment among multi-drug resistant (MDR) patients and blocking the spread of MDR and extensively drug-resistant tuberculosis. Molecular methods have the potency to provide accurate and rapid drug susceptibility results. We aimed to establish and evaluate the accuracy of a reverse dot blot hybridization (RDBH) assay to simultaneously detect the resistance of fluoroquinolones (FQs), kanamycin (KN), amikacin (AMK), capreomycin (CPM) and second-line injectable drugs (SLIDs) in Mycobacterium tuberculosis.
Methods: We established and evaluated the accuracy of the RDBH assay by comparing to the phenotypic drug susceptibility testing (DST) and sequencing in 170 M. tuberculosis, of which 94 and 27 were respectively resistant to ofloxacin (OFX) and SLIDs.
Results: The results show that, compared to phenotypic DST, the sensitivity and specificity of the RDBH assay for resistance detection were 63.8% and 100.0% for OFX, 60.0% and 100.0% for KN, 61.5% and 98.1% for AMK, 50.0% and 99.3% for CPM, and 55.6% and 100% for SLIDs, respectively; compared to sequencing, the sensitivity and specificity of the RDBH assay were 95.2% and 100.0% for OFX, 93.8% and 100.0% for SLIDs or KN (both based on mutations in rrs 1400 region and eis promoter), and 91.6% and 100.0% for AMK or CPM (both based on mutations in rrs 1400 region), respectively. The turnaround time of the RDBH assay was 7 h for testing 42 samples.
Conclusion: Our data suggested that compared to sequencing, the RDBH assay could serve as a rapid and reliable method for testing the resistance of M. tuberculosis against OFX and SLIDs, enabling early administration of appropriate treatment regimens among MDR tuberculosis patients.
Keywords: Mycobacterium tuberculosis, drug resistance, reverse dot blot hybridization, fluoroquinolones, second-line injectable drugs, ofloxacin, kanamycin, amikacin, capreomycin
Introduction
The prevalence of multi-drug resistant tuberculosis (MDR-TB) and the emergence of extensively drug resistant tuberculosis (XDR-TB) are major obstacles for controlling tuberculosis disease. In 2019, the World Health Organization (WHO) estimated that approximately 484,000 MDR-TB/rifampicin-resistant tuberculosis (RR-TB) cases occurred globally, however, only 38.6% MDR/RR-TB were detected and notified in 2018.1 A National Baseline Survey on Drug-resistant Tuberculosis during 2007–2008 in China2 showed that approximately 10.2% of all notified TB patients were MDR-TB, approximately 8% of the diagnosed MDR-TB patients were XDR-TB and most cases of MDR and XDR tuberculosis resulted from primary transmission. MDR-TB isolates are resistant to the two most effective and economical drugs, isoniazid (INH) and rifampicin (RIF), whereas XDR-TB isolates are also resistant to any fluoroquinolones (FQs) and at least one of the second-line injectable drugs (SLIDs) capreomycin (CPM), kanamycin (KN) and amikacin (AMK). MDR-TB is difficult to treat and requires less effective SLIDs, which are associated with more side-effects.3 Misuse or mismanagement of second-line drugs may develope XDR-TB. Treatment of XDR-TB is more difficult than MDR-TB and the prognosis is much worse. Therefore, rapid, reliable and user-friendly molecular assays for diagnosing the resistance to second-line drugs are urgently required to ensure timely and adequate adjustments in MDR-TB treatment, minimize the transmission of MDR-TB and prevent the development of XDR-TB.
Knowledge on the mutation profiles of the second-line drugs will be helpful to define markers of drug resistance and establish a molecular diagnosis test. Prior data showed that 80–100% FQs resistance of Mycobacterium tuberculosis is attributed to mutations in the quinolone resistance-determining region (QRDR) of gyrA,4,5 and approximately 60% of the resistance to CPM, KN and AMK is associated with mutations in rrs 1400 region.4,6 Mutations in eis promoter are reported to account for nearly 10–30% KN resistance.4,6 The most prevalent mutation loci in these genes or regions are gyrA94, gyrA90, gyrA91, rrs1401, eis (−10), eis (−12) and eis (−14).4,6,7 These resistance-associated mutations provide the basis for molecular diagnostic approaches. In 2016, the WHO recommended that for patients with confirmed RR/MDR-TB, a second-line lineprobe assay (the commercial MTBDRsl assay, Hain Lifescience, Nehren, Germany) may be used as the initial test, instead of phenotypic culture-based drug susceptibility testing (DST), to detect resistance to FQs and SLIDs.8 In this study, we established an in-house reverse dot blot hybridization (RDBH) assay based on multiplex PCRs for simultaneously detecting FQs and SLID resistance among 170 clinical isolates from China. Also, we evaluated the accuracy of the RDBH assay comparing it to sequencing and phenotypic DST (proportion method).
Methods
Mycobacterial Strains
Among the strain bank collected from 2001–2011 and with well-documented drug susceptibility or resistance profiles in our laboratory, a total of 170 M. tuberculosis isolates were chosen and included in this study, composed of all of the ofloxacin (OFX) and (or) SLID resistant isolates and relevant number of control isolates. These isolates were obtained from 170 adult patients with pulmonary tuberculosis from institutes for tuberculosis control and cure as well as tuberculosis hospitals distributed in 15 provinces of China. The numbers isolated from each province were as follows: Anhui, 3; Beijing, 7; Fujian, 86; Gansu, 1; Guangxi, 5; Guizhou, 4; Henan, 5; Hunan, 3; Inner Mongolia, 8; Sichuan, 9; Shanghai, 6; Shanxi, 1; Tibet, 16; Xinjiang, 12; and Zhejiang, 4. No epidemiological analysis had been done among these isolates. H37Rv (ATCC 27294) was used as the reference strain.
Phenotypic Drug Susceptibility Testing
The isolate profiles of drug susceptibility were evaluated in our laboratory (National Tuberculosis reference laboratory) by the proportion method using Lowenstein-Jensen (L-J) slants with the following: INH, 0.2 µg/mL; RIF, 40 µg/mL; streptomycin, 4 µg/mL; ethambutol, 2 µg/mL; KN, 30 µg/mL; OFX, 2 µg/mL; CPM, 40 µg/mL;9 and AMK, 30 µg/mL.10 All the strains were stored in physiological saline containing 50% glycerol at −70 °C. Prior to characterizing the drug susceptibility, the strains were recovered on L-J medium for 4 weeks at 37 °C. H37Rv was used as the control with each batch of DST.
Genomic DNA Extraction
M. tuberculosis genomic DNA was extracted from fresh cultures growing on L-J slants. The bacterial cells were harvested and transferred to microcentrifuge tubes containing 400 μL TE buffer (10 mM Tris-HCl and 1 mM EDTA, pH 8.0), then inactivated in a 95 °C water bath for 10 min and incubated at 85 °C for 30 min. After centrifugation for 5 min at 12,000 rpm, the supernatant containing DNA was collected and stored at −20 °C for further use.
Multiplex PCRs
The RDBH assay was designed based on multiplex PCRs. Three PCR primer pairs (gyrA, rrs 1400 region and eis promoter), biotinylated at the 5ʹ end were designed to work together in a multiplex reaction. The sequences, amplicon sizes and concentrations of primers used in multiplex PCRs are shown in Table 1. Amplifications were performed in a final volume of 50 μL containing 2×Hot Start Taq (Sinobio, Shanghai, China), 10–100 ng of genomic DNA, 2 μL dimethylsulfoxide, 200 μM of each deoxynucleoside triphosphate and each forward/reverse primer with concentrations shown in Table 1. The cycling condition was as follows: 5 min at 94 °C, 20 cycles of 40 sec at 94 °C, 50 sec at 58 °C and 60 sec at 68 °C, 20 cycles of 40 sec at 94 °C, 50 sec at 58 °C and 60 sec at 72 °C, then 10 min at 72 °C. The PCR products were analyzed by electrophoresis using 3% agarose gels.
 |
Table 1 Sequences, Amplicon Sizes and Concentrations of Primers Used in Multiplex PCRs |
RDBH Assay
By targeting the major gene mutation sites conferring resistance to FQs, KN, AMK and CPM, 3 probes detecting wild type (WT) sequences and 10 probes recognizing mutant type (MT) sequences (Table 2) were newly designed by using the Software Primer Premier v.5.0. The sizes of probes were adjusted to maintain the difference of the melting temperatures within 6 °C so that they could be processed under the same hybridization and washing conditions. Two identical WT probes of gyrA were used on the membrane to add more certainty when reading the results. All the probes were covalently bonded to the negatively charged nylon membrane (Biodyne C, Pall Corporation, Ann Arbor, MI, USA). Thirty μL of each PCR product was diluted in 140 μL 5× saline‒sodium phosphate‒ethylenediaminetetraacetic acid (SSPE)/0.1% sodium dodecyl sulfate (SDS) buffer, heat-denatured at 100 °C for 10 min then immediately cooled on ice. The denatured single-stranded DNA was applied on the membrane in the miniblotter slots (Immunetics, Cambridge, MA, USA) and hybridized with probes at 60 °C for 1 h. The membrane was washed twice with 2× SSPE/0.5% SDS buffer at 50 °C and subsequently incubated at 42 °C for 40 min in 20 mL 2× SSPE/0.5% SDS containing 1:4000 diluted peroxidase. Then the unbound conjugate was removed by washing twice in 2× SSPE/0.5% SDS for 10 min at 42 °C, and rinsed once in 2× SSPE for 5 min at room temperature. Finally the membrane was developed by incubating with 1 mL of 3,3ʹ,5,5ʹ-Tetramethylbenzidine (Beyotime, Shanghai, China) reagent for 5 min in the dark.
 |
Table 2 List of Oligonucleotide Probes Used in This RDBH Assay |
A clear visible blue-green spot was recorded as positive. Clinical isolate was considered to be susceptible to the drug when the WT probes reacted positively while the MT probes were negative. When the MT probe had a stronger or equal color than the corresponding WT probe, the strain was considered to be a mutant genotype and therefore resistant to the drug. A strain without positive spots of WT and MT probes was recognized to have a specific mutation, and interpreted as resistant to the drug. H37Rv was used as positive control and water was employed as negative control. The assay was performed and read in a double-blind way, the results were compared to those obtained by proportion method and sequencing.
Sequencing
Mutations in the key fragment of 3 genes or regions (gyrA, rrs 1400 region and eis promoter) were also determined by sequencing. The primer sequences used for separate PCR were equal to that used in multiplex-PCRs. The amplifications were performed in a 0.2-mL tube contained 10 µL 2×PCR mix, 3 pmol each primer, 10–200 ng DNA, and RNase-free water to a final volume of 20 µL. The PCR conditions were as follows: 10 min at 94 °C, 35 cycles of 1 min at 94 °C, 1 min at 63 °C and 1 min at 72 °C, and 10 min at 72 °C. The PCR products of each gene/region were characterized by sequencing using the forward primers on an ABI Prism 3730 automated DNA sequencer (ABI Prism, Carlsbad, CA, USA). The resulting DNA sequences were analyzed using the basic local alignment search tool (http://www.ncbi.nih.gov/BLAST), and the specific mutations in protein sequences of the individual isolates were identified.
Statistical Analysis
A Pearson’s chi-square test was used to determine the sensitivity and specificity of the RDBH assay in comparison with phenotypic DST and sequencing. The consistency analysis on the results of the different methods was conducted by kappa identity test. Kappa values below 0.41, 0.41–0.75 and above 0.75 are considered as limited consistency, moderate consistency and excellent agreement, respectively.11 All statistical analyses were performed using SPSS 18.0 software (SPSS Inc., Chicago, IL, USA).
Results
Drug-Resistant Profiles
Phenotypic DST showed that among 170 M. tuberculosis, 94 (55.3%), 25 (14.7%), 13 (7.6%) and 20 (11.8%) were resistant to OFX, KN, AMK and CPM, respectively. In total, 27 were SLID resistant and 93 were MDR, of which 26 were XDR. A full susceptibility profile for all strains is shown in Table 3.
 |
Table 3 Drug Susceptibility Patterns of 170 Clinical M. tuberculosis Isolates |
Sequencing Results
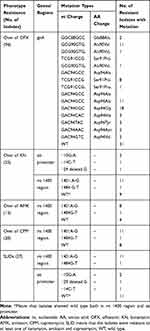
Sequencing for gyrA demonstrated that an AGC to ACC polymorphism at codon 95 was present in all 170 clinical isolates except for H37Rv. The mutation S95T in gyrA gene was known not to confer drug resistance and has been reported to be widely distributed in Beijing family or Euro-American lineage strains,5,6,12 and was excluded in the present study when mentioning mutations. The probes we set for the RDBH assay for FQs resistance detection did not cover this codon. Among 94 OFX resistant isolates, 63 (67.0%) carried mutations in the QRDR of gyrA at codons 88, 90, 91 and 94. The most common mutation was observed at codon 94 (42 isolates, 44.7%), where the codon Asp was replaced with Gly (18 isolates, 19.1%), Ala (13 isolates, 13.8%), His (3 isolates, 3.2%), Tyr (3 isolates, 3.2%), Asn (2, 2.1%) and Val (2 isolates, 2.1%), respectively (Table 4). The Asp94 mutations were found in two combinations, once with an Ala90Val mutation and once with a Ser91Pro mutation. Ala90Val was the next most predominant mutation (13 isolates, 13.8%), of which one combined with Ser91Pro and one with Asp94Ala. Other mutations within the gyrA included Ser91Pro (n = 10), and Gly88Ala (n = 2). The results are shown in Table 4. All of the OFX susceptible isolates were found to carry a wild type of gyrA.
Among 25 KN resistant isolates, 16 carried mutations in eis promoter or rrs 1400 region. Mutations in the eis promoter included (−10) G-A (n = 2), (−29) deleted G (n = 1) and (−14) C-T (n = 1), whilst mutations in rrs 1400 region included 1401 A-G (n = 11) and 1484 G-T (n = 1) (Table 4). None of the KN susceptible isolates were found carrying mutations in neither eis promoter nor rrs 1400 region.
It was reported that AMK and CPM resistance was mostly attributed to the mutations in rrs4,6 but not in the eis promoter, which was concordant with our results that none of the AMK or CPM resistant isolates carried mutations in this region (Table 5). In the present study, we only studied the relationship between AMK, CPM and mutations in rrs 1400 region, but not with that in eis promoter. Sequencing results showed that 9 out of 13 AMK resistant and 12 out of 20 CPM resistant isolates carried mutations in rrs 1400 region, the most frequent mutation site in AMK or CPM resistant isolates both was rrs 1401 A-G (n was 8 and 11, respectively) (Table 4). Three out of 157 AMK susceptible and one out of 150 CPM susceptible isolates carried mutation of rrs 1401 A-G.
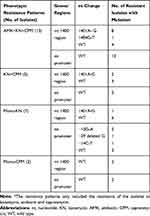 |
Table 5 Distributions of Mutations in rrs 1400 Region and eis Promoter by Sequencing in 27 Second-Line Injectable Drug Resistant M. tuberculosis Isolates According to the Resistance Patterns* |
In total, 16 out of 27 SLID resistant isolates (resistant to at least one of SLIDs KN, AMK or CPM) carried mutations in rrs 1400 region or eis promoter (Table 4).
Comparision of the RDBH Assay, Phenotypic DST and Sequencing
Our previous report had built a RDBH assay to simultaneously detect RIF, INH, ethambutol and streptomycin resistance.13 In the present study, a RDBH assay based on multiplex PCRs was built to detect FQs and SLIDs resistance and evaluated its performance by comparing to phenotypic DST and sequencing. The RDBH assay can test up to 42 M. tuberculosis DNA samples at a time, the turnaround time from the beginning of multiplex PCRs to provide resistance results was 7 h, as described in our previous report.13 The results of agarose gel electrophoresis show that each target fragment was successfully amplified. The RDBH results were determined by two readers according to the blot on the hybrid membrane. All of the interpretations on the susceptibility results of FQs and SLIDs of 170 isolates between two readers were identical. Figure 1 shows the hybridized image of the RDBH assay on 39 M. tuberculosis samples and Supplemental Table 1 shows the corresponding results of the RDBH assay, phenotypic DST and sequencing of each sample.
 |
Figure 1 The hybridized image detected with the reverse dot blot hybridization assay for ofloxacin and second-line injectable drug resistance conferring mutations. |
The sensitivity, specificity, positive predictive value (PPV), negative predictive value (NPV), concordance and Kappa values of the RDBH assay for detecting FQs and SLIDs resistance and XDR compared with phenotypic DST and sequencing are shown in Tables 6 and 7. The Kappa values and concordance between the RDBH assay and phenotypic DST varied from 0.61–0.72 and 80.0–95.3%, respectively, whilst between the RDBH assay and sequencing varied from 0.87–0.97 and 98.2–99.4%, respectively.
 |
Table 6 Sensitivity, Specificity, PPV, NPV and Agreement of the RDBH Assay Compared with Phenotypic DST Among M. tuberculosis |
 |
Table 7 Sensitivity, Specificity, PPV, NPV and Agreement of the RDBH Assay Compared with Sequencing Among M. tuberculosis Isolates |
Ofloxacin Resistance
In the present study, only one of the FQs was used to perform phenotypic DST among 170 isolates. However, mutations in gyrA were reported to be associated with OFX resistance, as well as levofloxacin and moxifloxacin resistance.14,15 So the performance of the RDBH assay based on mutations in gyrA can predict FQs resistance especially OFX resistance.
In two phenotypically OFX resistant isolates, both MT blots and WT blots were negative, and were interpreted as resistant. Additionally, 58 phenotypically OFX resistant isolates were correctly predicted as OFX resistant and the mutation patterns of each isolate identified by the RDBH assay were equal to that acquired by sequencing, among which three OFX resistant isolates carrying double mutations were all correctly identified by the RDBH assay. Two isolates carried mutations of gyrA 88 GGC-GCC (Gly-Ala) and one carried 94 (GAC-CAC) (Asp-His) acquired by sequencing which could not be identified by the RDBH assay. All of the phenotypically OFX susceptible isolates and all of the OFX resistant isolates with wild type of gyrA were identified as susceptible by the RDBH assay.
The Kappa value between the RDBH assay and phenotypic DST for OFX resistance detection was 0.61, indicating moderate agreement, whilst the Kappa value between the RDBH assay and sequencing was 0.96, indicating excellent agreement.
Kanamycin Resistance
In the present study, the RDBH assay predicted KN resistance based on mutations in the rrs 1400 region and eis promoter. Among 25 phenotypically KN resistant M. tuberculosis isolates, 14 displayed positive MT blots and negative WT blots and 1 only displayed a negative WT blot, so were interpreted as KN resistant. Of the 15 isolates, 14 showed the identical mutation patterns between the RDBH assay and sequencing, and 1 only displaying a negative WT blot was found carrying a mutation of eis (−29) deleted G by sequencing. None of the KN phenotypically susceptible isolates were misclassified as resistant by the RDBH assay. One simultaneously resistant to KN, AMK and CPM isolate carried a mutation of rrs 1484 G-T and could not be correctly identified by the RDBH assay. The Kappa value between the RDBH assay and phenotypic DST for KN resistance detection was 0.72, indicating moderate agreement, whilst the Kappa value between the RDBH assay and sequencing was 0.97, indicating excellent agreement.
Amikaxin/Capreomycin Resistance
In the present study, the probes designed for AMK and CPM resistance detection was aimed at mutations in the rrs 1400 region, which was identical to that used for KN resistance detection. Among 13 phenotypically AMK resistant and 20 phenotypically CPM resistant M. tuberculosis, eight and ten were correctly identified by the RDBH assay, respectively, whilst among 157 phenotypically AMK susceptible and 150 phenotypically CPM susceptible isolates, three and one were misclassified as AMK and CPM resistant, respectively. All of the AMK or CPM resistant isolates predicted by the RDBH assay showed specific mutation types, which were equal to that acquired by sequencing. One isolate carried a mutation of rrs 1484 G-T (as described in the Kanamycin Resistance section) and could not be identified by the RDBH assay.
SLID Resistance
For the SLID resistance determination, the results on detecting mutations in eis promoter and rrs 1400 region were combined to assess the performance of the RDBH assay. The isolates carrying mutations in rrs 1400 region or eis promoter acquired by sequencing were recognized as SLID genotypic resistant. Fifteen out of 27 SLID phenotypically resistant isolates and 15 out of 16 SLID genotypically resistant isolates were correctly predicted by the RDBH assay. All of the SLID phenotypically or genotypically susceptible isolates were correctly predicted by the RDBH assay (Tables 6 and 7).
Extensively Drug-Resistance
For the XDR determination, the results on detecting mutations in gyrA, eis promoter and rrs 1400 region were combined to assess the performance of the RDBH assay. The isolates carried mutations in gyrA, rrs 1400 region or eis promoter acquired by sequencing were recognized as genotypically XDR. Eleven out of 26 phenotypically XDR isolates and 11 out of 14 genotypically XDR isolates were correctly predicted by the RDBH assay. All of the phenotypically or genotypically non-XDR isolates were correctly predicted by the RDBH assay (Tables 6 and 7).
Discussion
An accurate and fast DST which can diagnose XDR from MDR is important to prevent the transmission of XDR and initiate proper treatments for MDR-TB patients. Most of the molecular and commercial diagnostic methods based upon identification of specific gene mutations associated with drug resistance were focused on INH and RIF.16–21 Only a commercial method, the MTBDRsl (Hain Lifescience, Nehren, Germany), was reported to detect the resistance of second-line antitubercular agents (FQs and SLIDs) simultaneously and has been used in many countries.22–26 It has been reported that patients with early available molecular DST results had a more rapid culture conversion and higher rate of treatment success.27,28 The RDBH assay we built took only 7 h from multiplex PCRs to provide results and allowed the simultaneous analysis of 42 clinical DNA samples for FQs and SLID susceptibility, which was comparable to that needed by the MTBDRsl v2.0 based on isolates and were obviously less than the time (6 weeks) needed for L-J slants based DST.29 The present study also provides detailed performance of the RDBH assay on predicting KN, AMK and CPM resistance and XDR by comparing to the phenotypic DST and sequencing.
A system review showed that globally 60.0–90.0% of M. tuberculosis isolates resistant to FQs presented mutations in codons between 88 to 94 in the QRDR of the gyrA gene,30,31 which was consistent with our results (67.0%). At the present study the most common mutation loci was at codon Asp94 which showed six substitutions (Gly, Ala, Tyr, Asn, His and Val), followed by Ala90Val, Ser91Pro and Gly88Ala, similar to the trends reported in previous studies from China4,6 and Korea.32 In previous studies from Brazil or Mozambique, the most frequent mutation in gyrA was Ala90Val.33,34 A study from Iran reported that 100% FQs resistant isolates carried mutations at codon 94 of gyrA.35 Previous studies show that mutations at codon 94 are related to the high level of in vitro phenotypic OFX resistance and clinical resistance,36,37 whilst mutations at codon 90 and 91 are related to the low level resistance of OFX and their implication with clinical outcomes is discussed.38,39 Specific substitutions Ala90Val, Asp94Ala, and Asp94Tyr are reported to be related with moxiflocaxin resistance.14 In 2019, moxifloxacin and levofloxacin instead of OFX were recommended for longer MDR-TB treatment.40 So it would be possible that the RDBH assay, which could provide specific mutation patterns, provides clues for clinicians to choose moxifloxacin in tuberculosis treatments.
Several molecular DST methods, ie, multiplex allele specific (MAS)-PCR,41 MTBDRsl Line probe assay (Hain Lifescience GmbH, Nehren, Germany),24 a high-throughput assay system based on allele-specific primer extension and MagPlex-TAG microspheres32 and whole-genome sequencing have been used in FQs42 resistance diagnosis. Mutations at codon 90 and 94 in gyrA were targeted by Liang et al, used in MAS-PCR, and the sensitivity and specificity of this method were 67.3% and 100%.41 MTBDRsl which targeted mutations in the QRDRs of gyrA and gyrB (not included in MTBDRsl version 1.0) were reported to have a sensitivity from 57–100% and specificity from 77–100% on detecting clinical isolates compared to phenotypic DST, whilst having a sensitivity from 33–100% and specificity from 83.6–100% on detecting clinical isolates compared to sequencing.22–26 In the present study, the RDBH assay used one WT probe and seven MT probes targeting mutations in codons 90-94 of gyrA and could detect 63.8% (60/94) of OFX phenotypically resistant isolates and 95.2% (60/63) gyrA mutated isolates. For the discrepancy caused by gyrA88, we did not design WT or MT probes targeting this codon in the RDBH assay, resulting in the misjudgment. There was still one out of three isolates carrying a mutation gyrA 94 (GAC-CAC) (Asp-His) which could not be correctly identified by the RDBH assay, one explanation may be that the strain was a mixed colony and colonies used for sequencing were different from the ones used for the RDBH assay. We also found that, compared with the control strain H37Rv, some strains showed weak signals in addition to the strong signals at some MT probes for gyrA 94 (Figure 1), we speculated that there were mixed infections and the weak signal was produced by the one with the concentration close to the detection limitation. To confirm the hypothesis, it is necessary to perform PCR with DNA from monoclonal strains. Sirgel et al reported finding two strains carried D94G plus D94N and they speculated there was a mixed population,39 which was similar to the present study though they did not meet the standard and were not reported as mutant.
Heteroresistant M. tuberculosis infections (defined as concomitant infection with drug-resistant and drug-susceptible strains) have been reported in some areas, eg China43,44 and Saudi Arabia.45 In the present study, one isolate had identical intensity of color both in the WT and MT probe (Figure 1, lane 31), and was reported as OFX resistant, the results of sequencing and phenotypic DST supported this judgement. However, we speculated that the strain was heteroresistant to OFX. Previous studies showed that 11.0% OFX resistant isolates and 10.9% AMK-resistant isolates were detected with heteroresistance.43,44 In phenotypic DST proportion method, a strain with 1% resistant clones was defined as resistant, however, Ng et al reported that four routine molecular DSTs including GenoscholarNTM+MDRTBII, GenoType MTBDRplus v2.0, XpertMTB/RIF, XpertMTB/RIF Ultra showed limit detection with more than 1% (ranged from 1% to 80%) for RIF heteroresistance.46 It is suggesting that a tuberculosis patient infected with heteroresistant strains is more likely to be misdiagnosed and get improper treatment according to the results of molecular DSTs, resulting in the emergence and spread of XDR-TB. Besides mixed infection and heteroresistance, the membrane probe assay was still challenged by its false negative and false positive results from technical defects that both lead to negative consequences.8 In the present study, we found that the primer concentrations in the multiplex PCRs, the nucleic acid at 5’ end and GC content of the probes, hybridization or elution temperatures and reagent proportions were the key factors for the RDBH assay.
Compared with the principle of GenoType MTBDRsl version 2.0, our RDBH assay did not contain probes targeted at mutations in QRDR (codons from 461-501) of gyrB because of their low frequencies of occurrence and combination with mutations in the QRDR of gyrA,6,47 but included two more MT probes to detect specific mutation forms in codon 94 of gyrA.17 More and more evidence showed that mutations in the QRDR of gyrB were associated with FQs resistance,4,6,35,47 and mutations at gyrA 88-89 accounting for 1–22% FQs resistance,4,6,48 so including the probes targeted at these mutations would be helpful to improve the sensitivity of the RDBH assay.
In the present study, among 27 SLID resistant isolates, the most frequent mutation detected was rrs A1401G (40.7%), which was consistent with the reports from China,4,6,49 Georgia America,50 eSwatini, Uganda and Somalia48 and Northwest Pakistan,51 while different from the report from India which showed that the most prevalent mutation was eis C (−12)T (46.1%).52 As for the SLID resistance, the RDBH assay showed 51.9% sensitivity and 100% specificity compared to the phenotypic DST. The inclusion of probes targeted at mutations in the eis promoter in the RDBH assay obtained an increased sensitivity with 11.1% than that based on probes targeted only in rrs 1400 region, which was comparable to the reported sensitivity differences between MTBDRsl v2.0, which added probes targeted at eis promoter, and MTBDRsl v1.0.24,25 In the present study, the RDBH assay showed an excellent sensitivity (87.5%) for SLID resistance detection in comparison to sequencing. One isolate simultaneously resistant to KN, AMK and CPM carried a mutation of rrs 1484 G-T which has been shown to be associated with KN, AMK and CPM resistance,53–55 the mutation was outside the targeted area and could not be correctly identified by the RDBH assay. Another SLID resistant isolate (resistant to KN but susceptible to AMK and CPM) was found to carry a mutation of eis (−29) deleted G which has never been reported, and was found by negative WT blot in the membrane and interpreted as resistant. However, the mutation was outside of the targeted detection area in the RDBH assay, we speculated that there may be a mixed infection or the changed spatial structure caused by the deletion in sequence affected the binding force with WT. In total, the RDBH assay showed an excellent agreement with sequencing (Kappa value = 0.97) for SLID resistance detection.
A report from Pakistan showed that only 30% AMK resistant isolates carried mutations in rrs 1400 region,53 which was apparently lower than that (69%) in the present study and other reports from China (50–86%).7,54 A previous report from China showed that AMK or CPM resistant isolates had lower mutation frequency in rrs 1400 region than KN resistant isolates,54 however, in the present study, AMK resistant isolates showed the highest mutation frequency in this region among three SLID resistant isolates, which was consistent with another report from China.7 A systematic review of 22 studies found that the A1401G mutation was present in 78% of AMK-resistant and in 76% of CPM-resistant isolates, but in only 56% of KN-resistant isolates,56 the trend of mutation prevalence was concordant with the present study which was 62%, 55% and 44% in AMK-resistant, CPM-resistant and KN-resistant isolates, respectively. We also found that mutations in the eis promoter were only found in four isolates susceptible to AMK and CPM, but resistant to KN, implying that mutations in the eis promoter do not always confer cross-resistance between KN and CPM or AMK, which was concordant with previous studies.57,58 However, a study showed that eis C (−14)T were also conferred with low level AMK resistance.59
In the present study, the total mutation frequency in rrs 1400 region and eis promoter of KN resistant strains was still lower than that in rrs 1400 region of AMK resistant isolates (64.0% vs 69.2%), but higher than that in rrs 1400 region of CPM resistant isolates (64.0% vs 60.0%) according to sequencing. Consequently, compared to the phenotypic DST, the sensitivity of the RDBH assay for predicting resistance of AMK was higher than KN and CPM (61.5%>60.0%>50.0%). The low sensitivity of the RDBH assay for CPM resistance detection strengthens the necessity to include more codons good for CPM resistance detection. It has been reported that tlyA is another gene confering CPM resistance.4,57,60 Li et al reported that mutations in this gene were found in 3 out of 4 CPM resistant but KN and AMK susceptible isolates.4 A system review showed that 1–3% CPM resistant isolates carried mutations in tlyA, but none of the CPM susceptible isolates carried them, suggesting that mutations in tlyA were potentially highly specific markers of CPM resistance.56 Miotto et al reported that mutations of N236K, pooled frameshifts and premature stop codons in tlyA were high-confidence mutations associated with CPM resistance.57 However, the gene tlyA was not sequenced and the specific codons or regions of this gene were not included in the RDBH assay in this study. The good news was that in the latest MDR-TB treatment guidelines released in 2019 by WHO, AMK instead of KN and CPM was recommended for use and listed as one of the group C medicines.40 The RDBH assay built in the present study provided a rapid and highly reliable method for AMK resistance detection compared to the phenotypic DST and sequencing, nevertheless, there is an urgency to explore the resistance mechanisms of SLIDs in M. tuberculosis to raise the diagnostic abilities of molecular DSTs.
In the present study, we combined the susceptibility results of OFX and SLIDs from the RDBH assay, and found that 42.3% phenotypically XDR isolates and 84.6% genotypically XDR isolates had been correctly identified. It has been reported that the sensitivity of MTBDRsl version 2.0 in detecting XDR among mycobacterial isolates was estimated ranging from 75% to 100%17,61 compared to culture-based DST. The ability of the RDBH assay for predicting XDR is directly affected by its ability for predicting FQs and SLIDs resistance. Consequently, the difference of mutation rates in varying regions will cause the sensitivity difference of the RDBH assay. The Guideline Development Group of WHO decided that the line probe assay can be used for the diagnosis of XDR-TB while acknowledging that the diagnostic accuracy is sub-optimal and felt that the assay could be used for surveillance of XDR-TB given statistical approaches to adjust for lower sensitivity and specificity during surveillance studies.16
The main limitation of this study was that the interpretation of the results may be affected by heterogeneous drug resistance, it would be better to use the vision equipment to get the blot intensity and set a cut-off value to diagnose resistance in the future, like the proportion method applies ≥1% as the limit of detection for resistance. While our study also needed to improve the signal intensity of some probes, eg, gyrA 90 GCG-GTG and eis promoter (−10) G-A, we speculated that the probes started with T or A were the major reason, second the low GC% of the probe meant the sequences on the membrane were more easily eluted, and then caused the weak signals. Third, the performance of the RDBH assay was only evaluated in cultures which may result in missed diagnoses of second-line drug resistant tuberculosis due to the decrease of minor mutated clones in culture-based strains, so performance in sputum samples should be specifically evaluated in the future.
Conclusions
The RDBH assay based on multiplex PCRs built in our study could determine FQs and SLID resistance of 1 to 42 M. tuberculosis samples within 7 h, and showed a high consistency to sequencing and a general consistency to the phenotypic DST method, suggesting that it could serve as a valid supplementary tool for simultaneously determining the resistance to FQs and SLIDs and emphasizing the urgency to explore the resistance mechanisms of FQs and SLIDs against M. tuberculosis.
Abbreviations
AMK, amikacin; CPM, capreomycin; DST, drug susceptibility testing; FQs, fluoroquinolones; INH, isoniazid; KN, kanamycin; L-J, Lowenstein-Jensen; MAS, multiplex allele specific; MDR-TB, multi-drug resistant tuberculosis; MT, mutant type; NPV, negative predictive value; OFX, ofloxacin; PPV, positive predictive value; QRDR, quinolone resistance-determining region; RDBH, reverse dot blot hybridization; RIF, rifampicin; RR-TB, rifampicin-resistant tuberculosis; SDS, sodium dodecyl sulfate; SLIDs, second-line injectable drugs; SSPE, saline‒sodium phosphate‒ethylenediaminetetraacetic acid; WHO, World Health Organization; WT, wild type; XDR-TB, extensively drug resistant tuberculosis.
Data Sharing Statement
The datasets used and/or analysed during the current study are available from the corresponding author on reasonable request.
Ethical Approval and Consent to Participate
The study obtained approval from the Ethics Committee of National Institute for Communicable Disease Control and Prevention, Chinese Center for Disease Control and Prevention. The patients with tuberculosis included in the present research were given a subject information sheet and they all gave written informed consent to participate in the study.
Acknowledgments
The authors thank all staff working in the tuberculosis hospitals or institutes for tuberculosis control of Anhui, Beijing, Fujian, Gansu, Guangxi, Guizhou, Henan, Hunan, Inner Mongolia, Sichuan, Shanghai, Shanxi, Tibet, Xinjiang and Zhejiang provinces for supplying the clinical M. tuberculosis isolates.
Funding
This study was supported by grants (Mega Projects of Research on the Prevention and Control of HIV/AIDS, Viral Hepatitis Infectious Diseases 2018ZX10103001-003-012 and 2013ZX10003002-001) from the Ministry of Science and Technology, China.
Disclosure
The authors declare that they have no competing interests.
References
1. World Health Organization. Global Tuberculosis Report 2019. Geneva, Switzerland: World Health Organization; 2019.
2. Zhao Y, Xu S, Wang L, et al. National survey of drug-resistant tuberculosis in China. N Engl J Med. 2012;366(23):2161–2170. doi:10.1056/NEJMoa1108789
3. Jain A, Dixit P. Multidrug-resistant to extensively drug resistant tuberculosis: what is next? J Biosci. 2008;33(4):605–616.
4. Li Q, Gao H, Zhang Z, et al. Mutation and transmission profiles of second-line drug resistance in clinical isolates of drug-resistant Mycobacterium tuberculosis from Hebei Province, China. Front Microbiol. 2019;10:1838. doi:10.3389/fmicb.2019.01838
5. Papaventsis D, Casali N, Kontsevaya I, Drobniewski F, Cirillo DM, Nikolayevskyy V. Whole genome sequencing of Mycobacterium tuberculosis for detection of drug resistance: a systematic review. Clin Microbiol Infect. 2017;23(2):61–68. doi:10.1016/j.cmi.2016.09.008
6. Zhao LL, Chen Y, Liu HC, et al. Molecular characterization of multidrug-resistant Mycobacterium tuberculosis isolates from China. Antimicrob Agents Chemother. 2014;58(4):1997–2005. doi:10.1128/AAC.01792-13
7. Zhang Z, Liu M, Wang Y, Pang Y, Kam KM, Zhao Y. Molecular and phenotypic characterization of multidrug-resistant Mycobacterium tuberculosis isolates resistant to kanamycin, amikacin, and capreomycin in China. Eur J Clin Microbiol Infect Dis. 2014;33(11):1959–1966. doi:10.1007/s10096-014-2144-5
8. World Health Organization.The use of molecular line probe assays for the detection of resistance to second-line anti-tuberculosis drugs, policy update. Geneva, Switzerland: World Health Organization; 2016, WHO/HTM/TB/2016.07.
9. World Health Organization. Policy guidance on drug-susceptibility testing (DST) of second-line antituberculosis drugs (WHO/HTM/TB/2008.392). Geneva: World Health Organization; 2008.
10. World Health Organization. Updated critical concentrations for first-line and second-line DST (as for May 2012); 2012.
11. Landis JR, Koch GG. The measurement of observer agreement for categorical data. Biometrics. 1977;33(1):159–174. doi:10.2307/2529310
12. An DD, Hong Duyen NT, Lan NTN, et al. Beijing genotype of Mycobacterium tuberculosis is significantly associated with high-level fluoroquinolone resistance in Vietnam. Antimicrob Agents Chemother. 2009;53(11):4835–4839. doi:10.1128/AAC.00541-09
13. Wan L, Guo Q, Wei JH, et al. Accuracy of a reverse dot blot hybridization assay for simultaneous detection of the resistance of four anti-tuberculosis drugs in Mycobacterium tuberculosis isolated from China. Infect Dis Poverty. 2020;9(1):38. doi:10.1186/s40249-020-00652-z
14. Willby M, Sikes RD, Malik S, Metchock B, Posey JE. Correlation between GyrA substitutions and ofloxacin, levofloxacin, and moxifloxacin cross-resistance in Mycobacterium tuberculosis . Antimicrob Agents Chemother. 2015;59(9):5427–5434. doi:10.1128/AAC.00662-15
15. Huo F, Zhang F, Xue Y, et al. Increased prevalence of levofloxacin-resistant Mycobacterium tuberculosis in China is associated with specific mutations within the gyrA gene. Int J Infect Dis. 2020;92:241–246. doi:10.1016/j.ijid.2020.01.021
16. Chen C, Kong W, Zhu L, et al. Evaluation of the GenoType((R)) MTBDRplus line probe assay on sputum-positive samples in routine settings in China. Int J Tuberc Lung Dis. 2014;18(9):1034–1039.
17. World Health Organization. The use of molecular line probe assays for the detection of resistance to isoniazid and rifampicin, policy update. Geneva, Switzerland: World Health Organization; WHO/HTM/TB/2016.12.
18. de Vos M, Derendinger B, Dolby T, et al. Diagnostic accuracy and utility of fluoroType MTBDR, a new molecular assay for multidrug-resistant tuberculosis. J Clin Microbiol. 2018;56(9). doi:10.1128/JCM.00531-18.
19. Makhado NA, Matabane E, Faccin M, et al. Outbreak of multidrug-resistant tuberculosis in South Africa undetected by WHO-endorsed commercial tests: an observational study. Lancet Infect Dis. 2018;18(12):1350–1359. doi:10.1016/S1473-3099(18)30496-1
20. Zar HJ, Workman LJ, Prins M, et al. Tuberculosis diagnosis in children using xpert ultra on different respiratory specimens. Am J Respir Crit Care Med. 2019;200(12):1531–1538. doi:10.1164/rccm.201904-0772OC
21. Boehme CC, Nabeta P, Hillemann D, et al. Rapid molecular detection of tuberculosis and rifampin resistance. N Engl J Med. 2010;363(11):1005–1015. doi:10.1056/NEJMoa0907847
22. Lacoma A, Garcia-Sierra N, Prat C, et al. GenoType MTBDRsl for molecular detection of second-line-drug and ethambutol resistance in Mycobacterium tuberculosis strains and clinical samples. J Clin Microbiol. 2012;50(1):30–36. doi:10.1128/JCM.05274-11
23. Kambli P, Ajbani K, Nikam C, et al. Determination of MICs of levofloxacin for Mycobacterium tuberculosis with gyrA mutations. Int J Tuberc Lung Dis. 2015;19(10):1227–1229. doi:10.5588/ijtld.14.0277
24. Tagliani E, Cabibbe AM, Miotto P, et al. Diagnostic performance of the New Version (v2.0) of GenoType MTBDRsl assay for detection of resistance to fluoroquinolones and second-line injectable drugs: a multicenter study. J Clin Microbiol. 2015;53(9):2961–2969. doi:10.1128/JCM.01257-15
25. Theron G, Peter J, Richardson M, Warren R, Dheda K, Steingart KR. GenoType((R)) MTBDRsl assay for resistance to second-line anti-tuberculosis drugs. Cochrane Database Syst Rev. 2016;9:CD010705.
26. Chandak RJ, Malhotra B, Bhargava S, Goel SK, Verma D, Tiwari J. Evaluation of MTBDRsl for detecting resistance in Mycobacterium tuberculosis to second-line drugs. Int J Tuberc Lung Dis. 2019;23(12):1257–1262. doi:10.5588/ijtld.18.0562
27. Shi W, Davies Forsman L, Hu Y, et al. Improved treatment outcome of multidrug-resistant tuberculosis with the use of a rapid molecular test to detect drug resistance in China. Int J Infect Dis. 2020;96:390–397. doi:10.1016/j.ijid.2020.04.049
28. Sun F, Li Y, Chen Y, et al. Introducing molecular testing of pyrazinamide susceptibility improves multidrug-resistant tuberculosis treatment outcomes: a prospective cohort study. Eur Respir J. 2019;53(3):1801770. doi:10.1183/13993003.01770-2018
29. World health organization. Technical manual for drug susceptibility testing of medicines used in the treatment of tuberculosis. 2018. Licence: CC BY-NC-SA 3.0 IGO.
30. Avalos E, Catanzaro D, Catanzaro A, et al. Frequency and geographic distribution of gyrA and gyrB mutations associated with fluoroquinolone resistance in clinical Mycobacterium tuberculosis isolates: a systematic review. PLoS One. 2015;10(3):e0120470. doi:10.1371/journal.pone.0120470
31. Malik S, Willby M, Sikes D, Tsodikov OV, Posey JE. New insights into fluoroquinolone resistance in Mycobacterium tuberculosis: functional genetic analysis of gyrA and gyrB mutations. PLoS One. 2012;7(6):e39754. doi:10.1371/journal.pone.0039754
32. Kim K, Yang JS, Choi HB, Lee SH. Detection of resistance to fluoroquinolones and injectable drugs among antituberculosis drugs by allele-specific primer extension on a microsphere-based platform. J Microbiol Methods. 2018;144:111–116. doi:10.1016/j.mimet.2017.11.007
33. Feliciano CS, Namburete EI, Rodrigues Placa J, et al. Accuracy of whole genome sequencing versus phenotypic (MGIT) and commercial molecular tests for detection of drug-resistant Mycobacterium tuberculosis isolated from patients in Brazil and Mozambique. Tuberculosis (Edinb). 2018;110:59–67. doi:10.1016/j.tube.2018.04.003
34. Namburete EI, Tivane I, Lisboa M, et al. Drug-resistant tuberculosis in Central Mozambique: the role of a rapid genotypic susceptibility testing. BMC Infect Dis. 2016;16:423. doi:10.1186/s12879-016-1766-x
35. Sakhaee F, Ghazanfari M, Ebrahimzadeh N, et al. A comparative study of phenotypic and genotypic first- and second-line drug resistance testing of Mycobacterium tuberculosis. Biologicals. 2017;49:33–38. doi:10.1016/j.biologicals.2017.07.003
36. Leung KL, Yip CW, Yeung YL, et al. Usefulness of resistant gene markers for predicting treatment outcome on second-line anti-tuberculosis drugs. J Appl Microbiol. 2010;109(6):2087–2094. doi:10.1111/j.1365-2672.2010.04840.x
37. Rigouts L, Coeck N, Gumusboga M, et al. Specific gyrA gene mutations predict poor treatment outcome in MDR-TB. J Antimicrob Chemother. 2016;71(2):314–323. doi:10.1093/jac/dkv360
38. Dominguez J, Boettger EC, Cirillo D, et al. Clinical implications of molecular drug resistance testing for Mycobacterium tuberculosis: a TBNET/RESIST-TB consensus statement. Int J Tuberc Lung Dis. 2016;20(1):24–42. doi:10.5588/ijtld.15.0221
39. Sirgel FA, Warren RM, Streicher EM, Victor TC, van Helden PD, Bottger EC. gyrA mutations and phenotypic susceptibility levels to ofloxacin and moxifloxacin in clinical isolates of Mycobacterium tuberculosis. J Antimicrob Chemother. 2012;67(5):1088–1093. doi:10.1093/jac/dks033
40. World health organization. WHO consolidated guidelines on drug-resistant tuberculosis treatment. Geneva: World Health Organization; 2019. Licence: CC BY-NC-SA 3.0 IGO.
41. Liang YP, Chen Y, Xiao TY, et al. Applied multiplex allele specific PCR to detect second-line drug resistance among multidrug-resistant tuberculosis in China. Tuberculosis (Edinb). 2017;107:1–4. doi:10.1016/j.tube.2017.07.010
42. Chen X, He G, Wang S, Lin S, Chen J, Zhang W. Evaluation of whole-genome sequence method to diagnose resistance of 13 anti-tuberculosis drugs and characterize resistance genes in clinical multi-drug resistance Mycobacterium tuberculosis isolates from China. Front Microbiol. 2019;10:1741. doi:10.3389/fmicb.2019.01741
43. Zhang X, Zhao B, Huang H, et al. Co-occurrence of amikacin-resistant and -susceptible Mycobacterium tuberculosis isolates in clinical samples from Beijing, China. J Antimicrob Chemother. 2013;68(7):1537–1542. doi:10.1093/jac/dkt082
44. Liu Q, Luo T, Li J, Mei J, Gao Q. Triplex real-time PCR melting curve analysis for detecting Mycobacterium tuberculosis mutations associated with resistance to second-line drugs in a single reaction. J Antimicrob Chemother. 2013;68(5):1097–1103. doi:10.1093/jac/dks509
45. Abakur EHA, Alnour TM, Abuduhier F, Albalawi FM, Alfifi KA. Emerging of heteroresistance Mycobacterium tuberculosis in Saudi Arabia. Infect Disord Drug Targets. 2019;19. doi:10.2174/1871526519666190326141550
46. Ng KCS, Supply P, Cobelens FGJ, et al. How well do routine molecular diagnostics detect rifampin heteroresistance in Mycobacterium tuberculosis? J Clin Microbiol. 2019;57(11). doi:10.1128/JCM.00717-19.
47. Pantel A, Petrella S, Veziris N, et al. Extending the definition of the GyrB quinolone resistance-determining region in Mycobacterium tuberculosis DNA gyrase for assessing fluoroquinolone resistance in M. tuberculosis. Antimicrob Agents Chemother. 2012;56(4):1990–1996. doi:10.1128/AAC.06272-11
48. Kateete DP, Kamulegeya R, Kigozi E, et al. Frequency and patterns of second-line resistance conferring mutations among MDR-TB isolates resistant to a second-line drug from eSwatini, Somalia and Uganda (2014–2016). BMC Pulm Med. 2019;19(1):124. doi:10.1186/s12890-019-0891-x
49. Zhang Z, Li T, Qu G, Pang Y, Zhao Y. In vitro synergistic activity of clofazimine and other antituberculous drugs against multidrug-resistant Mycobacterium tuberculosis isolates. Int J Antimicrob Agents. 2015;45(1):71–75. doi:10.1016/j.ijantimicag.2014.09.012
50. Campbell PJ, Morlock GP, Sikes RD, et al. Molecular detection of mutations associated with first- and second-line drug resistance compared with conventional drug susceptibility testing of Mycobacterium tuberculosis. Antimicrob Agents Chemother. 2011;55(5):2032–2041. doi:10.1128/AAC.01550-10
51. Jabbar A, Phelan JE, de Sessions PF, et al. Whole genome sequencing of drug resistant Mycobacterium tuberculosis isolates from a high burden tuberculosis region of North West Pakistan. Sci Rep. 2019;9(1):14996. doi:10.1038/s41598-019-51562-6
52. Ramakrishna V, Singh PK, Prakash S, Jain A. Second line injectable drug resistance and associated genetic mutations in newly diagnosed cases of multidrug-resistant tuberculosis. Microb Drug Resist. 2020;26(8):971–975. doi:10.1089/mdr.2019.0215
53. Ahmad K, Ahmad Z, Somayya R, Ali A, Rahat S. Analysis of rrs gene mutations in amikacin resistant clinical isolates of Mycobacterium tuberculosis from Khyber Pakhtunkhwa, Pakistan. Microb Pathog. 2017;108:66–70. doi:10.1016/j.micpath.2017.05.002
54. Du Q, Dai G, Long Q, et al. Mycobacterium tuberculosis rrs A1401G mutation correlates with high-level resistance to kanamycin, amikacin, and capreomycin in clinical isolates from mainland China. Diagn Microbiol Infect Dis. 2013;77(2):138–142. doi:10.1016/j.diagmicrobio.2013.06.031
55. Ogari CO, Nyamache AK, Nonoh J, Amukoye E. Prevalence and detection of drug resistant mutations in Mycobacterium tuberculosis among drug naive patients in Nairobi, Kenya. BMC Infect Dis. 2019;19(1):279. doi:10.1186/s12879-019-3911-9
56. Georghiou SB, Magana M, Garfein RS, Catanzaro DG, Catanzaro A, Rodwell TC. Evaluation of genetic mutations associated with Mycobacterium tuberculosis resistance to amikacin, kanamycin and capreomycin: a systematic review. PLoS One. 2012;7(3):e33275. doi:10.1371/journal.pone.0033275
57. Miotto P, Tessema B, Tagliani E, et al. A standardised method for interpreting the association between mutations and phenotypic drug resistance in Mycobacterium tuberculosis. Eur Respir J. 2017;50(6):1701354. doi:10.1183/13993003.01354-2017
58. Zaunbrecher MA, Sikes RD
59. Rodwell TC, Valafar F, Douglas J, et al. Predicting extensively drug-resistant Mycobacterium tuberculosis phenotypes with genetic mutations. J Clin Microbiol. 2014;52(3):781–789. doi:10.1128/JCM.02701-13
60. Maus CE, Plikaytis BB, Shinnick TM. Mutation of tlyA confers capreomycin resistance in Mycobacterium tuberculosis. Antimicrob Agents Chemother. 2005;49(2):571–577. doi:10.1128/AAC.49.2.571-577.2005
61. Rufai SB, Umay K, Singh PK, Singh S. Performance of Genotype MTBDRsl V2.0 over the Genotype MTBDRsl V1 for detection of second line drug resistance: an Indian perspective. PLoS One. 2020;15(3):e0229419. doi:10.1371/journal.pone.0229419
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.