Back to Archived Journals » Cell Health and Cytoskeleton » Volume 7
Current perspectives on mitochondrial inheritance in fungi
Received 23 April 2015
Accepted for publication 19 June 2015
Published 4 August 2015 Volume 2015:7 Pages 143—154
DOI https://doi.org/10.2147/CHC.S59508
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 4
Editor who approved publication: Professor Denis Wirtz
Jianping Xu,1,2 He Li2
1Department of Biology, McMaster University, Hamilton, Canada; 2The Key Laboratory for Non-Wood Forest Cultivation and Conservation of the Federal Ministry of Education, Central South University of Forestry and Technology, Changsha, People’s Republic of China
Abstract: The mitochondrion is an essential organelle of eukaryotes, generating the universal energy currency, adenosine triphosphate, through oxidative phosphorylation. However, aside from generation of adenosine triphosphate, mitochondria have also been found to impact a diversity of cellular functions and organ system health in humans and other eukaryotes. Thus, inheriting and maintaining functional mitochondria are essential for cell health. Due to the relative ease of conducting genetic and molecular biological experiments using fungi, they (especially the budding yeast Saccharomyces cerevisiae) have been used as model organisms for investigating the patterns of inheritance and intracellular dynamics of mitochondria and mitochondrial DNA. Indeed, the diversity of mitochondrial inheritance patterns in fungi has contributed to our broad understanding of the genetic, cellular, and molecular controls of mitochondrial inheritance and their evolutionary implications. In this review, we briefly summarize the patterns of mitochondrial inheritance in fungi, describe the genes and processes involved in controlling uniparental mitochondrial DNA inheritance in sexual crosses in basidiomycete yeasts, and provide an overview of the molecular and cellular processes governing mitochondrial inheritance during asexual budding in S. cerevisiae. Together, these studies reveal that complex regulatory networks and molecular processes are involved in ensuring the transmission of healthy mitochondria to the progeny.
Keywords: uniparental inheritance, biparental inheritance, mating type, actin cable, mitochore, mitochondrial partition
Introduction
Mitochondria are most commonly known as the powerhouse of eukaryotic cells, generating the universal energy currency, adenosine triphosphate (ATP), to support a diversity of cellular functions. It is believed that mitochondria were first described in 1856 by Swiss anatomist and physiologist Von Kölliker when he noted the arrangement of granules in the sarcoplasm of striated muscle tissues.1 These intracellular structures, later called sarcosomes by Retzius in 1890, were termed mitochondria by Benda in 1898. The term mitochondrion is a combination of the Greek words “mitos” (thread) and “chondros” (granule). Though much remains to be understood, over the past century, significant progress has been made with regard to the structure, function, origin, and inheritance of mitochondria. Indeed, among the diversity of intracellular organelles, the mitochondrion has arguably resulted in the most Nobel Prizes. For example, Warburg won a Nobel Prize in 1931 for revealing that mitochondria were associated with cellular respiration. Krebs was awarded the Nobel Prize in 1953 for localizing the enzymes in the citric acid cycle to mitochondria. Palade won the Nobel Prize in 1974 for developing a method to purify intact functional mitochondria. Mitchell, who won a Nobel Prize in 1978, developed the chemiosmotic theory for ATP synthesis that linked the electron transport chain, proton gradient formation, oxidative phosphorylation, and ATP biosynthesis. More recently, determining the composition and structures of the complexes involved in oxidative phosphorylation, in particular the ATP synthase complex, resulted in a Nobel Prize for Walker and Boyer in 1997.
While ATP generation is the function most commonly ascribed to mitochondria, mitochondria also provide a central platform for many other biological processes, including metabolite biosynthesis, ion homeostasis, and apoptosis. In recent years, mutations in mitochondrial DNA (mtDNA) and defects in mitochondria have been linked to a diversity of phenotypic traits including resistance to drugs,2 host defense,3 and virulence4 in fungal pathogens, nuclear genome stability and aging in the budding yeast Saccharomyces cerevisiae,5,6 and male sterility in plants.7 In humans, mitochondria have been linked to an increasing number of conditions, including diabetes, neurodegenerative disorders, cancers, and fertility.8–10
Unlike the majority of intracellular membrane structures, mitochondria contain their own genetic material. While most of the proteins in the mitochondria are encoded by the nuclear genome, the mtDNA in most eukaryotes encodes some of the key components of the translation machinery and the oxidative phosphorylation complexes. Thus, functional mtDNA is essential for ensuring functional mitochondria, and defects in mtDNA and/or in mtDNA inheritance can have severe detrimental consequences for cells. Indeed, mitochondrial genomes also have sophisticated interactions with the nuclear genome as the key protein complexes in mitochondria contain subunits encoded by both the nuclear and the mitochondrial genomes. Thus, understanding the mechanisms driving mtDNA transmission has far-reaching implications in many fields. Fungi, especially the model yeast S. cerevisiae, have served as excellent models for understanding the mechanisms of mitochondrial inheritance. Below we review our current understanding of mitochondrial inheritance in fungi. We first describe the general features of mtDNA inheritance in fungi, including the genes involved in controlling mtDNA inheritance. We then briefly review the molecular and cellular processes involved in mitochondrial inheritance in S. cerevisiae. We finish by describing the relevance of understanding fungal mtDNA inheritance to the health of humans and other animals.
Differences between mitochondrial and nuclear DNA inheritance
In typical plant and animal cells, the mitochondrial genes and genomes differ from their nuclear counterparts in several aspects.11–13 First, within each plant or animal cell, there is usually only one nucleus with one or a few copies of a haploid genome (depending on the ploidy of the organism). In contrast, each cell typically has multiple mitochondria, with each mitochondrion containing multiple mitochondrial genomes. Second, during each mitotic cell cycle, the nuclear genome is replicated exactly once, and each daughter cell receives half of the replicated nuclear DNA. In contrast, the replication and partitioning of the mitochondria and mitochondrial genomes are not as stringently controlled: some mitochondria and mitochondrial genomes replicate more often than others and they partition into daughter cells unevenly during cell division. Third, nuclear genes segregate, re-assort, and recombine predominantly during sexual reproduction and very rarely during somatic growth. In contrast, heterogeneous mitochondrial genomes within the cell often segregate quickly during mitotic division and somatic growth. Fourth, during sexual mating and meiosis, the inheritance of nuclear genes and genomes follow Mendelian laws, while mitochondrial genes and genomes do not obey these laws. In the great majority of sexual plants and animals, the mitochondrial genomes are inherited uniparentally, usually from the maternal parent.11–14
In many fungi, the structure and inheritance of both the nuclear and mitochondrial genomes differ slightly from those in plants and animals.12,13 For example, in many filamentous fungi, instead of having a single diploid nucleus per cell, each cell may contain two or multiple haploid nuclei. While the two nuclei in dikaryotic cells are typically synchronized in their replication and partitions,15 the nuclei in multinucleated cells may not be synchronized or stringently controlled.16 In addition, fungal mating typically involves vegetative cells that are morphologically indistinguishable from each other and are similar in size. This is different from the morphologically differentiated gametes in plants and animals where the male gametes are typically much smaller than female gametes.12,13 Furthermore, there is a diversity of sexual reproductive systems in fungi, from homothallism to heterothallism, from same-sex mating to bi-sex mating, and from bipolar to tetra-polar mating.17 The diversity of fungal mating systems also contributes to variation in mitochondrial inheritance patterns in fungi. For example, the basidiomycete yeast Cryptococcus neoformans shows uniparental mtDNA inheritance in opposite-sex mating but biparental mtDNA inheritance in same-sex matings.18,19 Indeed, compared with the relative uniformity of maternal mitochondrial inheritance in plants and animals, fungi exhibit a diversity of mtDNA inheritance patterns from strictly uniparental to biparental, a mixture of both uniparental and biparental, as well as recombinant mtDNA genotypes.12,13 Even within the same species, different strains, strain combinations, and environmental factors have also been found to influence mtDNA inheritance patterns.13,20,21
Due to the importance of mitochondria and their diverse patterns of inheritance, mitochondrial genetics have attracted significant attention from biologists in diverse fields such as evolutionary biology, genetics, cell biology, molecular biology, developmental biology, and medicine. In this review, our focus is on fungal mtDNA inheritance. The patterns of mtDNA inheritance among the diversity of fungi were summarized recently.12,13,22 Below, we briefly describe the main mtDNA inheritance patterns in sexual crosses in fungi.
Mitochondrial DNA inheritance in sexual crosses in fungi
As described above, while genetic difference between gametes is typically required for mating to occur in fungi, most often the gametes of fungal mating partners are morphologically indistinguishable. In both the ascomycete yeasts and the basidiomycetes (including both yeast and filamentous basidiomycetes), mating involves morphologically undifferentiated vegetative yeast cells or mycelia.12–15,22–24 In ascomycete yeasts, such as the model organisms S. cerevisiae and Schizosaccharomyces pombe, the zygotes inherit mtDNA from both parental cells. However, the parental mtDNA genotypes actively segregate during subsequent divisions through budding or fission.14 In basidiomycetes, sexual reproduction is most often achieved by the fusion of two haploid mycelia (in filamentous basidiomycetes) or two similar-sized yeast cells (in unicellular basidiomycetes). Interestingly, a diversity of mitochondrial inheritance patterns has been found, from uniparental to biparental in basidiomycetes.12,13 In basidiomycetes with uniparental mtDNA inheritance in sexual crosses, the mating type locus often plays an important role (for details of the known genes and regulatory mechanisms, see below).
Unlike the gametes in ascomycete yeasts and basidiomycetes, which lack morphological differentiation between the gamete types, the gametes of filamentous ascomycetes are morphologically differentiated but the differentiation is not associated with mating type.23 Instead, each mating type of a filamentous ascomycete can produce two types of gametes: the small dispersing “male” gametes called microconidia and the large, more complex and sessile “female” structures called ascogonia.23 In sexual crosses, mitochondria are inherited predominantly from the female-like structure, analogous to anisogamous mating and maternal mtDNA inheritance in plants and animals.13,23,25 However, mitochondrial plasmids in filamentous ascomycetes may show paternal inheritance, unlike the maternal inheritance of mitochondrial genomic DNA.13,26 Thus, in ascomycetes, there seems to be no stringent genetic mechanism controlling mtDNA inheritance during sexual mating. Below we briefly describe the genes and regulatory mechanisms that control uniparental mtDNA inheritance in basidiomycetes.
Genes and regulations of uniparental mtDNA inheritance in basidiomycetes
In filamentous basidiomycetes, mating involves genetically different but morphologically undifferentiated vegetative cells. Typically, genetic compatibility between the mating partners is not a prerequisite for the initial mycelial anastomosis and cell fusion.27,28 However, whether fertile mycelia can be established is determined after the initial cell fusion. In many filamentous basidiomycetes, such as the model species Schizophyllum commune and Coprinopsis cinerea, if their mating type alleles are different and genetically compatible, the haploid nuclei of the two mating partners within each fused cell replicate and spread throughout the cytoplasm of the reciprocal mating partner to establish a diploid or dikaryotic colony.29–31 However, the mitochondria do not migrate. The net result of this type of mating is that the mated mycelia are genetically identical with regard to their nuclear genome but their mitochondrial genomes may differ depending on the location of the mycelia within the colony from which samples are taken for analyses.29–31
While this general pattern holds true for many filamentous basidiomycete species, there are exceptions, and most of these exceptions can be explained by differences in the migration abilities of specific nuclei. Indeed, the nuclear migrating abilities seem to differ among species and sometimes even among strains within species.30–32 For example, there is no evidence of nuclear migration in mating between homokaryons in the button mushroom Agaricus bisporus.33 In a closely related species, the field mushroom Agaricus bitorquis, some homokaryotic strains show nuclear migration abilities while others do not, with mating partners having an important effect on the nuclear migration abilities of their mates.34 In the model species C. cinerea, five of 14 crosses examined showed unidirectional nuclear migration while the remaining nine showed bidirectional nuclear migration, with the directionality found to be strain-pair specific.30 In these cases, only the homokaryotic strains that are capable of receiving nuclei from mating partners become fertile and have their mitochondrial genomes inherited in sexual progeny. Interestingly, in these species, nuclear migration is controlled by the mating type locus,27 consistent with the mating type gene playing a significant role in mitochondrial inheritance in these species. At present, the molecular and cellular processes of how mating type genes control nuclear migration remain unknown.
The role of mating type locus in mtDNA inheritance has been very prominently demonstrated in several basidiomycete yeasts. Similar to mating in ascomycete yeasts, mating in basidiomycete yeasts involves genetically different but morphologically indistinguishable yeast cells. However, unlike biparental mitochondrial inheritance in the ascomycete yeasts S. cerevisiae and S. pombe, uniparental mitochondrial inheritance is very common in basidiomycete yeasts. In addition, there is increasing evidence that mating type genes play an important role in determining mtDNA inheritance in basidiomycete yeasts. For example, in the human yeast fungal pathogen C. neoformans, mtDNA is inherited almost exclusively from the MATa parent.24,35 Similarly, mtDNA inheritance in Ustilago maydis, Microbotryum violaceum (syn. Ustilago violacea), and Cryptococcus amylolentus are predominantly uniparental.36–38 However, variations in mtDNA inheritance patterns do exist among strains within a species and between closely related species. For example, in Cryptococcus gattii, a close relative of C. neoformans, both uniparental and biparental mtDNA inheritance have been found and the inheritance patterns were often strain/strain-pair specific, with environmental conditions also playing a role in some of the crosses.20 In the next section, we review the specific genes that have been identified in controlling uniparental mtDNA inheritance in basidiomycete yeasts.
Genetic factors controlling uniparental mtDNA inheritance in basidiomycete yeasts
Several recent studies identified the candidate genes involved in controlling uniparental mtDNA inheritance in two basidiomycete yeasts, C. neoformans and U. maydis. In C. neoformans, a homeodomain transcription factor called sex-identity (sxi) gene within the mating type locus (sxi1α in MATα and sxi2a in MATa) was found to control uniparental mtDNA inheritance.18,19 In a typical cross in this species, almost all progeny inherit mtDNA from the MATa parent. However, when the sex-determining gene in either or both mating type backgrounds was disrupted, mitochondrial inheritance became biparental and recombinant mtDNA genotypes were frequently recovered.18,19 Aside from the “sex”-determining genes, other independent genetic factors and environmental factors have also been shown to influence mtDNA inheritance in C. neoformans. For example, a key prezygotic transcription factor, Mat2p, which governs the pheromone sensing and response pathway, plays a critical role in mtDNA inheritance and its role is independent of the Sxi1α/Sxi2a complex.39 In addition, both ploidy and environmental stresses such as high temperature and ultraviolet irradiation could lead to the inheritance of MATα parental mtDNA as well as recombinant mtDNA genotypes.21,40 At present, the downstream targets of the Sxi1α/Sxi2a complex and the molecular processes by which they and other factors control mtDNA inheritance in C. neoformans are still unknown.
Similar to ascomycete yeasts such as S. cerevisiae and S. pombe, both C. neoformans and C. gattii have a bipolar mating system, with mating compatibility controlled by a single locus with two alternative alleles. However, the mating type loci in C. neoformans and C. gattii are much larger than those in ascomycetes.17 In the model smut fungus U. maydis, two unlinked loci, called the a and b loci, are involved in controlling mating compatibility, and different alleles at both loci are needed in order to establish a fertile diploid zygote. In this tetrapolar species, the a locus has two alleles (a1 and a2) while the b locus has multiple alleles.41 In typical crosses, mtDNA is inherited from the a2 parent. Fedler et al identified that uniparental mtDNA inheritance in U. maydis was controlled by the a2-specific genes lga2 and rga2.38 The absence of a functional lga2 in U. maydis strongly promoted biparental mtDNA inheritance and mtDNA recombination. In contrast, disruption of rga2 led to the inheritance of mtDNA from the a1 parent. The results suggest that Rga2p likely functions to protect mtDNA from Lga2-mediated elimination. The inheritance of the a1 mitochondrial genotype in the absence of rga2 also suggests that an rga2-independent pathway likely exists to suppress the selective a2-controlled mtDNA inheritance. Indeed, a constitutively expressed Mrb1p protein located in the mitochondrial matrix was found to antagonize the Lga2p-mediated loss of mtDNA, likely through indirect interaction with Lga2p.42 In both C. neoformans and U. maydis, there is increasing evidence for selective degradation of mtDNA from one parent at an early stage of sexual mating.35,43 However, the genes and molecular processes that mark the mitochondria from the different parents for inheritance or targeted elimination, either before or during mating, remain to be determined.
Molecular and cellular processes govern mitochondrial inheritance in S. cerevisiae
While the inheritance of mtDNA in sexual crosses has attracted attention from geneticists, molecular and cell biologists have also made significant headway in recent years in understanding the molecular and cellular processes involved in mitochondrial inheritance during mitotic division. During the life span of a somatic/vegetative cell, there can be significant metabolic activities, driven largely by energy production through oxidative phosphorylation in the mitochondria. Within the mitochondria, oxidative phosphorylation generates reactive oxygen species (ROS) that can damage the mitochondria, including mtDNA, mitochondrial proteins, and mitochondrial membranes. Some of this damage can be repaired but severely damaged, and nonfunctional mitochondria will be disposed of. In addition, during asymmetrical cell divisions such as budding, the damaged mitochondria can be selectively excluded from new daughter cells.
In the following sections, we review the genes and molecular processes that ensure the inheritance of functional mitochondria during budding in S. cerevisiae. We begin by first describing the yeast cell cycle.
Cell cycle in S. cerevisiae
The cell cycle is defined as the complete process of DNA replication, mitosis, and cytokinesis that leads to the production of two daughter cells from a single mother cell. A typical cell cycle includes four phases: G1 (gap 1), S (DNA synthesis/replication), G2 (gap 2), and M (mitosis). All cell types undergo some version of this basic cycle, although details of regulation and the size of the gap phases (G1 and G2) can differ significantly among species. Unlike the majority of cell divisions in higher eukaryotes where a mother cell typically undergoes mitosis to produce two similarly sized and genetically identical daughter cells, S. cerevisiae produces progeny cells through budding, generating an asymmetric mother-daughter cell relationship. Specifically, after initiation, the daughter bud grows continuously through the cell cycle and the nucleus migrates into the bud neck where it undergoes mitosis. This distinctive cellular architecture in S. cerevisiae requires an early duplication of the spindle pole bodies and a reorganization of the interphase cytoplasmic microtubules into a mitotic nuclear spindle, permitting bud formation and nuclear migration to the bud neck. An additional feature of the asymmetrical cell division in S. cerevisiae is the lack of clearly defined S, G2, and M phases. A variety of cell cycle control genes have been identified that impact different phases and transitions in the cell cycle.44 Below we focus on those that impact mitochondrial inheritance.
mtDNA replication and organization in S. cerevisiae
In a typical S. cerevisiae cell, there may be 1–10 mitochondria, forming a dynamic subcortical reticulum that may fuse and split continuously within the cell. Within each mitochondrion, there are typically multiple copies of the mitochondrial genome, with a total of up to approximately 200 copies in a single cell.45 Most of the mtDNA molecules are present as linear molecules of variable lengths with a few circular ones.45 Three models have been proposed for mtDNA replication in S. cerevisiae, ie, RNA-primed replication, rolling-circle replication, and recombination-dependent replication.45–48 These models are not mutually exclusive: each has experimental support and could contribute to mtDNA replication in yeast.45–48 Both rolling-circle replication and recombination-dependent replication require double-stranded breaks. Recombination-dependent replication followed by crossing over can generate recombinant mtDNA genotypes within the mitochondria. Table 1 summarizes the genes known to be involved in mtDNA maintenance and inheritance in S. cerevisiae. In both the rolling-circle and recombination-dependent models, two proteins, ie, Ntg1p and Mhr1p, play essential roles to ensure mtDNA replication to generate mtDNA concatemers. The newly replicated concatemers are selectively transmitted to daughter cells and are then processed into circular, unit-sized mtDNA.45–48
Aside from mtDNA molecules, many mitochondrial proteins are co-transmitted to daughter cells during cell replication. Indeed, within mitochondria, mtDNA interacts with mitochondrial proteins to form mt-nucleoids that act as units of inheritance. Each mt-nucleoid in S. cerevisiae typically contains three to four copies of mtDNA and approximately 30 proteins.49 The major mtDNA-binding protein in S. cerevisiae is the high-mobility group-like non-histone protein Abf2p, which bends the DNA backbone with a preference for GC-rich gene sequences, packing the double-stranded DNA molecules into 190 nm structures.50 The mt-nucleoid also includes other proteins such as the heat shock protein 60 (Hsp60p), which plays an essential role in the folding of mitochondrial proteins as well as folding and stabilizing the protein-single strand DNA complexes.51 Within each mitochondrion, the positions of mt-nucleoids are relatively stable and do not diffuse randomly. This behavior has been demonstrated by the mtDNA inheritance in sexual crosses where parental mitochondria from the two parents fuse but the mt-nucleoids do not completely mix, resulting in the position of buds playing a significant role in daughter cell mtDNA genotypes recovered from the mated zygotes.11 However, recombinant mtDNA molecules can also be recovered in progeny, consistent with some levels of mt-nucleoid mixing between parental mitochondria during sexual mating.11
During yeast budding, daughter cells receive mitochondria and mitochondrial genomes from mother cells prior to cytokinesis. Time-lapse microscopic analyses of mitochondria revealed that mitochondrial inheritance is accomplished through the following general processes.48,52 In the G1 phase prior to nuclear DNA replication, mitochondria polarize toward the site of bud emergence. During the S phase when the nuclear DNA are replicating, some of the polarized mitochondria move toward the developing bud (called anterograde movement) while others move away from the bud to the opposite end of the mother cell (called retrograde movement). During the G2 phase, the mitochondria are transiently immobilized at the opposite ends of cell division. Finally, during mitosis, the mitochondria are released and redistributed to the bud and mother cell.
Many proteins have been identified as involved in mitochondrial stability, movement, and inheritance from mother to bud cells (Table 1). These proteins coordinate mitochondria transport (the anterograde and retrograde movements) by interacting with the surface of the mitochondria, ensure proper segregation of the mt-nucleoids within the mitochondrial matrix, and partition the mitochondrial inner and outer membranes to the mother and bud cells. Below we describe two different models of how different proteins guide the anterograde movement for mitochondria from the mother cell to the bud in S. cerevisiae.
Models of mitochondrial partitioning during budding
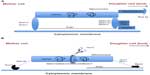
In the first model (Figure 1A), actin polymerization and dynamics by the Arp2/3 protein complex generate the forces to power anterograde mitochondrial movements.52–54 In this model, the actin cables within the cytoplasm interact with the mitochondrial outer membrane proteins, Mmm1p, Mdm10p and Mdm12p, with the Mmm1p–Mdm10p–Mdm12p complex functioning similarly to the kinetochore. The protein complex is also referred to as the mitochore, analogous to the kinetochore, linking mitochondria to the cytoskeleton.48 This model further suggests that two members of the Puf family of RNA-binding proteins, Jsn1p and Puf3p, recruit the Arp2/3 protein complex to the mitochondrial surface.55,56 These actin cables are assembled by formins, conserved proteins that are located from the bud neck to the bud tip and are associated with the plus ends of actin filaments. Thus, mitochores provide directionality to the Arp2/3-dependent mitochondrial movement, with the mitochore continuously mediating the reversible, cyclic binding of the mitochondria to the actin cables, which are used as tracks for mitochondrial movement. In contrast, retrograde movement of mitochondria toward the distal end of the mother cell is achieved as a passive byproduct of actin elongation at the plus end. In this model, two proteins, class V myosin Myo2p and the Rab-GTPase Ypt11p (that hydrolyzes GTP to GDP), are proposed as being required for the retention of mitochondria at the bud tip through an actin-independent mechanism.57 In contrast, proteins Num1p and Dnm1p are involved in retention of mitochondria in the mother cell through an actin-dependent mechanism.58
The alternative model states that the myosin motor, Myo2p, drives anterograde mitochondrial movement.59 In this model, Myo2p, together with the rab-type GTPase, Ypt11p, and the mitochondrial membrane protein Mmr1p, mediates bud-directed transport of mitochondria along the actin cables (Figure 1B). At the bud end, the cortical endoplasmic reticulum and Mmr1p together serve as the anchor for actin cables for mitochondrial movement. At the other end of the mother cell, Num1p, by interacting with Mdm36p, anchors the mitochondria to the cell cortex of the plasma membrane, forming the mitochondria-endoplasmic reticulum-cortex anchor. Together, the antagonistic activities of Myo2p and Num1p stretch the mitochondria to enhance mitochondrial fission by the dynamin-related protein, Dnm1p. In this model, proteins Mdm10p, Mdm12p, Mdm34p, and Mmm1p form the endoplasmic reticulum-mitochondria encounter structure to select the mitochondrial fission site and ensure that the healthy and aging portions of the mitochondria and mtDNA are respectively partitioned to the bud and the mother cell (Figure 1B).59
The current evidence suggests that the second model is likely more reflective of the structures involved in mitochondrial inheritance dynamics than the first model. The first model was built on the original observation that the key “mitochore” component Mmm1p was a mitochondrial outer membrane protein.60 However, Mmm1p was later identified to be located in the endoplasmic reticulum.61 In addition, several pieces of evidence suggested an essential and direct role of Myo2p in anterograde mitochondrial transport.62,63 First, defects in mitochondrial inheritance in myo2 mutants can be rescued by expressing a chimeric mitochondria-specific motor Myo2-Fis1 that carries a mitochondrial outer membrane anchor in place of its cargo-binding domain. Second, mitochondria-specific loss-of-function alleles of myo2 are synthetically lethal with ypt11Δ and that the synthetic lethality can be rescued by Myo2-Fis1. Third, Myo2p was detected on the surface of isolated mitochondria by immuno-electron microscopy. The authors62,63 suggested that Myo2p was likely the main driver of the anterograde mitochondrial movement in yeast, and that the anterograde movement requires both Ypt11p and Mmr1p (Table 1; Figure 1B). However, the role of Arp2/3 in this model remains to be determined.
Control and maintenance of mitochondrial health in yeast aging
There are two types of cellular lifespan and associated aging in yeasts and most other eukaryotes: the chronological lifespan, which refers to the survival time of stationary phase, non-dividing cells; and the replicative lifespan, which refers to the number of times that a cell can divide before senescence occurs (ie, before it is unable to divide despite permissive environmental conditions). In S. cerevisiae, mother cells age with each budding cycle; however, daughter cells are typically born with a full replicative lifespan, including having healthy mitochondria. In this asymmetric mother–daughter relationship, there are two major types of quality control mechanisms to ensure that the inherited mitochondria in the daughter bud cell are functional and healthy. The first type of quality control deals with mitochondrial repair and the second type is involved in identifying and selecting against damaged mitochondria that are beyond repair.
Three processes are known to be involved in itochondrial repair: mitochondrial fusion, protein refolding, and protease degradation of damaged mitochondrial proteins and replacement by functional versions.64 Specifically, mitochondrial fusion repairs low-functioning mitochondria by intraorganellar reorganization and functional complementation.65 Molecular chaperones can fold denatured and unfolded proteins to restore their functions.66,67 For mitochondrial proteins that are damaged beyond repair, those located on the outside of the mitochondrial membrane are degraded by the proteasome in the cytosol. In contrast, damaged proteins within the mitochondria can be degraded/rescued by the mitochondrial AAA+ proteases and Pim1p/Lon.66 Pim1p/Lon is a conserved ATP-dependent protease in the mitochondrial matrix with several functions, including chaperone activity for the folding and assembly of the respiratory complex and the turnover and refolding of misfolded proteins. In aged yeast cells, Pim1p activity is decreased, reducing the protein repair ability.6 Furthermore, the deletion of pim1 increases ROS within the mitochondrial cytosol and decreases the replicative lifespan in yeast, consistent with unrepaired damage to mitochondrial oxygen-handling proteins playing a significant role in reducing the replicative lifespan in yeast.6
Identification and elimination of non-functional mitochondria may be achieved by one of two interrelated processes: mitophagy; and mitochondrial fusion and fission.68,69 Here, the membrane potential in mitochondria serves as a key signal of mitochondrial health. Mitochondria with a low membrane potential are specifically tagged and selected against during bud formation and/or targeted for destruction through mitophagy.64,68 Similarly, mitochondrial fusion and fission are important processes safeguarding mitochondrial and cellular health. Specifically, inhibition of mitochondrial outer membrane fusion shortens the life span while inhibition of mitochondrial fission extends the lifespan of S. cerevisiae and two other ascomycete fungi.68–72 Several genes are known to be involved in the mitochondrial fusion/fission process. Fzo1p and Mgm1p, two mitochondrial GTPases, mediate outer and inner membrane fusion, respectively. In contrast, Dnm1p facilitates mitochondrial fission. Deletions of these genes negatively impact mitochondrial function and yeast lifespan. Together, these studies indicate that maintaining mitochondria as a continuous reticulum that can undergo regular fission and fusion promotes longer lifespan.
Mother-daughter asymmetry and mtDNA inheritance
Despite the safeguarding and repair mechanisms, mitochondria are constantly exposed to potentially damaging metabolic byproducts such as ROS that can negatively impact mitochondria proteins and mtDNA. During the S and G2 phases, there are significant energy requirements from the mitochondria to drive the synthesis of DNA, RNA, proteins, and many other cellular components, simultaneously producing ROS. Different parts of the mitochondria and mtDNA may be exposed to different amounts of ROS. There is increasing evidence indicating that during budding, mitochondria with more damage are selectively retained in the mother cells, while those without damage (or with less damage) are selectively passed on to daughter cells.73–75 Specifically, first, carbonylated proteins, which accumulate during the aging process, are more abundant in the mitochondria of mother cells than in those of daughter cells.76 Second, experiments employing fluorescent biosensors suggest that bud-localized mitochondria produce fewer ROS and are more reducing compared with mother-cell mitochondria.73–75 Third, mitochondria with a mutation in a gene encoding a subunit of the mitochondrial ATP synthase produced daughters that were born old.77 Cells with the mutation had significantly reduced mitochondrial membrane potential and were unable to partition healthy active mitochondria to daughter cells, ultimately resulting in the generation of cells totally lacking mitochondria.77 At present, the detailed mechanisms of control for the mother-daughter asymmetry are not known. One hypothesis is that the anchorage machinery in the bud tip preferentially binds to the healthier mitochondria (or portions of them) while that at the distal end in the mother cell binds to the less healthy mitochondria (or portions of them).
Relevance of fungal mtDNA inheritance to that in animals and humans
There are several major differences between fungal and animal mitochondria and mitochondrial inheritance. First, unlike the large variation in mitochondrial genome sizes among fungal species,13 the mitochondrial genome sizes are very similar among animal species, at approximately 16 kb.78 Second, mitochondrial inheritance in sexual crosses in animals is almost universally uniparental and maternal, unlike the diverse patterns observed among fungal species.12 Third, there is abundant evidence indicating that mtDNA evolves much faster than nuclear genes in animals.78 In contrast, there is little evidence for faster evolution of mtDNA than that of nuclear DNA in fungi.79 Despite these differences, heteroplasmy, a frequently reported condition for sexual crosses in fungi, was recently found to be almost universal in the blood and skeletal muscles in humans, mainly due to the high somatic mutation rate of mtDNA.80 Indeed, the high mutation rate in animal mtDNA suggests that ensuring the inheritance of healthy mitochondria is paramount in animals. Below we briefly summarize the main mechanisms that control mtDNA inheritance in animals.
In animals, the male gametes (the sperm) typically contain very few mitochondria while the female gametes (the egg) contain abundant mitochondria. In addition, the mitochondria in male gametes are clustered into the tail end of the sperm. During fertilization, mitochondria in the sperm are selectively excluded from entering the egg, ensuring a predominantly uniparental and maternal pattern of mtDNA inheritance. Even if mitochondria from the male gamete enter the egg, there is evidence for active degradation of the male mitochondria.81 The exclusion of male gamete mtDNA from the zygote is regarded as a response to its significantly higher mutation rate compared with female gamete mtDNA. Even during development of the female gamete, several mechanisms are known to be involved to ensure mitochondrial health in the egg. These mechanisms include mtDNA bottlenecks during germ cell development and selection against specific mtDNA mutation types during maternal transmission. For details of our current understandings of these processes in animals, please see the recent review by Stewart and Larsson.82 At present, the molecular controls of mtDNA bottlenecks and mtDNA selection in animals are largely unknown. However, the effects of these processes in animals are similar to those described earlier for mtDNA inheritance during budding in S. cerevisiae.
Conclusion and perspectives
Mitochondria are ubiquitous organelles in eukaryotes and are responsible for generating ATP, the universal cellular energy. Many biological phenomena, including human diseases, are related to mitochondria. Thus, the mechanisms governing the maintenance and inheritance of healthy mitochondria have attracted increasing attention from geneticists, molecular and cellular biologists, evolutionary biologists, and medical professionals. Fungi, especially the budding yeast S. cerevisiae, have served as excellent model organisms for understanding the fundamental mechanisms governing mitochondrial health and inheritance.
Researchers over the last 40 years have identified a great diversity of mitochondrial inheritance patterns in sexual mating in fungi, ranging from uniparental to biparental and to the frequent generation of recombinant genotypes. The two major groups of fungi, the ascomycetes and the basidiomycetes, have shown very different mtDNA inheritance patterns. In ascomycetes, there is no known genetic factor that ensures uniparental mtDNA inheritance. In contrast, genes involved in mating have increasingly been shown to be involved in mtDNA inheritance in basidiomycetes. However, the detailed molecular and cellular processes for uniparental mitochondrial inheritance require further investigations.
Similarly, though a number of genes have been identified as involved in mitochondrial maintenance and inheritance during budding in S. cerevisiae, much remains unknown. For example, what are the proteins and processes involved in polarizing the mitochondria during budding to ensure that the healthier portion will partition to the daughter bud? Is mitochondrial partitioning synchronized with chromosomal segregation, and if so, how? Unlike most eukaryotes, S. cerevisiae can grow under both aerobic and anaerobic conditions. Under extended anaerobic conditions, there is no oxidative phosphorylation and damage by ROS should be minimized. In this situation, is mitochondrial inheritance controlled similarly as that under aerobic conditions? More broadly, do genes identified in mitochondrial maintenance and inheritance in asexual budding of S. cerevisiae also function similarly in other fungi during asexual growth? Given the diversity of mtDNA inheritance patterns in sexual crosses in fungi, we expect that a diversity of novel genes will be identified for controlling mitochondrial maintenance and inheritance in fungi.
Acknowledgments
We thank Anne Howells for the invitation to contribute to this review and Anna Tozer for her reminders to keep us on track. We thank Heather Yoell for her critical reading of the paper. Research in our laboratories on fungal genetics has been supported by the Natural Sciences and Engineering Research Council of Canada and the National Natural Science Foundation of China (grant 31100479).
Disclosure
The authors report no conflicts of interest in this work.
References
Van der Giezen M. Mitochondria and the rise of eukaryotes. BioScience. 2011;61:594–601. | |
Brun S, Dalle F, Saulnier P, et al. Biological consequences of petite mutations in Candida glabrata. J Antimicrob Chemother. 2005;2: 307–314. | |
Cheng S, Clancy CJ, Zhang Z, et al. Uncoupling of oxidative phosphorylation enables Candida albicans to resist killing by phagocytes and persist in tissue. Cell Microbiol. 2007;9:492–501. | |
Olson A, Stenlid J. Plant pathogens: mitochondrial control of fungal hybrid virulence. Nature. 2001;411:438. | |
Dirick L, Bendris W, Loubiere V, Gostan T, Gueydon E, Schwob E. Metabolic and environmental conditions determine nuclear genomic instability in budding yeast lacking mitochondrial DNA. G3 (Bethesda). 2013;3:411–423. | |
Erjavec N, Bayot A, Gareil M, et al. Deletion of the mitochondrial Pim1/Lon protease in yeast results in accelerated aging and impairment of the proteasome. Free Radic Biol Med. 2013;56:9–16. | |
Saumitou-Laprade P, Cuguen J, Vernet P. Cytoplasmic male sterility in plants: molecular evidence and the nucleo-cytoplasmic conflict. Trends Ecol Evol. 1994;9:431–435. | |
Duchen MR, Szabadkai G. Roles of mitochondria in human disease. Essays Biochem. 2010;47:115–137. | |
Taylor RW, Turnbull DM. Mitochondrial DNA mutations in human disease. Nat Rev Genet. 2005;5:389–402. | |
Singh R, Pandey R, Abbas AM. Mitochondria, spermatogenesis, and male infertility. Mitochondrion. 2010;10:419–428. | |
Birky CW Jr. Relaxed and stringent genomes: why cytoplasmic genes don’t obey Mendel’s laws. J Hered. 1994;85:355–365. | |
Wilson A, Xu J. Mitochondrial inheritance: diverse patterns and mechanisms with an emphasis on fungi. Mycology. 2012;3:158–166. | |
Xu J, Wang P. Mitochondrial inheritance in basidiomycete fungi. Fungal Biol Rev. 2015 In press. | |
Birky CW Jr. The inheritance of genes in mitochondria and chloroplasts: laws, mechanisms, and models. Annu Rev Genet. 2001;35:125–148. | |
Casselton LA, Kües U. The origin of multiple mating types in the model mushrooms Coprinopsis cinerea and Schizophyllum commune. In: Heitman J, Kronstad JW, Taylor JW, Casselton LA, editors. Sex in Fungi: Molecular Determination and Evolutionary Implications. Washington, DC, USA: ASM Press; 2007. | |
James TY, Stenlid J, Olson A, Johannesson H. Evolutionary significance of imbalanced nuclear ratios within heterokaryons of the basidiomycete fungus Heterobasidion parviporum. Evolution. 2008;62: 2279–2296. | |
Heitman J, Kronstad JW, Taylor JW, Casselton LA. Sex in Fungi: Molecular Determination and Evolutionary Implications. Washington, DC, USA: ASM Press; 2007. | |
Yan Z, Hull CM, Heitman J, Sun S, Xu J. SXI1α controls uniparental mitochondrial inheritance in Cryptococcus neoformans. Curr Biol. 2004;14:R743–R744. | |
Yan Z, Hull CM, Sun S, Heitman J, Xu J. The mating type-specific homeodomain genes sxi1α and sxi2a coordinately control uniparental mitochondrial inheritance in Cryptococcus neoformans. Curr Genet. 2007;51:187–195. | |
Wang ZX, Wilson A, Xu J. Mitochondrial DNA inheritance in the human fungal pathogen Cryptococcus gattii. Fungal Genet Biol. 2015;75:1–10. | |
Yan Z, Sun S, Shahid M, Xu J. Environment factors can influence mitochondrial inheritance in the fungus Cryptococcus neoformans. Fungal Genet Biol. 2007;44:315–322. | |
Basse CW. Mitochondrial inheritance in fungi. Curr Opin Microbiol. 2010;13:712–719. | |
Debuchy R, Berteaux-Lecellier V, Silar P. Mating systems and sexual morphogenesis in ascomycetes. In: Borkovich KA, Ebbole DJ, Momany M, editors. Cellular and Molecular Biology of Filamentous Fungi. Washington, DC, USA: ASM Press; 2010. | |
Xu J, Ali RY, Gregory DA, et al. Uniparental mitochondrial transmission in sexual crosses in Cryptococcus neoformans. Curr Microbiol. 2000;40:269–273. | |
Billiard S, López-Villavicencio M, Devier B, Hood ME, Fairhead C, Giraud T. Having sex, yes, but with whom? Inferences from fungi on the evolution of anisogamy and mating types. Biol Rev. 2011;86:421–442. | |
May G, Taylor JW. Independent transfer of mitochondrial plasmids in Neurospora crassa. Nature. 1989;359:320–322. | |
Raper JR. Genetics of Sexuality in Higher Fungi. New York, NY, USA: Ronald Press; 1966. | |
Ku¨es U, James TY, Heitman J. Mating type in basidiomycetes: unipolar, bipolar, and tetrapolar patterns of sexuality. In: Pöggeler S, Wöstemeyer J, editors. Evolution of Fungi and Fungal-like Organisms, the Mycota XIV. Heidelberg, Germany: Springer-Verlag; 2011. | |
Casselton LA, Condit A. A mitochondrial mutant of Coprinus lagopus. J Gen Microbiol. 1972;72:521–527. | |
May G, Taylor JW. Patterns of mating and mitochondrial DNA inheritance in the agaric basidiomycete Coprinus cinereus. Genetics. 1988;118:213–220. | |
Specht CA, Novotny CP, Ullrich RC. Mitochondrial DNA of Schizophyllum commune: restriction map, genetic map, and mode of inheritance. Curr Genet. 1992;22:129–134. | |
Jin T, Horgen PA. Uniparental mitochondrial transmission in the cultivated button mushroom, Agaricus bisporus. Appl Environ Microbiol. 1994;60: 4456–4460. | |
Xu J, Kerrigan RW, Horgen PA, Anderson JB. Localization of the mating type gene in Agaricus bisporus. Appl Environ Microbiol. 1993;59:3044–3049. | |
Hintz WE, Anderson JB, Horgen PA. Nuclear migration and mitochondrial inheritance in the mushroom Agaricus bitorquis. Genetics. 1988;119:35–41. | |
Yan Z, Xu J. Mitochondria are inherited from the MATa parent in crosses of the basidiomycete fungus Cryptococcus neoformans. Genetics. 2003;163:1315–1325. | |
Wilch G, Ward S, Castle A. Transmission of mitochondrial DNA in Ustilago violacea. Curr Genet. 1992;22:135–140. | |
Findley K, Sun S, Fraser JA, et al. Discovery of a modified tetrapolar sexual cycle in Cryptococcus amylolentus and the evolution of MAT in the Cryptococcus species complex. PLoS Genet. 2012;8:e1002528. | |
Fedler M, Luh KS, Stelter K, Nieto-Jacobo F, Basse CW. The a2 mating-type locus genes lga2 and rga2 direct uniparental mitochondrial DNA (mtDNA) inheritance and constrain mtDNA recombination during sexual development of Ustilago maydis. Genetics. 2009;181: 847–860. | |
Gyawali R, Lin X. Prezygotic and postzygotic control of uniparental mitochondrial DNA inheritance in Cryptococcus neoformans. mBio. 2013;4:e00112–e00113. | |
Skosireva I, James TY, Sun S, Xu J. Mitochondrial inheritance in haploid x non-haploid crosses in Cryptococcus neoformans. Curr Genet. 2010;56:163–176. | |
Banuett F. History of the mating types in Ustilago maydis. In: Heitman J, Kronstad JW, Taylor JW, Casselton LA, editors. Sex in Fungi: Molecular Determination and Evolutionary Implications. Washington, DC, USA: ASM Press; 2007. | |
Bortfeld M, Auffarth K, Kahmann R, Basse CW. The Ustilago maydis a2 mating-type locus genes lga2 and rga2 compromise pathogenicity in the absence of the mitochondrial p32 family protein Mrb1. Plant Cell. 2004;16:2233–2248. | |
Nieto-Jacobo F, Pasch D, Basse CW. The mitochondrial Dnm1-like fission component is required for lgA2-induced mitophagy but dispensable for starvation-induced mitophagy in Ustilago maydis. Eukaryot Cell. 2012;11:1154–1166. | |
Bahler J. Cell-cycle control of gene expression in budding and fission yeast. Annu Rev Genet. 2005;39:69–94. | |
Williamson D. The curious history of yeast mitochondrial DNA. Nat Rev Genet. 2002;3:1–7. | |
Maleszka R, Skelly PJ, Clark-Walker GD. Rolling circle replication of DNA in yeast mitochondria. EMBO J. 1991;10:3923–3929. | |
Shibata FL. DNA recombination protein-dependent mechanism of homoplasmy and its proposed functions. Mitochondrion. 2007;7: 17–23. | |
Solieri L. Mitochondrial inheritance in budding yeasts: towards an integrated understanding. Trends Microbiol. 2010;18:521–530. | |
Kucej M, Butow RA. Evolutionary tinkering with mitochondrial nucleoids. Trends Cell Biol. 2007;17:586–592. | |
Brewer LR, Friddle R, Noy A, et al. Packaging of single DNA molecules by the yeast mitochondrial protein Abf2p. Biophys J. 2003;85: 2519–2524. | |
Kaufman BA, Kolesar JE, Perlman PS, Butow RA. A function for the mitochondrial chaperonin Hsp60 in the structure and transmission of mitochondrial DNA nucleoids in Saccharomyces cerevisiae. J Cell Biol. 2003;163:457–461. | |
Boldogh IR, Nowakowski DW, Yang HC, et al. A protein complex containing Mdm10p, Mdm12p, and Mmm1p links mitochondrial membranes and DNA to the cytoskeleton-based segregation machinery. Mol Biol Cell. 2003;14:4618–4627. | |
Boldogh IR, Fehrenbacher KL, Yang HC, Pon LA. Mitochondrial movement and inheritance in budding yeast. Gene. 2005;354:28–36. | |
Boldogh IR, Yang HC, Nowakowski WD, et al. Arp2/3 complex and actin dynamics are required for actin-based mitochondrial motility in yeast. Proc Natl Acad Sci U S A. 2001;98:3162–3167. | |
García-Rodríguez LJ, Gay AC, Pon LA. Puf3p, a Pumilio family RNA binding protein, localizes to mitochondria and regulates mitochondrial biogenesis and motility in budding yeast. J Cell Biol. 2007;176: 197–207. | |
Fehrenbacher KL, Boldogh IR, Pon LA. A role for Jsn1p in recruiting the Arp2/3 complex to mitochondria in budding yeast. Mol Biol Cell. 2005;16:5094–5102. | |
Boldogh IR, Ramcharan SL, Yang HC, Pon LA. A type V myosin(Myo2p) and a Rab-like G-protein (Ypt11p) are required for retention of newly inherited mitochondria in yeast cells during cell division. Mol Biol Cell. 2004;15:3994–4002. | |
Cerveny KL, Studer SL, Jensen RE, Sesaki H. Yeast mitochondrial division and distribution require the cortical num1 protein. Dev Cell. 2007;12:363–375. | |
Westermann B. Mitochondrial inheritance in yeast. Biochim Biophys Acta. 2014;1837:1039–1046. | |
Burgess SM, Delannoy M, Jensen RE. MMM1 encodes a mitochondrial outer membrane protein essential for establishing and maintaining the structure of yeast mitochondria. J Cell Biol. 1994;126: 1375–1391. | |
Kornmann B, Currie E, Collins SR, et al. An ER–mitochondria tethering complex revealed by a synthetic biology screen. Science. 2009;325: 477–481. | |
Förtsch J, Hummel E, Krist M, Westermann B. The myosin-related motor protein Myo2 is an essential mediator of bud-directed mitochondrial movement in yeast. J Cell Biol. 2011;194:473–488. | |
Chernyakov I, Santiago-Tirado F, Bretscher A. Active segregation of yeast mitochondria by Myo2 is essential and mediated by Mmr1 and Ypt11. Curr Biol. 2013;23:1818–1824. | |
Vevea JD, Swayne TC, Boldogh IR, Pon LA. Inheritance of the fittest mitochondria in yeast. Trends Cell Biol. 2014;24:53–60. | |
Youle RJ, van der Bliek AM. Mitochondrial fission, fusion, and stress. Science. 2012;337:1062–1065. | |
Weil A, Luce K, Dröse S, Wittig I, Brandt U, Osiewacz HD. Unmasking a temperature-dependent effect of the P. anserina i-AAA protease on aging and development. Cell Cycle. 2011;10:4280–4290. | |
Voos W. Chaperone-protease networks in mitochondrial protein homeostasis. Biochim Biophys Acta. 2013;1833:388–399. | |
Twig G, Elorza A, Molina AJ, et al. Fission and selective fusion govern mitochondrial segregation and elimination by autophagy. EMBO J. 2008;27:433–446. | |
Scheckhuber CQ, Wanger RA, Mignat CA, Osiewacz HD. Unopposed mitochondrial fission leads to severe lifespan shortening. Cell Cycle. 2011;10:3105–3110. | |
Kurashima K, Chae M, Inoue H, Hatakeyama S, Tanaka S. A uvs-5 strain is deficient for a mitofusin gene homologue, fzo1, involved in maintenance of long life span in Neurospora crassa. Eukaryot Cell. 2013;12:233–243. | |
Kato A, Kurashima K, Chae M, et al. Deletion of a novel F-box protein, MUS-10, in Neurospora crassa leads to altered mitochondrial morphology, instability of mtDNA and senescence. Genetics. 2010;185: 1257–1269. | |
Scheckhuber CQ, Erjavec N, Tinazli A, Hamann A, Nyström T, Osiewacz HD. Reducing mitochondrial fission results in increased life span and fitness of two fungal ageing models. Nat Cell Biol. 2007;9: 99–105. | |
McFaline-Figueroa JR, Vevea J, Swayne TC, et al. Mitochondrial quality control during inheritance is associated with lifespan and mother-daughter age asymmetry in budding yeast. Aging Cell. 2011;10: 885–895. | |
Erjavec N, Nyström T. Sir2p-dependent protein segregation gives rise to a superior reactive oxygen species management in the progeny of Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 2007;104: 10877–10881. | |
Klinger H, Rinnerthaler M, Lam YT, et al. Quantitation of (a)symmetric inheritance of functional and of oxidatively damaged mitochondrial aconitase in the cell division of old yeast mother cells. Exp Gerontol. 2010;45:533–542. | |
Aguilaniu H, Gustafsson L, Rigoulet M, Nyström T. Asymmetric inheritance of oxidatively damaged proteins during cytokinesis. Science. 2003;299:1751–1753. | |
Lai CY, Jaruga E, Borghouts C, Jazwinski SM. A mutation in the ATP2 gene abrogates the age asymmetry between mother and daughter cells of the yeast Saccharomyces cerevisiae. Genetics. 2002;162:73–87. | |
Xia XH. Rapid evolution of animal mitochondrial DNA. In: Singh RS, Xu J, Kulathinal RJ, editors. R apidly Evolving Genes and Genetic Systems. Oxford, UK: Oxford University Press; 2012. | |
Xu J, Yan Z, Guo H. Divergence, hybridization, and recombination in the mitochondrial genome of the human pathogenic yeast Cryptococcus gattii. Mol Ecol. 2009;17:2628–2642. | |
Payne BAI, Wilson IA, Yu-Wai-Man P. Universal heteroplasmy of human mitochondrial DNA. Hum Mol Genet. 2013;22:384–390. | |
Sutovsky P, Moreno RD, Ramalho-Santos J, Dominko T, Simerly C, Schatten G. Ubiquitinated sperm mitochondria, selective proteolysis, and the regulation of mitochondrial inheritance in mammalian embryos. Biol Reprod. 2000;63:582–590. | |
Stewart JB, Larsson NG. Keeping mtDNA in shape between generations. PLoS Genet. 2014;10:e1004670. |
 © 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.