Back to Journals » Drug Design, Development and Therapy » Volume 14
Curcumin Has Anti-Proliferative and Pro-Apoptotic Effects on Tongue Cancer in vitro: A Study with Bioinformatics Analysis and in vitro Experiments
Authors Ma C , Zhuang Z , Su Q , He J , Li H
Received 11 November 2019
Accepted for publication 22 January 2020
Published 4 February 2020 Volume 2020:14 Pages 509—518
DOI https://doi.org/10.2147/DDDT.S237830
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Anastasios Lymperopoulos
Chao Ma,1 Zongming Zhuang,1 Qisheng Su,2 Jianfeng He,1 Haoyu Li1
1Department of Ophthalmology, The First Affiliated Hospital of Guangxi Medical University, Nanning, People’s Republic of China; 2Department of Clinical Laboratory, The First Affiliated Hospital of Guangxi Medical University, Nanning, People’s Republic of China
Correspondence: Haoyu Li
Department of Ophthalmology, The First Affiliated Hospital of Guangxi Medical University, 6 Shuangyong Road, Nanning 530021, Guangxi Zhuang Autonomous Region, People’s Republic of China
Email [email protected]
Purpose: This study focused on the mechanism underlying the therapeutic effect of curcumin against tongue cancer (TC).
Methods: Target genes of TC and curcumin were identified, respectively. Three datasets of TC from Gene Expression Omnibus were included, and then the differentially expressed genes were collected. After combing the data from The Cancer Genome Atlas, bioinformatics analyses were performed to investigate hub genes in terms of the functions and correlations. The proliferation and migration of TC cells were evaluated with CCK-8 assay and scratch wound healing assay, respectively. Cell apoptosis was evaluated by TUNEL assay, flow cytometry and Western blot. Cell cycle was determined by flow cytometry.
Results: In this study, 15 hub genes were identified (TK1, TDRD3, TAGLN2, RNASEH2A, PDE2A, NCF2, MAP3K3, GPX3, GPD1L, GBP1, ENO1, CAT, ALDH6A1, AGPS and ACACB). They were mainly enriched in oxygen-related processes, such as oxidation-reduction process, reactive oxygen species metabolic process, hydrogen peroxide catabolic process, oxidoreductase activity and Peroxisome-related pathway. The expression levels of hub gene mRNAs were positively correlated with each other’s expression levels. None of the hub genes was correlated with prognosis (P > 0.05). Curcumin significantly inhibited CAL 27 cell proliferation and migration (P < 0.05), but significantly promoted cell apoptosis (P < 0.05).
Conclusion: Curcumin has potential therapeutic effect on treating TC by suppressing cell proliferation and migration, as well as promoting apoptosis through modulating oxygen-related signaling pathways.
Keywords: tongue neoplasms, curcumin, proliferation, apoptosis, migration, computational biology
Introduction
The incidence of tongue cancer (TC), namely oral squamous cell carcinoma, has increased significantly worldwide in the past few decades. Surveys in some areas have shown an increasing trend for the incidence of oral squamous cell carcinoma especially in women and young patients.1,2 TC is also the eighth most prevalent malignancy in the world, posing a major threat to human health. Studies have shown that the 5-year survival rate of TC patients was about 50% within 10 years.3 The main treatment of TC is surgical. However, the complications of surgical treatment can lead to the difficulties in speech, diet and respiration, as well as other dysfunctions and maxillofacial deformities. Meanwhile, the recurrence rate of single surgery is high with high probability of lymph node or distant organ metastasis.4,5 In addition, chemotherapy is a conventional method for the treatment of metastatic tumors. However, it also has serious side effects in clinical use, such as myelosuppression, gastrointestinal reaction, and liver and kidney damage.6 Furthermore, drug resistance often happens, leading to disease recurrence and treatment failure.7,8 The molecular mechanisms for TC are not completely clear, and the specific targeted drugs are not available in clinical practice to date. Therefore, the application of natural derivatives as an alternative treatment in the early stage may be a new research direction. There is growing evidence that natural derivatives have anticancer properties and may be developed as adjuvant drugs for cancer treatment.9–11 Moreover, the location of TC is easy to access, therefore, natural derivatives can be directly applied to the tumor area, which can avoid the first elimination and then enhance the bioavailability of natural derivatives.
Extracted from turmeric rhizome, curcumin (1,7-bis-[4-hydroxy-3-methoxyphenyl]-1,6-heptadiene-3,5-dione) is a bright yellow hydrophobic polyphenol. It is an easy-to-obtain, low-cost and safe drug.12,13 Previous reports have shown the effectiveness of curcumin as an anti-tumor chemo-preventive and chemo-sensitizing agent for many tumor types.14,15 Curcumin inhibits the progression and metastasis of cancer cells by blocking multiple signaling pathways including p53, PI3K, RAS, protein kinase B (Akt), Wnt-β, and MAPK.16 Furthermore, curcumin can inhibit the expression of many pathogenicity determinants, including tumor necrosis factor α (TNF-α), cyclin D1, epidermal growth factor receptor (EGFR), β-catenin, IKK and anti-apoptotic gene, B-cell lymphoma 2 (Bcl-2).17,18 Although curcumin has a strong effect on TC, which has been reported by precedent studies,19,20 the exact impacts of curcumin on TC cell apoptosis, cycle, proliferation and migration are still not clear. Besides, there is no systematic analysis on the possible acting site and potential mechanisms of curcumin on TC.
In this study, public databases and bioinformatics methods were adopted to analyze the targets and possible mechanisms of curcumin action on TC. The effects of curcumin at different concentrations on the proliferation, migration, apoptosis and cell cycle for TC cells were verified in vitro. The purpose of this study was to lay a foundation for further study on the role of curcumin on TC.
Methods
The Identification of Potential Target Genes
The molecular formula of curcumin was listed at the Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP, http://lsp.nwu.edu.cn/tcmsp.php, accessed by June 25, 2019),21 and the effective target genes were identified with the PharmMapper Server (http://www.lilab-ecust.cn/pharmmapper/, accessed by June 25, 2019)22–24 by using Druggable Pharmacophore Models. The microarray repositories of initial experimental studies on identifying different mRNA expressions between tumor tissues and non-tumor tissues from TC patients were included. The Gene Expression Omnibus (GEO) database was retrieved for obtaining the microarray data (GSE31056, GSE9844 and GSE13601, Table 1). The common differentially expressed genes (DEGs) were obtained using Venny v2.1 (https://bioinfogp.cnb.csic.es/tools/venny/index.html, accessed by September 30, 2019).25,26 The hub genes were obtained using Venny v2.1, and hub genes were considered as targets of curcumin on treating TC.
 |
Table 1 Detailed Information of Included Microarray Datasets from GEO Database |
Functional Analysis for Hub Genes
Depending on the identified hub genes, Gene ontology (GO) and Kyoto encyclopedia of genes and genomes (KEGG) analyses were conducted to explore the role of curcumin on TC using the Search Tool for the Retrieval of Interacting Genes/Proteins (STRING v11.0, https://string-db.org/, accessed by September 30, 2019).
Correlation Analysis and Survival Analysis
The Cancer Genome Atlas (TCGA) data about survival rate and clinical information of TC patients were extracted from OncoLnc (http://www.oncolnc.org/, accessed by September 30, 2019) and TCGA (https://cancergenome.nih.gov/, accessed by September 30, 2019), including these patients’ ID in TCGA, gender, race, age at index, survival time, vital status, treatment and tumor stage, as well as the hub gene mRNA expression profiles of TC patients. Data of TC were screened by limiting the site of onset from head and neck squamous carcinoma (HNSC) dataset. The relationship of co-expression between hub genes was evaluated using Pearson correlation coefficient analysis. TC patients were classified into high- or low-expression groups based on a 50th percentile cutoff for each hub gene mRNA. The survival analysis was based on overall survival (OS) using the log-rank test and Kaplan-Meier estimator, which were performed to obtain the log-rank P-value and evaluate the OS predictive potency for hub genes. The correlation heatmap and survival curves were performed by R v3.6. A working diagram for computational and bioinformatics analysis was shown (Figure 1).
 |
Figure 1 Working diagram showing computational and bioinformatics analysis. |
Cell and Drug Information
Human tongue cancer cell line (CAL 27) was provided by Procell Life Science & Technology (Wuhan, China) and was cultured in DMEM supplemented with 1 penicillin/streptomycin and 20% FBS at 37°C under one atmosphere with 5% CO2 in humidified air. Curcumin was produced by Solarbio Life Sciences Inc. (Beijing, China).
Cell Proliferation Assay
Cell Counting Kit-8 (CCK-8) assay was performed using CCK-8 Kit (Beyotime Biotechnology, Shanghai, China) according to the manufacturer’s instructions, followed by the spectrophotometric absorbance measurement at 450 nm.
Cell Migration Assay
TC cells at 100% confluence in 6-well plastic culture plates were scratched by a sterile pipette tip. Next, cells were rinsed with serum-free DMEM and photographed at different time points.
TUNEL Assay
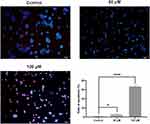
One Step TUNEL Apoptosis Assay Kit (Dalian Meilun Biotechnology, Dalian, China) was applied to evaluate cell apoptosis following the manufacturer’s instructions. The apoptotic index was evaluated depending on the TUNEL-positive cells in 5 non-overlapping fields for each treatment group under 200× magnification.
Flow Cytometry
TC cells were digested with trypsin and centrifuged at 500 g. Cell apoptosis was examined using Annexin V-APC/7-AAD apoptosis kit (Multisciences, Hangzhou, China). According to the manufacturer’s instructions. For cell cycle assay, TC cells were digested with trypsin and washed with ice-cold PBS, followed by resuspension in cold 75% ethanol. Cell cycle was evaluated using Cell Cycle Staining Kit (Multisciences, Hangzhou, China) according to the manufacturer’s instructions.
Western Blot
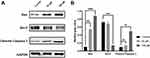
Equal amounts of denatured protein samples were subjected to 10% SDS-PAGE, proteins were then transferred onto polyvinylidene difluoride (PVDF) membranes (Solarbio Life Sciences, Beijing, China). After blocking with non-fat milk, PVDF membranes were incubated with appropriate antibodies (GAPDH [glyceraldehyde-3-phosphate dehydrogenase], Bax [Bcl-2-associated X protein], Bcl-2 and cleaved Caspase 3), all antibodies were obtained from Cell Signaling Technology Inc. (MA, USA). LI-COR automatic chemiluminescence image analysis system was applied for visualization. Quantification of Western blot signals was achieved with Odyssey Fc Imaging System.
Statistical Analysis
Statistical analysis was conducted by GraphPad Prism (v8.2.0) (GraphPad Software, San Diego, USA) using one-way and two-way ANOVA. P-values < 0.05 were considered statistical significant.
Results
Identification of Target Genes
After screening the TCMSP and PharmMapper severs, 158 potential target genes for curcumin were obtained. Three GEO datasets (GSE31056, GSE9844 and GSE13601) were analyzed in this study. These datasets included 80 TC samples and 62 normal samples. According to the adjusted P value, 541 DEGs were in common (Figure 2A). Fifteen hub genes were identified (Figure 2B) including thymidine kinase 1 (TK1), tudor domain containing 3 (TDRD3), transgelin 2 (TAGLN2), ribonuclease H2 subunit A (RNASEH2A), phosphodiesterase 2A (PDE2A), neutrophil cytosolic factor 2 (NCF2), mitogen-activated protein kinase kinase kinase 3 (MAP3K3), glutathione peroxidase 3 (GPX3), glycerol-3-phosphate dehydrogenase 1 like (GPD1L), guanylate binding protein 1 (GBP1), enolase 1 (ENO1), catalase (CAT), aldehyde dehydrogenase 6 family member A1 (ALDH6A1), alkylglycerone phosphate synthase (AGPS) and acetyl-CoA carboxylase beta (ACACB).
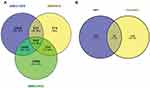 |
Figure 2 Venn diagrams showing the overlaps of the DEGs in common (A) and hub genes (B). |
GO Analysis and KEGG Pathway Enrichment Analysis on Hub Genes
To clarify the functional pathways for the hub genes, GO analysis and KEGG analysis were performed. The functional detection based on GO included biological process (BP), cellular component (CC) and molecular functionality (MF). As shown in Tables S1–S4, the hub genes were mainly enriched on oxidation-reduction process (GO:0055114), reactive oxygen species metabolic process (GO:0072593), hydrogen peroxide catabolic process (GO:0042744), oxidoreductase activity (GO:0016491) and peroxisome-related pathway (hsa04146), while CAT, GPX3, NCF2 and AGPS participated in most of the processes (Tables S1–S4).
Correlation Analysis and Survival Analysis on Hub Genes
The correlations among the expression levels of hub genes were measured by Pearson correlation coefficient analysis. The results indicated that hub gene mRNA expression levels were positively inter-correlated (Figure S1). Additionally, the prognostic significance of hub gene mRNA expression profiles was assessed. The survival curves indicated that none of the hub genes was significantly related to OS in TC patients (P > 0.05, Figure S2).
Curcumin Inhibits CAL 27 Cell Proliferation
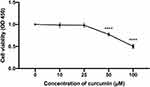
The anti-proliferative effect by curcumin on CAL 27 cells was investigated with CCK-8 assay in vitro. To this end, CAL 27 cells were treated with curcumin treatment at indicated concentrations. The results showed that curcumin inhibited CAL27 cell proliferation in a dose-dependent manner (P < 0.001, Figure 3). Therefore, curcumin has a potential anti-proliferative effect in CAL 27.
Curcumin Inhibits CAL 27 Cell Migration
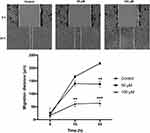
The anti-migration effect by curcumin on CAL 27 cells was assessed with the scratch wound assay in vitro. As shown in Figure 4, no significant difference between the 3 groups was observed at 6 h in terms of migration distance (P > 0.05). Curcumin at 100 μM significantly inhibited CAL 27 migration at 16 h (P < 0.01) and 24 h (P < 0.001) compared to the controls. Besides, CAL 27 migration was significantly suppressed by curcumin at 50 μM at 24 h compared to the controls (P < 0.01). Therefore, curcumin also has a potential anti-migration effect in CAL 27.
Curcumin Promotes CAL 27 Cell Apoptosis
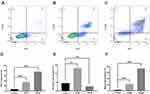
The pro-apoptosis effect by curcumin on CAL 27 cells was evaluated with TUNEL assay and flow cytometry. As demonstrated by flow cytometry (Figure 5). The results showed that curcumin treatment significantly promoted the late apoptosis as well as early apoptosis in TC cells (P < 0.0001). The effect of curcumin on TC cell apoptosis was further validated by TUNEL assay and Western blot (Figures 6 and 7).
Curcumin Induces S-Phase Cell Cycle Arrest in CAL 27 Cells
Finally, we evaluated the effect of curcumin on cell cycle using flow cytometry. The results showed that the proportion of S-phase cells in the curcumin-treated groups was significantly increased compared with that in the control group (Figure 8) (P < 0.0001). On the other hand, no difference was observed in the proportions of G1- and G2-phase cells between the curcumin-treated groups and the control group (P > 0.05).
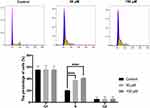 |
Figure 8 Cell cycle evaluated by flow cytometry. The CAL 27 cells were treated with curcumin at indicated concentrations, followed by flow cytometry assay. ****P < 0.0001, one-way ANOVA. |
Discussion
In this study, the intersection of three GEO databases was taken into investigation, followed by matching the intersection results with curcumin targets, and a total of 15 hub genes (TK1, TDRD3, TAGLN2, RNASEH2A, PDE2A, NCF2, MAP3K3, GPX3, GPD1L, GBP1, ENO1, CAT, ALDH6A1, AGPS, ACACB) were obtained. After GO and KEGG analysis of these hub genes, it was found that curcumin might had regulatory function on the apoptosis of TC cells via regulating oxidative stress-related pathways. In the following in vitro experiments, we found that curcumin could significantly upregulate the expressions of Caspase 3 and Bax while reducing the expression of Bcl-2 in CAL 27 cells. Based on flow cytometry, the apoptotic cells increased with the increase of curcumin concentration, and the change of cell cycle suggested that curcumin might regulate cell S phase by affecting DNA synthesis.
The GO/KEGG function of hub genes and the clinical prognosis of TCGA were verified by the intersection of GEO and curcumin targets. It was found that some of the hub genes were involved in the oxidative stress-related pathways, as well as apoptosis-related pathways. Studies have previously shown that increasing CAT could reduce the apoptosis of hematopoietic stem cells and progenitor cells induced by neuroblastoma and ionizing radiation.27,28 Additionally, in cardiomyocytes, GPX3 inhibition could significantly increase the level of reactive oxygen species (ROS), and then induce cardiomyocyte apoptosis by up-regulating the expression of caspase3.29 In Kashin-Beck disease, the down-regulation of GPX3 promoter methylation and GPX3 gene expression can induce the expression of PI3K/Akt/c-fos signaling pathway and lead to chondrocyte apoptosis.30 AGPS is an oncogene considered to be the target of antineoplastic drugs. The downregulation of its expression can promote tumor apoptosis.31,32 However, it had been found that NCF2 knockdown could significantly reduce the production of ROS and stimulate apoptosis, indicating that ROS produced by NOX2 has anti-apoptotic protective effect in cells.33 Therefore, according to the functions of these hub genes in cell apoptosis, further in-depth experiments were needed to illustrate the anti-TC effect of curcumin.
Here, the function of curcumin on the apoptosis of human TC cells was studied in vitro. The growth inhibition of CAL 27 cells was significantly enhanced by curcumin in a dosage-dependent manner, while TUNEL staining revealed a significant increase in cell apoptosis. Upon curcumin treatment, Western blot assay also showed that curcumin could promote the activation of cleaved Caspase 3 and downregulate the ratio of Bcl-2/Bax. This was consistent with the results of previous studies which reported the induction of apoptosis by curcumin in prostate and gastric cancer.34,35 Live cells have a higher level of Bcl-2, which inhibits apoptosis. Bcl-2 regulates a variety of cellular activities which involved proteins related to cell proliferation or apoptosis, for example, Caspase 3 and Bax.36 The initial steps of the intrinsic apoptosis pathway converge on the activity of Bax,37 which controls the commitment of cells to the apoptotic pathway by changing the physiology of mitochondria.38 Therefore, by detecting Caspase 3 expression and the ratio of Bcl-2 to Bax, the apoptosis-inducing effect of curcumin on CAL 27 cells could be explained. Based on the analyses on apoptosis and cell cycle, it was realized that the proportion of apoptotic cells increased with the increase of curcumin dose. In the cell cycle assay, it was found that S phase arrest occurred in the cell cycle with the intervention of curcumin. The drug or drug regulated protein enters the cell and binds to DNA, then leads to the arrest of cell replication bifurcation during replication. By activating the S phase checkpoint, the S phase is blocked and eventually leads to apoptosis.39 Accordingly, curcumin interferes with the biological activities of CAL 27 cells through the above pathways. In particular, curcumin or its regulatory protein ingredient enters the cell and binds to the cell DNA, resulting in an increase in apoptosis.
Based on bioinformatics analysis and in vitro experiments, curcumin could inhibit CAL 27 cells from proliferating, as well as promote the apoptosis. The main effects of curcumin on hub genes and possible pathway in TC were analyzed.
Conclusion
In conclusion, this study revealed the anti-proliferative and pro-apoptotic roles of curcumin in CAL 27 cells, probably due to its role in modulating oxygen production and metabolism. Therefore, curcumin might be explored as a promising therapeutic method to improve TC treatment. However, curcumin might not have the potential to predict the prognosis of TC.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Ng JH, Iyer NG, Tan MH, Edgren G. Changing epidemiology of oral squamous cell carcinoma of the tongue: a global study. Head Neck. 2017;39(2):297–304. doi:10.1002/hed.24589
2. Simard EP, Torre LA, Jemal A. International trends in head and neck cancer incidence rates: differences by country, sex and anatomic site. Oral Oncol. 2014;50(5):387–403. doi:10.1016/j.oraloncology.2014.01.016
3. Bello IO, Soini Y, Salo T. Prognostic evaluation of oral tongue cancer: means, markers and perspectives (I). Oral Oncol. 2010;46(9):630–635. doi:10.1016/j.oraloncology.2010.06.006
4. Michikawa C, Uzawa N, Kayamori K, et al. Clinical significance of lymphatic and blood vessel invasion in oral tongue squamous cell carcinomas. Oral Oncol. 2012;48(4):320–324. doi:10.1016/j.oraloncology.2011.11.014
5. Annertz K, Anderson H, Palmer K, Wennerberg J. The increase in incidence of cancer of the tongue in the Nordic countries continues into the twenty-first century. Acta Otolaryngol. 2012;132(5):552–557. doi:10.3109/00016489.2011.649146
6. Di Maio M, Basch E, Bryce J, Perrone F. Patient-reported outcomes in the evaluation of toxicity of anticancer treatments. Nat Rev Clin Oncol. 2016;13(5):319–325. doi:10.1038/nrclinonc.2015.222
7. Lu CS, Shieh GS, Wang CT, et al. Chemotherapeutics-induced Oct4 expression contributes to drug resistance and tumor recurrence in bladder cancer. Oncotarget. 2017;8(19):30844–30858. doi:10.18632/oncotarget.9602
8. Ji P, Zhang Y, Wang SJ, et al. CD44hiCD24lo mammosphere-forming cells from primary breast cancer display resistance to multiple chemotherapeutic drugs. Oncol Rep. 2016;35(6):3293–3302. doi:10.3892/or.2016.4739
9. Rashidi B, Malekzadeh M, Goodarzi M, Masoudifar A, Mirzaei H. Green tea and its anti-angiogenesis effects. Biomed Pharmacother. 2017;89:949–956. doi:10.1016/j.biopha.2017.01.161
10. Pavan AR, Silva GD, Jornada DH, et al. Unraveling the anticancer effect of curcumin and resveratrol. Nutrients. 2016;8:11. doi:10.3390/nu8110628
11. Goyal S, Gupta N, Chatterjee S, Nimesh S. Natural plant extracts as potential therapeutic agents for the treatment of cancer. Curr Top Med Chem. 2017;17(2):96–106. doi:10.2174/1568026616666160530154407
12. Teiten MH, Eifes S, Dicato M, Diederich M. Curcumin-the paradigm of a multi-target natural compound with applications in cancer prevention and treatment. Toxins (Basel). 2010;2(1):128–162. doi:10.3390/toxins2010128
13. Maheshwari RK, Singh AK, Gaddipati J, Srimal RC. Multiple biological activities of curcumin: a short review. Life Sci. 2006;78(18):2081–2087. doi:10.1016/j.lfs.2005.12.007
14. Momtazi AA, Shahabipour F, Khatibi S, Johnston TP, Pirro M, Sahebkar A. Curcumin as a microRNA regulator in cancer: a review. Rev Physiol Biochem Pharmacol. 2016;171:1–38.
15. Mirzaei H, Naseri G, Rezaee R, et al. Curcumin: a new candidate for melanoma therapy? Int J Cancer. 2016;139(8):1683–1695. doi:10.1002/ijc.30224
16. Kasi PD, Tamilselvam R, Skalicka-Wozniak K, et al. Molecular targets of curcumin for cancer therapy: an updated review. Tumour Biol. 2016;37(10):13017–13028. doi:10.1007/s13277-016-5183-y
17. Kunnumakkara AB, Anand P, Aggarwal BB. Curcumin inhibits proliferation, invasion, angiogenesis and metastasis of different cancers through interaction with multiple cell signaling proteins. Cancer Lett. 2008;269(2):199–225. doi:10.1016/j.canlet.2008.03.009
18. Shehzad A, Wahid F, Lee YS. Curcumin in cancer chemoprevention: molecular targets, pharmacokinetics, bioavailability, and clinical trials. Arch Pharm (Weinheim). 2010;343(9):489–499. doi:10.1002/ardp.v343:9
19. Hoornstra D, Vesterlin J, Parnanen P, et al. Fermented lingonberry juice inhibits oral tongue squamous cell carcinoma invasion in vitro similarly to curcumin. In Vivo. 2018;32(5):1089–1095. doi:10.21873/invivo.11350
20. Liao F, Liu L, Luo E, Hu J. Curcumin enhances anti-tumor immune response in tongue squamous cell carcinoma. Arch Oral Biol. 2018;92:32–37. doi:10.1016/j.archoralbio.2018.04.015
21. Ru J, Li P, Wang J, et al. TCMSP: a database of systems pharmacology for drug discovery from herbal medicines. J Cheminform. 2014;6:13. doi:10.1186/1758-2946-6-13
22. Liu X, Ouyang S, Yu B, et al. PharmMapper server: a web server for potential drug target identification using pharmacophore mapping approach. Nucleic Acids Res. 2010;38(WebServer issue):W609–W614. doi:10.1093/nar/gkq300
23. Wang X, Pan C, Gong J, Liu X, Li H. Enhancing the enrichment of pharmacophore-based target prediction for the polypharmacological profiles of drugs. J Chem Inf Model. 2016;56(6):1175–1183. doi:10.1021/acs.jcim.5b00690
24. Wang X, Shen Y, Wang S, et al. PharmMapper 2017 update: a web server for potential drug target identification with a comprehensive target pharmacophore database. Nucleic Acids Res. 2017;45(W1):W356–W360. doi:10.1093/nar/gkx374
25. Huang da W, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4(1):44–57. doi:10.1038/nprot.2008.211
26. Huang da W, Sherman BT, Lempicki RA. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009;37(1):1–13. doi:10.1093/nar/gkn923
27. Polito L, Bortolotti M, Pedrazzi M, Mercatelli D, Battelli MG, Bolognesi A. Apoptosis and necroptosis induced by stenodactylin in neuroblastoma cells can be completely prevented through caspase inhibition plus catalase or necrostatin-1. Phytomedicine. 2016;23(1):32–41. doi:10.1016/j.phymed.2015.11.006
28. Xiao X, Luo H, Vanek KN, LaRue AC, Schulte BA, Wang GY. Catalase inhibits ionizing radiation-induced apoptosis in hematopoietic stem and progenitor cells. Stem Cells Dev. 2015;24(11):1342–1351. doi:10.1089/scd.2014.0402
29. Gong Y, Yang J, Cai J, Liu Q, Zhang JM, Zhang Z. Effect of Gpx3 gene silencing by siRNA on apoptosis and autophagy in chicken cardiomyocytes. J Cell Physiol. 2019;234(6):7828–7838. doi:10.1002/jcp.v234.6
30. Han L, Yang X, Sun W, et al. The study of GPX3 methylation in patients with Kashin-Beck disease and its mechanism in chondrocyte apoptosis. Bone. 2018;117:15–22. doi:10.1016/j.bone.2018.08.017
31. Zhu Y, Liu A, Zhang X, et al. The effect of benzyl isothiocyanate and its computer-aided design derivants targeting alkylglycerone phosphate synthase on the inhibition of human glioma U87MG cell line. Tumour Biol. 2015;36(5):3499–3509. doi:10.1007/s13277-014-2986-6
32. Zhu Y, Han Y, Ma Y, Yang P. ADME/toxicity prediction and antitumor activity of novel nitrogenous heterocyclic compounds designed by computer targeting of alkylglycerone phosphate synthase. Oncol Lett. 2018;16(2):1431–1438. doi:10.3892/ol.2018.8873
33. Italiano D, Lena AM, Melino G, Candi E. Identification of NCF2/p67phox as a novel p53 target gene. Cell Cycle (Georgetown, Tex). 2012;11(24):4589–4596. doi:10.4161/cc.22853
34. Sha J, Li J, Wang W, et al. Curcumin induces G0/G1 arrest and apoptosis in hormone independent prostate cancer DU-145 cells by down regulating Notch signaling. Biomed Pharmacother. 2016;84:177–184. doi:10.1016/j.biopha.2016.09.037
35. Zhou X, Wang W, Li P, et al. Curcumin enhances the effects of 5-fluorouracil and oxaliplatin in inducing gastric cancer cell apoptosis both in vitro and in vivo. Oncol Res. 2016;23(1–2):29–34. doi:10.3727/096504015X14452563486011
36. Martinou JC, Youle RJ. Mitochondria in apoptosis: Bcl-2 family members and mitochondrial dynamics. Dev Cell. 2011;21(1):92–101. doi:10.1016/j.devcel.2011.06.017
37. Oltvai ZN, Milliman CL, Korsmeyer SJ. Bcl-2 heterodimerizes in vivo with a conserved homolog, Bax, that accelerates programmed cell death. Cell. 1993;74(4):609–619. doi:10.1016/0092-8674(93)90509-O
38. Maes ME, Schlamp CL, Nickells RW. BAX to basics: how the BCL2 gene family controls the death of retinal ganglion cells. Prog Retin Eye Res. 2017;57:1–25. doi:10.1016/j.preteyeres.2017.01.002
39. Ewald B, Sampath D, Plunkett W. H2AX phosphorylation marks gemcitabine-induced stalled replication forks and their collapse upon S-phase checkpoint abrogation. Mol Cancer Ther. 2007;6(4):1239–1248. doi:10.1158/1535-7163.MCT-06-0633
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.