Back to Archived Journals » Advances in Genomics and Genetics » Volume 4
Consequences of ongoing retrotransposition in mammalian genomes
Authors Maxwell P
Received 30 June 2014
Accepted for publication 24 July 2014
Published 26 September 2014 Volume 2014:4 Pages 129—142
DOI https://doi.org/10.2147/AGG.S57145
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr John Martignetti
Patrick H Maxwell
Department of Biological Sciences, Rensselaer Polytechnic Institute, Troy, NY, USA
Abstract: Retrotransposons can have significant influences on gene expression and genome stability through their ability to integrate reverse-transcript copies of their sequences at new genomic locations by retrotransposition. These elements have been long known to retrotranspose in mammalian germ cells to give rise to inherited insertion alleles, but more recent work has also shown that retrotransposition can occur in mammalian somatic cells, particularly in brain tissue and tumors. Retrotransposition makes appreciable contributions to spontaneous disease-causing alleles in humans and a more significant contribution to spontaneous mutations in mice. Genome-wide studies have found high levels of polymorphic retrotransposon insertions in human populations that are consistent with ongoing retrotransposition. Many insertions do not disrupt exons, but insertions into introns or flanking genes can alter gene expression patterns, generate truncated or antisense gene transcripts, alter splicing patterns, or result in premature polyadenylation of gene transcripts. Furthermore, the very high genomic copy numbers of these elements can lead to nonallelic homologous recombination events that produce gene deletions/duplications and genome rearrangements, and can also lead to evolution of particular insertions or types of elements to have cellular functions through exaptation. Mobility of these elements occurs despite multiple epigenetic mechanisms to restrict their expression. While the potential for retrotransposons to significantly influence mammalian health and cellular functions is clear, substantial research efforts will be needed to fully elucidate the actual contributions of natural levels of mobility of endogenous elements to the health and development of humans and other mammals.
Keywords: retrotransposon, human, mouse, mutations, epigenetics, exaptation
Introduction
Retrotransposons are abundant mobile DNA elements in eukaryotic genomes that have great potential to influence genome maintenance, genome organization, and gene expression through their duplicative replication cycle (retrotransposition). Recent studies indicate that there is significant ongoing retrotransposition in the human germ line and that somatic retrotransposition may have implications for mammalian brain development/function and cancer.1–3 Mammalian genomes harbor long terminal repeat (LTR) retrotransposons/endogenous retroviruses (ERVs) and non-LTR retrotransposons that account for approximately 43% and 37% of the human and mouse genomes, respectively, with non-LTR retrotransposons representing 35% and 27% of each genome, respectively.4,5 Retrotransposition involves reverse transcription of an RNA copy of an element to generate a complementary DNA (cDNA) molecule that is integrated into a new genomic site. Autonomous retrotransposons encode proteins to carry out this replication cycle, and their proteins also mobilize nonautonomous retrotransposons.
Non-LTR retrotransposons are mobile in mammals, including autonomous long interspersed nuclear element-1 (LINE-1 or L1), nonautonomous short interspersed nuclear elements (SINEs), the primate-specific SINE Alu, and hominid-specific SINE-variable number of tandem repeats-Alu (SVA) elements (Figure 1).1,6,7 Full-length L1 elements consist of a 5′ untranslated region (UTR) with an internal RNA polymerase II promoter, an antisense promoter, two open reading frames (ORF1 and ORF2), and a 3′ UTR that ends in a polyadenine (poly[A]) sequence (Figure 1). Approximately 100 and 3,000 active L1 elements are present in the human and mouse genomes, respectively, and are mobile through target-primed reverse transcription (Figure 2).8–11 Transcription of the L1 messenger RNA (mRNA) is followed by translation of ORF1 and ORF2 proteins (ORF1p and ORF2p). ORF1p forms a higher order ribonucleoprotein particle with the L1 RNA that includes ORF2p. The ribonucleoprotein particle gains access to a genomic site, ORF2p endonuclease activity cleaves one strand of the target with preference for a short adenine–thymine-rich consensus sequence,12,13 and ORF2p reverse transcriptase activity uses the exposed 3′-hydroxyl of the cleaved DNA to prime reverse transcription of L1 RNA, beginning at the 3′ poly(A) tail. A staggered cleavage of the second DNA strand at the target site provides another free 3′-hydroxyl for synthesis of the second strand of L1 DNA. Gaps at the 5′ end of each L1 DNA strand due to the staggered cleavage are filled and sealed by cellular proteins. Target-primed reverse transcription results in target-site duplications (from fill-in of gaps), the presence of a 3′ poly(A) sequence, and 5′ truncations if reverse transcription is not complete.
Human and mouse L1 mobilize human Alu and SVA elements and mouse SINEs (Figure 2) despite a preference for L1 proteins to mobilize the RNA from which they were translated.14–17 Mouse SINE B2 is derived from transfer RNA sequences while human Alu and mouse SINE B1 are derived from 7SL RNA sequences of the ribosome signal recognition particle (Figure 1).7 Sequences needed for association with the signal recognition particle are required for efficient Alu mobilization.18 SVA elements are variable in length and consist of four sequence elements: a variable number of CCCTCT repeats, an Alu-derived sequence, variable numbers of approximately 35–50 base pair repeat motifs (variable number of tandem repeats), and sequences from the 3′ end of a human ERV-K element (SINE-R) (Figure 1).19,20 SINEs contain internal RNA polymerase III promoters, while SVA elements are likely transcribed by RNA polymerase II.1,7 Retrotransposition of human Alu and SVA elements minimally requires L1 ORF2p,15,17 but ORF1p increases Alu mobilization and is needed to mobilize some SVA elements.17,21 L1 elements can also mobilize reporter gene transcripts, which requires both ORF1p and ORF2p, providing support for the role of L1 in generating retrocopies of cellular RNA that do not code for proteins (processed pseudogenes) or that do code for proteins and that are expressed from heterologous promoters at target sites (retrogenes).22
Mice, but not humans, also harbor mobile LTR retrotransposons/ERVs. This review will only focus on elements restricted to an intracellular retrotransposition cycle, rather than infectious endogenous mouse retroviruses.23 The autonomous mouse intracisternal A-particle (IAP) and MusD LTR retrotransposons are in the retrovirus family, are present in about 300 and ten active copies in the mouse genome, respectively, and consist of 5′ and 3′ LTR sequences that flank a central region with gag, pro, and pol genes (Figure 1).24,25 IAP and MusD are derived from ERVs through loss of the retroviral env gene required for infectivity, since IAP elements with an env gene (IAPE) are present in the mouse genome, and sequence alterations to a MusD retrotransposon were able to reconstitute an infectious retrovirus.26,27 These elements are often referred to as ERVs, based on their origin, but for the remainder of this review they will be referred to as LTR retrotransposons. Retrotransposition of these elements begins with initiation of RNA polymerase II transcription within the 5′ LTR and termination in the 3′ LTR to produce a terminally redundant mRNA translated into Gag and Gag-Pro-Pol fusion proteins (Figure 2).9 Gag protein forms a virus-like particle that encapsidates the mRNA and fusion protein. The Pro protease activity processes Gag and the fusion protein while Pol contains reverse transcriptase/RNase H and integrase activities. Within the virus-like particle, reverse transcriptase uses a transfer RNA primer, template switching between redundant sequences at the termini of the mRNA, and the associated RNase H activity to generate a double-stranded cDNA copy of the element. Integrase complexes with this cDNA, gains access to a genomic target site, and makes a staggered double-stranded cleavage to join the 3′ ends of each cDNA strand to the target site. Fill-in of the gaps at the 5′ ends and sealing of the nicks by cellular proteins generates target-site duplications, as for non-LTR elements. Mouse IAP elements mobilize nonautonomous internally deleted IAPΔ1 elements, which requires all three IAP gene functions despite a preference for IAP proteins to mobilize the mRNA used for their translation.24 MusD mobilizes the related nonautonomous early transposon (ETn) element (Figures 1 and 2), which also requires all three MusD gene functions and results from sequence similarity between MusD and ETn LTRs, primer binding sites, and polypurine tract sequences.25,28
Developmental timing and tissue specificity of mammalian retrotransposition
Proteins of autonomous retrotransposons are necessary for retrotransposition, so expression of these elements determines the potential times and cell types in which retrotransposition can occur. L1 RNA is present in human and rodent germ cells (male and female), embryonic stem (ES) cells, neural progenitor cells, cancer cells/tumors, in mouse preimplantation embryos, and in a variety of human somatic tissues.29–38 In addition, L1 RNA is more abundant in normal human fibroblasts late during senescence ex vivo and in skeletal muscle and liver of aged mice.39,40 However, many transcripts are partial-length, which can arise from premature termination at multiple internal polyadenylation signal sequences and from splicing.32,41,42 Full-length L1 RNA has been detected in human ovaries, several human somatic tissues, and in mouse zygotes, preimplantation embryos, and prepuberal testes at postnatal day 14.32,34,35,38 L1 ORF1p and ORF2p are present in somatic and germ cells of human fetal and adult testes, including vascular endothelial cells.43 Mouse ORF1p is present in somatic and germ cells from late embryonic ovaries and testes as well as prepuberal and adult testes.35 Multiple mouse and human cancer cell lines express ORF1p, and a large-scale survey detected human ORF1p in approximately half of cancers tested.33,44
Mouse LTR retrotransposons are typically expressed in germ cells and early development.23,45 IAP RNA is present in male germ cells, oocytes, ovulated eggs, very early embryos, and at reduced levels as preimplantation development progresses.28,38,46–48 Limited somatic expression of IAP may be restricted to particular individual elements based on their genomic context.23 MusD transcripts are expressed early in embryogenesis, at reduced levels as embryogenesis proceeds, in mouse ES cells, and at elevated levels in skeletal muscle and liver of aged mice.28,40
Retrotransposition in humans and mice occurs in primordial germ cells, germ cells, and/or early embryos, based on inheritance of spontaneous insertion mutations, but it also occurs to some extent in somatic cells. Identification that the mother of a patient with a disease-causing L1 insertion allele was a somatic and germ line mosaic for the insertion allele is consistent with retrotransposition during early development.49 Mouse or human L1 transgenes in mice and rats retrotranspose in ovaries and testes, at high levels in preimplantation embryos, in brain tissue, and at low levels in somatic cells of several other tissues (0.2% to 2% of cells).36,37 Increases in copy number measured by quantitative polymerase chain reaction are consistent with L1 retrotransposition in human brain tissue, senescent normal human fibroblasts, as well as L1 and MusD retrotransposition in skeletal muscle and livers of aged mice.30,39,40,50 IAP insertions have been observed during cultivation of mouse hematopoietic stem cell lines and in mouse tumor cells.51–54 Engineered L1 elements can retrotranspose when introduced into human ES cells, human and rat neural progenitor cells, human oocytes, primary human fibroblasts, a variety of cancer cell lines, and retrotranspose at higher levels in response to oxidative stress and ionizing radiation.29–31,36,55–59
The extent of ongoing retrotransposition in mammals
Retrotransposon insertions contribute to a modest proportion of spontaneous disease-causing mutations in humans, approximately 0.1%, but contribute to approximately 10% of spontaneous mutations in mice.23 All insertions in the mouse data set that was analyzed to calculate the value of approximately 10% of spontaneous mutations were IAP, ETn, or MusD LTR retrotransposons, but mice are also estimated to have many more active L1 elements than humans (~3,000 versus ~100).10,11,23 Approximately 100 human disease-causing retrotransposon insertions have been documented, nearly all of which were germ line insertions.1 Interestingly, most spontaneous mutations in humans and mice resulted from retrotransposition of nonautonomous human Alu and mouse IAPΔ1 or ETn elements, rather than autonomous elements.1,23
Detection of thousands of retrotransposon insertion polymorphisms in human samples using genome-wide approaches is consistent with substantial ongoing retrotransposition in humans.60–68 Insertion polymorphisms not present in the reference human genome identified in multiple individuals or multiple tissues from the same individual are considered germ line insertions, while insertion polymorphisms detected in one, but not all, tissue samples from one individual are considered somatic insertions. Of studies not biased for a particular type of retrotransposon, two identified >7,000 polymorphic germ line insertions each, and a third identified >24,000 insertion polymorphisms in brain tissues, most of which were somatic.65–67 Polymorphic insertions in these studies were most often Alu elements (~60%–80%), followed in descending order by L1, SVA, and very rare (1% or less of insertions) LTR elements.65–67 Interestingly, the percentage of L1 insertions was slightly higher (~32%) among the brain insertions65 than among the germ line insertions (~22% and ~14%, respectively).66,67 Estimates of L1 polymorphic insertions per individual human genome vary from over 100 to over 200.62,64,66,67 Retrotransposition rates per live births in humans have been estimated to be approximately 1 in 100, 1 in 20, and 1 in 900 for L1, Alu, and SVA elements, respectively.1
A genome-wide identification of LTR element insertion polymorphisms in three mouse strains relative to a well-sequenced fourth strain reported a high frequency of polymorphic insertions.69 Of ~5,500 IAP insertions present in at least one of the four strains, ~61% were polymorphic (not present in all four strains). Of ~1,100 ETn/MusD elements (these two elements were not distinguished) present in at least one of the four strains, ~26% were polymorphic.69
Genome-wide approaches have also been used to follow up on early observations of somatic L1 insertions in a few instances of cancer.70–72 Examples of tumor-specific retrotransposition include nine tumor-specific L1 insertions in six of 20 lung tumors;63 194 tumor-specific insertions (183 L1, ten Alu, one ERV) among 43 colorectal, glioblastoma, multiple myeloma, ovarian, and prostate tumors, though none were present in glioblastoma or multiple myeloma samples;66 69 tumor-specific L1 insertions in 16 colorectal tumors;73 and 12 tumor-specific L1 insertions in five of 19 hepatocellular carcinomas.67 Intriguingly, the two studies that surveyed all types of retrotransposons identified mostly or only L1 insertions,66,67 in contrast to the prevalence of Alu insertions among the germ line and brain tissue data sets just discussed.
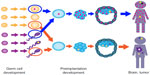
Overall, retrotransposition is primarily occurring in germ cells or early embryonic development, in brain tissue and some cancers and may be elevated during aging (Figure 3). Polymorphic insertions can be identified among small samples of human genomes, and elevated retrotransposon-induced mutations in mice relative to humans appear to result from active mouse LTR retrotransposons.
Consequences of ongoing retrotransposition in mammals
A major factor determining the consequences of new retrotransposition events is the site of insertion. Most polymorphic insertions are present in intergenic regions and introns, rather than exons, which is expected if integration is fairly random, since exons comprise a small portion of mammalian genomes.60–62,65,66 Polymorphic germ line insertions were depleted in genes and enriched in DNA regions hypomethylated in sperm, though Alu insertions were slightly overrepresented in introns.61,65,66,68 However, very rare Alu polymorphisms were equally frequent in exons and transcribed nonexonic sequences per base pair.68 Also, rare polymorphic insertions of IAP and ETn/MusD elements in mice were significantly more likely to be inserted within introns in the sense-orientation relative to the gene target, which is likely to alter gene expression, than were insertions present in all four strains analyzed.23,69 This indicates that some apparent insertion site preferences may result from selection against insertions that alter gene expression.68,69
Some differences between human polymorphic somatic and germ line insertions have been noted. Tumor- and/or brain-specific insertions were more frequently in genes, often present in introns and noncoding gene sequences, enriched in genes frequently mutated in cancer or expressed in brain, and enriched in DNA regions hypomethylated in tumors.63,65–67,73,74 Of particular note for cancer, one study concluded that many tumor-specific retrotransposition events occur after the initiation of tumorigenesis,73 which fits with the upregulation of ORF1p in many human tumors at later stages of tumorigenesis.44
Retrotransposition into introns or sites flanking genes can have mutagenic outcomes, even though coding sequences are not disrupted (Figure 4). Insertions of L1, SINE, and IAP elements upstream of genes or in introns can change gene expression levels and patterns or produce truncated transcripts that alter gene function, including production of oncogenic transcripts.7,36,54,67,75–78 IAP and L1 elements have bidirectional promoters that can produce sense and antisense transcripts of flanking sequences.79,80 Intronic insertions can decrease gene expression due to premature polyadenylation or aberrant splicing.23,41,42,76,81–86 Alu sequences in particular are relatively often incorporated as exons into transcripts, both for alternatively and constitutively spliced transcripts.84,86 In addition, Alu sequences in 5′ UTRs frequently increased or decreased translation levels.86 Inclusion of mouse SINE B1 elements or Alu sequences upstream of reporter gene promoters led to increased gene repression and epigenetic modifications with time, though the SINE copies were unmethylated.87 A particular mouse SINE B2 element and a subfamily of mouse SINE B1 were found to function as boundary elements that influenced dynamic changes in chromatin and transcriptional regulation, which required simultaneous RNA polymerase III sense transcription and RNA polymerase II antisense transcription of the SINE or the transition from RNA polymerase III transcription to RNA polymerase II transcription of the same DNA strand of the SINE, respectively.88,89 Recent high throughput sequencing of human and mouse transcription start sites (TSSs) identified that retrotransposon TSSs comprised a surprisingly large percentage of total TSSs, ~3%–8% and ~2%–4%, in different tissues from humans and mice, respectively.90 Numerous chimeric L1-flanking gene transcripts initiating from the L1 antisense promoter in humans and mice have been identified in various tissues and cells, and some transcripts have been proposed to influence cancer progression by altering gene expression.80,91–94
Retrotransposition can also produce additional changes in target site sequences (Figure 4). L1 retrotransposition can produce deletions or other large-scale sequence rearrangements in cultured cells and has deleted 18 kilobase pairs and 15 kilobase pairs of genomic sequence from the human and chimpanzee genomes, respectively.95–97 Transcription of L1 sequences can sometimes read through the L1 poly(A) signal sequence, resulting in transcription of flanking sequences and leading to insertion of those flanking sequences at new genomic sites.98,99 Intriguingly, overexpression of L1 ORF2 (but not an endonuclease mutant) promoted specific chromosomal translocations relevant to cancer that are mediated by intrachromosomal and interchromosomal interactions due to binding of ligand-activated androgen receptor to specific chromosome sites.100
Cleavage of genomic DNA during retrotransposition can also affect cell growth and survival. Expression of plasmid copies of L1 or of only ORF2 can generate double-strand DNA breaks that reduce entry into mitosis and cell viability, dependent on L1 ORF2 endonuclease activity.101,102 Cell lines with wild-type p53 tumor suppressor that were stably transfected with an engineered L1 exhibited reduced retrotransposition, elevated apoptosis, elevated gamma-H2AX foci, elevated BAX levels, and greater proportions of cells in G2-M phase relative to cell lines with mutant p53.56,103 Expression of full-length L1 or only ORF2 also increased expression of a senescence marker in human cell lines.32,102
L1 retrotransposons have produced thousands of retrocopies (processed pseudogenes and retrogenes) in mammalian genomes.104,105 Two notable examples are an insertion of a retrocopy that generated a fusion gene responsible for restriction of human immunodeficiency virus replication in owl monkeys and a retrogene associated with chondrodysplasia in certain breeds of dogs.106,107 Retrogenes give rise to proteins that can influence cellular functions, and it has been estimated that ~6% of human processed pseudogenes are transcribed.105 Pseudogene transcripts can sometimes lead to Dicer-dependent repression of genes by pairing with partner gene transcripts to give rise to short-interfering RNAs (siRNAs), can act as sinks for microRNAs (miRNAs) to derepress genes, and potentially recruit histone-modifying enzymes to gene promoters.108–110 A survey of 939 low pass human genomes and 85 deeply sequenced genomes identified 48 retrocopy germ line insertion polymorphisms and three tumor-specific insertions.111 Analysis of 17 inbred mouse strain genomes and ten chimpanzee genomes identified 755 and 19 polymorphic retrocopy insertions, respectively.111
Influence of the accumulation of retrotransposons in mammalian genomes
The presence of many dispersed retrotransposon copies can also influence cells and genomes through their transcription, nonallelic homologous recombination between retrotransposons at different genomic sites to generate mutations and genome rearrangements, spreading of repressive chromatin marks into flanking DNA, and evolution of such sequences to contribute to cellular functions (exaptation). For instance, increased Alu transcription during ex vivo aging of human adult stem cells inhibited recruitment of cohesin and condensin proteins to sites of DNA damage in pericentric heterochromatin, and stable knockdown of Alu RNA in senescent cells led to loss of markers of senescence in these cells and proliferation.112 Nonallelic homologous recombination involving Alu elements has caused gene deletion or duplication events producing mutations responsible for a variety of human diseases, including acute myeloid leukemia, familial hypercholesterolemia, familial breast cancer, von Hippel-Lindau disease, and multiple osteochondromas.113–117 On an evolutionary scale, comparison of the chimpanzee and human genomes identified >600 chimp-specific Alu-Alu recombination events that deleted >700 kilobase pairs of sequence from the chimpanzee genome.118 Additional reports have identified that non-LTR and LTR retrotransposon sequences are frequently present at breakpoint junctions of chromosome rearrangements, including rearrangements associated with human disease.119–123
There is the potential for spreading of heterochromatin from silenced retrotransposon sequences into flanking genes, but this has only occasionally been studied by comparing polymorphic insertion sites. One study of polymorphic insertions identified limited histone H3 lysine 9 trimethylation (H3K9me3) and H4K20me3 spreading from LTR retrotransposons in mouse ES cells, but a second study found no evidence for heterochromatin spreading from 29 LTR retrotransposons near TSSs into genes.124,125 Rather, evidence for potential spreading of euchromatin into the LTR retrotransposon sequences was obtained.125
Retrotransposon sequences can potentially evolve to contribute to cellular/organismal functions through exaptation. Exaptation occurs when genomic sequences originally acted upon by selection to provide a particular function are coopted through selection to provide a different function, or when apparently nonfunctional genomic sequences evolve to provide a function.126,127 This is distinct from adaptation, which involves evolution of genomic sequences through selection for the function that the sequences currently have.126 Sequences of particular types of retrotransposons or of particular inserted copies of retrotransposons could be acted upon by selection to acquire cellular functions, and examples include functions of SINE RNAs and influences of retrotransposons on patterns of gene expression. Transcription of SINE elements is increased by some cellular stresses, including heat shock.7 Human Alu and mouse SINE B2 RNAs are able to directly bind to RNA polymerase II after heat shock to inhibit transcription of genes repressed by heat shock.128–130 Alu RNA can also prevent general translational repression by binding to and inhibiting the double-stranded RNA-activated protein kinase PKR (EIF2AK2).131 Analysis of placental enhancer sequences in mouse and rat trophoblast stem cells identified a number of species-specific enhancers residing in retrotransposon sequences that could have contributed to interspecies placental diversification.132 Many species-specific binding sites for the CTCF protein that establishes chromatin/transcriptional boundaries are associated with species-specific SINE insertions.133 Overall, retrotransposon sequences have roles or potential roles in gene networks regulating early development, cell-fate, and responses to stress.127
Restriction of autonomous mammalian retrotransposons
Considering how retrotransposons can influence genome stability and gene expression, it is not surprising that their expression is negatively regulated by multiple mechanisms. These repressive mechanisms include transcriptional silencing through DNA methylation and repressive histone modifications, small RNA-based mRNA degradation and gene silencing, and inhibition by a family of cytidine deaminase proteins. DNA methylation at CpG dinucleotides is a common epigenetic mark for silencing gene expression, and promoters of mammalian L1 elements and mouse autonomous LTR retrotransposons are methylated in most tissues and most stages of development.33,45 Mouse L1 retrotransposons are demethylated in developing germ cells, modestly methylated in oocytes and zygotes, and demethylated during preimplantation development,134,135 which correlates with global demethylation and remethylation events during germ line and embryonic development.45,135 In contrast, IAP elements are less demethylated than L1 elements during germ cell development and preimplantation development.134–136 L1 and/or IAP RNA levels are elevated in mouse embryos deficient for activity of the maintenance DNA methyltransferase gene Dnmt1, mouse ES cells deficient for Dnmt1 and/or the de novo DNA methyltransferase genes Dnmt3a and Dnmt3b, and germ cells of male mice deficient for the DNA methyltransferase-like gene Dnmt3l.136–139 In addition, mice that developed tumors due to DNA hypomethylation caused by a hypomorphic allele of Dnmt1 frequently harbored oncogenic IAP insertions in their tumors.54
Silencing of retrotransposons is also associated with repressive histone modifications, though different marks appear to be important for different elements and/or developmental stages. L1 and IAP are enriched for the activating mark, histone H3 lysine 4 trimethylation (H3K4me3), and the repressive mark, H3K9me3, when they are expressed at the two-cell stage, but are only enriched for H3K9me3 at the eight-cell stage as expression begins to decrease.38 IAP and ETn/MusD LTRs are strongly enriched for H3K9me3 and H4K20me3 in mouse ES cells.140 H3K9me3 catalyzed by the SET-domain containing histone methyltransferase SETDB1 in mouse ES cells appears to be critical for silencing these LTR elements, but not L1, in ES cells.141,142 H3K9me3 of LTR elements is maintained in ES cells triply deficient for Dnmt1, Dnmt3a, and Dnmt3b, and retention of methylation at IAP elements in doubly deficient Dnmt3a and Dnmt3b ES cells is reduced by conditional knockout of Setdb1, indicating that H3K9me3 is acting upstream of DNA methylation in ES cells.141–143 In contrast, mouse LTR retrotransposons are not more highly expressed in mouse embryonic fibroblasts (MEFs) deficient for Setdb1, are enriched for H4K20me3, but not H3K9me3, in mouse fibroblast cell lines, and are silenced by Dnmt1 in postimplantation embryos.142,144 L1 promoters are silenced by H3K9me3 and H4K20me3 in MEFs dependent on binding of retinoblastoma protein family members to L1.145 However, L1 promoters in mouse fibroblast cell lines are enriched for H3K9me3, but not H4K20me3.144 Different histone marks may therefore be more or less important at different points in development to initiate silencing, direct DNA methylation, and/or maintain silencing of particular types of retrotransposons.
siRNA and miRNA pathways can regulate retrotransposons. Endogenous L1 siRNA has been observed in human cell lines, resulting from transcription from an antisense promoter in the L1 5′ UTR,80 and can inhibit L1 retrotransposition in a manner dependent on the small RNA-processing enzyme DICER.146 Inhibition of Dicer in mouse two-cell embryos resulted in elevated IAP RNA levels in mouse eight-cell embryos.147 Furthermore, the Drosha-DGCR8 microprocessor that is necessary for biogenesis of miRNA can negatively regulate L1 and Alu retrotransposition.148
Piwi proteins are animal-specific members of the Argonaute protein family that associate with piwi-interacting RNAs (piRNAs) to mediate small RNA-dependent gene regulation, particularly of retrotransposons in the germ line.149 Piwi proteins bind cleavage products of single-stranded RNA produced from genomic sites referred to as piRNA clusters and use their slicer activity to cleave complementary RNA molecules, which then generates piRNAs of the complementary sequences that can be used to process additional piRNAs from piRNA cluster transcripts.149 Mice with mutations in one of the three mouse Piwi genes, Mili, Miwi, or Miwi2, have similar phenotypes of male sterility due to defects at earlier (Mili and Miwi2) or later (Miwi) stages of spermatogenesis, elevated transcript levels of L1 and IAP elements, decreased methylation of L1 and IAP elements, and reduced levels of piRNAs.150–153 DNA methylation defects indicate that Piwi proteins and piRNA likely regulate epigenetic marks at retrotransposon sequences, in addition to cleaving retrotransposon RNA, though this role in DNA methylation has not been well defined.149 Conditional knockout of Mili later during spermatogenesis (in differentiating spermatogonia) demonstrated that elevated L1, but not IAP, expression resulted from reduced slicing of L1 RNA, even though L1 sequences were methylated.154 Interestingly, high PIWIL2 (homolog of mouse Mili) expression in human induced pluripotent stem cells (iPSC) compared to bonobo and chimpanzee iPSC contributes to reduced L1 mobility in human iPSC.155
Differences in silencing through epigenetic and piRNA-based mechanisms offer at least partial explanations for the increased mobility of certain nonautonomous retrotransposons, such as ETn and SINEs, compared to their partner autonomous elements, MusD and L1, respectively. ETn is expressed at 30-fold and 170-fold higher levels than MusD in mouse embryos or ES cells, respectively, which results at least partly from guanine–cytosine-rich sequences in the body of MusD elements, but not ETn, which direct greater DNA methylation, H3K9me3 deposition, and reduced H3K9 acetylation of MusD elements.28,156 In contrast to L1 elements, mouse SINE B1 methylation and RNA levels were unaltered in Mili mutants or MitoPLD mutants (deficient for a Piwi-interacting protein), indicating that SINEs may not be strongly regulated through Piwi proteins and piRNAs.157
An additional distinct mechanism of retrotransposon inhibition involves a family of proteins with roles in innate immunity. Mammalian apolipoprotein B mRNA editing catalytic peptide (APOBEC) and activation-induced deaminase (AID) proteins restrict the mobility of a variety of retroelements, including LTR and non-LTR retrotransposons.158 Seven human APOBEC3 genes inhibit L1 retrotransposition to varying degrees, at least three of these genes inhibit mouse LTR retrotransposons,159–161 APOBEC1 from multiple mammals can restrict IAP, MusD, and L1,162 and AID from multiple species inhibits mobility of L1 and MusD.163 Inhibition of LTR retrotransposons is frequently dependent on the cytidine deaminase activity of these proteins,159,160,162,163 which deaminates cytosines to uracils in newly synthesized retrotransposon DNA, leading to hypermutation and degradation of the DNA.158 In contrast, cytidine deaminase activity of these proteins is dispensable for inhibition of L1, and restriction appears to involve inhibition of cDNA synthesis through interactions with L1 ORF1p and/or mRNA.161,163–165 High APOBEC3B expression in human iPSC compared to iPSC from bonobos and chimpanzees contributes to reduced mobility of L1 in human iPSC.155
The existence of several mechanisms for inhibiting retrotransposons is consistent with the need for mammals to tightly restrict the mobility of these elements. Use of multiple mechanisms may further indicate a need for redundancy to achieve the necessary level of inhibition or that the different mechanisms act in restricted developmental stages and cell types. Observations regarding timing and requirement of particular epigenetic marks and silencing mechanisms have led to proposals that the contribution of different marks and mechanisms varies during development, but that the combined mechanisms provide an overall restriction of retrotransposons.142,154
Concluding remarks
Retrotransposons are mobile in germ and somatic cells of mammals at appreciable levels despite many mechanisms to restrict their expression. This mobility contributes to mutagenesis and genetic variation in human populations through a variety of influences on gene expression/function. Despite all the potential of these elements, defining the contribution of endogenous retrotransposons expressed at normal levels to the health and function of cells and organisms remains a significant challenge. This is partly due to the difficulties of characterizing the effects of many polymorphic insertions and experimentally manipulating DNA sequences present at thousands of copies per genome. During the search for such functions and consequences, consideration should be given to issues raised in a recent review of noncoding and repetitive DNA. These include analyses indicating that most genomic DNA sequences are unlikely to have functions, despite widespread transcription of genomic sequences, and that alleles with minor beneficial or detrimental effects may behave as neutral in effective population sizes for mammals.166 Furthermore, only a minority of retrotransposon insertions/elements have been demonstrated to have functions, and most retrotransposon-derived transcripts are present at very low levels in cells.166 Continued study of retrotransposons may uncover surprising and significant impacts on mammals, but may require consideration of these elements from new perspectives, and will require substantial effort.
Acknowledgments
The author would like to thank members of the Maxwell laboratory for feedback on the manuscript.
Disclosure
The author reports no conflicts of interest in this work.
References
Hancks DC, Kazazian HH. Active human retrotransposons: variation and disease. Curr Opin Genet Dev. 2012;22(3):191–203. | |
Rodić N, Burns KH. Long interspersed element-1 (LINE-1): passenger or driver in human neoplasms? PLoS Genet. 2013;9(3):e1003402. | |
Reilly MT, Faulkner GJ, Dubnau J, Ponomarev I, Gage FH. The role of transposable elements in health and diseases of the central nervous system. J Neurosci. 2013;33(45):17577–17586. | |
Lander ES, Linton LM, Birren B, et al; International Human Genome Sequencing Consortium. Initial sequencing and analysis of the human genome. Nature. 2001;409(6822):860–921. | |
Waterston RH, Lindblad-Toh K, Birney E, et al; Mouse Genome Sequencing Consortium. Initial sequencing and comparative analysis of the mouse genome. Nature. 2002;420(6915):520–562. | |
Boissinot S, Furano AV. The recent evolution of human L1 retrotransposons. Cytogenet Genome Res. 2005;110(1–4):402–406. | |
Ichiyanagi K. Epigenetic regulation of transcription and possible functions of mammalian short interspersed elements, SINEs. Genes Genet Syst. 2013;88(1):19–29. | |
Luan DD, Korman MH, Jakubczak JL, Eickbush TH. Reverse transcription of R2Bm RNA is primed by a nick at the chromosomal target site: a mechanism for non-LTR retrotransposition. Cell. 1993;72(4):595–605. | |
Beauregard A, Curcio MJ, Belfort M. The take and give between retrotransposable elements and their hosts. Annu Rev Genet. 2008;42:587–617. | |
Brouha B, Schustak J, Badge RM, et al. Hot L1s account for the bulk of retrotransposition in the human population. Proc Natl Acad Sci U S A. 2003;100(9):5280–5285. | |
Goodier JL, Ostertag EM, Du K, Kazazian HH. A novel active L1 retrotransposon subfamily in the mouse. Genome Res. 2001;11(10):1677–1685. | |
Feng Q, Moran JV, Kazazian HH, Boeke JD. Human L1 retrotransposon encodes a conserved endonuclease required for retrotransposition. Cell. 1996;87(5):905–916. | |
Jurka J. Sequence patterns indicate an enzymatic involvement in integration of mammalian retroposons. Proc Natl Acad Sci U S A. 1997;94(5):1872–1877. | |
Wei W, Gilbert N, Ooi SL, et al. Human L1 retrotransposition: cis preference versus trans complementation. Mol Cell Biol. 2001;21(4):1429–1439. | |
Dewannieux M, Esnault C, Heidmann T. LINE-mediated retrotransposition of marked Alu sequences. Nat Genet. 2003;35(1):41–48. | |
Dewannieux M, Heidmann T. L1-mediated retrotransposition of murine B1 and B2 SINEs recapitulated in cultured cells. J Mol Biol. 2005;349(2):241–247. | |
Hancks DC, Goodier JL, Mandal PK, Cheung LE, Kazazian HH. Retrotransposition of marked SVA elements by human L1s in cultured cells. Hum Mol Genet. 2011;20(17):3386–3400. | |
Bennett EA, Keller H, Mills RE, et al. Active Alu retrotransposons in the human genome. Genome Res. 2008;18(12):1875–1883. | |
Ostertag EM, Goodier JL, Zhang Y, Kazazian HH. SVA elements are nonautonomous retrotransposons that cause disease in humans. Am J Hum Genet. 2003;73(6):1444–1451. | |
Wang H, Xing J, Grover D, et al. SVA elements: a hominid-specific retroposon family. J Mol Biol. 2005;354(4):994–1007. | |
Wallace N, Wagstaff BJ, Deininger PL, Roy-Engel AM. LINE-1 ORF1 protein enhances Alu SINE retrotransposition. Gene. 2008;419(1–2):1–6. | |
Esnault C, Maestre J, Heidmann T. Human LINE retrotransposons generate processed pseudogenes. Nat Genet. 2000;24(4):363–367. | |
Maksakova IA, Romanish MT, Gagnier L, Dunn CA, van de Lagemaat LN, Mager DL. Retroviral elements and their hosts: insertional mutagenesis in the mouse germ line. PLoS Genet. 2006;2(1):e2. | |
Dewannieux M, Dupressoir A, Harper F, Pierron G, Heidmann T. Identification of autonomous IAP LTR retrotransposons mobile in mammalian cells. Nat Genet. 2004;36(5):534–539. | |
Ribet D, Dewannieux M, Heidmann T. An active murine transposon family pair: retrotransposition of “master” MusD copies and ETn trans-mobilization. Genome Res. 2004;14(11):2261–2267. | |
Ribet D, Harper F, Dewannieux M, Pierron G, Heidmann T. Murine MusD retrotransposon: structure and molecular evolution of an “intracellularized” retrovirus. J Virol. 2007;81(4):1888–1898. | |
Ribet D, Harper F, Dupressoir A, Dewannieux M, Pierron G, Heidmann T. An infectious progenitor for the murine IAP retrotransposon: emergence of an intracellular genetic parasite from an ancient retrovirus. Genome Res. 2008;18(4):597–609. | |
Baust C, Gagnier L, Baillie GJ, Harris MJ, Juriloff DM, Mager DL. Structure and expression of mobile ETnII retroelements and their coding-competent MusD relatives in the mouse. J Virol. 2003;77(21):11448–11458. | |
Garcia-Perez JL, Marchetto MC, Muotri AR, et al. LINE-1 retrotransposition in human embryonic stem cells. Hum Mol Genet. 2007;16(13):1569–1577. | |
Coufal NG, Garcia-Perez JL, Peng GE, et al. L1 retrotransposition in human neural progenitor cells. Nature. 2009;460(7259):1127–1131. | |
Georgiou I, Noutsopoulos D, Dimitriadou E, et al. Retrotransposon RNA expression and evidence for retrotransposition events in human oocytes. Hum Mol Genet. 2009;18(7):1221–1228. | |
Belancio VP, Roy-Engel AM, Pochampally RR, Deininger P. Somatic expression of LINE-1 elements in human tissues. Nucleic Acids Res. 2010;38(12):3909–3922. | |
Rosser JM, An W. L1 expression and regulation in humans and rodents. Front Biosci (Elite Ed). 2012;4:2203–2225. | |
Packer AI, Manova K, Bachvarova RF. A discrete LINE-1 transcript in mouse blastocysts. Dev Biol. 1993;157(1):281–283. | |
Branciforte D, Martin SL. Developmental and cell type specificity of LINE-1 expression in mouse testis: implications for transposition. Mol Cell Biol. 1994;14(4):2584–2592. | |
Muotri AR, Chu VT, Marchetto MC, Deng W, Moran JV, Gage FH. Somatic mosaicism in neuronal precursor cells mediated by L1 retrotransposition. Nature. 2005;435(7044):903–910. | |
Kano H, Godoy I, Courtney C, et al. L1 retrotransposition occurs mainly in embryogenesis and creates somatic mosaicism. Genes Dev. 2009;23(11):1303–1312. | |
Fadloun A, Le Gras S, Jost B, et al. Chromatin signatures and retrotransposon profiling in mouse embryos reveal regulation of LINE-1 by RNA. Nat Struct Mol Biol. 2013;20(3):332–338. | |
De Cecco M, Criscione SW, Peckham EJ, et al. Genomes of replicatively senescent cells undergo global epigenetic changes leading to gene silencing and activation of transposable elements. Aging Cell. 2013;12(2):247–256. | |
De Cecco M, Criscione SW, Peterson AL, Neretti N, Sedivy JM, Kreiling JA. Transposable elements become active and mobile in the genomes of aging mammalian somatic tissues. Aging (Albany NY). 2013;5(12):867–883. | |
Perepelitsa-Belancio V, Deininger P. RNA truncation by premature polyadenylation attenuates human mobile element activity. Nat Genet. 2003;35(4):363–366. | |
Belancio VP, Hedges DJ, Deininger P. LINE-1 RNA splicing and influences on mammalian gene expression. Nucleic Acids Res. 2006;34(5):1512–1521. | |
Ergün S, Buschmann C, Heukeshoven J, et al. Cell type-specific expression of LINE-1 open reading frames 1 and 2 in fetal and adult human tissues. J Biol Chem. 2004;279(26):27753–27763. | |
Rodić N, Sharma R, Sharma R, et al. Long interspersed element-1 protein expression is a hallmark of many human cancers. Am J Pathol. 2014;184(5):1280–1286. | |
Maksakova IA, Mager DL, Reiss D. Keeping active endogenous retroviral-like elements in check: the epigenetic perspective. Cell Mol Life Sci. 2008;65(21):3329–3347. | |
Pikó L, Hammons MD, Taylor KD. Amounts, synthesis, and some properties of intracisternal A particle-related RNA in early mouse embryos. Proc Natl Acad Sci U S A. 1984;81(2):488–492. | |
Dupressoir A, Heidmann T. Germ line-specific expression of intracisternal A-particle retrotransposons in transgenic mice. Mol Cell Biol. 1996;16(8):4495–4503. | |
Peaston AE, Evsikov AV, Graber JH, et al. Retrotransposons regulate host genes in mouse oocytes and preimplantation embryos. Dev Cell. 2004;7(4):597–606. | |
van den Hurk JA, Meij IC, Seleme MC, et al. L1 retrotransposition can occur early in human embryonic development. Hum Mol Genet. 2007;16(13):1587–1592. | |
Muotri AR, Marchetto MC, Coufal NG, et al. L1 retrotransposition in neurons is modulated by MeCP2. Nature. 2010;468(7322):443–446. | |
Ishihara H, Tanaka I. Detection and cloning of unique integration sites of retrotransposon, intracisternal A-particle element in the genome of acute myeloid leukemia cells in mice. FEBS Lett. 1997;418(1–2):205–209. | |
Pogue-Geile KL, Gott JA, Greenberger JS. The role of intracisternal A-type particles in the evolution of factor-independent murine hematopoietic cell lines. Leukemia. 1998;12(1):4–12. | |
Lee JS, Haruna T, Ishimoto A, Honjo T, Yanagawa S. Intracisternal type A particle-mediated activation of the Notch4/int3 gene in a mouse mammary tumor: generation of truncated Notch4/int3 mRNAs by retroviral splicing events. J Virol. 1999;73(6):5166–5171. | |
Howard G, Eiges R, Gaudet F, Jaenisch R, Eden A. Activation and transposition of endogenous retroviral elements in hypomethylation induced tumors in mice. Oncogene. 2008;27(3):404–408. | |
Moran JV, Holmes SE, Naas TP, DeBerardinis RJ, Boeke JD, Kazazian HH. High frequency retrotransposition in cultured mammalian cells. Cell. 1996;87(5):917–927. | |
Haoudi A, Semmes OJ, Mason JM, Cannon RE. Retrotransposition-competent human LINE-1 induces apoptosis in cancer cells with intact p53. J Biomed Biotechnol. 2004;2004(4):185–194. | |
Shi X, Seluanov A, Gorbunova V. Cell divisions are required for L1 retrotransposition. Mol Cell Biol. 2007;27(4):1264–1270. | |
Farkash EA, Kao GD, Horman SR, Prak ET. Gamma radiation increases endonuclease-dependent L1 retrotransposition in a cultured cell assay. Nucleic Acids Res. 2006;34(4):1196–1204. | |
Giorgi G, Marcantonio P, Del Re B. LINE-1 retrotransposition in human neuroblastoma cells is affected by oxidative stress. Cell Tissue Res. 2011;346(3):383–391. | |
Beck CR, Collier P, Macfarlane C, et al. LINE-1 retrotransposition activity in human genomes. Cell. 2010;141(7):1159–1170. | |
Ewing AD, Kazazian HH. High-throughput sequencing reveals extensive variation in human-specific L1 content in individual human genomes. Genome Res. 2010;20(9):1262–1270. | |
Huang CR, Schneider AM, Lu Y, et al. Mobile interspersed repeats are major structural variants in the human genome. Cell. 2010;141(7):1171–1182. | |
Iskow RC, McCabe MT, Mills RE, et al. Natural mutagenesis of human genomes by endogenous retrotransposons. Cell. 2010;141(7):1253–1261. | |
Ewing AD, Kazazian HH. Whole-genome resequencing allows detection of many rare LINE-1 insertion alleles in humans. Genome Res. 2011;21(6):985–990. | |
Baillie JK, Barnett MW, Upton KR, et al. Somatic retrotransposition alters the genetic landscape of the human brain. Nature. 2011;479(7374):534–537. | |
Lee E, Iskow R, Yang L, et al; Cancer Genome Atlas Research Network. Landscape of somatic retrotransposition in human cancers. Science. 2012;337(6097):967–971. | |
Shukla R, Upton KR, Muñoz-Lopez M, et al. Endogenous retrotransposition activates oncogenic pathways in hepatocellular carcinoma. Cell. 2013;153(1):101–111. | |
Witherspoon DJ, Zhang Y, Xing J, et al. Mobile element scanning (ME-Scan) identifies thousands of novel Alu insertions in diverse human populations. Genome Res. 2013;23(7):1170–1181. | |
Zhang Y, Maksakova IA, Gagnier L, van de Lagemaat LN, Mager DL. Genome-wide assessments reveal extremely high levels of polymorphism of two active families of mouse endogenous retroviral elements. PLoS Genet. 2008;4(2):e1000007. | |
Morse B, Rotherg PG, South VJ, Spandorfer JM, Astrin SM. Insertional mutagenesis of the myc locus by a LINE-1 sequence in a human breast carcinoma. Nature. 1988;333(6168):87–90. | |
Miki Y, Nishisho I, Horii A, et al. Disruption of the APC gene by a retrotransposal insertion of L1 sequence in a colon cancer. Cancer Res. 1992;52(3):643–645. | |
Liu J, Nau MM, Zucman-Rossi J, Powell JI, Allegra CJ, Wright JJ. LINE-I element insertion at the t(11;22) translocation breakpoint of a desmoplastic small round cell tumor. Genes Chromosomes Cancer. 1997;18(3):232–239. | |
Solyom S, Ewing AD, Rahrmann EP, et al. Extensive somatic L1 retrotransposition in colorectal tumors. Genome Res. 2012;22(12):2328–2338. | |
Bundo M, Toyoshima M, Okada Y, et al. Increased l1 retrotransposition in the neuronal genome in schizophrenia. Neuron. 2014;81(2):306–313. | |
Michaud EJ, van Vugt MJ, Bultman SJ, Sweet HO, Davisson MT, Woychik RP. Differential expression of a new dominant agouti allele (Aiapy) is correlated with methylation state and is influenced by parental lineage. Genes Dev. 1994;8(12):1463–1472. | |
Druker R, Bruxner TJ, Lehrbach NJ, Whitelaw E. Complex patterns of transcription at the insertion site of a retrotransposon in the mouse. Nucleic Acids Res. 2004;32(19):5800–5808. | |
Yang Z, Boffelli D, Boonmark N, Schwartz K, Lawn R. Apolipoprotein(a) gene enhancer resides within a LINE element. J Biol Chem. 1998;273(2):891–897. | |
Girardot M, Guibert S, Laforet MP, Gallard Y, Larroque H, Oulmouden A. The insertion of a full-length Bos taurus LINE element is responsible for a transcriptional deregulation of the Normande Agouti gene. Pigment Cell Res. 2006;19(4):346–355. | |
Christy RJ, Huang RC. Functional analysis of the long terminal repeats of intracisternal A-particle genes: sequences within the U3 region determine both the efficiency and direction of promoter activity. Mol Cell Biol. 1988;8(3):1093–1102. | |
Speek M. Antisense promoter of human L1 retrotransposon drives transcription of adjacent cellular genes. Mol Cell Biol. 2001;21(6):1973–1985. | |
Thien H, Rüther U. The mouse mutation Pdn (Polydactyly Nagoya) is caused by the integration of a retrotransposon into the Gli3 gene. Mamm Genome. 1999;10(3):205–209. | |
Michel D, Chatelain G, Mauduit C, Benahmed M, Brun G. Recent evolutionary acquisition of alternative pre-mRNA splicing and 3′ processing regulations induced by intronic B2 SINE insertion. Nucleic Acids Res. 1997;25(16):3228–3234. | |
Meischl C, Boer M, Ahlin A, Roos D. A new exon created by intronic insertion of a rearranged LINE-1 element as the cause of chronic granulomatous disease. Eur J Hum Genet. 2000;8(9):697–703. | |
Sorek R, Ast G, Graur D. Alu-containing exons are alternatively spliced. Genome Res. 2002;12(7):1060–1067. | |
Han JS, Szak ST, Boeke JD. Transcriptional disruption by the L1 retrotransposon and implications for mammalian transcriptomes. Nature. 2004;429(6989):268–274. | |
Shen S, Lin L, Cai JJ, et al. Widespread establishment and regulatory impact of Alu exons in human genes. Proc Natl Acad Sci U S A. 2011;108(7):2837–2842. | |
Estécio MR, Gallegos J, Dekmezian M, Lu Y, Liang S, Issa JP. SINE retrotransposons cause epigenetic reprogramming of adjacent gene promoters. Mol Cancer Res. 2012;10(10):1332–1342. | |
Lunyak VV, Prefontaine GG, Núñez E, et al. Developmentally regulated activation of a SINE B2 repeat as a domain boundary in organogenesis. Science. 2007;317(5835):248–251. | |
Román AC, González-Rico FJ, Moltó E, et al. Dioxin receptor and SLUG transcription factors regulate the insulator activity of B1 SINE retrotransposons via an RNA polymerase switch. Genome Res. 2011;21(3):422–432. | |
Faulkner GJ, Kimura Y, Daub CO, et al. The regulated retrotransposon transcriptome of mammalian cells. Nat Genet. 2009;41(5):563–571. | |
Mätlik K, Redik K, Speek M. L1 antisense promoter drives tissue-specific transcription of human genes. J Biomed Biotechnol. 2006; 2006(1):71753. | |
Wolff EM, Byun HM, Han HF, et al. Hypomethylation of a LINE-1 promoter activates an alternate transcript of the MET oncogene in bladders with cancer. PLoS Genet. 2010;6(4):e1000917. | |
Cruickshanks HA, Vafadar-Isfahani N, Dunican DS, et al. Expression of a large LINE-1-driven antisense RNA is linked to epigenetic silencing of the metastasis suppressor gene TFPI-2 in cancer. Nucleic Acids Res. 2013;41(14):6857–6869. | |
Li J, Kannan M, Trivett AL, et al. An antisense promoter in mouse L1 retrotransposon open reading frame-1 initiates expression of diverse fusion transcripts and limits retrotransposition. Nucleic Acids Res. 2014;42(7):4546–4562. | |
Symer DE, Connelly C, Szak ST, et al. Human l1 retrotransposition is associated with genetic instability in vivo. Cell. 2002;110(3):327–338. | |
Gilbert N, Lutz-Prigge S, Moran JV. Genomic deletions created upon LINE-1 retrotransposition. Cell. 2002;110(3):315–325. | |
Han K, Sen SK, Wang J, et al. Genomic rearrangements by LINE-1 insertion-mediated deletion in the human and chimpanzee lineages. Nucleic Acids Res. 2005;33(13):4040–4052. | |
Holmes SE, Dombroski BA, Krebs CM, Boehm CD, Kazazian HH. A new retrotransposable human L1 element from the LRE2 locus on chromosome 1q produces a chimaeric insertion. Nat Genet. 1994;7(2):143–148. | |
Moran JV, DeBerardinis RJ, Kazazian HH. Exon shuffling by L1 retrotransposition. Science. 1999;283(5407):1530–1534. | |
Lin C, Yang L, Tanasa B, et al. Nuclear receptor-induced chromosomal proximity and DNA breaks underlie specific translocations in cancer. Cell. 2009;139(6):1069–1083. | |
Gasior SL, Wakeman TP, Xu B, Deininger PL. The human LINE-1 retrotransposon creates DNA double-strand breaks. J Mol Biol. 2006;357(5):1383–1393. | |
Wallace NA, Belancio VP, Deininger PL. L1 mobile element expression causes multiple types of toxicity. Gene. 2008;419(1–2):75–81. | |
Belgnaoui SM, Gosden RG, Semmes OJ, Haoudi A. Human LINE-1 retrotransposon induces DNA damage and apoptosis in cancer cells. Cancer Cell Int. 2006;6:13. | |
Zhang Z, Carriero N, Gerstein M. Comparative analysis of processed pseudogenes in the mouse and human genomes. Trends Genet. 2004;20(2):62–67. | |
Pei B, Sisu C, Frankish A, et al. The GENCODE pseudogene resource. Genome Biol. 2012;13(9):R51. | |
Sayah DM, Sokolskaja E, Berthoux L, Luban J. Cyclophilin A retrotransposition into TRIM5 explains owl monkey resistance to HIV-1. Nature. 2004;430(6999):569–573. | |
Parker HG, VonHoldt BM, Quignon P, et al. An expressed fgf4 retrogene is associated with breed-defining chondrodysplasia in domestic dogs. Science. 2009;325(5943):995–998. | |
Tam OH, Aravin AA, Stein P, et al. Pseudogene-derived small interfering RNAs regulate gene expression in mouse oocytes. Nature. 2008;453(7194):534–538. | |
Poliseno L, Salmena L, Zhang J, Carver B, Haveman WJ, Pandolfi PP. A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature. 2010;465(7301):1033–1038. | |
Hawkins PG, Morris KV. Transcriptional regulation of Oct4 by a long non-coding RNA antisense to Oct4-pseudogene 5. Transcription. 2010;1(3):165–175. | |
Ewing AD, Ballinger TJ, Earl D, et al; Broad Institute Genome Sequencing and Analysis Program and Platform. Retrotransposition of gene transcripts leads to structural variation in mammalian genomes. Genome Biol. 2013;14(3):R22. | |
Wang J, Geesman GJ, Hostikka SL, et al. Inhibition of activated pericentromeric SINE/Alu repeat transcription in senescent human adult stem cells reinstates self-renewal. Cell Cycle. 2011;10(17):3016–3030. | |
Strout MP, Marcucci G, Bloomfield CD, Caligiuri MA. The partial tandem duplication of ALL1 (MLL) is consistently generated by Alu-mediated homologous recombination in acute myeloid leukemia. Proc Natl Acad Sci U S A. 1998;95(5):2390–2395. | |
Kim SH, Bae JH, Chae JJ, et al. Long-distance PCR-based screening for large rearrangements of the LDL receptor gene in Korean patients with familial hypercholesterolemia. Clin Chem. 1999;45(9):1424–1430. | |
Puget N, Stoppa-Lyonnet D, Sinilnikova OM, et al. Screening for germ-line rearrangements and regulatory mutations in BRCA1 led to the identification of four new deletions. Cancer Res. 1999;59(2):455–461. | |
Franke G, Bausch B, Hoffmann MM, et al. Alu-Alu recombination underlies the vast majority of large VHL germline deletions: molecular characterization and genotype-phenotype correlations in VHL patients. Hum Mutat. 2009;30(5):776–786. | |
Jennes I, de Jong D, Mees K, Hogendoorn PC, Szuhai K, Wuyts W. Breakpoint characterization of large deletions in EXT1 or EXT2 in 10 multiple osteochondromas families. BMC Med Genet. 2011;12:85. | |
Han K, Lee J, Meyer TJ, et al. Alu recombination-mediated structural deletions in the chimpanzee genome. PLoS Genet. 2007;3(10):1939–1949. | |
Abeysinghe SS, Chuzhanova N, Krawczak M, Ball EV, Cooper DN. Translocation and gross deletion breakpoints in human inherited disease and cancer I: nucleotide composition and recombination-associated motifs. Hum Mutat. 2003;22(3):229–244. | |
Florl AR, Schulz WA. Peculiar structure and location of 9p21 homozygous deletion breakpoints in human cancer cells. Genes Chromosomes Cancer. 2003;37(2):141–148. | |
Rockwood LD, Felix K, Janz S. Elevated presence of retrotransposons at sites of DNA double strand break repair in mouse models of metabolic oxidative stress and MYC-induced lymphoma. Mutat Res. 2004;548(1–2):117–125. | |
Robberecht C, Voet T, Zamani Esteki M, Nowakowska BA, Vermeesch JR. Nonallelic homologous recombination between retrotransposable elements is a driver of de novo unbalanced translocations. Genome Res. 2013;23(3):411–418. | |
Brueckner LM, Sagulenko E, Hess EM, et al. Genomic rearrangements at the FRA2H common fragile site frequently involve non-homologous recombination events across LTR and L1(LINE) repeats. Hum Genet. 2012;131(8):1345–1359. | |
Rebollo R, Karimi MM, Bilenky M, et al. Retrotransposon-induced heterochromatin spreading in the mouse revealed by insertional polymorphisms. PLoS Genet. 2011;7(9):e1002301. | |
Rebollo R, Miceli-Royer K, Zhang Y, Farivar S, Gagnier L, Mager DL. Epigenetic interplay between mouse endogenous retroviruses and host genes. Genome Biol. 2012;13(10):R89. | |
Brosius J, Gould SJ. On “genomenclature”: a comprehensive (and respectful) taxonomy for pseudogenes and other “junk DNA”. Proc Natl Acad Sci U S A. 1992;89(22):10706–10710. | |
Rebollo R, Romanish MT, Mager DL. Transposable elements: an abundant and natural source of regulatory sequences for host genes. Annu Rev Genet. 2012;46:21–42. | |
Allen TA, Von Kaenel S, Goodrich JA, Kugel JF. The SINE-encoded mouse B2 RNA represses mRNA transcription in response to heat shock. Nat Struct Mol Biol. 2004;11(9):816–821. | |
Espinoza CA, Allen TA, Hieb AR, Kugel JF, Goodrich JA. B2 RNA binds directly to RNA polymerase II to repress transcript synthesis. Nat Struct Mol Biol. 2004;11(9):822–829. | |
Mariner PD, Walters RD, Espinoza CA, et al. Human Alu RNA is a modular transacting repressor of mRNA transcription during heat shock. Mol Cell. 2008;29(4):499–509. | |
Chu WM, Ballard R, Carpick BW, Williams BR, Schmid CW. Potential Alu function: regulation of the activity of double-stranded RNA-activated kinase PKR. Mol Cell Biol. 1998;18(1):58–68. | |
Chuong EB, Rumi MA, Soares MJ, Baker JC. Endogenous retroviruses function as species-specific enhancer elements in the placenta. Nat Genet. 2013;45(3):325–329. | |
Schmidt D, Schwalie PC, Wilson MD, et al. Waves of retrotransposon expansion remodel genome organization and CTCF binding in multiple mammalian lineages. Cell. 2012;148(1–2):335–348. | |
Lane N, Dean W, Erhardt S, et al. Resistance of IAPs to methylation reprogramming may provide a mechanism for epigenetic inheritance in the mouse. Genesis. 2003;35(2):88–93. | |
Smith ZD, Chan MM, Mikkelsen TS, et al. A unique regulatory phase of DNA methylation in the early mammalian embryo. Nature. 2012;484(7394):339–344. | |
Walsh CP, Chaillet JR, Bestor TH. Transcription of IAP endogenous retroviruses is constrained by cytosine methylation. Nat Genet. 1998;20(2):116–117. | |
Okano M, Bell DW, Haber DA, Li E. DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell. 1999;99(3):247–257. | |
Liang G, Chan MF, Tomigahara Y, et al. Cooperativity between DNA methyltransferases in the maintenance methylation of repetitive elements. Mol Cell Biol. 2002;22(2):480–491. | |
Bourc’his D, Bestor TH. Meiotic catastrophe and retrotransposon reactivation in male germ cells lacking Dnmt3L. Nature. 2004;431(7004):96–99. | |
Mikkelsen TS, Ku M, Jaffe DB, et al. Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature. 2007;448(7153):553–560. | |
Matsui T, Leung D, Miyashita H, et al. Proviral silencing in embryonic stem cells requires the histone methyltransferase ESET. Nature. 2010;464(7290):927–931. | |
Leung DC, Lorincz MC. Silencing of endogenous retroviruses: when and why do histone marks predominate? Trends Biochem Sci. 2012;37(4):127–133. | |
Leung D, Du T, Wagner U, et al. Regulation of DNA methylation turnover at LTR retrotransposons and imprinted loci by the histone methyltransferase Setdb1. Proc Natl Acad Sci U S A. 2014;111(18):6690–6695. | |
Rangasamy D. Distinctive patterns of epigenetic marks are associated with promoter regions of mouse LINE-1 and LTR retrotransposons. Mob DNA. 2013;4(1):27. | |
Montoya-Durango DE, Liu Y, Teneng I, et al. Epigenetic control of mammalian LINE-1 retrotransposon by retinoblastoma proteins. Mutat Res. 2009;665(1–2):20–28. | |
Yang N, Kazazian HH. L1 retrotransposition is suppressed by endogenously encoded small interfering RNAs in human cultured cells. Nat Struct Mol Biol. 2006;13(9):763–771. | |
Svoboda P, Stein P, Anger M, Bernstein E, Hannon GJ, Schultz RM. RNAi and expression of retrotransposons MuERV-L and IAP in preimplantation mouse embryos. Dev Biol. 2004;269(1):276–285. | |
Heras SR, Macias S, Plass M, et al. The Microprocessor controls the activity of mammalian retrotransposons. Nat Struct Mol Biol. 2013;20(10):1173–1181. | |
Chuma S, Nakano T. piRNA and spermatogenesis in mice. Philos Trans R Soc Lond B Biol Sci. 2013;368(1609):20110338. | |
Aravin AA, Sachidanandam R, Girard A, Fejes-Toth K, Hannon GJ. Developmentally regulated piRNA clusters implicate MILI in transposon control. Science. 2007;316(5825):744–747. | |
Carmell MA, Girard A, van de Kant HJ, et al. MIWI2 is essential for spermatogenesis and repression of transposons in the mouse male germline. Dev Cell. 2007;12(4):503–514. | |
Kuramochi-Miyagawa S, Watanabe T, Gotoh K, et al. DNA methylation of retrotransposon genes is regulated by Piwi family members MILI and MIWI2 in murine fetal testes. Genes Dev. 2008;22(7):908–917. | |
Reuter M, Berninger P, Chuma S, et al. Miwi catalysis is required for piRNA amplification-independent LINE1 transposon silencing. Nature. 2011;480(7376):264–267. | |
Di Giacomo M, Comazzetto S, Saini H, et al. Multiple epigenetic mechanisms and the piRNA pathway enforce LINE1 silencing during adult spermatogenesis. Mol Cell. 2013;50(4):601–608. | |
Marchetto MC, Narvaiza I, Denli AM, et al. Differential L1 regulation in pluripotent stem cells of humans and apes. Nature. 2013;503(7477):525–529. | |
Maksakova IA, Zhang Y, Mager DL. Preferential epigenetic suppression of the autonomous MusD over the nonautonomous ETn mouse retrotransposons. Mol Cell Biol. 2009;29(9):2456–2468. | |
Ichiyanagi K, Li Y, Li Y, et al. Locus- and domain-dependent control of DNA methylation at mouse B1 retrotransposons during male germ cell development. Genome Res. 2011;21(12):2058–2066. | |
Koito A, Ikeda T. Intrinsic immunity against retrotransposons by APOBEC cytidine deaminases. Front Microbiol. 2013;4:28. | |
Esnault C, Heidmann O, Delebecque F, et al. APOBEC3G cytidine deaminase inhibits retrotransposition of endogenous retroviruses. Nature. 2005;433(7024):430–433. | |
Bogerd HP, Wiegand HL, Doehle BP, Lueders KK, Cullen BR. APOBEC3A and APOBEC3B are potent inhibitors of LTR- retrotransposon function in human cells. Nucleic Acids Res. 2006;34(1):89–95. | |
Kinomoto M, Kanno T, Shimura M, et al. All APOBEC3 family proteins differentially inhibit LINE-1 retrotransposition. Nucleic Acids Res. 2007;35(9):2955–2964. | |
Ikeda T, Abd El Galil KH, Tokunaga K, et al. Intrinsic restriction activity by apolipoprotein B mRNA editing enzyme APOBEC1 against the mobility of autonomous retrotransposons. Nucleic Acids Res. 2011;39(13):5538–5554. | |
MacDuff DA, Demorest ZL, Harris RS. AID can restrict L1 retrotransposition suggesting a dual role in innate and adaptive immunity. Nucleic Acids Res. 2009;37(6):1854–1867. | |
Metzner M, Jäck HM, Wabl M. LINE-1 retroelements complexed and inhibited by activation induced cytidine deaminase. PLoS One. 2012;7(11):e49358. | |
Horn AV, Klawitter S, Held U, et al. Human LINE-1 restriction by APOBEC3C is deaminase independent and mediated by an ORF1p interaction that affects LINE reverse transcriptase activity. Nucleic Acids Res. 2014;42(1):396–416. | |
Palazzo AF, Gregory TR. The case for junk DNA. PLoS Genet. 2014;10(5):e1004351. |
 © 2014 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2014 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.