Back to Journals » OncoTargets and Therapy » Volume 13
Circular RNA MYLK Promotes Hepatocellular Carcinoma Progression Through the miR29a/KMT5C Signaling Pathway
Authors Gao J , Li E, Liu W, Yang Q, Xie C , Ai J, Zhou F, Liao W, Wu L
Received 18 April 2020
Accepted for publication 4 August 2020
Published 25 August 2020 Volume 2020:13 Pages 8615—8627
DOI https://doi.org/10.2147/OTT.S258715
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Prof. Dr. Nicola Silvestris
Jun Gao,1,* Enliang Li,1,* Weiwei Liu,1,* Qingping Yang,2 Chunyan Xie,3 Jiyuan Ai,1 Fan Zhou,1 Wenjun Liao,1 Linquan Wu1
1Department of Hepatobiliary and Pancreatic Surgery, The Second Affiliated Hospital of Nanchang University, Nanchang, Jiangxi, People’s Republic of China; 2Department of Assisted Reproductive, The First Affiliated Hospital of Nanchang University, Nanchang, Jiangxi, People’s Republic of China; 3Jiangxi Medical College, Nanchang University, Nanchang, Jiangxi 330006, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Linquan Wu; Wenjun Liao
Department of Hepatobiliary and Pancreatic Surgery, The Second Affiliated Hospital of Nanchang University, No. 1, Minde Road, Nanchang, Jiangxi 330006, People’s Republic of China
Tel +86 791 86311529
Fax +86 791 86262262
Email [email protected]; [email protected]
Purpose: This study aimed to investigate the functions of the circular RNA circMYLK (hsa_circ_0002768) in the development of hepatocellular carcinoma (HCC) and to identify the underlying mechanisms of the circMYLK/miR29a/KMT5C axis.
Materials and Methods: Quantitative real-time polymerase chain reaction (qRT-PCR) was utilized to explore the expressions of circMYLK, miR-29a and KMT5C in HCC tissues and cells. A potential miRNA (miR-29a) regulated by circMYLK was also explored, and the target relationship between miR-29a and KMT5C was confirmed. FISH, qRT-PCR, Western blotting, and dual-luciferase reporter assays were used to examine the circMYLK/miR29a/KMT5C signaling pathways involved in HCC development. Additionally, HCC cells were implanted into nude mice subcutaneously to test the role of circMYLK in tumor growth.
Results: circMYLK was determined to be significantly upregulated in HCC tissues and cells. Suppression of circMYLK repressed HCC cell proliferation, migration, and invasion while increasing apoptosis. In addition, FISH, qRT-PCR, and Western blotting, as well as dual-luciferase reporter assays, revealed that circMYLK could bind to miR-29a. In rescue experiments, miR-29a had the potential to eliminate the inhibitory effect of circMYLK knockdown in HCC. Moreover, miR-29a was found to target the KMT5C gene, which was positively regulated by circMYLK. Finally, a nude mouse tumorigenicity assay showed that injection of circMYLK siRNA into nude mice drastically suppressed xenograft tumor formation in vivo.
Conclusion: Our current study demonstrated that circMYLK promotes HCC progression by acting as a competing endogenous RNA of miR-29a, which regulates the downstream oncogene KMT5C.
Keywords: hepatocellular carcinoma, circMYLK, miR-29a, KMT5C
Introduction
Hepatocellular carcinoma (HCC) is a common malignant gastrointestinal cancer and ranks fifth in terms of global cases and second in terms of deaths for males.1 The prognosis of HCC is unfavorable, and HCC often presents at a high recurrence rate due to the high frequency of intrahepatic and extrahepatic metastases.2 Despite many different known therapeutic interventions, including chemotherapy and surgery, the clinical efficacy and prognosis of treatments are not high.1,3,4 Therefore, elucidating the molecular mechanisms that drive HCC development is a necessity and is crucial for the development of better strategies and therapeutic methods for this disease.
Circular RNAs (circRNAs) are noncoding RNAs with a continuous closed loop structure due to the joining of their 5′ and 3′ RNA ends in a covalent manner.5 CircRNAs are tissue-specific, conserved, abundant, and normally stable.6 Multiple circRNAs have been found to be abnormally expressed in tumor tissues and participate in cancer progression and metastasis.7,8 Recent studies have shown that circRNAs can regulate gene expression by acting as microRNA (miRNA) sponges, transcription regulators, and even RNA-binding protein sequestering agents.9 However, it is well known that the ceRNA system represents the main means through which circRNAs achieve their biological functions. For example, CircFAT1 promotes invasion and tumorigenesis of osteosarcoma via sequestration of miR-375.10 Hsa_circ_001783 regulates breast cancer by modulating miR‐200c, ZEB1/2 or ETS1.11 CircSMARCA5 acts as a potential biomarker in the diagnosis of glioblastoma multiforme.12 CircMTO1 promotes HCC proliferation, invasion and metastasis by directly targeting miR‐9.13 Moreover, a recent study indicated that circMYLK is upregulated in HCC.14 Interestingly, through bioinformatics analysis, we found that circMYLK may specifically bind to miR-29a in HCC. miRNAs are non-coding small RNAs that can bind to the 3ʹUTR of an mRNA, thereby inhibiting the translation of this mRNA or reducing its stability.15,16 Studies have shown that various miRNAs play a vital role in the progression of HCC by regulating cell proliferation, invasion, and apoptosis.17,18 Our previous study indicated that miR-29a was down-regulated in HCC and further revealed that miR-29a could repress HCC cell growth and invasion.19 Moreover, bioinformatics analysis found that Histone lysine N methyltransferase 5C (KMT5C) was a target gene of miR-29a. Studies have demonstrated that KMT5C plays an important role in human cancer and is correlated with the progression of gastrointestinal tumors.20 Based on these studies, we have explored here the potential effects of circMYLK in HCC development and examined whether circMYLK affects HCC progression through the miR29a/KMT5C signaling pathway.
Materials and Methods
Clinical Samples
The Institutional Ethics Committee of the Second Affiliated Hospital of Nanchang University in China approved all the experiments that involved the use of humans as subjects. All experiments were conducted in accordance with the Helsinki Declaration of 1964. All patients received and signed a written informed consent prior to participation. Tumor tissues and associated paracancerous tissue specimens were collected from 60 HCC patients who underwent surgical treatment at the Second Affiliated Hospital of Nanchang University between the years 2015 and 2017. All samples were pathologically confirmed, and the patients did not undergo immunotherapy, chemotherapy, or radiotherapy before surgery. After collection, the samples were immediately preserved in liquid nitrogen at −80°C. Clinical data for the patients are summarized in Supplementary Table S1.
Cell Culture
HCC cell lines (MHCC-97H, HCC-LM3) and the human normal liver cell line HL-7702 were procured from the Shanghai Institute of Cell Biology (Shanghai, China). Cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM) containing 10% of fetal bovine serum (FBS) [Biological Industries, Israel] and maintained in an incubator with 5% CO2 at 37°C.
Cell Transfection
MHCC-97H and HCC-LM3 cell lines were transfected with corresponding plasmids and microRNA inhibitors (RiboBio, Guangzhou, China) using Lipofectamine 3000 (Invitrogen, Thermo Fisher Scientific, Inc., USA) according to the manufacturer’s protocol. The cells were harvested for further evaluation 48 hours post-transfection. The siRNA sequences used are listed in Supplementary Table S2.
qRT-PCR and Western Blotting
Quantitative RT-PCR was conducted following methods described previously.19 The 2−ΔΔCt method was used for the calculation of relative mRNA levels. Primers for U6, circMYLK, miR-29a, and glyceraldehyde-3-phosphate dehydrogenase (GAPDH) were synthesized by RiboBio (Primer sequences for qRT-PCR are shown in Supplementary Table S3). For Western blotting, cells were treated with radioimmunoprecipitation assay buffer (RIPA) [Solarbio, Beijing, China], and cell lysates were separated via 10% SDS-PAGE. Blocking was performed using 5% Difco™ Skim Milk, and membranes were subsequently incubated with primary antibodies (targeting KMT5C or GAPDH; Abcam). Finally, horseradish peroxidase-conjugated secondary antibody (TransGen Biotech, Beijing, China) was added, and protein levels were normalized to total GAPDH levels.
Cell Proliferation Assay
A CCK8 kit was used for the analysis of cell viability. Briefly, transfected cells (5×103 cells/well) were seeded into 96‐well plates and cultured in complete medium for different times. Cells were then prepared using CCK8 reagents (Dojindo Molecular Technologies, Tokyo, Japan), and their density was measured after different times using a microplate reader (ELx808) at an absorbance of 450 nm.
Cell Wound Healing Assay
The cell migration ability was evaluated by seeding MHCC-97H and HCC-LM3 cells in 6-well plates, followed by incubation at 37°C in DMEM supplemented with 10% FBS. After cells reached 90–100% confluence, a 200-μL pipette tip was used to generate small artificial wounds over the cell surface. The cell debris was washed three times with phosphate-buffered saline (PBS), and then the cells were cultured in fresh medium for 24 hours. After that, micrographs of cell migration areas were captured and wound areas were compared.
Transwell Assay
Transwell chambers (Corning Inc., Corning, USA) with 8-μm pores were used to assess the migration and invasion abilities of cells. The cells were reconstituted in a serum-free medium and then added to the upper chamber in the presence or absence of Matrigel 24 hours post-transfection. The lower chambers were filled with 10% FBS-containing medium, and then the cells were cultured for 24 hours. Cells from the top chamber were harvested and fixed in polyoxymethylene, followed by staining with 0.1% crystal violet. Examination and counting of the cells was conducted using a microscope.
Flow Cytometry Analysis
Cell apoptosis was assessed via flow cytometry according to the manufacturer’s instructions. To achieve this, cells were first dissociated with trypsin and then collected and washed twice with PBS. Next, the cells were stained with 5 µL of Annexin V-FITC and 5 µL of a Propidium Iodide (PI) solution and incubated in the dark at room temperature. Subsequently, the cell apoptosis rate was analyzed using a flow cytometer (BD Biosciences).
5-Ethynyl-2′-Deoxyuridine (EdU) Assay
The rate of cell proliferation was determined using an EdU test (RiboBio), which was performed via fluorescence-activated cell sorting.
Tumorigenicity Assay
In vivo tumorigenicity tests were performed by suspending HCC-LM3 cells (1x106 cells/100 μL) in phosphate-buffered saline. The cells were subsequently injected (subcutaneously) into the right flank of 6-week-old nude mice. A caliper was used to measure the tumor size at three-day intervals based on the following formula: Tumor volume = (shortest diameter 2 × longest diameter)/2. After five weeks, all the nude mice were euthanized, and the tumors were extracted and weighed. To confirm lung metastasis, 1×106 cells/100 μL PBS were injected into nude mice through the tail vein. The mice were then sacrificed after four weeks, and the lungs were processed and embedded in paraffin. All animal experiment procedures were carried out following the ethical standards under a protocol approved by the Animal Experimental Ethics Committee of the Second Affiliated Hospital of Nanchang University, and were executed conforming to the “Guide for the Care and Use of Laboratory Animals” (revised 1985).
Luciferase Reporter Assay
Dual-luciferase reporter gene assays were performed to determine the binding sites of circMYLK and miR-29a and between miR-29a and the 3′-UTR of KMT5C. RiboBio designed the mutant-type MYLK (Mut-circMYLK) and wild-type circMYLK (Wt-MYLK) vectors. Using Lipofectamine 3000, 293T cells were co-transfected with a mimic control or miR-29a mimic. Luciferase activity was determined using a Dual-Luciferase Assay System (Promega Corp., Madison, WI, USA) 48 hours post transfection. Furthermore, the effect of miR-29a expression on the luciferase activity of the 3′-UTR of KMT5C was determined as described above, and the relative luciferase activity was standardized to that of Renilla luciferase.
Fluorescence in situ Hybridization (FISH)
Fluorescence in situ hybridization (FISH) experiments were conducted using a Cy5-labeled probe (5ʹ-DIG-ACATCCCCCATGGTCTTCTACTGTCAACT-3ʹ) to examine the expression of circMYLK and a Cy3-labeled probe (5ʹ-DIG-TAACCGATTTCAGATGGTGCTA-3ʹ) to examine miR-29a expression. After prehybridization (1×PBS/0.5% Triton X-100), samples were hybridized in hybridization buffer (1000 mg/mL sheared salmon sperm DNA, 1000 mg/mL yeast transfer RNA, 10 mM DDT, 4×SSC, 1×Denhardt’s solution, 10% dextran sulfate, and 40% formamide) with specific probes at 60°C overnight and then imaged.
Hematoxylin and Eosin (H&E) Staining, Immunohistochemistry, and Immunofluorescence Assays
After specimens were fixed with 10% formalin, they were embedded in paraffin blocks and sectioned. Subsequently, some sections were deparaffinized and stained with H&E (Hematoxylin and Eosin) for morphological examination, whereas others were immunostained. For immunohistochemistry and immunofluorescence assays, after being dewaxed, dehydrated, and rehydrated, the sections were treated with 5 μg/mL anti-KMT5C antibody (ab91224; Abcam) overnight at 4°C. Then, a biotinylated secondary antibody (TransGen Biotech, Beijing, China) was applied based on an SP-IHC assay. All experiments were performed according to the methods described previously.21
Statistical Analyses
Data were analyzed using GraphPad Prism 7.0 (San Diego, CA) and SPSS 22.0 (SPSS, Inc., Chicago, IL) software. Measurement data are reported as the mean ± the standard deviation. A Student’s t-test was used to compare the means of two groups, whereas analysis of variance (ANOVA) was used to compare the means of three or more groups. A comparison with P < 0.05 was considered statistically significant.
Results
circMYLK Overexpressed in HCC Tissues and Cell Lines
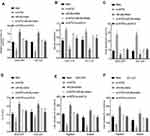
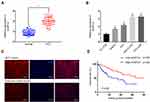
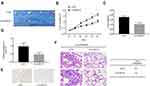
To assess the expression levels and functions of circMYLK in HCC, 60 HCC specimens and adjacent non-tumor tissues were analyzed via qRT-PCR. The results indicated that circMYLK expression was significantly higher in HCC tissues compared to paired normal specimens (Figure 1A). Similarly, qRT-PCR revealed that the expression of circMYLK in four HCC cell lines (HepG2, PLC, MHCC-97H, HCC-LM3) was higher than the human normal liver cell line HL-7702, and the two HCC cell lines with the highest expression of circMYLK were selected for follow-up experiments (Figure 1B). In addition, a dose–response experiment was conducted to determine the amount of RNA circMYLK that could achieve the maximum effect, and we then used this optimal dose to transfect liver cancer cells (Supplementary Figure 1A–B). Next, we conducted a FISH experiment, which also revealed that circMYLK was overexpressed in HCC tissues relative to non-tumor samples (Figure 1C; Supplementary Figure 1C). In addition, the expression of circMYLK was closely associated with clinical features, including tumor size, differentiation, and tumor-node-metastasis classification (Supplementary Table S1). Finally, a Kaplan-Meier survival curve was plotted based on each patient’s follow-up situation, which showed that elevated circMYLK expression was related to poor overall survival rate in HCC patients (Figure 1D). Generally speaking, then, circMYLK was overexpressed in HCC tissue samples and cell lines.
circMYLK Promotes the Development and Progression of HCC
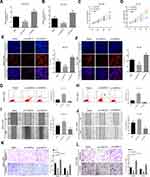
The effects of circMYLK on HCC cell proliferation, migration, invasion, and apoptosis were next determined by transfecting HCC cell lines with circMYLK siRNA (si-circMYLK) or a circMYLK overexpression vector (p-circMYLK). Subsequently, qRT-PCR was performed 48 hours post transfection to assess the transfection efficiency of p-circMYLK and si-circMYLK in HCC cell lines (Figure 2A and B). CCK8 and EdU assays showed that silencing of circMYLK led to a significant inhibition of cell viability and proliferation, whereas circMYLK overexpression significantly promoted cell viability and proliferation (Figure 2C–F). Flow cytometry indicated that the apoptotic rate increased with suppression of circMYLK, while the apoptotic rate decreased with circMYLK overexpression (Figure 2G–H). Wound healing and transwell assays demonstrated that circMYLK knockdown hindered HCC cell migration and invasion, while circMYLK overexpression promoted HCC cell migration and invasion (Figure 2I–L). These results revealed that circMYLK was involved in and facilitated the progression of HCC.
circMYLK Specifically Binds to miR-29a
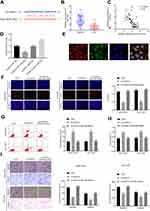
According to miRbase, there was a miR-29a seed region within the circMYLK complementary sequence (Figure 3A). Subsequently, qRT-PCR results from 60 pairs of HCC tissues and adjacent normal tissues demonstrated that miR-29a expression was downregulated in HCC tissues (Figure 3B). Correlation analysis revealed that there was a significant negative correlation between miR-29a and circMYLK (Figure 3C). Dual luciferase reporter gene assays indicated that circMYLK (WT) was blocked by a miR-29a mimic, whereas no similar reduction in luciferase activity was observed with the circMYLK 3′-UTR after mimic NC treatment, suggesting that circMYLK could competitively bind to miR-29a (Figure 3D). Additionally, FISH analysis revealed that circMYLK and miR-29a were colocalized in the cytoplasm of HCC cells (Figure 3E). qRT-PCR results showed that in both cell lines, circMYLK knockdown led to a significant increase in miR-29a expression, while circMYLK overexpression displayed the exact opposite phenotype (Supplementary Figure 2A). Next, we performed qRT-PCR to verify the transfection efficiency of our miR-29a inhibitor in MHCC-97H and HCC-LM3 cell lines. MiR-29a expression was significantly reduced in both cell lines, indicating that it had good transfection efficiency (Supplementary Figure 2B). To determine whether circMYLK promoted HCC progression via interaction with miR-29a, rescue experiments were conducted by co-transfection with a miR-29a inhibitor and circMYLK siRNA. Edu assays indicated that the si-circMYLK+miR29a inhibitor group showed a higher growth rate than the si-circMYLK group, and no apparent differences were observed in comparison to the si-NC group (Figure 3F). In addition, compared with the si-NC group, anoikis in the si-circMYLK+miR-29a inhibitor group was not significantly different (Figure 3G). Wound healing, migration, and invasion assays revealed that HCC cells in the si-circMYLK+miR-29a inhibitor group exhibited faster migration and greater invasion ability than those in the si-circMYLK group, and no apparent differences were observed in comparison with the si-NC group (Figure 3H–I; Supplementary Figure 2C). Thus, the results demonstrated that circMYLK can specifically bind to miR-29a.
circMYLK Regulates KMT5C Expression via miR-29a
The downstream target genes of miR-29a were predicted using three ellipses: TargetScan, microT, and Pictar, and the results are presented in a Venn diagram. Target genes were found in the intersection of the three databases (Figure 4A). It is worth noting that because KMT5C is related to the metastasis of digestive tract cancer,20 and its function in HCC remains unclear, it was selected for further analysis. As per the TCGA database, date obtained demonstrated that KMT5C was highly expressed in HCC (Figure 4B). Subsequently, we conducted a dual luciferase reporter gene assay to confirm the regulatory relationship between miR‑29a and KMT5C. Our results showed that miR‑29a mimics could reduce the luciferase activities of the wild‑type KMT5C reporter vector (KMT5C‑WT) rather than a mutant KMT5C reporter vector (KMT5C‑MUT), indicating that KMT5C was the target gene of miR‑29a (Figure 4C). Additionally, immunofluorescence assay revealed that KMT5C was overexpressed in HCC tissues (Figure 4D; Supplementary Figure 3A). Moreover, a total of 60 HCC specimens and adjacent non-tumor tissues were subjected to qRT-PCR, and the results indicated increased expression levels of KMT5C mRNA in HCC tissues (Figure 4E). Next, Western blot analysis was carried out to detect the protein expression levels of KMT5C in a human normal liver cell line and two HCC cell lines. Not surprisingly, the results revealed that the KMT5C protein expression levels in HCC cell lines were higher than that in our human normal liver cell line (Supplementary Figure 3B). Correlation analysis showed that KMT5C was negatively correlated with miR-29a expression (Figure 4F). In addition, there was a remarkable positive correlation between KMT5C and circMYLK expression in HCC (Figure 4G). Finally, Western blot analysis was used to assess changes in KMT5C protein quantity after circMYLK knockdown or a reduction in miR-29a expression, and the results indicated that a miR-29a inhibitor increased the elevation of KMT5C protein, while si-circMYLK decreased the KMT5C protein level through a sponge effect on miR‐29a (Figure 4H). The obtained findings indicated that circMYLK regulated KMT5C expression via miR-29a.
Role of KMT5C and miR‑29a in HCC Progression Stimulated by circMYLK
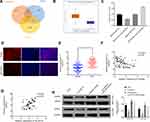
We next turned to explore how circMYLK exerts its biological function in HCC. Si-KMT5C, miR-29a inhibitor and p-circMYLK were transfected into HCC cell lines, and we then divided the cells into five transfection groups: Blank, si-KMT5C, miR-29a inhibitor, si-KMT5C+miR-29a inhibitor, and si-KMT5C+p-circMYLK. Moreover, the expression of KMT5C was examined through qRT-PCR experiments. In cells transfected with miR-29a inhibitors, or p-circMYLK, the mRNA level of KMT5C was upregulated in MHCC-97H and HCC-LM3 cells (Figure 5A). Wound healing assays showed that silencing of KMT5C significantly inhibited cell migration in HCC cell lines, whereas a miR-29a inhibitor enhanced cell migration and weakened the inhibitory effect of si-KMT5C on cell migration. In addition, upregulation of circMYLK eliminated the inhibitory effect of si-KMT5C on cell migration (Figure 5B; Supplementary Figure 4A). Flow cytometry results showed that si-KMT5C increased the apoptotic rate of HCC cells, while miR-29a inhibitor resulted in decreased apoptosis. In addition, miR-29a inhibitor eliminated the role of si-KMT5C in HCC cells and decreased the level of cell apoptosis. Up-regulation of circMYLK in HCC cells further abolished the role of si-KMT5C on cell apoptosis (Figure 5C; Supplementary Figure 4B). EdU assays demonstrated that HCC cell growth was significantly reduced in this si-KMT5C group, while the miR-29a inhibitor group showed increased cell proliferation. miR-29a inhibitor restrained the effect of si-KMT5C on cell proliferation. Moreover, overexpression of circMYLK in HCC cells eliminated the effect of si-KMT5C on cell proliferation (Figure 5D; Supplementary Figure 4C). Finally, migration and invasion assays yielded the same results, based on the number of migratory and invasive cells (Figure 5E–F; Supplementary Figure 4D). Thus, our study results revealed that circMYLK competitively combines with miR-29a to upregulate the expression of KMT5C to promote HCC progression.
circMYLK Knockdown Suppresses the Tumorigenicity of HCC Cells in vivo
The effect of circMYLK in vivo was determined by stably transducing HCC-LM3 cells with negative control (si-NC) and si-circMYLK, followed by subcutaneous administration in nude mice. The results show that injection of si-circMYLK into mice drastically repressed the tumorigenicity of HCC cells (Figure 6A–B). Evaluation of tumor growth 35 days after administration showed that si-circMYLK diminished the average tumor weight relative to negative controls (Figure 6C). qRT-PCR analysis further revealed that circMYLK expression in tumors from the group administered si-circMYLK was significantly reduced compared to that in NC group tumors (Figure 6D). Moreover, immunohistochemical staining demonstrated that si-circMYLK reduced KMT5C expression in nude mouse tumor tissues (Figure 6E). In addition, si-circMYLK decreased lung metastasis in nude mice relative to mice in the control group (Figure 6F). These results show that silencing of circMYLK reduced KMT5C expression and suppresses xenograft tumor formation in vivo.
Discussion
HCC is one of the most common malignant tumors throughout the world.22 There are few noticeable symptoms in the early stages of HCC. When symptoms emerge, patients are often already at an advanced disease stage and thus have lost the opportunity for radical surgery.23 In addition, HCC is not very sensitive to radiotherapy or chemotherapy and has a poor prognosis.24 If the pathogenesis of HCC can be elucidated, a great progress can be made in liver cancer treatment. Our current study aimed to identify the role of circMYLK in regulating miR-29a and KMT5C in HCC progression. In general, our study revealed that suppression of circMYLK upregulated miR-29a to repress HCC cell proliferation, migration, and invasion abilities and promoted apoptosis through the inhibition of KMT5C in HCC.
circMYLK, derived from the MYLK gene, is among several circRNAs that have been implicated in malignant tumor development.25,26 Previous studies have revealed that circMYLK is upregulated and plays key roles in some solid tumors, such as bladder cancer and prostate cancers.25,27 The initial results obtained from our current study indicated that circMYLK was highly expressed in HCC specimens and cell lines and was related to the clinical features of HCC. With regard to function, upregulation of circMYLK enhanced HCC cell migration, proliferation, and invasion and subsequently inhibited apoptosis as evidenced by cell functional tests. Studies have reported that circRNAs acting as competing endogenous RNAs for miRNAs are a prime example of a cancer development mechanism. circRNAs are ncRNAs with closed-loop structures28 and are widely involved in carcinogenesis and cancer progression, including in HCC.29,30 For instance, Liu et al10 reported that CircFAT1 is anomalously transcribed in osteosarcoma cell lines and tissues and is closely related to cell metastasis and invasion in osteosarcoma. In addition, Han et al13 demonstrated that circMTO1 was an ideal biomarker in HCC through the regulation of miR-9. Similarly, Yu et al31 reported that circ-cSMARCA5 was a competing endogenous RNA of miR-17-3P and miR-181b-5p, thereby greatly decreasing HCC cell migration and proliferation. The results of the current study demonstrated that circMYLK influenced KMT5C levels through miR-29a, indicating that circMYLK was an oncogene involved in HCC pathogenesis and development.
Another significant finding of the present study was that circMYLK functioned as a ceRNA to regulate KMT5C via sequestration of miR-29a in HCC. Various studies have reported that miR-29a is intensively involved in different types of tumors, including breast cancer32 and Burkitt lymphoma,33 as well as some solid tumors.34,35 In addition, other reports have also indicated aberrant function and expression of miR-29a in HCC. For instance, Mahati et al36 showed that miR-29a was decreased in HCC and functioned as a tumor suppressor by inhibiting HCC cell migration and growth through regulation of CLDN1. Similarly, Xu et al37 demonstrated that miR-29a acts as a tumor suppressor gene in metastatic HCC through regulation of SPARC. Our previous studies have shown that miR-29a reduced migration, proliferation and invasion of HCC via IFITM3.19 This was corroborated by the findings of the current study, which showed that miR-29a inhibited HCC progression by inhibiting KMT5C. Histone lysine N methyltransferase 5C (KMT5C), also referred to as suppressor of variegation 4–20 homolog 2 (SUV420H2), plays an important role in human cancer and is highly associated with invasive activity.38 A number of studies have demonstrated the role played by KMT5C in tumorigenicity using both in vitro and in vivo assays. In human pancreatic cancer, it was shown that KMT5C was highly expressed in tumor tissues compared with adjacent normal tissues.20 Furthermore, the epigenetic orchestrator activity of KMT5C is fundamental to pancreatic cancer development because it maintains epithelial/mesenchymal cell states.39 Apart from this, KMT5C is known to exert a vital impact on the motility and invasiveness of breast cancer cells, further emphasizing its potential as a target for breast cancer therapy.40 Our findings further indicate that circMYLK rescued miR-29a-induced suppression of the KMT5C gene. Thus, it is reasonable to conclude that circMYLK promotes HCC progression through the miR29a/KMT5C signaling pathway.
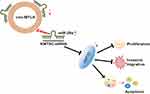
In conclusion, the current study suggests that circMYLK is upregulated in HCC tissues and cell lines. Suppression of circMYLK restores miR-29a to repress HCC cell proliferation, migration, and invasion while increasing apoptosis via down-regulation of KMT5C levels (Figure 7), suggesting that the circMYLK/miR-29a/KMT5C axis has potential as a treatment target for HCC and should be explored. However, further research is needed in other HCC cell lines to understand the regulatory mechanisms underlying the occurrence and development of HCC.
Acknowledgments
This research was funded by the National Natural Science Foundation of China (No. 91180335) and the Key foundation of Jiangxi Provincial Science and Technology Department (No. 20181BBG70025).
Author Contributions
All authors contributed to data analysis, drafting or revising the article, have agreed on the journal to which the article will be submitted, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Disclosure
The authors report no conflicts of interest for this work.
References
1. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global Cancer Statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. doi:10.3322/caac.21492
2. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2017. CA Cancer J Clin. 2017;60(5):277–300.
3. Toshiya O, Hisashi I, Kazuma O, Yukihito K, Teruhisa S, Eigo O. Clinicopathologic features and risk factors for extrahepatic recurrences of hepatocellular carcinoma after curative resection. World J Surg. 2012;36(1):136–143. doi:10.1007/s00268-011-1317-y
4. Yang Y, Hiroaki N, Hideo O, et al. Patterns and clinicopathologic features of extrahepatic recurrence of hepatocellular carcinoma after curative resection. Surgery. 2007;141(2):196–202. doi:10.1016/j.surg.2006.06.033
5. Chen L, Yang L. Regulation of circRNA biogenesis. RNA Biol. 2015;12(4):381–388. doi:10.1080/15476286.2015.1020271
6. Xiao M, Ai Y, Wilusz J. Biogenesis and functions of circular RNAs come into focus. Trends Cell Biol. 2020;30(3):226–240. doi:10.1016/j.tcb.2019.12.004
7. Zhong Y, Du Y, Yang X, et al. Circular RNAs function as ceRNAs to regulate and control human cancer progression. Mol Cancer. 2018;17(1):79. doi:10.1186/s12943-018-0827-8
8. Salzman J, Chen RE, Olsen MN, Wang PL, Brown PO, Moran JV. Cell-type specific features of circular RNA expression. PLoS Genet. 2013;9(9):e1003777. doi:10.1371/journal.pgen.1003777
9. Qu S, Yang X, Li X, et al. Circular RNA: A new star of noncoding RNAs. Cancer Lett. 2015;365(2):141–148. doi:10.1016/j.canlet.2015.06.003
10. Liu G, Huang K, Jie Z, et al. CircFAT1 sponges miR-375 to promote the expression of Yes-associated protein 1 in osteosarcoma cells. Mol Cancer. 2018;17(1):170. doi:10.1186/s12943-018-0917-7
11. Liu Z, Zhou Y, Liang G, et al. Circular RNA hsa_circ_001783 regulates breast cancer progression via sponging miR-200c-3p. Cell Death Dis. 2019;10(2):55. doi:10.1038/s41419-018-1287-1
12. Barbagallo D, Caponnetto A, Brex D, et al. CircSMARCA5 regulates VEGFA mRNA splicing and angiogenesis in glioblastoma multiforme through the binding of SRSF1. Cancers. 2019;11(2):194. doi:10.3390/cancers11020194
13. Han D, Li J, Wang H, et al. Circular RNA circMTO1 acts as the sponge of microRNA-9 to suppress hepatocellular carcinoma progression. Hepatology. 2017;66(4):1151–1164. doi:10.1002/hep.29270
14. Li Z, Hu Y, Zeng Q, et al. Circular RNA MYLK promotes hepatocellular carcinoma progression by increasing Rab23 expression by sponging miR-362-3p. Cancer Cell Int. 2019;19(1):211. doi:10.1186/s12935-019-0926-7
15. Otsuki T, Ishikawa M, Hori Y, Goto G, Sakamoto A. Volatile anesthetic sevoflurane ameliorates endotoxin-induced acute lung injury via microRNA modulation in rats. Biomed Rep. 2015;3(3):408–412. doi:10.3892/br.2015.428
16. Iwakawa HO, Tomari Y. The functions of MicroRNAs: mRNA decay and translational repression. Trends Cell Biol. 2015;25(11):651–665. doi:10.1016/j.tcb.2015.07.011
17. Ge X, Gong L. MiR-590-3p suppresses hepatocellular carcinoma growth by targeting TEAD1. Tumour Biol. 2017;39(3):1010428317695947. doi:10.1177/1010428317695947
18. Wang X, Zhou Z, Zhang T, et al. Overexpression of miR-664 is associated with poor overall survival and accelerates cell proliferation, migration and invasion in hepatocellular carcinoma. Onco Targets Ther. 2019;12:2373–2381. doi:10.2147/OTT.S188658
19. Liang Y, Li E, Min J, et al. miR‑29a suppresses the growth and metastasis of hepatocellular carcinoma through IFITM3. Oncol Rep. 2018;40(6):3261–3272. doi:10.3892/or.2018.6745
20. Viotti M, Wilson C, Mccleland M, Koeppen H, Mellman I. SUV420H2 is an epigenetic regulator of epithelial/mesenchymal states in pancreatic cancer. J Cell Bio. 2017;217(2):763–777. doi:10.1083/jcb.201705031
21. Ai J, Gong C, Wu J, et al. MicroRNA181c suppresses growth and metastasis of hepatocellular carcinoma by modulating NCAPG. Cancer Manag Res. 2019;11:3455–3467. doi:10.2147/CMAR.S197716
22. Huang J, Deng Q, Wang Q, et al. Exome sequencing of hepatitis B virus–associated hepatocellular carcinoma. Nat Genet. 2012;44(10):1117–1121. doi:10.1038/ng.2391
23. Villanueva A. Hepatocellular carcinoma. N Engl J Med. 2019;380(15):1450–1462. doi:10.1056/NEJMra1713263
24. Bruix J, Gores G, Mazzaferro V. Hepatocellular carcinoma: clinical frontiers and perspectives. Gut. 2014;63(5):844–855. doi:10.1136/gutjnl-2013-306627
25. Zhong Z, Huang M, Lv M, et al. Circular RNA MYLK as a competing endogenous RNA promotes bladder cancer progression through modulating VEGFA/VEGFR2 signaling pathway. Cancer Lett. 2017;403:305–317. doi:10.1016/j.canlet.2017.06.027
26. Chen J, Li Y, Zheng Q, et al. Circular RNA profile identifies circPVT1 as a proliferative factor and prognostic marker in gastric cancer. Cancer Lett. 2017;388:208–219. doi:10.1016/j.canlet.2016.12.006
27. Dai Y, Li D, Chen X, Tan X, Zhang X. Circular RNA Myosin Light Chain Kinase (MYLK) promotes prostate cancer progression through modulating Mir-29a expression. Med Sci Monit. 2018;24:3462–3471. doi:10.12659/MSM.908009
28. Memczak S, Jens M, Elefsinioti A, et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013;495(7441):333–338. doi:10.1038/nature11928
29. Ning B, Peng E, Qiu X, et al. circFBLIM1 act as a ceRNA to promote hepatocellular cancer progression by sponging miR-346. J Exp Clin Canc Res. 2018;37(1):172. doi:10.1186/s13046-018-0838-8
30. Li X, Ding J, Wang X, Cheng Z, Zhu Q. NUDT21 regulates circRNA cyclization and ceRNA crosstalk in hepatocellular carcinoma. Oncogene. 2020;39(4):891–904. doi:10.1038/s41388-019-1030-0
31. Yu J, Xu Q, Wang Z, et al. Circular RNA cSMARCA5 inhibits growth and metastasis in hepatocellular carcinoma. J Hepatol. 2018;68(6):1214–1227. doi:10.1016/j.jhep.2018.01.012
32. Wu Y, Shi W, Tang T, et al. miR-29a contributes to breast cancer cells epithelial-mesenchymal transition, migration, and invasion via down-regulating histone H4K20 trimethylation through directly targeting SUV420H2. Cell Death Dis. 2019;10(3):176. doi:10.1038/s41419-019-1437-0
33. Mazzoccoli L, Robaina MC, Bacchi CE, Soares Lima SC, Klumb CE. miR29 promoter and enhancer methylation identified by pyrosequencing in Burkitt lymhoma cells: interplay between MYC and miR29 regulation. Oncol Rep. 2019;775–784.
34. Muluhngwi P, Krishna A, Vittitow S, et al. Tamoxifen differentially regulates miR-29b-1 and miR-29a expression depending on endocrine-sensitivity in breast cancer cells. Cancer Lett. 2017;388:230–238. doi:10.1016/j.canlet.2016.12.007
35. Yang Y, Peng W, Tang T, et al. MicroRNAs as promising biomarkers for tumor-staging: evaluation of MiR21 MiR155 MiR29a and MiR92a in predicting tumor stage of rectal cancer. Asian Pac J Cancer Prev. 2014;15(13):5175–5180. doi:10.7314/APJCP.2014.15.13.5175
36. Mahati S, Xiao L, Yang Y, Mao R, Bao Y. miR-29a suppresses growth and migration of hepatocellular carcinoma by regulating CLDN1. Biochem Bioph Res Co. 2017;486(3):732–737. doi:10.1016/j.bbrc.2017.03.110
37. Xu C, Dong Q, Zhang X, et al. microRNA-29a suppresses cell proliferation by targeting SPARC in hepatocellular carcinoma. Inter J Mol Med. 2012;30(6):1321–1326. doi:10.3892/ijmm.2012.1140
38. Souza PP, Völkel P, Trinel D, Vandamme J, Angrand PO. The histone methyltransferase SUV420H2 and heterochromatin proteins HP1 interact but show different dynamic behaviours. BMC Cell Biol. 2009;10(1):41. doi:10.1186/1471-2121-10-41
39. Wu H, Siarheyeva A, Zeng H, et al. Crystal structures of the human histone H4K20 methyltransferases SUV420H1 and SUV420H2. FEBS Lett. 2013;587(23):3859–3868. doi:10.1016/j.febslet.2013.10.020
40. Shinchi Y, Hieda M, Nishioka Y, et al. SUV420H2 suppresses breast cancer cell invasion through down regulation of the SH2 domain-containing focal adhesion protein tensin-3. Exp Cell Res. 2015;334(1):90–99. doi:10.1016/j.yexcr.2015.03.010
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.