Back to Journals » Cancer Management and Research » Volume 12
Circ-ATP5H Induces Hepatitis B Virus Replication and Expression by Regulating miR-138-5p/TNFAIP3 Axis
Authors Jiang W, Wang L, Zhang Y, Li H
Received 20 July 2020
Accepted for publication 1 October 2020
Published 2 November 2020 Volume 2020:12 Pages 11031—11040
DOI https://doi.org/10.2147/CMAR.S272983
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Sanjeev K. Srivastava
Wenxiu Jiang,1,* Lili Wang,2,* Yajuan Zhang,1 Hongliang Li1
1Department of Infectious Diseases, The People’s Hospital of Danyang, Affiliated Danyang Hospital of Nantong University, Danyang City, Jiangsu Province, People’s Republic of China; 2Department of Clinical Research, The Second Hospital of Nanjing, Nanjing University of Chinese Medicine, Nanjing City, Jiangsu Province, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Hongliang Li
Department of Infectious Diseases, The People’s Hospital of Danyang, Affiliated Danyang Hospital of Nantong University, No. 2 Xinmin West Road, Danyang 212300, Jiangsu Province, People’s Republic of China
Tel/Fax +86 511-86553015
Email [email protected]
Background: Circular RNAs (circRNAs) play an important regulatory role in various cancers, including hepatocellular carcinoma (HCC). This study aimed to investigate the function of hsa_circ_0006942 (circ-ATP5H) in hepatitis B virus (HBV)-associated HCC and its underlying mechanism.
Methods: The levels of circ-ATP5H, miR-138-5p and tumor necrosis factor alpha-induced protein 3 (TNFAIP3) were determined using quantitative real-time polymerase chain reaction (qRT-PCR) or Western blot assay. The copies of HBV DNA were examined using qRT-PCR. The levels of hepatitis B surface antigen (HBsAg) and hepatitis B e antigen (HBeAg) were detected via enzyme-linked immunosorbent assay (ELISA). Dual-luciferase reporter assay was used to analyze the interactions among circ-ATP5H, miR-138-5p and TNFAIP3.
Results: Circ-ATP5H and TNFAIP3 levels were increased, while miR-138-5p level was reduced in HBV-positive HCC tissues and cells. Knockdown of circ-ATP5H hindered HBV DNA replication and decreased HBsAg and HBeAg levels in HBV-infected cells. Circ-ATP5H silencing suppressed HBV replication and expression by regulating miR-138-5p. Moreover, miR-138-5p blocked HBV replication and expression via targeting TNFAIP3. Furthermore, circ-ATP5H up-regulated TNFAIP3 via absorbing miR-138-5p.
Conclusion: Circ-ATP5H promoted HBV replication and expression through modulating miR-138-5p/TNFAIP3 axis, suggesting a new biomarker for HBV-related HCC treatment.
Keywords: hepatitis B virus, hepatocellular carcinoma, circ-ATP5H, miR-138-5p, TNFAIP3
Introduction
Hepatocellular carcinoma (HCC) is one of the most frequent malignancies, and its five-year survival rate is as low as 18%.1 Chronic hepatitis B virus (HBV) infection is a common etiology of liver diseases, including liver failure, cirrhosis and HCC.2,3 To date, 3.5% of the global population is suffering from HBV infection.4 Although antiviral therapy effectively inhibits HBV replication, the existence of HBV covalently closed circular DNA (cccDNA) prevents the virus from being completely eliminated.5,6 Therefore, exploring the potential mechanism of HBV replication is crucial to discovering new strategies for HBV treatment.
Circular RNAs (circRNAs) possess a covalent closed-loop structure formed by joining 5ʹend and 3ʹend.7 Mechanically, circRNAs participate in gene regulation by serving as microRNA (miRNA) sponges.8 Besides, mounting evidence has demonstrated that circRNAs are involved in the biological and pathological processes of various diseases, including cancer.9,10 In addition, Yu et al revealed that some plasma circRNAs had important diagnostic value in HBV-associated HCC.11 In 2018, Cui et al found a large number of aberrantly expressed circRNAs in HBV-infected HCC patients and verified that hsa_circ_0006942 derived from ATP synthase peripheral stalk subunit d (ATP5H) was up-regulated in HBV-positive HCC tissues.12 Nevertheless, the role and potential mechanism of circ-ATP5H in HBV-related HCC have not been studied.
Moreover, abnormal expression of miRNAs plays a critical role in a variety of tumors.13 Furthermore, accumulating evidence has manifested that miRNAs are implicated in the pathogenesis of HBV-related HCC via modulating gene expression and pathway activity.14,15 For example, miR-325-3p attenuated cell viability and triggered apoptosis in HBV-specific HCC cells by down-regulating aquaporin 5 (AQP5).16 Also, miR-3928v facilitated the development of HBV-positive HCC via repressing VDAC3 and regulating NF-κB/EGR1 pathway.17 Moreover, miR-302c-3p impeded tumor metastasis in HBV-associated HCC by binding to TRAF4.18 In this research, we unveiled that miR-138-5p level was overtly reduced in HBV-producing HCC tissues and cells. However, the function of miR-138-5p in HBV-positive HCC remains unknown.
Tumor necrosis factor alpha-induced protein 3 (TNFAIP3) is a critical regulator in immune and inflammatory diseases.19 Additionally, TNFAIP3 is strongly related to malignant diseases by modulating ubiquitin-dependent signaling.20 In the current study, TNFAIP3 was inversely related to miR-138-5p in HBV-producing HBV tissues, so we speculated that TNFAIP3 might be a target of miR-138-5p.
In this research, we investigated the abundance of circ-ATP5H in HBV-positive HCC patients and cells. Further, we explored the function of circ-ATP5H in HBV replication and expression. Also, we verified that the circ-ATP5H/miR-138-5p/TNFAIP3 regulatory axis might provide a novel approach for HBV-related HCC treatment.
Materials and Methods
Specimen Collection
HBV-associated HCC tissues (n=31) and adjacent non-tumor tissues (n=31) were obtained from HBV-related HCC patients enrolled in The People’s Hospital of Danyang, Affiliated Danyang Hospital of Nantong University. This research was approved by the Ethics Committee of The People’s Hospital of Danyang, Affiliated Danyang Hospital of Nantong University. All participants carefully read and signed the written informed consent. The demographic data of HBV-related HCC patients are shown in Table 1.
 |
Table 1 Clinicopathological Features of HBV-Related HCC Patients (n=31) |
Cell Culture
HCC cell lines (Huh7 and HepG2) and HBV-infected cell line HepG2.2.15 were bought from China Center for Type Culture Collection (CCTCC; Wuhan, China). Huh7 and HepG2 cells were cultured in Dulbecco’s modified Eagle medium (DMEM; Hyclone, Logan, UT, USA) supplemented with 10% fetal bovine serum (FBS; Hyclone) and 1% penicillin/streptomycin. HepG2.2.15 cells were incubated in minimum Eagle’s medium (MEM; Hyclone) containing 10% FBS and 300 µg/mL G418 (Solarbio, Beijing, China). Huh7-1.3 cells were constructed by transfecting Huh7 cells with recombinant pcDNA 3.0–1.3 mer containing HBV genomic DNA 1.3 mer fragment. All cells were cultured in a 5% CO2 incubator at 37°C.
Cell Transfection
Small interfering RNA (siRNA) against circ-ATP5H (si-circ-ATP5H) and the negative control (si-NC), miR-138-5p mimics (miR-138-5p) and the control (miR-NC), circ-ATP5H or TNFAIP3 overexpression vector (circ-ATP5H or TNFAIP3) and the control vector (pcDNA), miR-138-5p inhibitor (anti-miR-138-5p) and the negative control (anti-miR-NC) were purchased from Ribobio (Guangzhou, China). When cells were cultured to ~80% confluence, oligonucleotides and vectors (50 nM) were transfected into cells using Lipofectamine 3000 (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions.
Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR)
RNA was isolated using Trizol Reagent (Solarbio). Then, the complementary DNA was synthesized via the specific Reverse Transcription kit (Qiagen, Frankfurt, Germany). Subsequently, quantitative PCR was carried out using SYBR Green PCR Master Mix (LMAI Bio, Shanghai, China), and the relative expression was calculated via the 2−ΔΔCt method. The primers were exhibited below: circ-ATP5H-F: 5ʹ-TAATGCGCTGAAGGTTCCCG-3ʹ, circ-ATP5H-R: 5ʹ-GAGAGACACCCACTCAGCAC-3ʹ; miR-138-5p-F: 5ʹ-GCGAGCTGGTGTTGTGAATC-3ʹ, miR-138-5p-R: 5ʹ-AGTGCAGGGTCCGAGGTATT-3ʹ; TNFAIP3-F: 5ʹ-TCCTCAGGCTTTGTATTTGAGC-3ʹ, TNFAIP3-R: 5ʹ-TGTGTATCGGTGCATGGTTTTA-3ʹ; GAPDH-F: 5ʹ-GGTCTCCTCTGACTTCAACA-3ʹ, GAPDH-R: 5ʹ-GTGAGGGTCTCTCTCTTCCT-3ʹ; U6-F: 5ʹ-CGCTTCGGCAGCACATATAC-3ʹ, U6-R: 5ʹ-AAATATGGAACGCTTCACGA-3ʹ. GAPDH or U6 was employed as an internal control.
HBV Replication Analysis
After 48 h of transfection, the culture medium was collected and centrifuged to obtain the supernatant. Subsequently, Column Viral DNAout kit (Zhen Shanghai and Shanghai Industrial Co., Ltd., Shanghai, China) was used to extract HBV DNA from the supernatant. Then, HBV DNA levels were detected using a careHBV PCR Assay (Qiagen). The HBV primers used were: HBV-F: 5ʹ-ATCCTGCTGCTATGCCTCATCTT-3ʹ, HBV-R: 5ʹ-ACAGTGGGGGAAAGCCTACGAA-3ʹ.
Enzyme-Linked Immunosorbent Assay (ELISA)
The supernatant was separated from the culture medium after transfection for 48 h. Then, the levels of hepatitis B surface antigen (HBsAg) and hepatitis B e antigen (HBeAg) in the supernatant were examined using the corresponding ELISA kit (CUSABIO, Wuhan, China) following the manufacturer’s requirements.
Dual-Luciferase Reporter Assay
The fragment of circ-ATP5H or TNFAIP3 3ʹUTR containing miR-138-5p binding sites was cloned to pmirGLO vector (LMAI Bio) to generate WT-circ-ATP5H or TNFAIP3 3ʹUTR-WT. To form the mutant reporters (MUT-circ-ATP5H or TNFAIP3 3ʹUTR-MUT), the binding sequence was mutated. Then, the constructed plasmids were transfected into HBV-infected cells together with miR-NC or miR-138-5p. Final, the luciferase intensity was examined via Dual-Lucy Assay Kit (Solarbio).
Western Blot Assay
After extracting protein with RIPA buffer (Beyotime, Shanghai, China), the equal amount of protein samples were separated with 12% polyacrylamide gel electrophoresis and transferred to polyvinylidene fluoride (PVDF) membranes (Millipore, Billerica, MA, USA). Subsequently, the membranes were incubated with primary antibodies against TNFAIP3 (1:1000, ab74037; Abcam, Cambridge, UK) or GAPDH (1:2500, ab9485; Abcam) overnight at 4°C after reacting with 5% skim milk. Then, the membranes were combined with secondary antibody conjugated with horseradish peroxidase (1:20,000, ab205718; Abcam). The protein intensity was measured using an ECL reagent (Millipore).
Statistical Analysis
Data were expressed as mean ± standard deviation in three independent experiments by using Graphpad Prism 7.0 software (GraphPad, San Diego, CA, USA). Student’s t-test was used to compare the differences between two groups. One-way analysis of variance was utilized to assess differences between more than two groups. The linear relationship between miR-138-5p and circ-ATP5H or TNFAIP3 was detected using Spearman correlation coefficient. P < 0.05 was considered statistically significant.
Results
Circ-ATP5H Was Up-Regulated, and miR-138-5p Was Down-Regulated in HBV-Related HCC Tissues and Cells
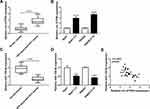
Firstly, we examined the abundance of circ-ATP5H in HBV-associated HCC tissues and adjacent non-tumor tissues. As depicted in Figure 1A, circ-ATP5H level was remarkably increased in HBV-associated HCC tissues relative to adjacent non-cancer tissues (P<0.0001). In addition, the expression of circ-ATP5H was significantly elevated in HBV-specific cells compared to HCC cells (P<0.0001, P<0.0001, Figure 1B). Secondly, we detected miR-138-5p level in HBV-related HCC tissues and adjacent normal tissues. As shown in Figure 1C, miR-138-5p was markedly down-regulated in HBV-associated HCC tissues compared with adjacent non-tumor tissues (P<0.0001). Moreover, miR-138-5p level in Huh7-1.3 and HepG2.2.15 cells was strikingly declined in comparison with Huh7 and HepG2 cells (P=0.0003, P<0.0001, Figure 1D). Furthermore, circ-ATP5H expression was negatively correlated with miR-138-5p expression in HBV-related HCC tissues (P<0.001, Figure 1E).
Circ-ATP5H Silencing Inhibits HBV Replication and Expression in HBV-Infected Cells
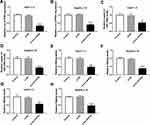
To investigate the effect of circ-ATP5H on HBV replication and expression, Huh7-1.3 and HepG2.2.15 cells were transduced with si-NC or si-circ-ATP5H. First, qRT-PCR analysis was used to determine the knockdown efficiency of circ-ATP5H (P<0.0001, P<0.0001, Figure 2A and B). Next, qRT-PCR and ELISA assays were performed to assess HBV DNA replication and the levels of HBsAg and HBeAg. The results indicated that circ-ATP5H silencing resulted in a marked decrease in HBV DNA replication (P=0.001, P=0.0003, Figure 2C and D). Additionally, ELISA analysis suggested that circ-ATP5H down-regulation overtly reduced the levels of HBsAg and HBeAg relative to the controls (P=0.0003, P<0.0001, P=0.0031, P<0.0001, Figure 2E–H). Overall, these data demonstrated that depletion of circ-ATP5H suppressed HBV replication and expression in HBV-infected cells.
Circ-ATP5H Directly Interacts with miR-138-5p
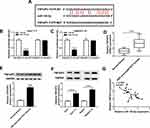
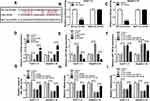
The online database starBase v2.0 predicted that circ-ATP5H and miR-138-5p had a putative binding sequence (Figure 3A). Subsequently, dual-luciferase reporter analysis exhibited that miR-138-5p mimics strikingly reduced the luciferase activity of WT-circ-ATP5H reporter, but did not affect MUT-circ-ATP5H reporter (P<0.0001, P<0.0001, Figure 3B and C). Besides, transfection with si-circ-ATP5H remarkably decreased circ-ATP5H level, and introduction of circ-ATP5H overtly increased circ-ATP5H level, indicating a significant knockdown and overexpression efficiency (P<0.0001, P<0.0001, P<0.0001, P<0.0001, Figure 3D). In addition, down-regulation of circ-ATP5H elevated miR-138-5p level, and overexpression of circ-ATP5H repressed miR-138-5p expression (P<0.0001, P<0.0001, P<0.0001, P<0.0001, Figure 3E). To explore whether circ-ATP5H affected HBV replication and expression via regulating miR-138-5p, Huh7-1.3 and HepG2.2.15 cells were transfected with si-NC, si-circ-ATP5H, si-circ-ATP5H+anti-miR-NC or si-circ-ATP5H+anti-miR-138-5p, respectively. First, transfection with anti-miR-138-5p abated the increase in miR-138-5p level caused by circ-ATP5H knockdown (P<0.0001, P<0.0001, P<0.0001, P<0.0001, Figure 3F). Moreover, silence of circ-ATP5H suppressed HBV DNA replication and the levels of HBsAg and HBeAg, while these impacts were reversed after introduction of anti-miR-138-5p (P<0.0001, P<0.0001, P<0.0001, P<0.0001, P<0.0001, P<0.0001, P<0.0001, P<0.0001, P<0.0001, P=0.0001, P<0.0001, P<0.0001, Figure 3G–I). Collectively, these results indicated that circ-ATP5H knockdown impeded HBV replication and expression via sponging miR-138-5p.
TNFAIP3 is a Target of miR-138-5p
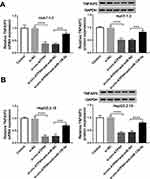
Next, the target genes of miR-138-5p were predicted using the online database starBase v2.0, and TNFAIP3 was selected as the candidate gene (Figure 4A). To illuminate the relationship between miR-138-5p and TNFAIP3, dual-luciferase reporter analysis was carried out. As illustrated in Figure 4B and C, miR-138-5p mimics significantly decreased the luciferase activity of TNFAIP3 3ʹUTR-WT reporter (P<0.0001, P<0.0001). Compared with adjacent non-tumor tissues, TNFAIP3 mRNA and protein levels were prominently increased in HBV-associated HCC tissues (P<0.0001, P=0.0002, Figure 4D and E). Moreover, we examined the protein expression of TNFAIP3 in HCC cells with or without HBV infection, and the results showed that TNFAIP3 protein level in Huh7-1.3 and HepG2.2.15 cells was remarkably raised in comparison with Huh7 and HepG2 cells (P<0.0001, P<0.0001, Figure 4F). Additionally, miR-138-5p and TNFAIP3 levels were negatively correlated in HBV-related HCC tissues (P<0.001, Figure 4G). These data evidenced that miR-138-5p directly interacted with TNFAIP3.
MiR-138-5p Hinders HBV Replication and Expression by Modulating TNFAIP3
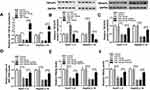
First, the inhibition and overexpression efficiency of miR-138-5p was verified by transfecting anti-miR-138-5p or miR-138-5p into HBV-specific cells (P=0.0086, P<0.0001, P=0.0035, P<0.0001, Figure 5A). Then, suppression of miR-138-5p elevated TNFAIP3 protein level, and overexpression of miR-138-5p reduced TNFAIP3 protein level (P<0.0001, P<0.0001, P<0.0001, P<0.0001, Figure 5B). To elucidate the roles of miR-138-5p and TNFAIP3 in HBV replication and expression, a series of rescue experiments were performed in Huh7-1.3 and HepG2.2.15 cells introduced with miR-NC, miR-138-5p, miR-138-5p+pcDNA or miR-138-5p+TNFAIP3. As displayed in Figure 5C, introduction of TNFAIP3 alleviated the reduction of TNFAIP3 protein expression induced by miR-138-5p up-regulation (P<0.0001, P<0.0001, P<0.0001, P<0.0001). Furthermore, miR-138-5p overexpression restrained HBV DNA replication and the levels of HBsAg and HBeAg, whereas these effects were attenuated by up-regulating TNFAIP3 (P<0.0001, P<0.0001, P<0.0001, P<0.0001, P<0.0001, P=0.0004, P<0.0001, P<0.0001, P<0.0001, P=0.0002, P<0.0001, P<0.0001, Figure 5D–F). Thus, these data indicated that TNFAIP3 overturned the inhibition of miR-138-5p on HBV replication and expression.
Circ-ATP5H Regulates TNFAIP3 Expression by Sponging miR-138-5p
To explore the association among circ-ATP5H, miR-138-5p and TNFAIP3, TNFAIP3 expression was determined in Huh7-1.3 and HepG2.2.15 cells introduced with si-NC, si-circ-ATP5H, si-circ-ATP5H+anti-miR-NC or si-circ-ATP5H+anti-miR-138-5p. The results revealed that TNFAIP3 mRNA and protein levels were significantly declined in the si-circ-ATP5H group relative to the si-NC group, while transfection with anti-miR-138-5p abolished the decrease in TNFAIP3 expression caused by circ-ATP5H silencing (P<0.0001, P=0.0007, P<0.0001, P=0.0007, P<0.0001, P=0.0001, P<0.0001, P<0.0001, Figure 6A and B). Overall, these results evidenced that circ-ATP5H sponged miR-138-5p to up-regulate TNFAIP3.
Discussion
HBV replication is the primary driving force for HBV-associated diseases, leading to the ultimate goal of HBV treatment is to continuously inhibit or eliminate HBV replication.21 Hepatitis B surface antigen (HBsAg) is a hallmark of HBV infection, and hepatitis B e antigen (HBeAg) represents the activity of HBV replication.22 Nevertheless, the molecular mechanisms of HBV replication and expression are still poorly studied. In the present study, we revealed a new regulatory mechanism for HBV-associated HCC.
Recently, a growing number of studies have elaborated that circRNAs serve as biomarkers for monitoring tumor development based on its stable structure.23 In HBV-positive HCC, Zhu et al suggested that plasma hsa_circ_0027089 was a promising marker for tumor diagnosis.24 Our research validated that circ-ATP5H level was strikingly elevated in HBV-positive HCC tissues and cells. More importantly, interference of circ-ATP5H blocked HBV replication and expression in HBV-specific cells. Thereafter, we further studied the potential basis of circ-ATP5H. According to bioinformatics analysis, we conjectured that circ-ATP5H was a sponge for miR-138-5p. Compelling evidence has highlighted that circRNAs mediate the pathogenesis of various diseases by acting as competing endogenous RNAs (ceRNAs) for miRNA.25 For instance, up-regulation of circRNA-5692 ameliorated the malignancy of HCC via functioning as a ceRNA for miR-328-5p to elevate DAB2IP level.26 Besides, circ-TCF4.85 depletion impeded HCC progression by regulating miR-486-5p/ABCF2 pathway.27 In the current research, circ-ATP5H sponged miR-138-5p to regulate HBV replication and expression.
Mounting evidence has illuminated that miR-138-5p plays a pivotal regulatory role in multifarious cancers by mediating a series of biological processes. For example, up-regulation of miR-138-5p restrained cell growth and metastasis in prostate cancer via combining with FOXC1.28 In breast carcinoma, miR-138-5p hindered tumor development by binding to RHBDD1.29 In hepatocellular carcinoma, miR-138 functioned as a tumor-suppressing factor.30 Shiu et al disclosed that miR-138 accelerated HCC cell senescence via repressing telomerase reverse transcriptase (TERT) in hepatitis C virus (HCV)-associated HCC.31 In addition, Wang et al unveiled a marked reduction of miR-138 in HBV-related HCC tissues through Taqman low-density miRNA array.32 Herein, we verified that miR-138-5p level was declined in HBV-producing HCC. Furthermore, rescue experiments manifested that knockdown of miR-138-5p reversed the suppression of circ-ATP5H depletion on HBV replication and expression.
Moreover, increasing evidence has demonstrated that miRNAs widely affect gene expression by base-pairing with the 3ʹUTR of mRNA.33 In this research, dual-luciferase reporter analysis evidenced that miR-138-5p targeted TNFAIP3. Previous studies corroborated that TNFAIP3 (also known as A20) participates in tumorigenesis induced by uncontrolled ubiquitination by modulating inflammatory signaling cascade.34,35 Feng et al revealed that down-regulation of TNFAIP3 induced HCC progression via interacting with PFKL.36 In HBV-related HCC, miR-29c blocked HBV DNA replication by targeting TNFAIP3.37 In this report, we discovered that miR-138-5p impaired HBV replication and expression by down-regulating TNFAIP3. Additionally, circ-ATP5H enhanced TNFAIP3 expression via sponging miR-138-5p.
In short, these findings concluded that circ-ATP5H contributed to HBV replication and expression by mediating miR-138-5p/TNFAIP3 regulatory pathway. Therefore, circ-ATP5H might be a potential therapeutic target for HBV-associated HCC.
Abbreviations
HCC, hepatocellular carcinoma; HBV, hepatitis B virus; TNFAIP3, tumor necrosis factor alpha-induced protein 3; qRT-PCR, quantitative real-time polymerase chain reaction; HbsAg, hepatitis B surface antigen; ELISA, enzyme-linked immunosorbent assay; AQP5, aquaporin 5; PVDF, polyvinylidene fluoride.
Ethics Approval and Consent Participate
Written informed consent was obtained from patients with approval by the Institutional Review Board in The People’s Hospital of Danyang, Affiliated Danyang Hospital of Nantong University.
Funding
There is no funding to report.
Disclosure
The authors declare that they have no financial or non-financial conflicts of interest.
References
1. Chino F, Stephens SJ, Choi SS, et al. The role of external beam radiotherapy in the treatment of hepatocellular cancer. Cancer. 2018;124(17):3476–3489. doi:10.1002/cncr.31334
2. Fanning GC, Zoulim F, Hou J, Bertoletti A. Therapeutic strategies for hepatitis B virus infection: towards a cure. Nat Rev Drug Discov. 2019;18(11):827–844. doi:10.1038/s41573-019-0037-0
3. Seto WK, Lo YR, Pawlotsky JM, Yuen MF. Chronic hepatitis B virus infection. Lancet. 2018;392(10161):2313–2324. doi:10.1016/S0140-6736(18)31865-8
4. Yuen MF, Chen DS, Dusheiko GM, et al. Hepatitis B virus infection. Nat Rev Dis Primers. 2018;4:18035. doi:10.1038/nrdp.2018.35
5. Tang LSY, Covert E, Wilson E, Kottilil S. Chronic hepatitis B infection: a review. JAMA. 2018;319(17):1802–1813. doi:10.1001/jama.2018.3795
6. Liu S, Zhou B, Valdes JD, Sun J, Guo H. Serum hepatitis B virus RNA: a new potential biomarker for chronic hepatitis B virus infection. Hepatology. 2019;69(4):1816–1827. doi:10.1002/hep.30325
7. Qian L, Yu S, Chen Z, Meng Z, Huang S, Wang P. The emerging role of circRNAs and their clinical significance in human cancers. Biochim Biophys Acta Rev Cancer. 2018;1870(2):247–260. doi:10.1016/j.bbcan.2018.06.002
8. Panda AC. Circular RNAs act as miRNA sponges. Adv Exp Med Biol. 2018;1087:67–79. doi:10.1007/978-981-13-1426-1_6
9. Lei M, Zheng G, Ning Q, Zheng J, Dong D. Translation and functional roles of circular RNAs in human cancer. Mol Cancer. 2020;19(1):30. doi:10.1186/s12943-020-1135-7
10. Su M, Xiao Y, Ma J, et al. Circular RNAs in cancer: emerging functions in hallmarks, stemness, resistance and roles as potential biomarkers. Mol Cancer. 2019;18(1):90. doi:10.1186/s12943-019-1002-6
11. Yu J, Ding WB, Wang MC, et al. Plasma circular RNA panel to diagnose hepatitis B virus-related hepatocellular carcinoma: a large-scale, multicenter study. Int J Cancer. 2020;146(6):1754–1763. doi:10.1002/ijc.32647
12. Cui S, Qian Z, Chen Y, Li L, Li P, Ding H. Screening of up- and downregulation of circRNAs in HBV-related hepatocellular carcinoma by microarray. Oncol Lett. 2018;15(1):423–432. doi:10.3892/ol.2017.7265
13. Rupaimoole R, Calin GA, Lopez-Berestein G, Sood AK. miRNA deregulation in cancer cells and the tumor microenvironment. Cancer Discov. 2016;6(3):235–246. doi:10.1158/2159-8290.CD-15-0893
14. Petrini E, Caviglia GP, Abate ML, Fagoonee S, Smedile A, Pellicano R. MicroRNAs in HBV-related hepatocellular carcinoma: functions and potential clinical applications. Panminerva Med. 2015;57(4):201–209.
15. Sadri Nahand J, Bokharaei-Salim F, Salmaninejad A, et al. microRNAs: key players in virus-associated hepatocellular carcinoma. J Cell Physiol. 2019;234(8):12188–12225. doi:10.1002/jcp.27956
16. Zhang Z, Han Y, Sun G, Liu X, Jia X, Yu X. MicroRNA-325-3p inhibits cell proliferation and induces apoptosis in hepatitis B virus-related hepatocellular carcinoma by down-regulation of aquaporin 5. Cell Mol Biol Lett. 2019;24:13. doi:10.1186/s11658-019-0137-1
17. Zhang Q, Song G, Yao L, et al. miR-3928v is induced by HBx via NF-kappaB/EGR1 and contributes to hepatocellular carcinoma malignancy by down-regulating VDAC3. J Exp Clin Cancer Res. 2018;37(1):14. doi:10.1186/s13046-018-0681-y
18. Yang L, Guo Y, Liu X, et al. The tumor suppressive miR-302c-3p inhibits migration and invasion of hepatocellular carcinoma cells by targeting TRAF4. J Cancer. 2018;9(15):2693–2701. doi:10.7150/jca.25569
19. Vereecke L, Beyaert R, van Loo G. The ubiquitin-editing enzyme A20 (TNFAIP3) is a central regulator of immunopathology. Trends Immunol. 2009;30(8):383–391. doi:10.1016/j.it.2009.05.007
20. Ma A, Malynn BA. A20: linking a complex regulator of ubiquitylation to immunity and human disease. Nat Rev Immunol. 2012;12(11):774–785. doi:10.1038/nri3313
21. Liaw YF. Impact of therapy on the long-term outcome of chronic hepatitis B. Clin Liver Dis. 2013;17(3):413–423. doi:10.1016/j.cld.2013.05.005
22. Kao JH. Diagnosis of hepatitis B virus infection through serological and virological markers. Expert Rev Gastroenterol Hepatol. 2008;2(4):553–562. doi:10.1586/17474124.2.4.553
23. Ma Z, Shuai Y, Gao X, Wen X, Ji J. Circular RNAs in the tumour microenvironment. Mol Cancer. 2020;19(1):8. doi:10.1186/s12943-019-1113-0
24. Zhu K, Zhan H, Peng Y, et al. Plasma hsa_circ_0027089 is a diagnostic biomarker for hepatitis B virus-related hepatocellular carcinoma. Carcinogenesis. 2019. doi:10.1093/carcin/bgz154
25. Zhong Y, Du Y, Yang X, et al. Circular RNAs function as ceRNAs to regulate and control human cancer progression. Mol Cancer. 2018;17(1):79. doi:10.1186/s12943-018-0827-8
26. Liu Z, Yu Y, Huang Z, et al. CircRNA-5692 inhibits the progression of hepatocellular carcinoma by sponging miR-328-5p to enhance DAB2IP expression. Cell Death Dis. 2019;10(12):900. doi:10.1038/s41419-019-2089-9
27. Gao J, Dai C, Yu X, Yin XB, Zhou F. Circ-TCF4.85 silencing inhibits cancer progression through microRNA-486-5p-targeted inhibition of ABCF2 in hepatocellular carcinoma. Mol Oncol. 2020;14(2):447–461. doi:10.1002/1878-0261.12603
28. Huang H, Xiong Y, Wu Z, et al. MIR-138-5P inhibits the progression of prostate cancer by targeting FOXC1. Mol Genet Genomic Med. 2020;8(4):e1193. doi:10.1002/mgg3.1193
29. Zhao C, Ling X, Li X, Hou X, Zhao D. MicroRNA-138-5p inhibits cell migration, invasion and EMT in breast cancer by directly targeting RHBDD1. Breast Cancer. 2019;26(6):817–825. doi:10.1007/s12282-019-00989-w
30. Xiao JX, Xu W, Fei X, et al. Anillin facilitates cell proliferation and induces tumor growth of hepatocellular carcinoma via miR-138/SOX4 axis regulation. Transl Oncol. 2020;13(10):100815. doi:10.1016/j.tranon.2020.100815
31. Shiu TY, Shih YL, Feng AC, et al. HCV core inhibits hepatocellular carcinoma cell replicative senescence through downregulating microRNA-138 expression. J Mol Med (Berl). 2017;95(6):629–639. doi:10.1007/s00109-017-1518-4
32. Wang W, Zhao LJ, Tan YX, Ren H, Qi ZT. Identification of deregulated miRNAs and their targets in hepatitis B virus-associated hepatocellular carcinoma. World J Gastroenterol. 2012;18(38):5442–5453. doi:10.3748/wjg.v18.i38.5442
33. Fabian MR, Sonenberg N, Filipowicz W. Regulation of mRNA translation and stability by microRNAs. Annu Rev Biochem. 2010;79:351–379. doi:10.1146/annurev-biochem-060308-103103
34. Hymowitz SG, Wertz IE. A20: from ubiquitin editing to tumour suppression. Nat Rev Cancer. 2010;10(5):332–341. doi:10.1038/nrc2775
35. Malynn BA, Ma A. A20 takes on tumors: tumor suppression by an ubiquitin-editing enzyme. J Exp Med. 2009;206(5):977–980. doi:10.1084/jem.20090765
36. Feng Y, Zhang Y, Cai Y, et al. A20 targets PFKL and glycolysis to inhibit the progression of hepatocellular carcinoma. Cell Death Dis. 2020;11(2):89. doi:10.1038/s41419-020-2278-6
37. Wang CM, Wang Y, Fan CG, et al. miR-29c targets TNFAIP3, inhibits cell proliferation and induces apoptosis in hepatitis B virus-related hepatocellular carcinoma. Biochem Biophys Res Commun. 2011;411(3):586–592. doi:10.1016/j.bbrc.2011.06.191
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.