Back to Journals » Infection and Drug Resistance » Volume 12
Characterization of a carbapenem- and colistin-resistant Enterobacter cloacae carrying Tn6901 in blaNDM-1 genomic context
Authors Le-Ha TD , Le L, Le-Vo H , Anda M, Motooka D, Nakamura S, Tran LK, Tran PTB, Iida T, Cao V
Received 16 November 2018
Accepted for publication 8 February 2019
Published 3 April 2019 Volume 2019:12 Pages 733—739
DOI https://doi.org/10.2147/IDR.S194495
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Sahil Khanna
Tam-Duong Le-Ha,1 Lien Le,1 Hong-Ngoc Le-Vo,1 Mizue Anda,2 Daisuke Motooka,3 Shota Nakamura,3 Linh Khanh Tran,1 Phuong Thi-Bich Tran,1 Tetsuya Iida,2 Van Cao1
1Department of Immunology and Microbiology, Pasteur Institute in Ho Chi Minh City, Ho Chi Minh City, Vietnam; 2Department of Bacterial Infections, Research Institute for Microbial Diseases, Osaka University, Osaka, Japan; 3Genome Information Research Center, Research Institute for Microbial Diseases, Osaka University, Osaka, Japan
Abstract: We report a clinical strain of Enterobacter cloacae, PIMB10EC27, isolated in Vietnam in 2010 that was resistant to 21 of 26 tested antibiotics, including carbapenems (MICs >64 μg/mL) and colistin (MIC >128 μg/mL). The complete genome of strain PIMB10EC27 was sequenced by PacBio RSII and the Illumina Miseq system. Whole-genome analysis revealed that PIMB10EC27 contains a chromosome of the ST513 group (PIMBEC27, length 5,272,177 bp) and two plasmids, pEC27-1 of the IncX3 group (length 62,470 bp) and pEC27-2 of the IncHI1 group (length 84,602 bp). It also revealed that strain PIMB10EC27 carries 15 genes that confer resistance to at least 10 antibiotic groups. Particularly, the insertion of ISKpn19 and Tn6901 into the genomic context of blaNDM-1 was first identified and described. In another context, amino acid mutations G273D in PmrB and F515S in PmrC were first identified on the chromosome of PIMB10EC27, which may confer resistance to colistin in this strain.
Keywords: blaNDM-1, colistin, Enterobacter cloacae, multidrug-resistance, Tn6901
Carbapenems are one of the broad-spectrum groups of β-lactam antibiotics and have been considered the best choice for treatment of infections caused by multidrug-resistant bacteria,1 however, the recent increase in the rate of carbapenem resistance has been a cause for concern.2 Encoded by the blaNDM-1 gene, New Delhi metallo-β-lactamase 1 (NDM-1), one of the most active and transmissible carbapenemases among the carbapenem-hydrolyzing β-lactamases, was first characterized by Yong et al in 20093 and has rapidly spread globally. To date, at least 17 NDM alleles have been characterized4 and the Tn125 composite transposon bracketed by two copies of ISAba125 appears to be the main vehicle for dissemination of the blaNDM-1 gene.5 The lack of new generations of antibiotics has positioned colistin, a decades-old antibiotic, as one of the treatments of last resort against multidrug-resistant bacteria, particularly carbapenem-resistant Gram-negative bacteria.6,7 Unfortunately, colistin resistance has been reported and is increasing.8–11
Enterobacter cloacae belongs to the Enterobacteriaceae family. These Gram-negative bacteria have been a frequent cause of nosocomial multidrug-resistant bacterial infections in the last decade.12,13 Carbapenem-resistant E. cloacae has been commonly reported in Vietnam and many other countries in the world.14–16 However, colistin resistance in E. cloacae has not been reported widely, particularly carbapenemase-producing colistin-resistant E. cloacae has very recently been reported from some countries including India,10 the United States,17 and China.18–20
In this study, the whole genome of a clinical carbapenem- and colistin-resistant E. cloacae strain, PIMB10EC27, was sequenced and analyzed for its genetic characteristics that may associate with its resistance phenotypes.
PIMB10EC27 was recovered from the urine of a 72-year-old male patient admitted to Binh Dan Hospital in Vietnam in December 2010 with diagnoses of invasive prostate cancer and urosepsis, he had undergone prostatectomy at the same hospital 15 days earlier. The patient was discharged at the request of the family in a state of acute urinary retention and respiratory failure 4 days after hospitalization. During hospitalization, the patient was treated with meropenem, cefoperazone, sulbactam, clavulanic acid, and amoxicillin, but not with colistin. The strain was isolated as a part of the routine hospital laboratory procedures and identified as E. cloacae using a Vitek II kit (BioMérieux, USA).
The in vitro antibiotic susceptibility of PIMB10EC27 was tested by the Kirby-Bauer (KB) method, and the minimum inhibitory concentration (MIC) using the agar dilution method was also determined according to recommendations of the Clinical and Laboratory Standards Institute (CLSI 2018)19 except for colistin, which was performed with broth microdilution method and interpreted by the guidelines of European Union Committee for Antimicrobial Susceptibility Testing (EUCAST 2019).20 The results showed that PIMB10EC27 was resistant to 21 of 26 tested antibiotics, particularly imipenem (MIC >64 µg/mL), ertapenem (MIC =128 µg/mL), meropenem (MIC =128 µg/mL), and colistin (MIC >128 µg/mL); susceptible to amikacin, levofloxacin nitrofurantoin; and showed intermediate resistance to ciprofloxacin, oxfloxacin (Table 1).
 | Table 1 Antibiotic susceptibility of the E. cloacae PIMB10EC27 |
A Nextera XT DNA Library Prep Kit (Illumina Inc., USA) and SMRTbell Template Prep Kit (Pacific Biosciences, USA) were used to prepare libraries for the whole genome sequencing of PIMB10EC27 using simultaneously the MiSeq System (Illumina Inc.) with MiSeq Reagent Kit v.2 (2×150 cycles), and the PacBio RSII (Pacific Biosciences) together with Sequel Sequencing Kit 2.0 (8 rxn).
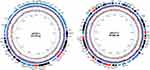
In total, 3.8Gb of filtered subreads were obtained from PacBio RSII sequencing platform. Using PacBio’s pre-patching program, we obtained 353Mb of 35,932 reads with the sizes of 500–27,847 bp. The pre-assemble reads were further packaged with HGAP program, which yielded a 5,272,177 bp chromosome and two plasmids of 62,318 bp and 84,480 bp. MiSeq data was used for the error correction of PacBio consensus sequences. By integrating these two platforms, we were able to obtain the complete genome which consisted of a chromosome (5,272,177 bp) belonging to the ST513 group and two plasmids, pEC27-1 of the IncX3 group (62,470 bp) and pEC27-2 of the IncHI1 group (84,602 bp) (Figure 1). The complete genome sequence was annotated with NCBI Prokaryotic Genome Annotation Pipeline servers and analyzed by bioinformatics programs or software, including ResFinder 2.1,21 MLST,22 Isaga,23 Galaxy,24 and Plasmid Finder25 for antibiotic resistance genes identification, chromosome classification, sequences of insertion sequence (IS) determination, integrons finding, and plasmids classification, respectively. In total, 15 coding genes conferring resistance to 10 antibiotic groups were found on PIMB10EC27 (Table 2). Among these genes, blaNDM-1, blaSHV-12, blaCMH, aac(3)-ID, strA, strB, dfrA-14, sul2, catA2, and tet(D) were associated with resistance phenotypes of PIMB10EC27.On the other hand qnrS1 equivalent to the intermediate fluoroquinolone-resistance phenotype of PIMB10EC27 has been proven to reduce the antibiotic susceptibility of bacteria to quinolones or fluoroquinolone.26–28 In addition, 160 putative transposase open reading frames (tORFs) were identified in which the number of tORFs located on the chromosome, pEC27-1 and pEC27-2 were 113, 10 and 37, respectively. Antibiotic resistance genes closely located to these transposase sequences may have contributed to the accumulation and increase in the antibiotic resistance of PIMB10EC27. A 1722-bp class 1 integron with gene cassette dfrA14 flanked by IS6 sequences was also found on plasmid pEC27-2.
 | Table 2 Distribution of coding genes conferring to antibiotic resistance in PIMB10EC27 |
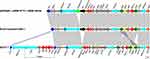
We further analyzed the genes in PIMB10EC27 that confer resistance to carbapenem and colistin. As a result, a blaNDM-1 gene was found on pEC27-1 of PIMB10EC27 along with a blaSHV-12 and bleMBL. A comparison of the nucleotide sequence of pEC27-1 and those of other IncX3 plasmids carrying blaNDM-1 and blaSHV-12, including pKPN5047 (KC311431), pNDM-HF727 (KF976405), pNDM-HN380 (JX104760), and RJA274 plasmid NDM-1 (KF877335) was carried out in order to identify if any unique structure characteristics contributed to the multidrug-resistance of PIMB10EC27. Comparative results of the genomic context surrounding the resistance genes revealed that all of the plasmids had a conserved sequence carrying ΔISAba125, blaNDM-1, bleMBL, trpF, dsbC, cutA1, groES, groEL, and ISCR21 (Figure 2). This conserved sequence is also a Tn125 composite transposon, in which the blaNDM-1 gene lies downstream of a truncated ISAba125 element that provides the −35 region for the promoter of blaNDM-1.29 A 93-bp sequence separates the right-hand inverted repeat of ISAba125 from the start codon of blaNDM-1, and the deletion of an IS5 was identified in the plasmid pEC27-1. An ISCR21-like element was also found downstream of blaNDM-1 and it is suggested that this element may be responsible for initial gene capture.5 The IS26-bounded region of pEC27-1 containing ygbI, ygbJ, and blaSHV-12 was found to be very distinct from the other plasmids. This region carried three copies of IS26, and the local ygbJ gene was split into two parts, whereas other plasmids carried only two copies of IS26 and the full ygbJ gene. The isolation and reverse with a copy of IS26 in the plasmid pEC27-1 of the 424-bp region of the ygbJ gene might propose there is an IS26-mediated re-organization. Interestingly, an insertion of ISKpn19 and Tn6901 into the mpr gene (encoding zinc metalloproteinase) lying further upstream of blaNDM-1 was also observed (Figure 2). Belonging to ISKra4 family, ISKpn19 is 2851 bp in length and has been found to lie downstream of blaOXA-181 in an IncX3-type plasmid.30 Tn6901, first described in plasmid Rts1 of Proteus vulgaris,31 is 6.9 kb in length and harbors 6 genes, including a transposase gene (tnpA), a resolvase gene (tnpR), an alcohol dehydrogenase gene (adh), a glyoxalase/bleomycin resistance gene (glo), an S-formylglutathione hydrolase gene (fgh), and a regulatory protein gene (frmR). We report here the first case, to our knowledge, of a 9.8-kb insertion sequence harboring ISKpn19 and Tn6901 in the genomic context of blaNDM-1. This insertion made the genomic context of blaNDM-1 in PIMB10EC27 more complex with 6 resistance genes, including adh, glo, fgh, blaNDM-1, bleMBL, and blaSHV-12, which may also affect the dissemination and expression of blaNDM-1. Further research is required to clarify these assumptions.
Genome analysis was also performed to identify the genes that confer resistance to colistin of PIMB10EC27. In this research, we did not find any mcr gene that recently characterized as a novel mobile gene responsible for colistin resistance among Gram negative bacteria.32 In addition, our transformation experiments revealed that none of the plasmids of PIMB10EC27 was associated with colistin resistance (data not shown). In another context, encoding genes involved in the attachment of L-Ara-4N and P-EtN to LPSs of PIMB10EC27, including phoP, phoQ, pmrB, pmrA, pmrC, mgrB, lpx, and arn were examined, and amino acid replacements G273D in PmrB and F515S in PmrC of PIMB10EC27 were recorded. The role of these mutations in colistin resistance remains to be investigated.
In summary, we successfully constructed the complete genome of multidrug-resistant E. cloacae strain PIMB10EC27 carrying the novel genomic context of blaNDM-1. To our knowledge, this is the first report in Vietnam of a carbapenemase-producing E. cloacae strain that was also resistant to colistin. PIMB10EC27 was isolated from a patient not being treated with colistin and not identified as a case of a plasmid-mediated colistin-resistance mechanism.
In the battle against multidrug-resistant bacteria, particularly carbapenem-resistant bacteria, colistin has been considered to remain effective. In such context, this report of a clinical isolate that is resistant to both colistin and carbapenem raises a high concern over future clinical management and infection control.
Nucleotide sequence accession numbers. The complete nucleotide sequences of PIMB10EC27 have been deposited in GenBank under accession numbers CP020089-CP020091.
(
Abbreviation list
MIC, minimum inhibitory concentration; E. cloacae, Enterobacter cloacae; CLSI, Clinical and Laboratory Standards Institute; NDM-1, New Delhi metallo-β-lactamase 1; ORF, Open Reading Frame.
Data availability
The datasets used and/or analysed during the current study are available from the corresponding author on reasonable request.
Acknowledgments
We thank Binh Dan Hospital in Ho Chi Minh City, Vietnam for providing the isolated bacteria for the research. We greatly appreciate Dr. James W. Jones for his valuable review. This work was supported by the US Armed Forces Research Institute of Medical Sciences (AFRIMS), the National Institute of Health (NIH), and the Grant for Joint Research Project of the Research Institute for Microbial Diseases, Osaka University.
Author contributions
VC and TI conceived and designed the study. TDLH, PTBT, LL, and LKT collected samples and performed experiments, TDLH, HNLV, PTBT, LL, DM, SN, and MA performed data analysis, TDLH and HNLV wrote the paper. All authors contributed to drafting and revising the article, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Disclosure
Dr Tam-Duong Le-Ha reports grants from the US Armed Forces Research Institute of Medical Sciences (AFRIMS), grants from the National Institute of Health (NIH), and grants from the Research Institute for Microbial Diseases, Osaka University, during the conduct of the study. Ms Lien Le reports grants from the US Armed Forces Research Institute of Medical Sciences (AFRIMS), grants from the National Institute of Health (NIH), and grants from the Research Institute for Microbial Diseases, Osaka University, during the conduct of the study. Miss Hong-Ngoc Le-Vo reports grants from US Armed Forces Research Institute of Medical Sciences, grants from National Institute of Health, and grants from Grant for Joint Research Project of the Research Institute for Microbial Diseases, Osaka University, during the conduct of the study. Dr Mizue Anda has nothing to disclose. Dr Daisuke Motooka has nothing to disclose. Dr Shota Nakamura has nothing to disclose. Dr Linh Khanh Tran reports grants from the US Armed Forces Research Institute of Medical Sciences (AFRIMS), grants from the National Institute of Health (NIH), and grants from the Grant for Joint Research Project of the Research Institute for Microbial Diseases, Osaka University, during the conduct of the study. Dr Phuong Thi-Bich Tran reports grants from The US Armed Forces Research Institute of Medical Sciences (AFRIMS), grants from The National Institute of Health (NIH), and grants from The Research Institute for Microbial Diseases, Osaka University, during the conduct of the study. Dr Tetsuya Iida has nothing to disclose. Dr Van Cao reports grants from the US Armed Forces Research Institute of Medical Sciences (AFRIMS), grants from the National Institute of Health (NIH), and grants from the Research Institute for Microbial Diseases, Osaka University, during the conduct of the study.
References
1. Papp-Wallace KM, Endimiani A, Taracila MA, Bonomo RA. Carbapenems: past, present, and future. Antimicrob Agents Chemother. 2011;55(11):4943–4960. doi:10.1128/AAC.00296-11
2. Nordmann P, Poirel L. The difficult-to-control spread of carbapenemase producers among Enterobacteriaceae worldwide. Clin Microbiol Infect. 2014;20. doi:10.1111/1469-0691.12742
3. Yong D, Toleman MA, Giske CG, et al. Characterization of a new metallo-beta-lactamase gene, bla(NDM-1), and a novel erythromycin esterase gene carried on a unique genetic structure in Klebsiella pneumoniae sequence type 14 from India. Antimicrob Agents Chemother. 2009;53(12):5046–5054. doi:10.1128/AAC.00774-09
4. Liu Z, Wang Y, Walsh TR, et al. Plasmid-mediated novel blaNDM-17 gene encoding a carbapenemase with enhanced activity in a sequence type 48 Escherichia coli strain. Antimicrob Agents Chemother. 2017;61(5). doi:10.1128/AAC.02233-16.
5. Poirel L, Bonnin RA, Boulanger A, Schrenzel J, Kaase M, Nordmann P. Tn125-related acquisition of bla(NDM)-like genes in Acinetobacter baumannii. Antimicrob Agents Chemother. 2012;56(2):1087–1089. doi:10.1128/AAC.05620-11
6. American Thoracic Society; Infectious Diseases Society of America. Guidelines for the management of adults with hospital-acquired, ventilator-associated, and healthcare-associated pneumonia. Am J Respir Crit Care Med. 2005;171(4):388–416. doi: 10.1164/rccm.200405-644ST.
7. Falagas ME, Kasiakou SK. Colistin: the revival of polymyxins for the management of multidrug-resistant gram-negative bacterial infections. Clin Infect Dis. 2005;40(9):1333–1341. doi:10.1086/429323
8. Catry B, Cavaleri M, Baptiste K, et al. Use of colistin-containing products within the European Union and European Economic Area (EU/EEA): development of resistance in animals and possible impact on human and animal health. Int J Antimicrob Agents. 2015;46(3):297–306. doi:10.1016/j.ijantimicag.2015.06.005
9. Bialvaei AZ, Kafil HS. Colistin, mechanisms and prevalence of resistance. Curr Med Res Opin. 2015;31(4):707–721. doi:10.1185/03007995.2015.1018989
10. Manohar P, Shanthini T, Ayyanar R, et al. The distribution of carbapenem- and colistin-resistance in Gram-negative bacteria from the Tamil Nadu region in India. J Med Microbiol. 2017;66(7):874–883. doi:10.1099/jmm.0.000508
11. Poirel L, Jayol A, Nordmann P. Polymyxins: antibacterial activity, susceptibility testing, and resistance mechanisms encoded by plasmids or chromosomes. Clin Microbiol Rev. 2017;30(2):557–596. doi:10.1128/CMR.00064-16
12. Musil I, Jensen V, Schilling J, Ashdown B, Kent T. Enterobacter cloacae infection of an expanded polytetrafluoroethylene femoral-popliteal bypass graft: a case report. J Med Case Rep. 2010;4:131. doi:10.1186/1752-1947-4-131
13. Sanders WE, Sanders CC. Enterobacter spp.: pathogens poised to flourish at the turn of the century. Clin Microbiol Rev. 1997;10(2):220–241.
14. Lee Y, Choi H, Yum JH, et al. Molecular mechanisms of carbapenem resistance in Enterobacter cloacae clinical isolates from Korea and clinical outcome. Ann Clin Lab Sci. 2012;42(3):281–286.
15. Kiedrowski LM, Guerrero DM, Perez F, et al. Carbapenem-resistant Enterobacter cloacae isolates producing KPC-3, North Dakota, USA. Emerg Infect Dis. 2014;20(9):1583–1585. doi:10.3201/eid2009.140344
16. Wilson BM, El Chakhtoura NG, Patel S, et al. Carbapenem-resistant Enterobacter cloacae in patients from the US Veterans Health Administration, 2006–2015. Emerg Infect Dis. 2017;23(5):878–880. doi:10.3201/eid2305.162034
17. Norgan AP, Freese JM, Tuin PM, Cunningham SA, Jeraldo PR, Patel R. Carbapenem- and colistin-resistant Enterobacter cloacae from Delta, Colorado, in 2015. Antimicrob Agents Chemother. 2016;60(5):3141–3144. doi:10.1128/AAC.03055-15
18. Huang L, Wang X, Feng Y, Xie Y, Xie L, Zong Z. First identification of an IMI-1 carbapenemase-producing colistin-resistant Enterobacter cloacae in China. Ann Clin Microbiol Antimicrob. 2015;14:51. doi:10.1186/s12941-015-0113-1
19.
20.
21. Zankari E, Hasman H, Cosentino S, et al. Identification of acquired antimicrobial resistance genes. J Antimicrob Chemother. 2012;67(11):2640–2644. doi:10.1093/jac/dks261
22. Belén A, Pavón I, Maiden MCJ. Multilocus sequence typing. Methods Mol Biol (Clifton, NJ). 2009;551:129–140.
23. Varani AM, Siguier P, Gourbeyre E, Charneau V, Chandler M. ISsaga is an ensemble of web-based methods for high throughput identification and semi-automatic annotation of insertion sequences in prokaryotic genomes. Genome Biol. 2011;12(3):R30. doi:10.1186/gb-2011-12-3-r30
24. Afgan E, Baker D, van den Beek M, et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2016 update. Nucleic Acids Res. 2016;44(W1):W3–W10. doi:10.1093/nar/gkw343
25. Carattoli A, Zankari E, Garcia-Fernandez A, et al. In silico detection and typing of plasmids using PlasmidFinder and plasmid multilocus sequence typing. Antimicrob Agents Chemother. 2014;58(7):3895–3903. doi:10.1128/AAC.02412-14
26. Martinez-Martinez L, Pascual A, Jacoby GA. Quinolone resistance from a transferable plasmid. Lancet. 1998;351(9105):797–799. doi:10.1016/S0140-6736(97)07322-4
27. Tran JH, Jacoby GA, Hooper DC. Interaction of the plasmid-encoded quinolone resistance protein Qnr with Escherichia coli DNA gyrase. Antimicrob Agents Chemother. 2005;49(1):118–125. doi:10.1128/AAC.49.1.118-125.2005
28. Jacoby GA, Walsh KE, Mills DM, et al. qnrB, another plasmid-mediated gene for quinolone resistance. Antimicrob Agents Chemother. 2006;50(4):1178–1182. doi:10.1128/AAC.50.4.1178-1182.2006
29. Poirel L, Lagrutta E, Taylor P, Pham J, Nordmann P. Emergence of metallo-β-lactamase NDM-1-producing multidrug-resistant Escherichia coli in Australia. Antimicrob Agents Chemother. 2010;54(11):4914–4916. doi:10.1128/AAC.00878-10
30. Zurfluh K, Poirel L, Nordmann P, Klumpp J, Stephan R. First detection of Klebsiella variicola producing OXA-181 carbapenemase in fresh vegetable imported from Asia to Switzerland. Antimicrob Resist Infect Control. 2015;4(1):38. doi:10.1186/s13756-015-0080-5
31. Murata T, Ohnishi M, Ara T, et al. Complete nucleotide sequence of plasmid Rts1: implications for evolution of large plasmid genomes. J Bacteriol. 2002;184(12):3194–3202.
32. Liu YY, Wang Y, Walsh TR, et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect Dis. 2016;16(2):161–168. doi:10.1016/S1473-3099(15)00424-7
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.