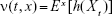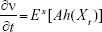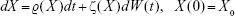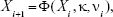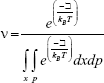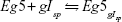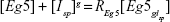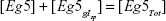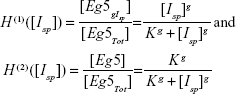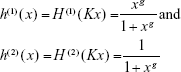Back to Journals » International Journal of Nanomedicine » Volume 13
Cancer drug therapy and stochastic modeling of “nano-motors”
Authors Sherin L, Farwa S, Sohail A , Li Z , Bég OA
Received 10 July 2018
Accepted for publication 30 August 2018
Published 15 October 2018 Volume 2018:13 Pages 6429—6440
DOI https://doi.org/10.2147/IJN.S168780
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Thomas Webster
Lubna Sherin,1 Shabieh Farwa,2 Ayesha Sohail,3 Zhiwu Li,4,5 O Anwar Bég6
1Department of Chemistry, COMSATS University Islamabad, Lahore 54000, Pakistan; 2Department of Mathematics, COMSATS University Islamabad, Wah Cantt, Pakistan; 3Department of Mathematics, COMSATS University Islamabad, Lahore 54000, Pakistan; 4Institute of Systems Engineering, Macau University of Science and Technology, Taipa, Macau; 5School of Electro-Mechanical Engineering, Xidian University, Xi’an 710071, China; 6Fluid Mechanics, Spray Research Group, Mechanical and Petroleum Engineering, School of Computing, Science and Engineering, G77, University of Salford, Manchester M54WT, UK
Background: Controlled inhibition of kinesin motor proteins is highly desired in the field of oncology. Among other interventions, there exists “targeted chemotherapeutic regime/options” of selective Eg5 competitive and allosteric inhibitors, inducing cancer cell apoptosis and tumor regression with improved safety profiles.
Research question: Though promising, such studies are still under clinical trials, for the discovery of efficient and least harmful Eg5 inhibitors. The aim of this research was to bridge the computational modeling approach with drug design and therapy of cancer cells.
Methods: A computational model, interfaced with the clinical data of “Eg5 dynamics” and “inhibitors” via special functions, is presented in this article. Comparisons are made for the drug efficacy, and the threshold values are predicted through numerical simulations.
Results: Results are obtained to depict the dynamics induced by ispinesib, when used as an inhibitor of kinesin Eg5, on cancer cell lines.
Keywords: drug efficient model, bipolar spindle, Eg5, cancer, mitotic arrest, Eg5 inhibitors
Introduction
The motor proteins are also termed as the “nano-motors” due to their nanoscale functionality and integrity in a sophisticated manner. These motors support the fundamental cellular processes and act as nanometer-sized machines, converting chemical energy into mechanical work. For the past few years, the fundamental principles and mechanisms of molecular motor dynamics have remained a topic of debate in the field of nanomedicine.1,2
In recent years, mitotic catastrophe (activated during mitosis) has been employed as promising onco-suppressive therapeutic strategy. In current clinical strategies, various antimitotic cancer drugs such as vinca alkaloids, taxanes, and epothilone derivatives, targeting microtubules during cell division, have proved to be clinically most successful anticancer agents in bladder, ovarian, breast, lung, and head malignancies.3 Mitosis is a fundamental cell division process that ensures accurate division of sister chromatids to the daughter cells. This vital function, coordinated by mitotic spindle made up of microtubules, has emerged as validated chemotherapeutic target. Microtubules are dynamic polymers of α/β tubulin dimer where chromosomes attach and segregate on mitotic spindle during cell division.4,5 Antimitotic drugs perturb it by suppressing MT dynamics, thus ensuing an inappropriate chromosome arrangement. Consequently spindle assembly check points are activated that trigger cancer cell death via mitotic arrest.6 Though MT targeting drugs are currently cancer-specific with good clinical outcome but devastating side effects, dose limiting toxicities are the main challenges associated with them as they hit tubulin which is involved in multiple intracellular processes. Moreover, due to innate or acquired resistance, patient relapse is a common condition encountered while administering these drugs.3 This has led to seek for novel alternative drug targets and to develop new generation mitosis-specific agents.
Kinesins spindle motor proteins and the microtubules work together and are considered potential therapeutic target in tumors. Being central in driving bipolar spindle and subsequent chromosome separation, they provide a chance for the development of specific antimitotic agents that may overcome resistance along with better side effects profile. Kinesins constitute a super family of >650 known motor proteins. These motor proteins move unidirectionally along the microtubule to perform exclusive functions and harness energy by ATP hydrolysis in all eukaryotic cells.7 The key cellular function that they perform within intracellular terrain includes microtubule remodeling, vesicular trafficking, mitotic spindle assembly, and chromosome segregation in dividing cells.8 The structure of all kinesins consists of a head, stalk, and tail domain. Head of kinesins contains 340 amino acid motor domain provided with an ATP-binding pocket and microtubule-binding interface. Being ATPases, kinesins hydrolyze ATP via motor domain to release energy required for the movement along MT. The stalk is used for dimerization and oligomerization while tail is involved in interaction with cargo.9 Each 8-nm-long tubulin dimer of polymer microtubule serves as one binding site for translocation of kinesin motor domain along microtubule powered by ATP hydrolysis.10
The human kinesins contribute significantly in cancer development, progression and drug resistance. High expression in various malignancies has directed huge amount of work toward targeting these motor proteins through chemotherapeutic intervention. Abnormal proliferation is the fundamental feature of all tumors. Proliferating neoplasmic cells strictly follow well-ordered events of cell cycle comprising growth S-phase where DNA replication takes place followed by mitosis for cell division. Orderly progression of cell cycle phases is mediated by checkpoints and is stalled under unsatisfied conditions leading to cell death. First identified motor protein was Eg5, a member of kinesin-5 family involved in the establishment of bipolar spindle. Downregulation of this protein results in a defected monopolar phenotype. They are involved in interconnection of chromosomes with spindle, movement of MY dynamics and spindle length maintenance.
Eg5: An explicit mitotic kinesin motor
Eg5, also known as kinesin spindle protein, KIF11, or Kinesin-5, is the simplest yet key player of the mitotic apparatus. Eg5, a human gene product encoded by KIF11 gene located at 10q24.1, is of particular interest because of its potential as a target for therapeutic intervention.11 In humans it is expressed in bone marrow, thymus, testis, and tonsils while is absent in adult post-mitotic central nervous system.12 It is a homotetrameric plus end directed N-terminal microtubule-based motor protein with a catalytic motor/ATPase domain to interact with ATP and microtubules to modulate the dynamics and organization of MT arrays. It can cross-link and slide antiparallel MTs to congregate mitotic spindle while it resides along parallel MT at both poles (Figure 1). A defective spindle hampers normal chromosomal segregation leading to mitotic arrest via check point protein activation.13 Metaphase spindle equator comprises antiparallel overlapped MTs, while microtubules with parallel orientation dominate near spindle poles. During bipolar spindle fabrication, Eg5 slides apart antiparallel microtubule filaments by pacing toward plus ends of each microtubule. It generates outward pushing force on centrosomes via cross-linking antiparallel microtubules to slide them away from each other. Eg5 inhibition arrests cells in mitosis with unseparated centrosomes, thus suppressing bipolar spindle production. It is found that Eg5 may also act as molecular brakes and restrict the movement of overlapping antiparallel filaments. Elongated anaphase spindle has been reported in the case of loss of Eg5 activity during cell division.14 Nevertheless, the magnitude of these braking force has not been directly calculated, and scaling of these forces with respect to relative velocity, orientation or overlap is unclear yet. Bipolar architecture of mitotic spindle is critical for proper segregation of chromosomes in daughter cells. The work of Shimamoto et al15 provides a conceptual framework toward mitotic apparatus complexity and an insight into self-organization of mitotic spindle. After probing feedback mechanism between Eg5 function and microtubule architecture (velocity and geometry) through experimentations, Shimamoto et al15 suggested that spindle assembly can regulate force generation for self-organization. The forces generated by Eg5 ensembles scale linearly with respect to motor number and length of microtubule overlap. They proposed that Eg5 ensembles can operate as a “force converter,” decoding microtubule regular features such as orientation and overlap length into a distinct force signature. Equator of metaphase spindle is a dynamic region where antiparallel microtubules keep gliding unceasingly at a speed of 23 mm/min (30–50 nm/s) while their minus ends are directed toward opposite spindle poles. This dynamic antiparallel setup ensembles of Eg5 generate either pushing or mechanical resistance, magnitude of that is directly proportional to MT sliding velocity, different from unloaded state, and length of overlaid microtubule. A linear integration of force output is attained by plus end movement of Eg5 ensembles in antiparallel geometry while stochastic force output is produced by Eg5 stepping across parallel filaments.15
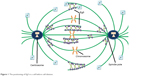  | Figure 1 The positioning of Eg5 in a cell before cell division. |
Eg5 is assumed to be a promising therapeutic target as it is overexpressed in actively proliferating solid tumors in pancreatic, lung, bladder, ovarian, and breast cancers.11,16 Elevated levels of Eg5 are considered tumorous as overexpression results in blast crisis chronic myeloid leukemia, activation in mouse B-cell leukemia, and triggering of genomic instability in transgenic mice.17,18 Its oncogenic potential is further verified by the observation that it elicits anti-proliferation of all-trans-retinoic acid in pancreatic cancer cell lines.19 Experimental data reveal that Eg5 promotes active cancer cell proliferation, colonization, and tumorigenesis in pancreatic malignancy in mice. It was found that Eg5 overexpression affected spindle morphology and resulted in multipolar spindle formation. Moreover, a significant increase in multinucleate interphase cell population was observed, resulting in the accumulation of polyploid cells, a condition strongly linked with genomic instability leading to cancer. Recent studies have shown that kinesin-5 also contributes in evolution of cancer toward metastases via spindle length scaling.20 Studies of many cancer cell lines along with in vivo human xenograft models have shown that Eg5 inhibition ceases bipolar mitotic spindle morphogenesis and ends up in loss-of-function phenotype as a monopolar spindle monoester with chromosome distribution in rosette-like configuration. This mitotic spindle disaster activates checkpoint proteins and induces mitotic arrest and subsequent apoptosis of cancer cell.21,22
Upregulation of Eg5, a potent biomarker in various malignancies, needs to be targeted for timely and efficacious treatment. In contrast to traditional antimitotic drugs targeting MT in actively proliferating as well as normal cells, severe side effects are not expected from Eg5 inhibitors for being target-specific for mitotically active cells. Small-molecule inhibitors targeting Eg5 signify a new generation target-specific anti-cancer agents that are currently undergoing Phase I and Phase II clinical trials.23 Mitotic arrest has been induced by these inhibitors by obstructing Eg5-dependent MT motility resulting in aberrant monoastral form. The first reported potent Eg5 inhibitor was monastrol that caused mitotic arrest without interfering with MT dynamics. Afterwards many other allosteric inhibitors have been reported, belonging to different chemical classes such as quinazoline, imidazoles, thiadiazoles, carbolines, dihydropyrazoles, isoquinolines, and benzimidazoles.
In silico biology for representing stochastic intracellular changes
Stochastic description of intracellular changes is well-demonstrated in computational systems biology. However, several models are usually generalization of the actual phenomena and the corresponding parameters may be imprecise. In such cases, the numerical tools can help to investigate the analysis results and their level of sensitivity corresponding to the parametric perturbations. In this paper, a stochastic simulation algorithm is adopted to analyze the variation in dynamics of the kinesin motor proteins in three cases (normal, cancerous, and drug-treated cells). We aim to provide the reader a simple yet dynamic model to demonstrate the self-organization of stochastic time stepping of the kinesin motor proteins under aforementioned conditions.
Problem statement
The study of motor proteins and microtubules is a rich field of research with a long list of open problems. Different approaches are available in the recent literature to model the stochastic transport, mitotic spindle dynamics in cancerous cells, and the self-organization of subcellular structures, the receptor trafficking, protein–DNA interactions, nuclear transport, membrane diffusion, and virus trafficking. This motivates us to study and compare change in dynamics of Eg5 motor proteins in normal and highly proliferating cancerous cells in addition to change exhibited by diseased cells in the presence of obstacles and transient traps, ie, inhibitors. In normal cells, metaphase spindle is a highly dynamic entity. Being made up of microtubules and associated kinesin motor proteins, it experiences huge fluctuations and directed fluxes in both physical and chemical processes. However, average number, position, and functions of all spindle constituents remain steady over time by virtue of its ability to correct transient changes. In cancerous cells, firmness of this steady-state is disturbed, and it experiences huge physical and chemical perturbations with evident incapability to recover or to correct transient fluctuations in morphology and position. Consequently an aggressive, unchecked mitotic division takes place leading to high proliferation rate of tumor cells.
This entire process is really rapid, and the laboratory experiments alone are not enough to capture the rapid alterations caused by randomness. The literature provides an evidence that the computational biology (in silico research) has always served successfully to model the inter and intracellular dynamics of malignant cells.24 This motivates us to use a computational model with some advanced conditions, in order to demonstrate the “Eg5 motor proteins based malignant cell’s intra-dynamics.” The research methodology encompasses the stochastic behavior of Eg5 proteins and its response to both the tumor disorders (proliferations) and the drug therapy. In this article, after careful analysis of the clinical trials, we have selected a special type of inhibitor “Ispinesib ()” which is believed to control the overexpression of the motor protein Eg520,23,25 in tumor cells. After the selection, we have extended the stochastic model available in the literature26–28 inline to the given biological problem. We have considered the Hill function formalism to demonstrate the addition of drug in the model as a reflection of its role in the laboratory-generated experiments.25 In this article, we have emphasized on the fact that the stochastic modeling of the dynamical behavior of motor proteins can help predict the outcomes of cancer drug therapy. The inhibition of Eg5 motor protein, a key mitotic kinesin motor protein involved in aggressive mitotic activity of cancerous cells, is targeted in this study to elaborate this statement. We have presented some results based on the mathematical model and the parametric values to demonstrate the efficiency of the computational tool to make important predictions. This will help improve the ongoing clinical trials on the control of Eg5 via inhibitors29,30 in cancerous cells.
Kolmogorov backward equation
The kinesin motor proteins are involved in the mitotic cell division. The modeling and simulation of kinesin dynamics and their mobility along the microtubules is really challenging in a normal (case 1), tumor (case 2), and treated cell (case 3). Several factors are involved, such as a variation in the frequency of the release of the proteins hence change in force and motion produced via catalysis in three different cases, there can be various time scales in operation, and the entire process is inherently stochastic. The Markov process (which is also famous as “Chemical Master Equation (CME)” framework) is appropriate to conduct such an analysis.
For the clear understanding of the molecular motor protein dynamics, in this article, a discrete approach is adopted and a stochastic model is considered. The probability of the molecule in a certain ith state at a time t can help to demonstrate the dynamics, provided that the backward Kolmogorov equation governs its transient dynamics.
Let Xt represent an I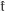 o diffusion in R
o diffusion in R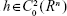 and taking τ=t in Dynkin’s formula, we have
and taking τ=t in Dynkin’s formula, we have
|
|
It is obvious that ν(t, x) is differentiable with respect to t and
|
|
In the above equation, E
Setting and definitions
Taking into account an n-dimensional stochastic differential equation
|
|
Let E be the space that may be specified as either E=R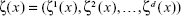 where each of ζ
where each of ζ
Consider a discrete numerical approximation of Equation 3 given as follows
|
|
where Xi∈E, ∀i ≥ 0 and Φ(., κ, νi):E→E is the discrete numerical flow, κ stands for the time-step size and νi represents a random vector.
In vivo and in vitro studies and the theoretical model
In this section, we discuss some of the experimental studies and interface with our theoretical model.
Data-based stochastic effects of EG5 inhibitors
Tumor cells and Eg5 data
Oncogenic role of Eg5 has been established by a large body of experiments, and it is identified as a potent prognostic biomarker in various malignancies. Lu et al31 investigated the correlation between upregulated expressions of Eg5 and clinicopathological characteristics in laryngeal squamous cell carcinoma (LSCC) patients. For that purpose immunohistochemistry (IHC) analysis in 137 LSCC cases along with one-step qPCR test with 20 fresh-frozen LSCC samples was done. Significant higher Eg5 protein level in comparison with corresponding non-cancerous tissues were correlated with lymph node metastasis and TNM stage, independent factors to envisage critical prognosis for LSCC patients.31 Similarly correlation between clinicopathological characteristics and Eg5 expression in non-muscle invasive urothelial carcinoma was investigated by analyzing large number of IHC specimens including grade: G1, 32 cases; G2, 92 cases; and G3, 39 cases; Stage: pTa, 49 cases and pT1, 114 cases. A total of 163 non-muscle invasive cases were analyzed via survival analysis to find out the prognostic significance of Eg5 immunoreactivity. A strong imperative connection between Eg5 overexpression and tumor grade (P=0.006) and tumor stage (P=0.057) was discovered. It was concluded that Eg5 overexpression signifies a self-regulating prognostic factor in envisaging initial intravesical recurrence in non-muscle invasive bladder cancer patients.16 Sun et al32 evaluated the prognostic significance of Eg5 upregulation in renal cell carcinoma (RCC) patients via IHC and correlated clinicopathological parameters using the univariate and multivariate analyses of 164 patients and 164 tissue specimens, regularly followed from 5 to 80 months. They suggested that Eg5 may serve as a prognostic factor for renal cancer prediction, evolution, and appropriate treatments. Experimental data indicated a significant relation between tumor nuclear grade (P=0.019), stage (P=0.007), and size (P=0.033) with Eg5 expression for recurrence-free RCC patient survival.32 Eg5 expression has also been associated with occurrence and rapid metastasizing ability of pancreatic cancer. In pancreas cancer cell lines, high expression of Eg5 promotes tumorigenesis by boosting cell proliferation in an ATPase activity-dependent manner. It stimulates multipolar spindle formation and consequent multi-nucleation leading to polyploid cell production. It was discovered that high expression of Eg5 encourages anchorage independent cell growth and tumor metastases in mice model.11
Eg5 data after inhibition
Due to close association of Eg5 with tumorigenesis, metastasis, and tumor drug resistance, Eg5/KSP has emerged as a promising target for anticancer therapeutic agents, and various KSP inhibitors are under clinical trials as monotherapy or adjuvant therapy candidate. Eg5 inhibitors cause cell cycle arrest during mitosis by targeting KSP via contrasting mechanisms. ATP-uncompetitive inhibitors bind near loop L5 to stabilize the bound nucleotide and trap the motor in a weak binding state. Some potent inhibitors of this category exhibiting clinical efficacy include monastrol, filanesib, litronesib, K858, S-trityl-L-cysteine (STLC), and ispinesib ()25,33–35 (in Figure 2, the kinesin-5 inhibitor complex is presented). Whereas thiazole FCPT act as an ATP competitive inhibitor and bind directly to the nucleotide binding active site.36 Some biaryl inhibitors such as PVZB1194 and GSK-133 act as allosteric competitive inhibitors of ATP binding and bind near the α4–α6 interface.35,37 BRD9876 is a similar type of ATP non-competitive inhibitor that preferentially bind to microtubule bound Eg5.38 These inhibitors may act as a chemical probe to understand and modulate the Eg5 activity within a cell.
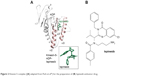  | Figure 2 Kinesin-5 complex ([A] adapted from Park et al39) for the preparation of (B) Ispinesib antitumor drug. |
Detailed analysis depicted different distinct effects of aforementioned inhibitors on MT stability and spindle integrity. L5 inhibitors obliterate Eg5 microtubule stabilizing ability causing metaphase spindle down fall in contrast to rigor inhibitor BRD9876 that stabilizes microtubule and hence metaphase spindle by suppressing MT depolymerization.
Computational model interfaced with real data
In the recent literature,40 the motor proteins and their associated stochastic dynamics are studied extensively due to their crucial role, especially in mitotic cell division. These motor proteins are mostly believed to obey periodic motion. Such a motion usually takes place along the potential, which actually describes the distinct biochemical states of the motor. There are several types of motor proteins involved in mitosis process in human cell. Among these, EG5 motor proteins play a dominant role in the cancer invasion. It has been verified through laboratory generated experiments, (as reported by Wakui et al25) that EG5 is overexpressed in malignant cells. We have focused on the dynamics of this special type of motor proteins using a theoretical model (that is explained in detail in “Kolmogorov backward equation” and “Setting and definitions” sections), where ν(x, t) represents the probability (or more precisely the probability density) for EG5 to be found at location x at time t. The conditions to solve the Kolmogorov backward equation were considered to be periodic. Since at the micro level, the probability density function can be written as a function of mean square displacement, therefore as the mean square displacement of EG5 proteins changed, we obtained a variation in the conditions interfaced in the numerical solver. This can be elaborated with the aid of the probability density function
|
|
where p is the momentum of the motor protein (depending on the rate of change of the displacement),  is the sum of kinetic and potential energies (as a function of displacement), kB is the Boltzmann constant and T is the absolute temperature. In this article, the parametric values were adapted from the data presented by Chen et al.41
is the sum of kinetic and potential energies (as a function of displacement), kB is the Boltzmann constant and T is the absolute temperature. In this article, the parametric values were adapted from the data presented by Chen et al.41
Another important feature of this study is that the drug therapy is synchronized in the model through the Hill’s function formulation.42 The EG5 molecules were considered as the enzymatic receptors and the drug molecules were considered as the inhibiting ligands. The simple manipulation of the chemical equations leads to an important formula, which is used extensively in the literature,43 but for the very first time has been synchronized with the sinusoidal drift (ϱ in Equation 3) of the Kolmogorov backward equation. The Hill function for our model is derived using the following formalism:
Hill functions
The reaction, in which g ligand molecules Isp bind the receptor Eg5, is given as follows:
|
|
In case of chemical equilibrium, the following relation is satisfied
|
|
In the above equation, K represents the reaction dissociation constant and [Eg5] and [Isp] stand for the concentration of the chemical species Eg5 and Isp, respectively. If we consider a constant number of receptors,
|
|
Equations 6 and 7 then lead to the following fractions:
|
|
where both H(1) and H(2) are known as Hill functions, representing the fraction of occupied and free molecules Eg5 respectively, with K
|
|
This non-dimensional entity h(1)(x) when incorporated with the sinusoidal drift (ϱ in Equation 3) of the Kolmogorov backward equation works as a useful tool to trace the effect of the drug inhibitors on the Eg5 enzymes. This impact is demonstrated with the aid of graphical interpretation in Figure 3.
  | Figure 3 Drug therapy – theoretical strategy and the impact on drift. |
We have solved the Kolmogorov backward equation to understand the three cases, kinesin motor proteins dynamics in normal cell, tumor cell, and treated cell. Each case was differentiated from the other case, based on the initial dynamics (interfaced in the numerical computations). The finite element algorithm was applied, and the mass and stiffness matrices were obtained. The matrix exponential is computed using a scaling and squaring algorithm with a Pade approximation.
Results and discussion
Mathematical modeling of the stochastic intracellular perturbations in the expression and function of Eg5 in malignancy and on inhibition may prove to be fruitful in understanding the current challenges of cellular scale studies. In this model, an elevated expression of Eg5 generates excess mechanical forces for the movement of anti-parallel microtubules and perturbs the balance of forces normally required for bipolar spindle formation, thereby inducing multipolar spindle assembly and impairing chromosome segregation. The cells may undergo mitotic slippage due to the inability to correct errors and satisfy the spindle assembly checkpoint, resulting in cytokinesis arrest and abnormal proliferation. These events in turn lead to genomic instability and ultimately tumorigenesis. This model displays the aggressive stochastic behavior of kinesin-5 in cancerous cells and tamed behavior upon inhibition. To demonstrate this behavior, in this section, we have presented some results based on the experimental findings and numerical simulations.
Figures 4 and 5 depict the probability density function for the motor proteins in a normal cell and in a cell with malignancy. We can see that the shape changes from regular soliton-like structure to frequent humps.
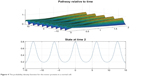  | Figure 4 The probability density function for the motor proteins in a normal cell. |
  | Figure 5 The probability density function for the motor proteins in a tumor cell when no drug was injected. |
From Figures 6 and 7 it is obvious that for lower values of g (that controls the drug concentration in our model), there is a decline in amplitude of the pathway and for increased values (as shown in Figure 8) it reduces asymptotically, the straight line after x=12 in Figure 8 shows apoptosis, that is, no motion of the motor protein.
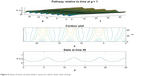  | Figure 6 States of motor protein inside a cancerous cell for lower dose of drug. |
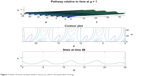  | Figure 7 States of motor protein inside a cancerous cell for increased dose of drug. |
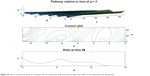  | Figure 8 States of motor protein of a treated cell, for enhanced drug dose; the straight line refers to no movement (ie, apoptosis). |
Conclusion
Different therapeutics have been designed to date to control the rapid proliferation of cancerous cells. These includes the cytotoxic antimitotic treatments, of which antitubulin compounds have received remarkable attention; however, such treatments are responsible for dose-limiting side effects. This has led the oncologists and biochemical engineers to design new mechanisms of action to reduce such “chemoresistance.” Mitosis-specific kinesin “Eg5,” which is a fundamental in mitotic progression, is considered as a target for designing such new anticancer agents.37,44 The recent clinical trials have proved that the kinesin Eg5 inhibitors have antiproliferative effect and induced apoptosis. Therefore, motivated by the stochastic nature of the process,45,46 we analyzed the effects induced by ispinesib, which is an inhibitor of kinesin Eg5, on cancer cell lines, with the aid of a theoretical model. The mathematical model is physically motivated by the existing models as well as the laboratory outcomes (depicted from the work of Chen et al41 specifically for ispinesib) and thus presents the dynamics to a better accuracy.
It is worth mentioning that no single drug designed for the Eg5 inhibition has progressed beyond clinical trials to date.47 Evidence is available in the literature, supporting the fact that different inhibitors target Eg5 differently,48 in normal, aberrant, and degenerative manner; furthermore, novel drug development strategies are under clinical trials.49,50 Chen et al41 investigated the effects of inhibitors on Eg5-mediated stabilization of dynamic microtubules and concluded that different Eg5 inhibitors inhibit spindle formation through contrasting mechanisms. Our future research will focus on the theoretical investigation of the contrasting effects of a group of inhibitors on Eg5 motor proteins.
In short, nanomotors are not always “the same but smaller.” Instead, sometimes “small is different” (Uzi Landman) applies, which creates opportunities to target their functionality for treatment and medication. Therefore, the sheer availability of data obtained from the clinical trials of “inhibition processes” of the nano-sized motor proteins is a great opportunity that calls for theoretical exploration.
Acknowledgments
The authors would like to acknowledge the support provided by NNSF and the Library staff (especially Ms Nasira Munir) of Comsats Institute of Information Technology, Lahore, Pakistan for their continued support. This work was supported in part by the National Natural Science Foundation of China under Grant Nos 61673309 and 61873342, and the Fundamental Research Funds for the Central Universities under Grant Nos JB160401 and JBG160415.
Disclosure
The authors report no conflicts of interest in this work.
References
Shenoy D, Fu W, Li J, et al. Surface functionalization of gold nanoparticles using hetero-bifunctional poly(ethylene glycol) spacer for intracellular tracking and delivery. Int J Nanomedicine. 2006;1(1):51–58. | ||
Banerjee A, Paluh JL, Mukherjee A, Kumar KG, Ghosh A, Naskar MK. Modeling the neuron as a nanocommunication system to identify spatiotemporal molecular events in neurodegenerative disease. Int J Nanomedicine. 2018;13:3105–3128. | ||
Jordan MA, Wilson L, Ann JM, Leslie W. Microtubules as a target for anticancer drugs. Nat Rev Cancer. 2004;4(4):253–265. | ||
Hayden JH, Bowser SS, Rieder CL, Bowser Samuel S, Rieder Conly L. Kinetochores capture astral microtubules during chromosome attachment to the mitotic spindle: direct visualization in live newt lung cells. J Cell Biol. 1990;111(3):1039–1045. | ||
Zhai Y, Kronebusch PJ, Simon PM, Borisy GG, Ye Z, Simon Patrick M. Microtubule dynamics at the G2/M transition: abrupt breakdown of cytoplasmic microtubules at nuclear envelope breakdown and implications for spindle morphogenesis. J Cell Biol. 1996;135(1):201–214. | ||
Rieder CL, Maiato H, Helder M. Stuck in division or passing through: what happens when cells cannot satisfy the spindle assembly checkpoint. Dev Cell. 2004;7(5):637–651. | ||
Rath O, Kozielski F, Oliver R, Frank K. Kinesins and cancer. Nat Rev Cancer. 2012;12(8):527–539. | ||
Endow SA, Endow Sharyn A. Microtubule motors in spindle and chromosome motility. Eur J Biochem. 1999;262(1):12–18. | ||
Miki H, Okada Y, Hirokawa N, Harukata M, Yasushi O, Nobutaka H. Analysis of the kinesin superfamily: insights into structure and function. Trends Cell Biol. 2005;15(9):467–476. | ||
Vale RD, Milligan RA, Vale Ronald D, Milligan Ronald A. The way things move: looking under the hood of molecular motor proteins. Science. 2000;288(5463):88–95. | ||
Liu M, Wang X, Yang Y, et al. Ectopic expression of the microtubule-dependent motor protein Eg5 promotes pancreatic tumourigenesis. J Pathol.2010;221(2):221–228. | ||
Ferhat L, Cook C, Chauviere M, et al. Expression of the mitotic motor protein Eg5 in postmitotic neurons: implications for neuronal development. J Neurosci. 1998;18(19):7822–7835. | ||
Compton DA. Spindle assembly in animal cells. Annu Rev Biochem. 2000;69:95–114. | ||
Rozelle DK, Hansen SD, Kaplan KB, Rozelle Daniel K, Hansen Scott D, Kaplan Kenneth B. Chromosome passenger complexes control anaphase duration and spindle elongation via a kinesin-5 brake. J Cell Biol. 2011;193(2):285–294. | ||
Shimamoto Y, Forth S, Kapoor TM, Yuta S, Scott F, Kapoor Tarun M. Measuring pushing and braking forces generated by ensembles of kinesin-5 crosslinking two microtubules. Dev Cell. 2015;34(6):669–681. | ||
Ding S, Xing N, Lu J, et al. Overexpression of Eg5 predicts unfavorable prognosis in non-muscle invasive bladder urothelial carcinoma. Int J Urol. 2011;18(6):432–438. | ||
Hansen GM, Justice MJ. Activation of Hex and mEg5 by retroviral insertion may contribute to mouse B-cell leukemia. Oncogene. 1999;18(47):6531–6539. | ||
Castillo A, Morse HC, Godfrey VL, Naeem R, Justice MJ. Overexpression of Eg5 causes genomic instability and tumor formation in mice. Cancer Res. 2007;67(21):10138–10147. | ||
Kaiser A, Brembeck FH, Nicke B, Wiedenmann B, Riecken EO, Rosewicz S. All-trans-retinoic acid-mediated growth inhibition involves inhibition of human kinesin-related protein HsEg5. J Biol Chem. 1999;274(27):18925–18931. | ||
Yang CF, Tsai WY, Chen WA, et al. Kinesin-5 contributes to spindle-length scaling in the evolution of cancer toward metastasis. Sci Rep. 2016;6:35767. | ||
Masuda A, Maeno K, Nakagawa T, Saito H, Takahashi T. Association between mitotic spindle checkpoint impairment and susceptibility to the induction of apoptosis by anti-microtubule agents in human lung cancers. Am J Pathol. 2003;163(3):1109–1116. | ||
Anne S, Lane Heidi A, Pierre D’Hérin, Maryannick H, Michel K, Niggt Erich A. Phosphorylation by p34cdc2 regulates spindle association of human Eg5, a kinesin-related motor essential for bipolar spindle formation in vivo. Cell. 1995;83:1159–1169. | ||
Duhl DM, Renhowe PA. Inhibitors of kinesin motor proteins – research and clinical progress. Curr Opin Drug Discov Devel. 2005;8(4):431–436. | ||
Sohail A, Li Z. Computational Nano-Bio-Technology. Hoboken (NJ): Wiley. 2018. | ||
Wakui H, Yamamoto N, Kitazono S, et al. A phase 1 and dose-finding study of LY2523355 (litronesib), an Eg5 inhibitor, in Japanese patients with advanced solid tumors. Cancer Chemother Pharmacol. 2014;74(1):15–23. | ||
Giuseppe DP. Kolmogorov Equations for Stochastic PDE’s with Multiplicative Noise in Stochastic Analysis and Applications. New York: Springer; 2007:235–263. | ||
Tabony J, James T. Morphological bifurcations involving reaction-diffusion processes during microtubule formation. Science. 1994;264(5156):245–249. | ||
Goychuk I, Kharchenko VO, Metzler R, Igor G, Ralf M. How molecular motors work in the crowded environment of living cells: coexistence and efficiency of normal and anomalous transport. PLoS One. 2014;9(3):e91700. | ||
Evi GP, Giuseppe G. Microtubules in Prostate Cancer in Precision Molecular Pathology of Prostate Cancer. New York: Springer; 2018:439–453. | ||
Lang Patrick Y, Gershon Timothy R. A new way to treat brain tumors: targeting proteins coded by microcephaly genes? Brain tumors and microcephaly arise from opposing derangements regulating progenitor growth. Drivers of microcephaly could be attractive brain tumor targets. BioEssays. 2018;40:1700243. | ||
Lu M, Zhu H, Wang X, et al. The prognostic role of Eg5 expression in laryngeal squamous cell carcinoma. Pathology. 2016;48(3):214–218. | ||
Sun D, Lu J, Ding K, et al. The expression of Eg5 predicts a poor outcome for patients with renal cell carcinoma. Med Oncol. 2013;30(1):476. | ||
Debonis S, Skoufias DA, Lebeau L, et al. In vitro screening for inhibitors of the human mitotic kinesin Eg5 with antimitotic and antitumor activities. Mol Cancer Ther. 2004;3(9):1079–1090. | ||
Nakai R, Iida S, Takahashi T, et al. K858, a novel inhibitor of mitotic kinesin Eg5 and antitumor agent, induces cell death in cancer cells. Cancer Res. 2009;69(9):3901–3909. | ||
Luo L, Carson JD, Molnar KS, et al. Conformation-dependent ligand regulation of ATP hydrolysis by human KSP: activation of basal hydrolysis and inhibition of microtubule-stimulated hydrolysis by a single, small molecule modulator. J Am Chem Soc. 2008;130(24):7584–7591. | ||
Rickert KW, Schaber M, Torrent M, et al. Discovery and biochemical characterization of selective ATP competitive inhibitors of the human mitotic kinesin KSP. Arch Biochem Biophys. 2008;469(2):220–231. | ||
Yokoyama H, Sawada J, Katoh S, et al. Structural basis of new allosteric inhibition in Kinesin spindle protein Eg5. ACS Chem Biol. 2015;10(4):1128–1136. | ||
Chattopadhyay S, Stewart AL, Mukherjee S, et al. Niche-based screening in multiple myeloma identifies a kinesin-5 inhibitor with improved selectivity over hematopoietic progenitors. Cell Rep. 2015;10:755–770. | ||
Park HW, Ma Z, Zhu H, Jiang S, Robinson RC, Endow SA. Structural basis of small molecule ATPase inhibition of a human mitotic kinesin motor protein. Sci Rep. 2017;7(1):15121. | ||
Bressloff Paul C. Stochastic Processes in Cell Biology. New York: Springer; 2014:41. | ||
Geng-Yuan C, You-Jung K, Sophia GA, et al. Eg5 inhibitors have contrasting effects on microtubule stability and spindle integrity depending on their modes of action. Biophys J. 2017;112:427a–428a. | ||
Weiss JN, Weiss James N. The Hill equation revisited: uses and misuses. FASEB J. 1997;11(11):835–841. | ||
Wender PA, Verma VA, Paxton TJ, Pillow TH. Function-oriented synthesis, step economy, and drug design. Acc Chem Res. 2008;41(1):40–49. | ||
Tao W, South VJ, Zhang Y, et al. Induction of apoptosis by an inhibitor of the mitotic kinesin KSP requires both activation of the spindle assembly checkpoint and mitotic slippage. Cancer Cell. 2005;8(1):49–59. | ||
Kwok BH, Kapitein LC, Kim JH, Peterman EJ, Schmidt CF, Kapoor TM. Allosteric inhibition of kinesin-5 modulates its processive directional motility. Nat Chem Biol. 2006;2(9):480–485. | ||
Kunwar A, Mogilner A, Ambarish K, Alexander M. Robust transport by multiple motors with nonlinear force–velocity relations and stochastic load sharing. Phys Biol. 2010;7(1):016012. | ||
Bojan M, Anirban C, Kyuho H, Bassik Michael C, Block Steven M. KIF15 nanomechanics and kinesin inhibitors, with implications for cancer chemotherapeutics. Proc Natl Acad Sci U S A. 2018;115:E4613–E4622. | ||
Bigley Alison L, Klein Stephanie K, Barry D, Leigh W, Rudmann Daniel G. Using automated image analysis algorithms to distinguish normal, aberrant, and degenerate mitotic figures induced by Eg5 inhibition. Toxicol Pathol. 2016;44:663–672. | ||
Talapatra SK, Schüttelkopf AW, Kozielski F. The structure of the ternary Eg5-ADP-ispinesib complex. Acta Crystallogr D Biol Crystallogr. 2012;68(Pt 10):1311–1319. | ||
Talapatra SK, Tham CL, Guglielmi P, et al. Crystal structure of the Eg5K858 complex and implications for structure-based design of thiadiazole-containing inhibitors. Eur J Med Chem. 2018;156:641–651. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms.php
and incorporate the Creative Commons Attribution
- Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms.php
and incorporate the Creative Commons Attribution
- Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.