Back to Journals » International Journal of Nanomedicine » Volume 17
bFGF-Loaded Mesoporous Silica Nanoparticles Promote Bone Regeneration Through the Wnt/β-Catenin Signalling Pathway
Authors Shen M , Wang L , Feng L, Gao Y, Li S, Wu Y, Xu C, Pei G
Received 18 March 2022
Accepted for publication 1 June 2022
Published 7 June 2022 Volume 2022:17 Pages 2593—2608
DOI https://doi.org/10.2147/IJN.S366926
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Phong A Tran
Mingkui Shen,* Lulu Wang,* Li Feng, Yi Gao, Sijing Li, Yulan Wu, Chuangye Xu, Guoxian Pei
School of Medicine, Southern University of Science and Technology, Shenzhen, 518055, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Chuangye Xu; Guoxian Pei, Email [email protected]; [email protected]
Background: Bone defects remain an unsolved clinical problem due to the lack of effective osteogenic induction protocols. Nanomaterials play an important role in bone defect repair by stimulating osteogenesis. However, constructing an effective bioactive nanomaterial remains a substantial challenge.
Methods: In this study, mesoporous silica nanoparticles (MSNs) were prepared and used as nanocarriers for basic fibroblast growth factor (bFGF). The characteristics and biological properties of the synthetic bFGF@MSNs were tested. The osteogenic effects of the particles on the behavior of MC3T3-E1 cells were investigated in vitro. In addition, the differentially expressed genes during induction of osteogenesis were analyzed by transcriptomic sequencing. Radiological and histological observations were carried out to determine bone regeneration capability in a distal femur defect model.
Results: Achieving bFGF sustained release, bFGF@MSNs had uniform spherical morphology and good biocompatibility. In vitro osteogenesis induction experiments showed that bFGF@MSNs exhibited excellent osteogenesis performance, with upregulation of osteogenesis-related genes (RUNX2, OCN, Osterix, ALP). Transcriptomic sequencing revealed that the Wnt/β-catenin signalling pathway could be activated in regulation of biological processes. In vivo, bone defect repair experiments showed enhanced bone regeneration, as indicated by radiological and histological analysis, after the application of bFGF@MSNs.
Conclusion: bFGF@MSNs can promote bone regeneration by activating the Wnt/β-catenin signalling pathway. These particles are expected to become a potential therapeutic bioactive material for clinical application in repairing bone defects in the future.
Keywords: mesoporous silica, basic fibroblast growth factor, osteogenesis, transcriptomics sequencing, Wnt/β-catenin signalling pathway, bone regeneration
Introduction
Clinical bone defects frequently result from congenital disorders, trauma, bone tumors, and osteomyelitis, etc,1–3 afflicting the physical and mental health of patients. Current clinical strategies typically use autografts, allografts, xenografts and inorganic grafts at the bone defects site for tissue repair and bone regeneration.4,5 However, these approaches have limitations, such as limited donor mass, poor osteoinductivity, infection, host immunoreaction and unsatisfactory biological activity, limiting their practical application.2,5,6 In addition, many previous studies have demonstrated that the direct injection of bioactive agents (IGF, TGF-β, and BMP) into the defect site may lead to uncontrolled release of drugs or factors.7 Therefore, clinical healing of bone defects remains a major challenge for surgeons, and synthetic biomaterials with the ability to stimulate osteogenesis for promoting bone healing have become the focus of research. To overcome the above challenges, the combination of nanomaterials with various bioactive agents may be a promising approach for bone defect repair.
Because of their high specific surface area, good biocompatibility and biodegradability, mesoporous materials have been widely studied in basic research and real-world applications,8 and mesoporous silica has attracted increasing attention in the biomedical field. As a typical nanocarrier, MSNs are widely used for the loading of drugs and biological macromolecules such as RNA and peptides, and they achieve sustained and controlled release of bioactive factors. Si ions released during the degradation of MSNs can upregulate the expression of osteogenesis-related genes (OCN, RUNX2 and OPN) in osteoblasts to promote bone repair.9 The mesoporous structure of MSNs contributes to the deposition of hydroxyapatite (HA) and promotes mineralization and osteogenesis. MSNs can also induce an immune microenvironment conducive to guiding immune-mediated osteogenesis, and macrophage uptake of nanomaterials plays an important role in the formation of this immune microenvironment.10 However, the biological activity of MSNs is relatively low, and the nanomaterial–cell interaction is weak. Therefore, the combination of drugs or growth factors with MSNs will provide great potential to enhance the bioactivity and osteogenic performance of bone repair materials.
Basic fibroblast growth factor (bFGF) is mainly secreted by endothelial cells, smooth muscle cells and macrophages and is a heparin-binding growth factor that is dysregulated in a variety of tissues.11 Previous studies have shown that bFGF plays a role in wound healing and tissue regeneration by stimulating neovascularization.12 bFGF can significantly promote the proliferation of stem cells from the apical papilla, maintain the stemness characteristics of mesenchymal stem cells and enhance collagen deposition,13,14 but the specific mechanism is still unclear. In addition, bFGF can induce the proliferation of bone marrow stem cells15,16 and promote the synthesis of collagen fibers and early vascularization, accelerating the process of bone formation.17 These studies have shown that bFGF has good biological characteristics, but the specific mechanism underlying the osteogenic effect of bFGF on osteoblasts remains unclear. More importantly, the biological properties of bFGF cannot be effectively exploited due to its instability. Therefore, how to improve the effective utilization of bFGF and clarify the associated mechanism of action are key research and questions that can provide a foundation for clinical application.
In this study, bFGF-loaded MSN nanocomposites (bFGF@MSNs) were synthesized to achieve bFGF-sustained release. Does the combination bFGF with MSNs have a better osteogenic effect than the use of bFGF or MSNs alone? If so, what is the exact mechanism of osteogenesis? In vitro experiments were conducted to evaluate the osteogenic performance of bFGF@MSNs. Then, RNA sequencing was used to analyse the associated mechanism of action and the differences in RNA expression levels. Finally, we injected bFGF@MSNs into distal femur defects to observe bone regeneration. It is expected that this study will propose a new concept of combining bFGF with MSNs to induce osteogenesis for bone regeneration (Scheme 1).
 |
Scheme 1 Schematic illustration of synthetic bFGF@MSNs-mediated induction of bone defect repair and the mechanism of osteogenic induction. |
Materials and Methods
Synthesis of MSNs
MSNs were synthesized according to the method reported in a previous study.18 Briefly, 0.4 g of cetyltrimethylammonium bromide (CTAB) was dissolved in 20 mL of distilled water at 70°C for 10 min. Then, 180 mL of 15.6 mM sodium hydroxide solution was added to the above solution and mixed thoroughly. Next, 2 mL of tetraethyl orthosilicate (TEOS) and 2 mL of ethyl acetate were carefully added dropwise to the above mixture, and stirred at 70°C for 2 h. After cooling at room temperature, the synthesized compounds were collected by centrifugation at 10000 rpm for 30 min and washed with distilled water and ethanol. The supernatant was discarded, the precipitate was resuspended in ethyl alcohol, and 0.48 g of ammonium nitrate was added to the above alcohol mixture, and then stirred and sonicated for 2 h. Finally, the synthesized MSNs were obtained by centrifugation, washing with ethyl alcohol, and drying under a vacuum.
bFGF Loading
Recombinant human bFGF protein was purchased from ThermoFisher Scientific (Gibco, USA). bFGF was loaded into MSNs following these steps. First, 15 mg of bFGF was dissolved in 1 mg/mL ethanol solution to obtain a 1.5 mg/mL bFGF solution. Second, 45 mg of MSNs were added to the above bFGF solution and stirred for 24 h in the dark. Third, bFGF@MSNs were obtained by centrifugation (10,000 rpm, 30 min), washed with ethyl alcohol, and then dried under a vacuum. Finally, the supernatants were pooled together, and the bFGF content was determined using a UV-Vis spectrophotometer at 525 nm.
Characterization of bFGF@MSNs
The morphologies of MSNs and bFGF@MSNs were evaluated by scanning electron microscopy (Nova NanoSem450; FEI, USA) at source voltage of 5 kV and magnification ranging from 10 × 104x to 50 × 104x. The distribution of particle diameter and zeta potential were measured by dynamic light scattering using a NanoDrop system (Thermo Scientific, USA).
Embedding Ratio and Drug Loading of bFGF@MSNs
To investigate the embedding ratio (ER%) and drug loading (DL%) of bFGF@MSNs, bFGF@MSNs were synthesized according to the above method, recording the content (W1) of added bFGF. Then, bFGF@MSNs were dissolved in distilled water. The absorbance of bFGF was detected by a UV spectrophotometer at 450 nm, and the content (W2) of bFGF in the solution was determined. After drying, the specimens were weighed (W3) again. The ER% and DL% were calculated according to the following formulas: ER% = (W1-W2)/W1×100% and DL%= (W1-W2)/ (W3-W1)×100%.
bFGF Release Profile in vitro
To investigate bFGF release, 25 mg of bFGF@MSNs were dissolved in 2.5 mL of PBS (pH = 7.4), and then the suspension was placed in a dialysis bag with a molecular weight cut-off of 10 kDa. After that, the dialysis bag was completely soaked in a 25 cm2 cell culture flask (430,168, Corning) filled with 90 mL of PBS, which was immersed in a 37°C water bath for the bFGF release test. The content of bFGF was detected by a UV spectrophotometer at 450 nm. At various timepoints (1 h, 2 h, 4 h, 8 h, 16 h, 32 h, 48 h, 60 h, 72 h, etc.), 1 mL of the release medium was removed for the absorbance test to calculate the cumulative release rate.
Degradation of MSNs and bFGF@MSNs
The rates of degradation of the MSNs and bFGF@MSNs were assessed in 0.1 mmol/L Tris-HCl solution (pH = 7.4). MSNs and bFGF@MSNs were immersed in Tris- HCl solution at concentrations of 100 μg/L and 400 μg/L at room temperature. At 1, 3, 7, 14 and 21 days, samples were centrifuged and half of the supernatant was replaced with Tris- HCl solution. The release of Si ions was measured by electron-coupled plasma atomic emission spectrometry.
Cell Culture
The preosteoblastic MC3T3-E1 cell line was obtained from the China Center for Type Culture Collection (CCTCC). MC3T3-E1 cells were cultured in standard medium (high-glucose DMEM supplemented with 10% v/v fetal bovine serum, 100 U/mL penicillin and 100 μg/mL streptomycin) with an atmosphere of 5% CO2 at 37°C. The MC3T3-E1 cells were passaged at 80% confluency and used for experiments through passage 3.
Cell Survival Assay
MC3T3-E1 cells were seeded at 2000 cells per well in 96-well plates and then cultured in different bFGF@MSN concentrations (0, 50 μg/L, 100 μg/L, 200 μg/L, 400 μg/L and 800 μg/L) for 48 hours. Then, 10 μL of CCK8 was added to each well. After incubation for 1 hour, the absorbance was detected by a microplate reader at 450 nm. The appropriate bFGF@MSN concentration was chosen according to the cell survival assay.
Cell Proliferation, Morphology and Viability
A CCK8 assay was used to evaluate cell proliferation. MC3T3-E1 cells were seeded at 2000 cells per well in 96-well plates and then cultured in appropriate bFGF@MSN concentrations for 1 day, 3 days, 5 days and 7 days and the absorbance was measured at 450 nm.
For evaluation of cell morphology and viability, MC3T3-E1 cells were seeded at a density of 2.45×105 cells/mL with nanosphere samples in 4-chamber confocal dishes (Cellvis, USA) and cultured in standard medium at 37°C with 5% CO2 in a cell incubator for 48 hours. After removing the medium and washing with PBS, the MC3T3-E1 cells were fixed with 4% paraformaldehyde (PFA; G-CLONE, China) at 4°C for 20 minutes. Afterwards, MC3T3-E1 cells were stained with AbFluor™ 488-Phalloidin (Abbkine, China) for 30 minutes and DAPI for 10 minutes. A confocal fluorescence microscope (Nikon, Japan) was used to observe the morphology of MC3T3-E1 cells. In addition, live and dead MC3T3-E1 cells were evaluated using a live/dead cell imaging kit (Invitrogen, USA), and representative images were acquired.
Alizarin Red S (ARS) Staining
MC3T3-E1 cells were seeded at a density of 2×104 cells/cm2 in prepared 6-well plates covered with gelatin. The standard medium was replaced with osteogenic differentiation medium (Cyagen, USA) when the cells reached 60% confluence and was refreshed every 3 days. After osteogenic induction for 14 and 21 days, samples were fixed with 4% PFA for 30 minutes. Alizarin Red dye solution was added to each well for 5 minutes. After washing with PBS, images were captured using a camera and microscope. The mineral deposits were dissolved in 10 mM cetylpyridinium chloride quantified (Macklin, China), and the optical density (OD) was measured with a Tecan Sunrise instrument at 570 nm.
Alkaline Phosphatase (ALP) Activity and Staining
MC3T3-E1 cells were seeded at a density of 1×104 cells per well in prepared 24-well plates covered with 0.1% gelatin. After osteogenic induction for 7 and 14 days, ALP was stained with an ALP staining kit (Beyotime, China). In addition, the cells were lysed with 0.3% Triton X-100 to release ALP protein and evaluated with an ALP Assay Kit (Beyotime, China).
Transcriptome Sequencing Analysis
After in vitro osteogenic induction for 7 days, the RNA of single MC3T3-E1 cells was extracted with a FastPure Cell/Tissue Total RNA Isolation Kit V2 (Vazyme, China). RNA quality and integrity were evaluated with an Agilent 2100 RNA Nano 6000 Assay kit (Agilent Technologies, CA, USA). The transcriptome sequencing service was provided by Annoroad Gene Technology Anoroad Inc. (Annoroad, China) and performed using the Illumina sequencing platform. To filter out differentially expressed genes, |log2 fold-change| ≥1, p value <0.05 and Q vvalue <0.05 were used as cut-offs. For further analysis, a volcano map and Gene Ontology (GO) bar chart were generated, and GO enrichment analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis of differentially expressed genes were performed. Differentially expressed genes related to osteogenesis were screened out in the biological process category.
In vitro Real-Time Polymerase Chain Reaction (RT-PCR)
The expression levels of genes related to both osteogenesis and Wnt/β-catenin signalling were detected by RT-PCR. After in vitro osteogenic induction for 7 days, the total RNA of single cells was extracted with a kit. Complementary DNA (cDNA) was synthesized by reverse transcription of pre-extracted RNA using a reverse transcription kit (Cyagen, China). The mixtures of synthesized cDNA, gene primers (Forward primers + Reverse primers) and Taq Pro Universal SYBR qPCR Master Mix (Vazyme, China) were put into PCR microplates. The RT-PCR process was performed on a StepOnePlus Real-Time PCR system (Biosystems, USA). The gene expression levels were normalized to those of glyceraldehyde-3-phosphate dehydrogenase (GAPDH). The gene primers are listed in Table 1.
 |
Table 1 Primers for Real-Time PCR |
Western Blotting
Western blot assays were performed to analyse the expression of related proteins. After in vitro osteogenic induction for 7 days, the total proteins were extracted with RIPA lysis buffer with phosphatase inhibitor and protease inhibitor (Epizyme, China). Then, the samples were placed in a 100°C metal bath for 15 minutes leading to protein denaturation. The extracted proteins were quantified with a protein quantification kit (Abbkine, China). Twenty micrograms of protein from each group was added to SDS-PAGE gels (GenScript, China) for protein electrophoresis. Afterwards, the proteins on the gel were transferred to preactivated polyvinylidene fluoride (PVDF) membranes (Millipore, USA). After blocking for 2 hours with 3% bovine serum albumin solution, the membranes were trimmed according to the molecular weight and then incubated with primary antibodies against GAPDH, RUNX2, Osterix, β-catenin, lipoprotein-receptor-related protein 5 (LRP5) and glycogen synthase kinase-3β (GSK-3β) overnight on a shaker at 4°C. The next day, after rewarming to room temperature for 30 minutes, the membranes were washed three times in tris buffered saline with tween (TBST) solution for 5 minutes each time and then incubated with the secondary antibodies for 2 hours. After washing again, the protein bands were visualized using a chemiluminescence kit on an automatic chemiluminescence system (Sage Creation, China).
Animal Experiments
Effects on in vivo osteogenic repair were monitored in a distal femur defect model. All animal experiments followed the guidelines of the Animal Experiment Center and the Laboratory Animal Guide for Ethical Review of Animal Welfare (GB/T 35892-2018) and were approved by the Ethics Committee of Southern University of Science and Technology (Ethics Resolution Number: SUSTech-JY2020215). Twenty-seven male C57/BL6-mice were purchased from the Laboratory Animal Center and divided into three groups (control, MSNs and bFGF@MSNs). To reduce intraoperative excretion, mice were fasted for 12 hours and subjected to water restriction for 4 hours preoperatively. After successful isoflurane inhalation anesthesia, the mice were immobilized in the prone position with subcutaneous injection of meloxicam (0.1 mg/kg). The hair was removed with a razor, and the exposed skin was disinfected with iodophor. A 10 mm incision was made, and anatomical exposure of the distal femur was performed using a subperiosteal dissection of soft tissue. The femur defect was produced via a 1 mm diameter drill bit. After injection of sterilized MSNs and bFGF@MSNs solutions (400 μg/L) which were prepared by dissolving dried powder samples in a 0.9% sodium chloride solution, the opening of the bone defect was sealed with bone wax to prevent fluid outflow (Figure 1). The wound was closed with a 4–0 silk suture. During postoperative care, prophylactic penicillin sodium (4 IU/kg, intramuscular) together with meloxicam (0.1 mg/kg, subcutaneous) was administered for 3 days.
Micro-Computed Tomography (Micro-CT) Evaluation
Bone repair was assessed by micro-CT as described previously.19 Three mice in each group were euthanized with carbon dioxide at 0, 2nd and 4th weeks after surgery. Femur specimens were obtained and fixed in 4% PFA for 48 hours. Then, femur specimens were scanned by a micro-CT (SkyScan 1276, Bruker, Belgium) at a 60 kV voltage and 100 mA electric current. NRecon, CTAn and CTvol of the supplied software were used to process the data and reconstitute 3D images. The bone mineral density (BMD) and the bone volume fraction, calculated as the percentage of bone volume to tissue volume (BV/TV), were analysed at 2nd and 4th weeks.
Histologic Analysis
After fixation in 4% PFA for approximately 48 hours, the harvested femur specimens were subsequently decalcified in EDTA decalcified fluid (OKA, China) for a total of 3 weeks. Then, the samples were completely dehydrated in a tissue dehydrator and embedded in paraffin wax. Paraffin wax slices (4.5 μm) were prepared by using a microtome. According to standard protocols, tissue slices were subjected to hematoxylin and eosin (H&E) staining. Through H&E staining, new bone tissue was evaluated in the defect area. Digital images of these sections were captured using a scanner (Leica, Germany).
Statistical Analysis
Statistical analysis was carried out with GraphPad Prism 6.0 software. Data were recorded as the mean ± standard deviation (SD). Student’s t test was used to evaluate the statistical significance of differences between two groups. The statistical significance of differences among multiple groups was analysed by one-way ANOVA for multiple comparisons. For all tests, a p value <0.05 was considered statistically significant.
Results and Discussion
MSNs were loaded with basic fibroblast growth factor to synthetize bFGF@MSNs. SEM images showed that MSNs and bFGF@MSNs were spherical nanoparticles with a uniform size distribution and good dispersion, as shown in Figure 2A and B. The morphology of nanoparticles significantly affects their interactions with cells and their behavior in biological processes.20 Spherical nanoparticles are more likely to be taken up and have a better biodistribution than nanoparticles of other shapes.21,22 Therefore, bFGF@MSNs were prone to cell uptake prior to exerting their effects. The characterization of diameter (Figure 2C) and zeta potential (Figure 2D) indicated that the size and zeta potential of MSNs were 135.6±1.473 nm and 37.0±1.338 (mV), respectively, and those of bFGF@MSNs were 140.4±1.683 nm and 36.3±0.829 (mV), respectively. The results obtained for the two measurements were basically consistent. The size of nanoparticles was suitable for drug loading and could enhance cell adhesion and biological activity.23 bFGF could be stored into the MSNs by exploiting the surface aperture to achieve sustained release. More importantly, bFGF@MSNs had a typical porous structure that favored cell growth and nutrient delivery.24 Thus, we thoroughly demonstrated the potential of bFGF@MSNs as carrier nanomaterials for bone tissue engineering.
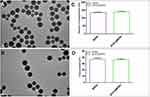 |
Figure 2 Characterization of bFGF@MSNs: (A and B) SEM characteristics, (C) diameter and (D) zeta potential of MSNs and bFGF@MSNs. |
As determined by UV Spectrophotometry, the drug loading of bFGF@MSNs was 28.3%, and the embedding ratio was 84.9%. In previous studies, MSNs have been proved to be capable of loading with various drugs, such as adriamycin, camptothecin, cisplatin and other small molecule chemotherapy drugs25–27 and large molecule drugs such as peptides, proteins and DNA.28–30 MSN loading with drugs is related to the mesoporous aperture, through which proteins can enter the interior of MSNs. Generally, MSNs with negative surface charges can be loaded with positively charged drugs.31,32 MSN loading with drugs can be improved by altering the encapsulation rate and drug loading rate by electrostatic adsorption. In our study, MSNs were loaded with bFGF by means of physical adsorption while stirring the mixture. In addition, the isoelectric point of recombinant human bFGF protein (Gibco, ThermoFisher Scientific, USA) was alkaline (pI = 9.6). In the bFGF loading section, we reported that “ … bFGF was dissolved in 1 mg/mL ethanol solution … ”, since the ethanol solution is neutral (pH = 7.0), the protein molecules of bFGF would have a positive charge when dissolved in ethanol, and pH is < pI under these conditions. After that, the negatively charged MSNs absorbed the positively charged bFGF to achieve the synthesis of bFGF@MSNs. As described in previous studies, bFGF could also enter the interior of MSNs through the aperture. The results indicated that the prepared nanoparticles had a high entrapment efficiency, which may be related to their unique mesoporous structure. bFGF could be adsorbed in the mesoporous channels and internal space, resulting in increased transport stability and sustained release ability.
Since the pH value of blood is 7.4, the release of bFGF in an environment with a pH of 7.4 was determined. The results are shown in Table 2 and Figure 3A. The cumulative bFGF release from bFGF@MSNs was 21.8% at 1 hour and 81.1% at 360 hours, indicating that the release of bFGF in vivo was relatively slow. The porous structure on the surface could avoid the premature release of internal bFGF and ensure the stability of bFGF. Conventionally, direct application of bioactive factors may cause uncontrolled release or loss of biological activity.33,34 Even after delivering factors to the target site, maintaining their biological activity is especially important for regulating cellular processes.20 In addition, the degradation analysis showed no significant difference between the MSNs and bFGF@MSNs groups, which had similar rates of Si icon release after 1, 3, 7, 14 and 21 days of soaking (Figure 3B). Meanwhile, the samples of MSNs and bFGF@MSNs achieved sustained release of Si ions, and over 21 days in the soaking experiment, the released concentrations of Si ions were 34.9 μg/L and 36.1 μg/L, respectively. Hence, it is important to develop approaches for the sustained release of bFGF and Si ions to meet the needs of different bone repair stages.
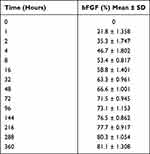 |
Table 2 Cumulative Release of bFGF in vitro |
 |
Figure 3 (A) The profile of cumulative bFGF release from bFGF@MSNs. (B) Si ions released from MSNs and bFGF@MSNs. |
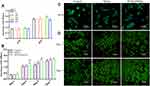
The preosteoblastic MC3T3-E1 cells grew well in standard medium with different bFGF@MSN concentrations (0, 50 μg/L, 100 μg/L, 200 μg/L, 400 μg/L and 800 μg/L) after 24 hours and 48 hours (Figure 4A). This result indicated the absence of cytotoxicity associated with the bFGF@MSNs. Compared to other concentrations, a concentration of 400 μg/L bFGF@MSNs was selected as the appropriate concentration because it obviously promoted cell proliferation. However, the cell survival ability gradually decreased with increasing concentrations. As shown in Figure 4A, 800 μg/L bFGF@MSNs diminished cell proliferation at 48 hours. This phenomenon was likely caused by the release of a large amount of bFGF from high concentrations of bFGF@MSNs, with toxic side effects.
The proliferation of MC3T3-E1 cells was evaluated at 1st, 3rd, 5th and 7th days, as shown in Figure 4B. Compared to the control and MSN-treated groups, the bFGF@MSNs-treated group showed an obvious increase in cell proliferation. Excellent cell proliferation provides the basis for osteogenic differentiation.35–37 As shown in Figure 3, the release of bFGF appeared to peak after 5 days. As a result, no significant difference in cell proliferation was found at 7th day. The morphologies of MC3T3-E1 cells cultured without or with MSNs or bFGF@MSNs are shown in Figure 4C. No detectable differences were found in terms of cell adhesion and cell stretching compared with the control and MSN groups. In addition, there were no dead cells after culture for 3 days and 5 days (Figure 4D). Meanwhile, the number of MC3T3-E1 cells in the bFGF@MSNs group was higher than that in the control and MSNs groups, confirming that bFGF@MSNs promoted cell proliferation. In summary, bFGF@MSNs had good biocompatibility.
The in vitro evaluation of the osteogenic differentiation of MC3T3-E1 cells is shown in Figure 5. Osteoblast differentiation is the basis of bone formation.38,39 During the process of osteogenic differentiation, the basic biological features are bone matrix synthesis, secretion, mineralization and maturation. Mineralized nodules are a marker of osteoblast maturation and the main morphological manifestation of osteoblast-driven osteogenesis. Observing the mineralized nodules of osteoblasts is a common method for evaluating osteoblast differentiation. Evaluation of mineralized matrix formation by ARS staining40 showed calcium deposition and nodule formation after osteogenic induction for 14 days and 21 days (Figure 5A). The intensity of the red color observed for the bFGF@MSNs group was visibly stronger than that observed for the control and MSNs groups, which was consistent with microscopy images revealing more calcium in bFGF@MSNs group. Evaluation of the amount of calcium deposition (Figure 5B) also supported the conclusion that a better osteogenic effect is achieved after bFGF@MSNs treatment. In addition, alkaline phosphatase (ALP) is an early enzyme marker that regulates the differentiation of bone marrow stem cells or preosteoblasts into osteoblastic cells.41,42 ALP production was detected through ALP staining and ALP activity assays (Figure 5C and D). The results showed that ALP products increased with the extension of culture time. After osteogenic induction for 7 days and 14 days, the ALP activity (Figure 5D) in the bFGF@MSNs group was significantly higher than that in the other groups, which was similar to the ALP staining pattern (Figure 5C). These results indicated that bFGF@MSNs could significantly facilitate osteogenic differentiation in vitro.
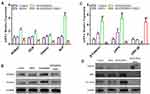
After MC3T3-E1 cells were cultured with osteogenic induction for 7 days, the total RNA of single cells in the control group and bFGF@MSNs group was extracted and subjected to transcriptomic analysis. The results of transcriptomic sequencing analysis are shown in Figure 6. There was a large amount of differential gene expression between the two groups, as shown in the volcano map (Figure 6A). This result suggested that bFGF@MSNs altered the gene expression of MC3T3-E1 cells during the process of osteogenic induction culture. Gene ontology (GO) analysis of differentially expressed genes showed that the expression of genes related to the biological processes (cell proliferation, biological adhesion and biological regulation) was upregulated (Figure 6B). bFGF@MSNs upregulated proliferation-related gene expression, resulting in the promotion of MC3T3-E1 cell proliferation (Figure 4B). Given that differentiation from preosteoblasts to osteoblasts is a biological process,38,39 the expression of osteogenesis-related genes (ALP, RUNX2, OCN, Osterix and Col1a1) was upregulated in the biological process category (Figure 6C). In the in vitro real-time PCR analysis, the mRNA levels of the osteogenesis-related genes (RUNX2, OCN, Osterix and ALP) were upregulated (Figure 7A). Meanwhile, Western blotting revealed higher levels of the osteogenesis-related proteins RUNX2 and Osterix (Figure 7B). Hence, bFGF@MSNs significantly promoted osteogenesis (Figure 5) by regulating biological processes (Figure 6D).
Most importantly, KEGG pathway analysis revealed that the Wnt/β-catenin signalling pathway was obviously altered, as shown in Figure 6E. In the in vitro real-time PCR analysis, the mRNA levels of the Wnt/β-catenin signalling-related factors β-catenin and LRP5 were upregulated, and those of GSK-3β were downregulated (Figure 7C). Western blotting revealed higher levels of the Wnt/β-catenin signalling-related proteins β-catenin and LRP5 and lower levels of the Wnt/β-catenin signalling-related proteins GSK-3β (Figure 7D). The Wnt signalling pathway is a complex protein interaction network. The Wnt/β-catenin pathway is a Wnt pathway that causes β-catenin to accumulate in the cytoplasm and eventually translocate to the nucleus as a transcription factor. Without Wnt, β-catenin cannot accumulate in the cytoplasm. The intracellular destruction complex that degrades the β-catenin consists of the following proteins: axin, adenomatous polyposis coli (APC), protein phosphatase 2A (PP2A), glycogen synthase kinase 3 (GSK3) and casein kinase 1α (CK1α). These proteins degrade β-catenin by via β-catenin ubiquitination and targeting to the proteasome for digestion.43
The Wnt/β-catenin signalling pathway is a fundamental and widespread regulatory mechanism44 that plays an important role in embryonic development and bone healing. In recent years, with increasing research on signalling molecules and pathways, it has been found that the Wnt/β-catenin pathway can increase the proliferation, differentiation and functional activity of osteoblasts45–47 and thus plays an important role in bone tissue homeostasis. A previous study indicated that β-catenin significantly enhanced ALP activity while promoting osteogenesis.48 Runx2 is an osteoblast-specific transcription factor that is involved in osteoblast proliferation, differentiation, and bone formation.49,50 Runx2 expression was reduced leading to the inhibition of bone regeneration via inhibition of the activity of the Wnt/β-catenin signalling pathway.51 Osterix is an osteoblast-specific transcription factor that is essential for osteoblast proliferation and differentiation and a downstream gene of Runx2 in osteoblast differentiation signalling.52 The dickkopf-1 (DKK1) protein is the main inhibitor53,54 of the Wnt/β-catenin signalling pathway, reducing the activity of osteoblasts and leading to reduced bone formation. In this study, MC3T3-E1 cells were cultured with bFGF@MSNs and DKK1-containing osteogenesis inducing medium. The mRNA and protein levels of the Wnt/β-catenin signalling-related factors β-catenin and LRP5 were upregulated, and those of GSK-3β were downregulated (Figure 7C and D). Compared with the control group, MSNs group and bFGF@MSNs group, the inhibitor-treated group showed markedly poor osteogenesis performance, as shown in Figure 5. The mRNA and protein levels of osteogenesis-related factors were significantly decreased (Figure 7A and B). Therefore, the Wnt/β-catenin signalling pathway plays an important role in osteogenic differentiation, and blocking this pathway might lead to bone regeneration disorder and delayed bone healing.55 These results indicated that bFGF@MSNs could significantly facilitate osteogenic differentiation by activating the Wnt/β-catenin signalling pathway in vitro.
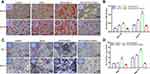
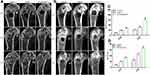
Given the success of our in vitro osteogenic differentiation studies, bone regenerative capacity was evaluated in vivo by micro-CT of distal femur defects at 2 and 4 weeks (Figure 8). Bone regeneration was observed macroscopically in two-dimensional (Figure 8A) and three-dimensional (Figure 8B) reconstructed images. The trend of bone regeneration involved growth from the edge of the defect to the center. Compared to that in the control group and MSN-treated group, visibly accelerated bone defect repair was observed in the bFGF@MSNs-treated group. In addition, the BMD (Figure 8C) and BV/TV (Figure 8D) values of the bFGF@MSNs group were superior to those of the control group and MSNs group. The bone regeneration capability of the bFGF@MSNs group was better than that of the other groups at both 2 and 4 weeks, which suggested that the sustained release of bFGF met the needs of different bone repair stages. Furthermore, new bone formation was detected by histological analysis after H&E staining at 2 and 4 weeks (Figure 9). There was no obvious inflammatory response in the sections among the three groups. We observed more newly formed bone and fibrous tissue in the bFGF@MSNs group than in the other groups. In contrast, few regenerative tissues were found in the control group, which was attributed to lack of inducers to support cell adhesion and differentiation.18 MSNs promoted osteogenesis moderately due to a lack of support from bioactive factors. Hence, the application of bFGF@MSNs is a promising alternative to promote bone regeneration.
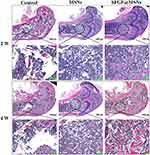 |
Figure 9 Histomorphological analysis of newly formed tissue by hematoxylin-eosin staining. |
Conclusion
In this study, we successfully synthesized bFGF@MSNs with an appropriate size, stable properties and good biocompatibility. With bFGF sustained release, the effective concentration of bFGF@MSNs was determined to be 400 μg/L which was beneficial for cell proliferation and adhesion. More importantly, bFGF@MSNs have the ability to significantly enhance osteogenic differentiation by upregulating osteogenesis-related gene and protein expression. Transcriptomic sequencing analysis demonstrated that bFGF@MSNs regulate the biological processes of MC3T3 cells in terms of cell proliferation, biological adhesion and biological regulation. In addition, bFGF@MSNs stimulated osteogenesis by activating the Wnt/β-catenin signalling pathway, and the inhibitor DDK1 significantly reduced osteogenic differentiation after bFGF@MSNs stimulation. Furthermore, an in vivo study also confirmed that bFGF@MSNs obviously promote bone regeneration in distal femur defects. Therefore, bFGF@MSNs may become a potential therapeutic for clinical application to repair bone defects in the future.
Acknowledgments
This work was supported by the Research Startup Fund of Southern University of Science and Technology (Y01416214).
Disclosure
The authors declare that they have no competing interests related to this work.
References
1. Wang M, Wu H, Li Q, et al. Novel aptamer-functionalized nanoparticles enhances bone defect repair by improving stem cell recruitment. Int J Nanomedicine. 2019;14:8707–8724. doi:10.2147/IJN.S223164
2. Zhou K, Yu P, Shi X, et al. Hierarchically porous hydroxyapatite hybrid scaffold incorporated with reduced graphene oxide for rapid bone ingrowth and repair. ACS Nano. 2019;13(8):9595–9606. doi:10.1021/acsnano.9b04723
3. Cyril M, Thomas BB, Wade S. Management of segmental bone defects. J Am Acad Orthop Surg. 2015;23(3):143–153.
4. Agarwal R, García AJ. Biomaterial strategies for engineering implants for enhanced osseointegration and bone repair. Adv Drug Deliv Rev. 2015;94:53–62. doi:10.1016/j.addr.2015.03.013
5. Ho-Shui-Ling A, Bolander J, Rustom LE, Johnson AW, Luyten FP, Picart C. Bone regeneration strategies: engineered scaffolds, bioactive molecules and stem cells current stage and future perspectives. Biomaterials. 2018;180:143–162. doi:10.1016/j.biomaterials.2018.07.017
6. Hu C, Ashok D, Nisbet DR, Gautam V. Bioinspired surface modification of orthopedic implants for bone tissue engineering. Biomaterials. 2019;219:119366. doi:10.1016/j.biomaterials.2019.119366
7. Zhu M, Zhu Y, Ni B, et al. Mesoporous silica nanoparticles/hydroxyapatite composite coated implants to locally inhibit osteoclastic activity. ACS Appl Mater Interfaces. 2014;6(8):5456–5466. doi:10.1021/am405013t
8. de Weerd C, Gomez L, Capretti A, et al. Efficient carrier multiplication in CsPbI(3) perovskite nanocrystals. Nat Commun. 2018;9(1):4199. doi:10.1038/s41467-018-06721-0
9. Shi M, Zhou Y, Shao J, et al. Stimulation of osteogenesis and angiogenesis of hBMSCs by delivering Si ions and functional drug from mesoporous silica nanospheres. Acta Biomater. 2015;21:178–189. doi:10.1016/j.actbio.2015.04.019
10. Liang H, Jin C, Ma L, et al. Accelerated bone regeneration by gold-nanoparticle-loaded mesoporous silica through stimulating immunomodulation. ACS Appl Mater Interfaces. 2019;11(44):41758–41769. doi:10.1021/acsami.9b16848
11. Zhang H, Wang K, Gao T, et al. Controlled release of bFGF loaded into electrospun core-shell fibrous membranes for use in guided tissue regeneration. Biomed Mater. 2020;15(3):035021. doi:10.1088/1748-605X/ab7979
12. Zhang X, Kang X, Jin L, Bai J, Liu W, Wang Z. Stimulation of wound healing using bioinspired hydrogels with basic fibroblast growth factor (bFGF). Int J Nanomedicine. 2018;13:3897–3906. doi:10.2147/IJN.S168998
13. Wu J, Huang GT, He W, et al. Basic fibroblast growth factor enhances stemness of human stem cells from the apical papilla. J Endod. 2012;38(5):614–622. doi:10.1016/j.joen.2012.01.014
14. Fortino VR, Chen RS, Pelaez D, Cheung HS. Neurogenesis of neural crest-derived periodontal ligament stem cells by EGF and bFGF. J Cell Physiol. 2014;229(4):479–488. doi:10.1002/jcp.24468
15. Lim J, Park EK. Effect of fibroblast growth factor-2 and retinoic acid on lineage commitment of bone marrow mesenchymal stem cells. Tissue Eng Regen Med. 2016;13(1):47–56. doi:10.1007/s13770-016-9102-0
16. D’Mello S, Elangovan S, Salem AK. FGF2 gene activated matrices promote proliferation of bone marrow stromal cells. Arch Oral Biol. 2015;60(12):1742–1749. doi:10.1016/j.archoralbio.2015.09.005
17. Abazari MF, Soleimanifar F, Enderami SE, et al. Incorporated-bFGF polycaprolactone/polyvinylidene fluoride nanocomposite scaffold promotes human induced pluripotent stem cells osteogenic differentiation. J Cell Biochem. 2019;120(10):16750–16759. doi:10.1002/jcb.28933
18. Qiu K, Chen B, Nie W, et al. Electrophoretic deposition of dexamethasone-loaded mesoporous silica nanoparticles onto Poly(L-Lactic Acid)/Poly(ε-Caprolactone) composite scaffold for bone tissue engineering. ACS Appl Mater Interfaces. 2016;8(6):4137–4148. doi:10.1021/acsami.5b11879
19. Ho MH, Yao CJ, Liao MH, Lin PI, Liu SH, Chen RM. Chitosan nanofiber scaffold improves bone healing via stimulating trabecular bone production due to upregulation of the Runx2/osteocalcin/alkaline phosphatase signalling pathway. Int J Nanomedicine. 2015;10:5941–5954. doi:10.2147/IJN.S90669
20. Chen L, Zhou X, He C. Mesoporous silica nanoparticles for tissue-engineering applications. Wiley Interdiscip Rev Nanomed Nanobiotechnol. 2019;11(6):e1573. doi:10.1002/wnan.1573
21. Huang X, Teng X, Chen D, Tang F, He J. The effect of the shape of mesoporous silica nanoparticles on cellular uptake and cell function. Biomaterials. 2010;31(3):438–448. doi:10.1016/j.biomaterials.2009.09.060
22. Huang X, Li L, Liu T, et al. The shape effect of mesoporous silica nanoparticles on biodistribution, clearance, and biocompatibility in vivo. ACS nano. 2011;5(7):5390–5399. doi:10.1021/nn200365a
23. Lu F, Wu SH, Hung Y, Mou CY. Size effect on cell uptake in well-suspended, uniform mesoporous silica nanoparticles. Small. 2009;5(12):1408–1413. doi:10.1002/smll.200900005
24. Aghayan HR, Hosseini MS, Gholami M, et al. Mesenchymal stem cells’ seeded amniotic membrane as a tissue-engineered dressing for wound healing. Drug Deliv Transl Res. 2022;12(3):538–549. doi:10.1007/s13346-021-00952-3
25. Wang Y, Shi W, Song WS, et al. Tumor cell targeted delivery by specific peptide-modified mesoporous silica nanoparticles. J Mater Chem. 2012;22(29):14608–14616. doi:10.1039/c2jm32398b
26. Lu J, Li Z, Zink JI, Tamanoi F. In vivo tumor suppression efficacy of mesoporous silica nanoparticles-based drug-delivery system: enhanced efficacy by folate modification. Nanomedicine. 2012;8(2):212–220. doi:10.1016/j.nano.2011.06.002
27. Ahn B, Park J, Singha K, Park H, Kim WJ. Mesoporous silica nanoparticle-based cisplatin prodrug delivery and anticancer effect under reductive cellular environment. J Mater Chem B. 2013;1(22):2829–2836. doi:10.1039/c3tb20319k
28. Shen D, Yang J, Li X, et al. Biphase stratification approach to three-dimensional dendritic biodegradable mesoporous silica nanospheres. Nano Lett. 2014;14(2):923–932. doi:10.1021/nl404316v
29. Sardan M, Yildirim A, Mumcuoglu D, Tekinay AB, Guler MO. Noncovalent functionalization of mesoporous silica nanoparticles with amphiphilic peptides. J Mater Chem B. 2014;2(15):2168–2174. doi:10.1039/C4TB00037D
30. Yang H, Zheng K, Zhang Z, et al. Adsorption and protection of plasmid DNA on mesoporous silica nanoparticles modified with various amounts of organosilane. J Colloid Interface Sci. 2012;369(1):317–322. doi:10.1016/j.jcis.2011.12.043
31. Liu J, Stace-Naughton A, Jiang X, Brinker CJ. Porous nanoparticle supported lipid bilayers (protocells) as delivery vehicles. J Am Chem Soc. 2009;131(4):1354–1355. doi:10.1021/ja808018y
32. Liu J, Jiang X, Ashley C, Brinker CJ. Electrostatically mediated liposome fusion and lipid exchange with a nanoparticle-supported bilayer for control of surface charge, drug containment, and delivery. J Am Chem Soc. 2009;131(22):7567–7569. doi:10.1021/ja902039y
33. Kim TH, Eltohamy M, Kim M, et al. Therapeutic foam scaffolds incorporating biopolymer-shelled mesoporous nanospheres with growth factors. Acta Biomater. 2014;10(6):2612–2621. doi:10.1016/j.actbio.2014.02.005
34. Hu C, Liu S, Zhang Y, et al. Long-term drug release from electrospun fibers for in vivo inflammation prevention in the prevention of peritendinous adhesions. Acta Biomater. 2013;9(7):7381–7388. doi:10.1016/j.actbio.2013.03.040
35. Mehrasa M, Asadollahi MA, Nasri-Nasrabadi B, et al. Incorporation of mesoporous silica nanoparticles into random electrospun PLGA and PLGA/gelatin nanofibrous scaffolds enhances mechanical and cell proliferation properties. Mater Sci Eng C Mater Biol Appl. 2016;66:25–32. doi:10.1016/j.msec.2016.04.031
36. Zhang Q, Qin M, Zhou X, et al. Porous nanofibrous scaffold incorporated with S1P loaded mesoporous silica nanoparticles and BMP-2 encapsulated PLGA microspheres for enhancing angiogenesis and osteogenesis. J Mater Chem B. 2018;6(42):6731–6743. doi:10.1039/C8TB02138D
37. Wang J, Zhang B, Lu W, et al. Cell proliferation stimulation ability and osteogenic activity of low molecular weight peptides derived from bovine gelatin hydrolysates. J Agric Food Chem. 2020;68(29):7630–7640. doi:10.1021/acs.jafc.0c02717
38. Yang L, Liu S, Mu S, et al. Leonurine hydrochloride promotes osteogenic differentiation and increases osteoblastic bone formation in ovariectomized mice by Wnt/β-catenin pathway. Biochem Biophys Res Commun. 2018;504(4):941–948. doi:10.1016/j.bbrc.2018.09.008
39. Pan FF, Shao J, Shi CJ, Li ZP, Fu WM, Zhang JF. Apigenin promotes osteogenic differentiation of mesenchymal stem cells and accelerates bone fracture healing via activating Wnt/β-catenin signalling. Am J Physiol Endocrinol Metab. 2021;320(4):E760–e771. doi:10.1152/ajpendo.00543.2019
40. Liu Y, Miao YL, Qin F, et al. Electrospun poly (Aspartic Acid)-modified Zein Nanofibers for promoting bone regeneration. Int J Nanomedicine. 2019;14:9497–9512. doi:10.2147/IJN.S224265
41. Peng H, Yin Z, Liu H, et al. Electrospun biomimetic scaffold of hydroxyapatite/chitosan supports enhanced osteogenic differentiation of mMSCs. Nanotechnology. 2012;23(48):485102. doi:10.1088/0957-4484/23/48/485102
42. Nakamura T, Nakamura-Takahashi A, Kasahara M, Yamaguchi A, Azuma T. Tissue-nonspecific alkaline phosphatase promotes the osteogenic differentiation of osteoprogenitor cells. Biochem Biophys Res Commun. 2020;524(3):702–709. doi:10.1016/j.bbrc.2020.01.136
43. Lie DC, Colamarino SA, Song HJ, et al. Wnt signalling regulates adult hippocampal neurogenesis. Nature. 2005;437(7063):1370–1375. doi:10.1038/nature04108
44. Á D, Gyula P, Bálint J, Szittya G, Havelda Z. AGO-unbound cytosolic pool of mature miRNAs in plant cells reveals a novel regulatory step at AGO1 loading. Nucleic Acids Res. 2019;47(18):9803–9817. doi:10.1093/nar/gkz690
45. Yu W, Zhang Y, Xu L, Sun S, Jiang X, Zhang F. Microarray-based bioinformatics analysis of osteoblasts on TiO2 nanotube layers. Colloids Surf B Biointerfaces. 2012;93:135–142. doi:10.1016/j.colsurfb.2011.12.025
46. Zhang R, Oyajobi BO, Harris SE, et al. Wnt/β-catenin signalling activates bone morphogenetic protein 2 expression in osteoblasts. Bone. 2013;52(1):145–156. doi:10.1016/j.bone.2012.09.029
47. Chen S, Feng J, Bao Q, et al. Adverse effects of osteocytic constitutive activation of ß-Catenin on bone strength and bone growth. J Bone Miner Res. 2015;30(7):1184–1194. doi:10.1002/jbmr.2453
48. Li B, Zhang H, Zeng M, et al. Bone marrow mesenchymal stem cells protect alveolar macrophages from lipopolysaccharide-induced apoptosis partially by inhibiting the Wnt/β-catenin pathway. Cell Biol Int. 2015;39(2):192–200. doi:10.1002/cbin.10359
49. Komori T. Regulation of proliferation, differentiation and functions of osteoblasts by Runx2. Int J Mol Sci. 2019;20(7):1694. doi:10.3390/ijms20071694
50. Komori T. Runx2, an inducer of osteoblast and chondrocyte differentiation. Histochem Cell Biol. 2018;149(4):313–323. doi:10.1007/s00418-018-1640-6
51. Lu XM, Zhao H, Wang EH. A high-fat diet induces obesity and impairs bone acquisition in young male mice. Mol Med Rep. 2013;7(4):1203–1208. doi:10.3892/mmr.2013.1297
52. Liu Q, Li M, Wang S, Xiao Z, Xiong Y, Wang G. Recent advances of osterix transcription factor in osteoblast differentiation and bone formation. Front Cell Dev Biol. 2020;8:601224. doi:10.3389/fcell.2020.601224
53. Mao B, Wu W, Davidson G, et al. Kremen proteins are Dickkopf receptors that regulate Wnt/beta-catenin signalling. Nature. 2002;417(6889):664–667. doi:10.1038/nature756
54. Mao B, Wu W, Li Y, et al. LDL-receptor-related protein 6 is a receptor for Dickkopf proteins. Nature. 2001;411(6835):321–325. doi:10.1038/35077108
55. Glass DA 2nd, Karsenty G. In vivo analysis of Wnt signalling in bone. Endocrinology. 2007;148(6):2630–2634. doi:10.1210/en.2006-1372
 © 2022 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms.php
and incorporate the Creative Commons Attribution
- Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2022 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms.php
and incorporate the Creative Commons Attribution
- Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.