Back to Journals » Cancer Management and Research » Volume 12
Associations Between miRNAs and Two Different Cancers: Breast and Colon
Authors Kundaktepe BP , Sozer V , Papila C, Durmus S , Kocael PC, Simsek G, Gelisgen R, Zengin K, Ulualp K, Uzun H
Received 17 August 2019
Accepted for publication 22 January 2020
Published 7 February 2020 Volume 2020:12 Pages 871—879
DOI https://doi.org/10.2147/CMAR.S227628
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Antonella D'Anneo
Berrin Papila Kundaktepe,1 Volkan Sozer,2 Cigdem Papila,3 Sinem Durmus,4 Pinar Cigdem Kocael,1 Gonul Simsek,5 Remise Gelisgen,4 Kagan Zengin,1 Kenan Ulualp,1 Hafize Uzun4
1Department of General Surgery, Cerrahpasa Faculty of Medicine, Istanbul University-Cerrahpasa, Istanbul, Turkey; 2Department of Biochemistry, Yildiz Technical University, Istanbul, Turkey; 3Department of Internal Medicine, Division of Oncology, Cerrahpasa Faculty of Medicine, Istanbul University-Cerrahpasa, Istanbul, Turkey; 4Department of Medical Biochemistry, Cerrahpasa Faculty of Medicine, Istanbul University-Cerrahpasa, Istanbul, Turkey; 5Department of Physiology, Cerrahpasa Faculty of Medicine, Istanbul University-Cerrahpasa, Istanbul, Turkey
Correspondence: Hafize Uzun
Department of Medical Biochemistry, Cerrahpasa Faculty of Medicine, Istanbul University Cerrahpasa, Cerrahpasa, İstanbul 34303, Turkey
Tel +90 212 414 30 56
Fax +90 212 633 29 87
Email [email protected]
Objective: Screening approaches using microRNAs (miRNAs) have been gaining increased attention owing to their potential applications in the diagnosis, prognosis, and monitoring of cancer, because aberrant miRNA expression plays a role in the development and advancement of malignancies. The objectives of this study were to characterize mir21, miR31, mir143, mir145, and control RNU43, which are differentially expressed in peripheral blood mononuclear cells (PBMCs) of breast and colorectal cancer patients, compared to that in controls and to establish whether this is specific to breast and colon cancer for use as tumor markers.
Methods: Thirty newly diagnosed patients with breast cancer and 30 patients with colorectal cancer were enrolled together with 30 healthy controls. PBMCs were isolated from venous blood samples of individuals. Next, miRNA expression analysis was performed by a two-step method of reverse transcription and qPCR.
Results: The expression levels of miR-143 and miR-31 were significantly decreased, whereas the expression levels of miR-145 and miR-21 were significantly increased in breast cancer patients compared to those in healthy subjects. Moreover, the expression levels of miR-143, miR-145, and miR-21 were significantly increased and, in contrast, the changes in the expression levels of miR-31 were not statistically significant in colon cancer compared to those in healthy subjects. miR-21 exhibited the highest increase in both breast and colon cancers. There was a weak positive correlation between miR-145 and CA-15.3 in patients with breast cancer (r = 0.451; p = 0.012). miR-143 was positively correlated with the TNM stage in colon cancer patients (r = 0.568; p = 0.001).
Conclusion: A biomarker panel composed of miR-21, miR-31, miR-143, and miR-145 in PBMC may provide a new diagnostic approach for the early detection of breast and colon cancer. As miR-21 expression was found to be the highest among all the miRNAs evaluated, it may represent a new tumor biomarker and a candidate therapeutic drug or gene target in colon and breast cancer.
Keywords: breast cancer, colon cancer, peripheral blood mononuclear cells, mir21, mir31, mir143, mir145
Introduction
Breast cancer is the most frequent malignancy in women worldwide, and is also the primary cause of death from cancer in all European countries. The 5-year survival rate for breast cancer varies between 69% and 84%, depending on the geographic region1. Several risk factors for breast cancer have been well documented: some mutations (e.g. involving BRCA1, BRCA2, p53); prolonged exposure to endogenous estrogens by early menarche, late menopause, late age at first childbirth; exogenous hormones; oral contraceptive use and hormone replacement therapy; alcohol consumption, overweight and obesity, and physical inactivity.2
The US National Cancer Institute has stated that colon cancer is the second-leading cause of deaths from cancer in the USA. Risk factors that increase the risk of colon cancer include: age; family history of colorectal cancer; personal history; inherited risk; alcohol consumption; cigarette smoking; race; obesity.3
MicroRNAs (miRNAs) are RNA molecules that do not encode proteins4 and which form a class of endogenous small RNAs (small RNAs). Single-stranded miRNAs in humans are approximately 21–23 nucleotides long.5 These molecules have been determined to have crucial roles in certain biological processes and pathological conditions. It has been considered that miRNAs act in the regulation of cellular gene expression at the transcriptional and post-transcriptional levels.6 Since they can bind to mRNAs with low specificity, and lead to mRNA degradation and translational inhibition, miRNAs have a significant regulatory role regarding gene expression.7,8 miRNAs are associated with numerous physiological processes, especially cell type identification and cell differentiation. In some cancers, some miRNAs function as oncogenes, others function as tumor suppressor genes, indicating that miRNAs regulate tumor progression, metastasis, and invasion.4,9 Recent studies have revealed increased miR-21 expression in breast cancer, suggesting a significant effect.10 This relationship between mir-21 and breast cancer has been investigated in detail, and it is reported that enhanced miR-21 expression is associated with serum levels of follicle-stimulating hormone, estradiol, β-human chorionic gonadotropin, testosterone and prolactin in patients with breast cancer. Furthermore, cell proliferation, colony formation, migration and invasion are increased after overexpression of miR-21 in breast cancer cells, and reduced by miR-21 suppression.10 Conversely, it was determined that miR-31, miR-143, and miR-145 are downregulated in breast cancer, inhibiting proliferation, migration, and invasion of breast cancer cells.11–13 Among those miRNAs, miR-21 and miR-31 have been shown to promote inflammation-associated colon tumorigenesis in animal models and are upregulated in colon cancer cells by targeting different genes.14,15 Moreover, the expression of miR-21 is greatly increased and that of miR-145 is decreased in chemo-resistant colon cancer cells, which are highly enriched in cancer stem cells (CSCs), indicating a role of these miRNAs in regulating CSCs.16 These studies show that miRNAs should be examined together in cancer to examine the diagnostic value of miRNAs known to be important in cancer pathogenesis as biomarkers and to clarify the mechanisms behind cancer progression.
In our miRNA selection, the oncogenic and tumor suppressor properties of miRNAs were considered. It is also known that dysregulation of miRNAs, and regulation of levels and effects on target mRNAs, are influenced by other miRNAs. Therefore, to demonstrate the effects of miRNAs in tumor pathogenesis, it is necessary to determine their synergistic effects, not just the examination of a single miRNA. Considering all of this information, miR-21 and miR-31, which exhibit oncogenic activity; and mir-143 and mir-145, which showed tumor suppressor properties, were selected in order to elucidate tumor epigenetics.
Our goal in this study was to evaluate using circulating miRNAs for diagnosing breast and colon cancer. For this, mir21, miR31, mir143, mir145 and control RNU43 miRNAs in peripheral blood mononuclear cells (PBMCs) were planned to be evaluated.
Materials and Methods
Study Population and Samples
Approval from the Ethics Committee of Cerrahpasa Medical Faculty for this study was received, and the study was conducted in conformity with the Declaration of Helsinki. Written informed consent of all participants was obtained before sampling. A prospective, cross-sectional study was performed on 60 newly diagnosed and histologically confirmed patients (30 patients with breast cancer and 30 with colon cancer) who had presented to the Cerrahpasa Medical Faculty, Oncology and general surgery Outpatient Clinic, Istanbul. The complete medical history, physical examination, laboratory investigation, and clinicopathological features were obtained and recorded for all patients. Tumor staging was performed in conformity with the American Joint Committee on Cancer (AJCC) system, 7th edition Tumor, Lymph nodes, Metastasis (TNM) staging classification.17 All patients were examined by the same medical oncologist and general surgeon.
Subjects
The controls were healthy individuals with no known malignancy, chronic disease or active inflammatory condition who provided equivalent whole blood samples. Patients who had undergone surgery, chemo/radiotherapy, and patients with underlying acute or chronic disease (acute infection, diabetes mellitus, hypertension, kidney diseases, cardiovascular disorders, rheumatological diseases, etc.) were excluded. Patients with normal leukocyte levels and control subjects were included in the study in order not to affect miRNA results. Patients with leukocytosis were not included in the study.
Venous blood samples of control subjects and patients with breast and colon cancer were collected to perform routine, complete blood counts and for further laboratory analyses. Thirty patients with breast cancer (30 females (100%), mean age: 52.73 ± 11.30), 30 patients with colon cancer (12 female (40%), 18 males (60%), mean age: 59.30 ± 11.57) and 30 healthy people as a control group (21 females (70%), 9 males (30%), mean age: 48.97 ± 4.74) were included in this study. For all individuals, standard protocols were also utilized for the evaluation of clinical parameters involving routine biochemical tests.
Although not specific, the most common tumor marker for breast cancer is CA 15.3; CA 19.9 for colon cancer, and carcinoembryonic antigen (CEA).18 Measurement of these markers, which are among the routine tests performed, was also performed by the ECLIA method, and values were obtained from patients’ files in the study. The coefficients of intra- and inter-assay variation (% CVs) of all tumor markers were less than 10%.
Isolation of PBMCs
Venous blood samples were collected into tubes which were coated inside with ethylenediaminetetraacetic acid (EDTA). Human peripheral blood cells were isolated immediately from leukocyte concentrates (buffy coats) by Ficoll-Paque density gradient centrifugation.19 Briefly, 3 mL Histopaque (Ficoll) was placed in a 15 mL Falcon tube. With the leaking method, the blood (approximately 4 mL) was added to the scallop so that it would not mix with the Ficoll. It was centrifuged at 1600 rpm for 30 min. The buffy coat was transferred to a new Falcon tube, and 8 mL of PBS was added. The supernatant was discarded after centrifugation for 12 min at 1600 rpm. If the red appearance persisted, it was washed once with PBS and kept at −80°C until RNA isolation.
Total RNA Isolation, miRNA Analysis, and RT-qPCR
Total RNA, including small RNAs, from PBMCs, was isolated by the mirVana RNA Isolation Kit (miRNeasy kit (Qiagen, Valencia CA)). All isolation protocols were conducted according to the manufacturers’ instructions without further modifications.
cDNA was synthesized from total RNA isolated from PBMCs in all subjects using the miScript Reverse Transcription kit (Qiagen). The PCR thermal cycling procedure was as follows: 60 min at 37°C and 5 min at 95°C. Concentration and quality of nucleic acids were evaluated using Qubit assays and a Qubit fluorometer. Then, using the miScript SYBR® Green PCR kit (Qiagen) and miRNA-specific primers, the expression of hsa-miR31, hsa-miR21, hsa-mir143, hsa-mir145 (hsa: homo sapiens) and RNU43 was analyzed with the StepOnePlus™ Real-Time PCR System (Applied Biosystems, Carlsbad, CA). RNU43 was used as an endogenous control. Template cDNA ≤ 2.5 μL was added to each well (for each patient and control) and the total volume was adjusted to 25 μL. The real-time thermal cycling parameters were as follows: 40 cycles, with a single initial activation step (15 min 95°C), denaturation (15 s 94°C), annealing (30 s 55°C), and extension (30 s 70°C). The respective expression levels of miR-21, miR-31, miR-143 and miR-145 were calculated using the 2−ΔΔCT method. Each sample was tested in triplicate.
Statistical Analysis
All statistical evaluations were performed using SPSS 22.0 software. The variable distributions were tested by visual (histograms/probability graphs) and analytic tests such as the Kolmogorov–Smirnov and Shapiro–Wilk tests. Continuous variables showing normal distribution were expressed as means ± standard deviation (SD) and were evaluated by Student’s t-test. Because miRNA levels had not been shown to be normally distributed, evaluations were performed using the Mann Whitney U-test in groups of 2 and using the Kruskal Wallis test in groups larger than 2. Spearman’s rank-order correlation coefficient was used for correlation analysis. A p value below 0.05 was considered as statistically significant.
Results
Demographic Changes, Tumor Marker Levels and Clinicopathological Features
The demographic features and tumor marker levels of all subjects included in the study are shown in Table 1. In our study, the mean age of patients with breast cancer and colon cancer was significantly higher than the controls (p < 0.05 and p ≤ 0.001, respectively); also, the mean age of colon cancer patients was significantly higher than patients with breast cancer (p < 0.05) (Table 1). CA 15.3 levels in breast cancer patients were very significantly elevated compared to those in the controls (p ≤ 0.001) and in patients with colon cancer (p < 0.001); CA 15.3 levels in the patients with colon cancer were significantly higher than those in the controls (p < 0.05) (Table 1). CA 19.9 levels in both breast cancer and colon cancer patients were significantly elevated compared to those in the controls (p < 0.05 and p < 0.001, respectively); also, CA 19.9 levels in colon cancer patients were significantly higher than those in breast cancer patients (p < 0.05) (Table 1). CEA levels in patients with breast and colon cancer were significantly elevated compared to those in the controls (p < 0.05 and p < 0.001, respectively). Furthermore, CEA levels in colon cancer patients were very significantly higher than in breast cancer patients (p < 0.01).
 |
Table 1 Demographic Features and Tumor Marker Levels in Control, Breast Cancer, and Colon Cancer Cases (Mean ± SD) |
Clinicopathological features of patients with breast or colon cancer are shown in Tables 2 and 3, respectively.
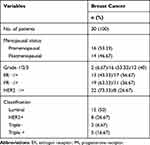 |
Table 2 Clinicopathological Features of Patients with Breast Cancer |
 |
Table 3 Clinicopathological Features of Patients with Colon Cancer |
miRNA Expression Levels in Patients and Controls
RT-qPCR returns the number of cycles that the samples underwent before they were detected, reported as a threshold value known as the Cycle Threshold (CT). CT values vary logarithmically with target expression levels. Mean CT values and standard deviations are used in the 2−ΔΔCT calculations. In breast cancer patients, miR-143 and miR-31 expression were found to be decreased by 0.89- (p < 0.001) and 0.66- (p < 0.05) fold, respectively, whereas the expression of mir-145 and miR-21 was found to be increased by 1.10- (p < 0.001) and 1.11- (p < 0.001) fold, respectively, compared to that in controls (Table 4).
 |
Table 4 miRNA Expression Levels in Control, Breast Cancer, and Colon Cancer Cases (Mean ± SD) |
When expression data from colon cancer patients were compared to those of control samples, mir-143, miR-145 and miR-21 were found to be increased by 2.30- (p < 0.001), 2.15- (p < 0.001), 2.68- (p < 0.001) fold, respectively; while there was no substantive change in expression of mir-31 (1.02 fold) (p = 0.393) (Table 4).
There was a weak positive correlation between miR-145 and CA 15.3 levels in patients with breast cancer (r = 0.451; p = 0.012) (Table 5). miR-31 was weakly positively correlated with PR (+) in breast cancer patients (r = 0.392; p = 0.032, Table 5). According to menopausal status in breast cancer group patients, there was no significant difference in miRNA abundance between pre- and postmenopausal individuals. No correlation was established involving the expression levels of miRNAs and CA 15.3, CA 19.9, and CEA in colon cancer (Table 6); however, miR-143 was positively correlated with TNM stage in colon cancer patients (r = −0.568; p = 0.001, Table 6).
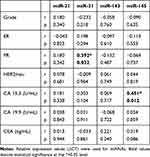 |
Table 5 Correlations of miRNAs with Clinicopathologic Data in Breast Cancer Cases |
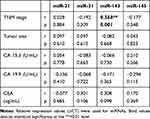 |
Table 6 Correlations of miRNAs with Clinicopathologic Data in Colon Cancer Cases |
Discussion
Inflammation in cancer is an important finding. The primary cells involved in inflammation are PBMCs. In the current study, mir-21 expression levels in PBMCs exhibited the greatest increase in breast cancer. mir-21 expression is present in breast cancer cells; thus, it may have an effect in cancer biology. When breast cancer cells were compared to control samples, miR-21 was found to be up-regulated by 1.11-fold. Hence, miRNA-21 may be a prognostic factor in breast cancer. However, we were not able to determine any correlations between the analyzed miRNAs and serum levels of various tumor markers. The results of Berber et al20 identified 1.8-fold over-expression of miR-21 in tumor tissue compared to that in benign breast tissue. However, such levels were also found to be elevated in metastatic and non-metastatic tumors. We did not identify any differences between the two groups regarding the levels of expression. Chan et al21 first reported that mir-21 significantly increased in patients with glioblastoma, and apoptosis is directed by caspase activation of mir-21-silenced glioblastoma cell lines. Yan et al22 investigated miRNAs that contribute to malignant progression of a tumor in primary breast cancer. Interestingly, in the case of 113 breast cancers, the most significantly upregulated miRNA was miR-21, and the high expression of this miRNA was unquestionably correlated with progressive clinical stage, lymph node metastasis, and shortened survival of patients. Multivariate Cox regression analysis revealed this predictive influence to be independent of the stage and the histological grade of the disease. These findings suggest that miR-21 may be used as a predictive molecular indicator regarding disease progression in breast cancer. Iyevleva et al23 reported that miR-21 expression levels were higher in tumors obtained from patients with bilateral breast cancer than in those from patients with unilaterally located neoplasms. In a study conducted by Hafez et al,24 it was determined that patients having breast cancer with tumor mass > 2 cm had significantly higher miR-21 expression than those with tumors smaller than 2 cm. A study conducted by De Mattos-Arruda et al25 revealed that either baseline assessment of miR-21 expression or its evaluation in residual cancer following neoadjuvant treatment might be used to realize the effectiveness of therapeutic responses, and also for the selection of patients with drug-resistant, HER2-positive tumors in whom novel pharmacological agents might be beneficial. Our results, together with the findings of others, suggest that circulating miR-21 levels, as a novel minimally invasive biomarker, can predict poor prognosis in patients with breast cancer, and its clinical application thus warrants further investigation.26–35
Knudsen et al36 asserted that, in the early stages of the normal-adenoma-adenocarcinoma sequence, the expression levels of miR-17, miR-21, and miR-145 are altered. In our study, when colon cancer cells were compared with control samples, miR-21 expression was found to be up-regulated 2.68-fold. Thus, miR-21, as an oncomiR, may act in advancement of colon cancer. Schetter et al37 found that miR-21 has a prominent role regarding initiation, progression, and metastatic events in colorectal cancer. Link et al38 found that miR-21 levels were upregulated in the stool of patients with colorectal adenomas and cancer compared to control levels. In another study conducted on patients with colorectal cancer, miR-21 expression was found to correlate positively with clinical stage, lymph node positivity, and distant metastatic events.39 Eslamizadeh et al40 found that miR-21 is significantly upregulated in both plasma and colorectal cancer samples of patients when compared to that of corresponding healthy control subjects. Studies concerning miR-21 conducted in larger patient groups with more severe colorectal cancers have clearly demonstrated that miR-21 is localized in the stroma.41–43
miR-31, a common oncomiR, is widely reported to enhance various cancer risks, and has been found to be dysregulated in various solid tumors.23,44,45 Our results also showed that when breast cancer cells were compared with control samples, miR-31 expression was found to be down-regulated 0.66-fold. miR-31 was weakly positively correlated with PR (+). Berber et al20 found that miR-31 is the most strongly down-regulated miRNA in triple-negative breast cancer (TNBC) tissues. miR-31 expression in metastatic tumor tissues was found to be lower than that in non-metastatic tumor tissues in their study, but this decline was not statistically significant. These findings supported the tumor-suppressor and inhibitory effects of miR-31 in TNBC regarding the invasion-metastasis event cascade. Iyevleva et al23 found that miR-31, as an anti-oncogenic agent, exhibits elevated expression levels in tumors of patients with bilateral breast cancer when compared to patients with unilateral neoplasms. Another study showed that the expression level of miR-21-3p is up-regulated, while the level of miR-31-5p is down-regulated in TNBC tissues.46 Our results and those of other studies have shown that miR-31 is downregulated and has an anti-metastatic role in breast cancer. It appears to serve as a tumor suppressor miRNA.
miR-31 expression is upregulated in tumor tissue compared to that in adjacent normal tissue.39,47,48 Overexpression of miR-31 correlates with clinical staging in colon cancer. In our study, when colon cancer patients were compared to controls, no alteration of miR-31 expression was determined. It can be suggested that the phenotype caused by abnormal expression of miR-31 is heavily reliant on endogenous expression levels.49 Bandrés et al47 determined the upregulation of miR-31 in colorectal tumors together with its positive correlation with colorectal cancer stage. Wang et al48 found that miR-31 expression is positively correlated with advanced TNM score together with deep tumoral invasion, asserting that the over-expression of miR-31 might contribute to the progression of colorectal cancer. Slaby et al39 were not able to determine any relationship of miR-31 expression levels with clinical or pathological staging. However, their results justified the significance of other miRNAs, particularly miR-31, regarding the regulation of colorectal tumor differentiation. Laurila and Kallioniemi49 reported that loss of miR-31 expression is correlated with a high risk of metastases in breast cancers, whereas elevated miR-31 expression is correlated with advanced staging in colon cancer. miR-31 plays dual roles: while it is clearly oncogenic in various cancers, it has a distinct suppressive role in certain others.49
miR-143 is reported to be a tumor suppressor in various tumors.50–53 Kirsten rat sarcoma viral oncogene homolog (KRAS) is a gene that acts as an on/off switch in cell signaling. It has been shown that it suppresses cell proliferation in cervical cancer, and directly inhibits KRAS and the down-signal pathway of KRAS in colorectal cancer cells.50,51 Johannessen et al54 found that miR-143 and miR-145 expression are lower in malignant tissues than in benign breast tissue and, additionally, lower in more severe tumors, with ER loss, and basal-like phenotypes in accordance with their in vitro tumor suppressive roles. miR-143 and miR-145 comprise an miRNA cluster, and they appear to have suppressor roles in various organ systems, either as individual miRNAs or by acting in clusters.54 Akao et al55 found that transfection of miR-143 and miR-145 precursors into cells resulted in serious inhibition of growth in human colon cancer. Slaby et al39 reported that miR-143 and miR-145, which are thought to be suppressors, are generally down-regulated. However, they may participate in the early stages of colorectal carcinogenesis because levels of these miRNAs increase with the clinical stage, even if this increase is non-significant. Tumors showing downregulation of miR-143 and miR-145 are more commonly cumbersome lumps and found in the proximal colon. miR-143 and miR-145 levels have high value as predictive markers for therapeutic responses to chemotherapeutic drugs in breast and colon cancer. Previously, these parameters had not been studied in either breast or colon cancer patients, so our study will fill an important gap in the literature. Motoyama et al56 found that miR-31 expression was significantly higher in tumor tissue compared to that in normal tissue; however, the expression levels of miR-143 and miR-145 were significantly lower in cancer tissues than in normal tissues. In our study, the expression levels of miR-143 were significantly lower in breast cancer patients than in healthy controls, whereas the expression of miR-145 was significantly higher. In addition, the expression levels of miR-143 and miR-145 in colon cancer cells were significantly higher than in those of healthy subjects. Contrary to the results we found in breast cancer, Yan et al57 reported that the miR143/miR145 cluster in breast cancer shows tumor suppressor activity by inhibiting ERBB3 translation; they were compliant with the suggestion that various miRNAs acting in one cluster could harmoniously suppress certain target mRNAs. However, in colon cancer, we have observed the miR-143/145 cluster effect.
The absence or overexpression of a given miRNA in a cancerous cell allows the miRNA to play its specific role in the onset and development of cancer. A biomarker panel including miR-21, miR-31, miR-143, and miR-145 in PBMC may provide a new diagnostic approach for the early detection of breast and colon cancer. This study demonstrated that since miR-21 in PBMCs exhibits the greatest increase among all miRNAs, it may be a new tumor biomarker, and a potential candidate drug or gene therapeutic target in colon and breast cancer.
Limitations
Since our funding was limited, our sample numbers were low. However, we are planning to further studies with higher numbers of patients in future studies.
Acknowledgments
This work was supported by the Istanbul University Research Fund (Project No: 42782).
Disclosure
The authors declare that they have no conflicts of interest.
References
1. Baeyens-Fernández JA, Molina-Portillo E, Pollán M, et al. Trends in incidence, mortality and survival in women with breast cancer from 1985 to 2012 in Granada, Spain: a population-based study. BMC Cancer. 2018;18(1):781. doi:10.1186/s12885-018-4682-1
2. WHO. Breast Cancer: Prevention and Control. WHO; 2016. Available from: https://www.who.int/cancer/detection/breastcancer/en/index2.html.
3. PDQ Screening and Prevention Editorial Board. Colorectal Cancer Prevention (PDQ®): patient Version; 2002. Available from: http://www.ncbi.nlm.nih.gov/pubmed/26389376.
4. Zhang L, Xu Y, Jin X, et al. A circulating miRNA signature as a diagnostic biomarker for non-invasive early detection of breast cancer. Breast Cancer Res Treat. 2015;154:423–434. doi:10.1007/s10549-015-3591-0
5. Felekkis K, Touvana E, Stefanou CH, Deltas C. microRNAs: a newly described class of encoded molecules that play a role in health and disease. Hippokratia. 2010;14(4):236.
6. Zhang Y, Lin C, Liao G, et al. MicroRNA-506 suppresses tumor proliferation and metastasis in colon cancer by directly targeting the oncogene EZH2. Oncotarget. 2015;6:32586–32601. doi:10.18632/oncotarget.5309
7. Tian Y, Pan Q, Shang Y, et al. MicroRNA-200 (miR-200) cluster regulation by achaete scute-like 2 (Ascl2): impact on the epithelial-mesenchymal transition in colon cancer cells. J Biol Chem. 2014;289:36101–36115. doi:10.1074/jbc.M114.598383
8. Kanematsu S, Tanimoto K, Suzuki Y, Sugano S. Screening for possible miRNA-mRNA associations in a colon cancer cell line. Gene. 2014;533:520–531. doi:10.1016/j.gene.2013.08.005
9. Al-Khanbashi M, Caramuta S, Alajmi AM, et al. Tissue and serum miRNA profile in Locally Advanced Breast Cancer (LABC) in Response to Neo-Adjuvant Chemotherapy (NAC) treatment. PLoS One. 2016;11:e0152032. doi:10.1371/journal.pone.0152032
10. Zhang C, Liu K, Li T, et al. miR-21: a gene of dual regulation in breast cancer. Int J Oncol. 2016;48:161–172. doi:10.3892/ijo.2015.3232
11. Luo LJ, Yang F, JJ D, et al. MiR-31 inhibits migration and invasion by targeting SATB2 in triple negative breast cancer. Gene. 2016;594:47–58. doi:10.1016/j.gene.2016.08.057
12. Xia C, Yang Y, Kong F, Kong Q, Shan C. MiR-143-3p inhibits the proliferation, cell migration and invasion of human breast cancer cells by modulating the expression of MAPK7. Biochimie. 2018;147:98–104. doi:10.1016/j.biochi.2018.01.003
13. Ding Y, Zhang C, Zhang J, et al. miR-145 inhibits proliferation and migration of breast cancer cells by directly or indirectly regulating TGF-beta1 expression. Int J Oncol. 2017;50:1701–1710. doi:10.3892/ijo.2017.3945
14. Li T, Luo W, Liu K, Li X, Xi T. miR-31 promotes proliferation of colon cancer cells by targeting E2F2. Biotechnol Lett. 2015;37:523–532. doi:10.1007/s10529-014-1715-y
15. Li C, Zhao L, Chen Y, et al. MicroRNA-21 promotes proliferation, migration, and invasion of colorectal cancer, and tumor growth associated with down-regulation of sec23a expression. BMC Cancer. 2016;16:605. doi:10.1186/s12885-016-2628-z
16. Yu Y, Nangia-Makker P, Farhana L, Rajendra SG, Levi E, Majumdar AP. miR-21 and miR-145 cooperation in the regulation of colon cancer stem cells. Mol Cancer. 2015;14:98. doi:10.1186/s12943-015-0372-7
17. Edge, S.B. & Compton, C.C. Ann Surg Oncol (2010) 17: 1471. https://doi.org/10.1245/s10434-010-0985-4
18. Malati T. Tumour markers: an overview. Indian J Clin Biochem. 2007;22(2):17–31. doi:10.1007/BF02913308
19. GE Healthcare. Isolation of Mononuclear Cells Methodology and Applications. Available from: https://www.sigmaaldrich.com/content/dam/sigma-aldrich/docs/Sigma-Aldrich/General_Information/1/ge-isolation-of-mononuclear-cells.pdf. Accessed February 4, 2020.
20. Berber U, Yilmaz I, Narli G, Haholu A, Kucukodaci Z, Demirel D. miR-205 and miR-200c: predictive micro RNAs for lymph node metastasis in triple negative breast cancer. J Breast Cancer. 2014;17:143–148. doi:10.4048/jbc.2014.17.2.143
21. Chan JA, Krichevsky AM, Kosik KS, Krichevsky AM, Kosik KS. microRNA-21 is an antiapoptotic factor in human glioblastoma cells. Cancer Res. 2005;65:6029–6033. doi:10.1158/0008-5472.CAN-05-0137
22. Yan LX, Huang XF, Shao Q, et al. MicroRNA miR-21 overexpression in human breast cancer is associated with the advanced clinical stage, lymph node metastasis, and poor patient prognosis. RNA. 2008;14:2348–2360. doi:10.1261/rna.1034808
23. Iyevleva AG, Kuligina E, Mitiushkina NV, Togo AV, Miki Y, Imyanitov EN. High level of miR-21, miR-10b, and miR-31 expression in bilateral vs. unilateral breast carcinomas. Breast Cancer Res Treat. 2012;131:1049–1059. doi:10.1007/s10549-011-1845-z
24. Hafez MM, Hassan ZK, Zekri AR, et al. MicroRNAs and metastasis-related gene expression in Egyptian breast cancer patients. Asian Pac J Cancer Prev. 2012;13:591–598. doi:10.7314/APJCP.2012.13.2.591
25. De Mattos-arruda L, Bottai G, Nuciforo PG, et al. MicroRNA-21 links epithelial-to-mesenchymal transition and inflammatory signals to confer resistance to neoadjuvant trastuzumab and chemotherapy in HER2-positive breast cancer patients. Oncotarget. 2015;6:37269–37280. doi:10.18632/oncotarget.v6i35
26. Jinling W, Sijing S, Jie Z, Guinean W. Prognostic value of circulating microRNA-21 for breast cancer: a systematic review and meta-analysis. Artif Cells Nanomed Biotechnol. 2017;45:1–6. doi:10.1080/21691401.2016.1216856
27. Yadav P, Mirza M, Nandi K, et al. Serum microRNA-21 expression as a prognostic and therapeutic biomarker for breast cancer patients. Tumour Biol. 2016;37:15275–15282. doi:10.1007/s13277-016-5361-y
28. Li S, Yang X, Yang J, Zhen J, Zhang D. Serum microRNA-21 as a potential diagnostic biomarker for breast cancer: a systematic review and meta-analysis. Clin Exp Med. 2016;16:29–35. doi:10.1007/s10238-014-0332-3
29. Usmani A, Shoro AA, Memon Z, Hussain M, Rehman R. The diagnostic, prognostic and predictive value of microRNA-21 in breast cancer patients, their daughters, and healthy individuals. Am J Cancer Res. 2015;5:2484–2490.
30. Wang G, Wang L, Sun S, Wu J, Wang Q. Quantitative measurement of serum microRNA-21 expression in relation to breast cancer metastasis in Chinese females. Ann Lab Med. 2015;35:226–232. doi:10.3343/alm.2015.35.2.226
31. Shen L, Wan Z, Ma Y, et al. The clinical utility of microRNA-21 as a novel biomarker for diagnosing human cancers. Tumour Biol. 2015;36:1993–2005. doi:10.1007/s13277-014-2806-z
32. Gao J, Zhang Q, Xu J, Guo L, Li X. Clinical significance of serum miR-21 in breast cancer compared with CA153 and CEA. Chin J Cancer Res. 2013;25:743–748. doi:10.3978/j.issn.1000-9604.2013.12.04
33. Si H, Sun X, Chen Y, et al. Circulating microRNA-92a and microRNA-21 as novel minimally invasive biomarkers for primary breast cancer. J Cancer Res Clin Oncol. 2013;139:223–229.
34. Wang B, Zhang Q. The expression and clinical significance of circulating microRNA-21 in the serum of five solid tumors. J Cancer Res Clin Oncol. 2012;138:1659–1666. doi:10.1007/s00432-012-1244-9
35. Asaga S, Kuo C, Nguyen T, Terpenning M, Giuliano AE, Hoon DS. Direct serum assay for microRNA-21 concentrations in early and advanced breast cancer. Clin Chem. 2011;57:84–91. doi:10.1373/clinchem.2010.151845
36. Knudsen KN, Nielsen BS, Lindebjerg J, Hansen TF, Holst R, Sorensen FB. microRNA-17 is the most up-regulated member of the miR-17-92 cluster during early colon cancer evolution. PLoS One. 2015;10:e0140503. doi:10.1371/journal.pone.0140503
37. Schetter AJ, Okayama H, Harris CC. The role of microRNAs in colorectal cancer. Cancer J. 2012;18:244–252. doi:10.1097/PPO.0b013e318258b78f
38. Link A, Balaguer F, Shen Y, et al. Fecal MicroRNAs as novel biomarkers for colon cancer screening. Cancer Epidemiol Biomarkers Prev. 2010;19:1766–1774. doi:10.1158/1055-9965.EPI-10-0027
39. Slaby O, Svoboda M, Fabian P, et al. Altered expression of miR-21, miR-31, miR-143 and miR-145 is related to clinicopathologic features of colorectal cancer. Oncology. 2007;72:397–402. doi:10.1159/000113489
40. Eslamizadeh S, Heidari M, Agah S, et al. The role of microRNA signature as diagnostic biomarkers in different clinical stages of colorectal cancer. Cell J. 2018;20:220–230. doi:10.22074/cellj.2018.5366
41. Nielsen BS, Jorgensen S, Fog JU, et al. High levels of microRNA-21 in the stroma of colorectal cancers predict short disease-free survival in stage II colon cancer patients. Clin Exp Metastasis. 2011;28:27–38. doi:10.1007/s10585-010-9355-7
42. Kjaer-Frifeldt S, Hansen TF, Nielsen BS, et al. The prognostic importance of miR-21 in stage II colon cancer: a population-based study. Br J Cancer. 2012;107:1169–1174. doi:10.1038/bjc.2012.365
43. Kang WK, Lee JK, Oh ST, Lee SH, Jung CK. Stromal expression of miR-21 in T3-4a colorectal cancer is an independent predictor of early tumor relapse. BMC Gastroenterol. 2015;15:2. doi:10.1186/s12876-015-0227-0
44. Gao W, Liu L, Xu J, et al. A systematic analysis of predicted MiR-31-targets identifies a diagnostic and prognostic signature for lung cancer. Biomed Pharmacother. 2014;68(4):419–427. doi:10.1016/j.biopha.2014.03.009
45. Yang X, Xu X, Zhu J, et al. miR-31 affects colorectal cancer cells by inhibiting autophagy in cancer-associated fibroblasts. Oncotarget. 2016;7(48):79617. doi:10.18632/oncotarget.v7i48
46. Ouyang M, Li Y, Ye S, et al. MicroRNA profiling implies new markers of chemoresistance of triple-negative breast cancer. PLoS One. 2014;9:e96228. doi:10.1371/journal.pone.0096228
47. Bandres E, Cubedo E, Agirre X, et al. Identification by Real-time PCR of 13 mature microRNAs differentially expressed in colorectal cancer and non-tumoral tissues. Mol Cancer. 2006;5:29. doi:10.1186/1476-4598-5-29
48. Wang CJ, Zhou ZG, Wang L, et al. Clinicopathological significance of microRNA-31, −143 and −145 expression in colorectal cancer. Dis Markers. 2009;26:27–34. doi:10.1155/2009/921907
49. Laurila EM, Kallioniemi A. The diverse role of miR-31 in regulating cancer-associated phenotypes. Genes Chromosomes Cancer. 2013;52:1103–1113. doi:10.1002/gcc.v52.12
50. Deftereos G, Corrie SR, Feng Q, et al. Expression of mir-21 and mir-143 in cervical specimens ranging from histologically normal through to invasive cervical cancer. PLoS One. 2011;6:e28423. doi:10.1371/journal.pone.0028423
51. Orang AV, Barzegari A. MicroRNAs in colorectal cancer: from diagnosis to targeted therapy. Asian Pac J Cancer Prev. 2014;15(17):6989–6999. doi:10.7314/APJCP.2014.15.17.6989
52. Liu X, Zhao W, Wang X, Zhu Y, Zhou Z, Shi B. Expression of mir-143 in serum of bladder cancer patients and its correlation with clinical features and prognosis. J BU ON. 2019;24(2):791–796.
53. Zhang J, Ma D, Liu H, et al. MicroRNA-143 shows tumor suppressive effects through inhibition of oncogenic K-Ras in pituitary tumor. Int J Clin Exp Pathol. 2017;10(11):10969–10978.
54. Johannessen C, Moi L, Kiselev Y, et al. Expression and function of the miR-143/145 cluster in vitro and in vivo in human breast cancer. PLoS One. 2017;12:e0186658. doi:10.1371/journal.pone.0186658
55. Akao Y, Nakagawa Y, Naoe T. MicroRNAs 143 and 145 are possible common onco-microRNAs in human cancers. Oncol Rep. 2006;16:845–850.
56. Motoyama K, Inoue H, Takatsuno Y, et al. Over- and under-expressed microRNAs in human colorectal cancer. Int J Oncol. 2009;34:1069–1075. doi:10.3892/ijo_00000233
57. Yan X, Chen X, Liang H, et al. miR-143 and miR-145 synergistically regulate ERBB3 to suppress cell proliferation and invasion in breast cancer. Mol Cancer. 2014;13:220. doi:10.1186/1476-4598-13-220
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
