Back to Journals » Pharmacogenomics and Personalized Medicine » Volume 14
Association of CYP3A5 Gene Polymorphisms and Amlodipine-Induced Peripheral Edema in Chinese Han Patients with Essential Hypertension
Authors Liang H, Zhang X, Ma Z, Sun Y, Shu C, Zhu Y, Zhang Y, Hu S, Fu X, Liu L
Received 13 November 2020
Accepted for publication 11 January 2021
Published 2 February 2021 Volume 2021:14 Pages 189—197
DOI https://doi.org/10.2147/PGPM.S291277
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Martin H Bluth
Hao Liang,1,2,* Xinru Zhang,3,* Zhuo Ma,4,* Yan Sun,1,2 Chang Shu,1 Yihua Zhu,5,6 Yanwei Zhang,6 Songnian Hu,1,2 Xiujuan Fu,3 Lihong Liu4
1State Key Laboratory of Microbial Resources, Institute of Microbiology, Chinese Academy of Sciences, Beijing, People’s Republic of China; 2University of Chinese Academy of Sciences, Beijing, People’s Republic of China; 3Department of Pharmacy, The Second Hospital of Jilin University, Changchun, Jilin Province, People’s Republic of China; 4Department of Pharmacy, Beijing Chao-Yang Hospital, Capital Medical University, Beijing, People’s Republic of China; 5College of Information Science and Technology, Nanjing Agricultural University, Nanjing, Jiangsu Province, People’s Republic of China; 6E-Seq Medical Technology Co. Ltd., Beijing, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Songnian Hu
State Key Laboratory of Microbial Resources, Institute of Microbiology, Chinese Academy of Sciences, Beijing, People’s Republic of China
Tel +86-1064806939
Email [email protected]
Lihong Liu
Department of Pharmacy, Beijing Chao-Yang Hospital, Capital Medical University, Beijing, People’s Republic of China
Tel +86-1085231000
Email [email protected]
Background: Amlodipine is one of the most used members of calcium channel blockers (CCB), available to treat hypertension. It is mainly metabolized by the Cytochrome P450 3A4/5 (CYP3A4/5) in the liver. Peripheral edema emerges as the major adverse drug reaction to amlodipine and is the primary reason for discontinuation of amlodipine therapy. However, genetic changes in CYP3A5 may lead to changes in the tolerability of amlodipine.
Purpose: In this study, we were interested whether variants in CYP3A5 have a role to play in amlodipine-induced peripheral edema.
Methods: A total number of 240 Chinese Han patients that have experienced hypertension were included in the study. Sixty-four patients had experienced amlodipine-induced peripheral edema, while the remaining 176 patients with no history of edema formed the control group. Twenty-four single-nucleotide polymorphisms (SNPs) of CYP3A5 gene were sequenced by targeted region sequencing method. The relationship of these genetic variants with amlodipine-induced peripheral edema risk was assessed using logistic regression.
Results: The allele frequencies of CYP3A5*1D (rs15524), CYP3A5*1E (rs4646453) and CYP3A5*3 (rs776746) were significantly different between cases and controls (P< 0.05). The CYP3A5 *3/*3 (CC) or CYP3A5 *1D/*1D (AA) carriers showed an increased risk of amlodipine-induced peripheral edema in dominant model. Meanwhile, patients carrying CYP3A5 *1E (AC/AA) showed a reduced risk of peripheral edema. Furthermore, we found a strong linkage disequilibrium among rs15524, rs4646453 and rs776746.
Conclusion: Our study reveals for the first time that CYP3A5 *1D, *1E and *3 were associated with amlodipine-induced peripheral edema in Chinese Han patients with hypertension. However, further studies comprising larger number of samples, more related genes and other factors are wanted.
Keywords: pharmacogenomics, amlodipine, edema, CYP3A5, SNP
Introduction
Calcium channel blockers (CCBs) are widely used in the treatment of hypertension. In addition to their antihypertensive effects, CCBs may also lead to many adverse reactions, including peripheral edema, dizziness, flushing, fatigue, headache, palpitations, and gingival enlargement.1–4 Peripheral edema, particularly of lower limbs, is the most common side effect of CCBs. CCBs-induced peripheral edema is more common in women and is related to age, upright posture and duration of CCBs’ therapy.2,5 Although it has been known for decades that CCBs could cause peripheral edema, there are still very few genetic markers that could be used in clinical treatment.
Amlodipine is a first-line antihypertensive drug and a long-acting 1,4-dihydropyridine calcium channel blocker.6 It became the fifth most prescribed medication in the United States in 2018.7 However, therapy trials report that the incidence of amlodipine-induced peripheral edema is about 19%, which also probably serves as the main reason behind the limited use of this drug.8,9 To our knowledge, factors that increase the serum concentration of drugs may also increase the risk of side effect. Accordingly, genetic factors affecting the metabolism of amlodipine will consequently influence its clearance, thus possibly resulting in peripheral edema. In recent years, numerous studies have focused on identifying genetic factors determining the responses to amlodipine therapy, including aspects of efficacy and safety.10–12 Nevertheless, the genetic predisposition to amlodipine-induced peripheral edema remains unclear. However, some studies shine light on CYP genes, which are involved in the metabolism of amlodipine.13,14
CYP3A is a subfamily of cytochrome P450 and is involved in the metabolism of many compounds.10,15–17 CYP3A4 and CYP3A5 are the two major enzymes of CYP3A family and play important roles in the dehydrogenation of amlodipine.18 However, it has been reported that CYP3A4, rather than CYP3A5, plays a more important role in amlodipine clearance in vivo.16 However, according to our observations, almost all SNPs of CYP3A4 are very rare in Chinese Han or East Asian.19–21 A recent study evaluated the influence of CYP3A polymorphisms on the pharmacokinetic (PK) parameters of 10 CYP3A substrates including amlodipine, and it turned out CYP3A4 polymorphisms did not show a pronounced influence on PK of amlodipine.22 So, as far as we know, CYP3A4 does not suffice as an explanation towards the incidence of amlodipine-induced adverse responses. Additionally, the contribution of CYP3A5 in amlodipine metabolism is still controversial. CYP3A5 expression varies among individuals due to gene polymorphisms that result in non-productive mRNA splicing and lower or undetectable expression of the protein.14,23,24 Pharmacogenomics studies have found that CYP3A5 gene polymorphisms have significant impact on drug metabolism, efficacy, and toxicity.25,26 Recent studies have reported a possible relationship between CYP3A5 and amlodipine. CYP3A5*3/*3 genotypes might be associated with blood pressure response to amlodipine,27 and CYP3A5*3 may affect the disposition of amlodipine.13 In Chinese hypertensive patients who underwent renal transplantation, researchers have demonstrated that CYP3A5*3 polymorphism affects the antihypertensive efficacy of amlodipine. In such patients with CYP3A5*3/*3, the reduction in diastolic blood pressure (DBP) was significantly higher than those with other genotypes.28 Additionally, compared with CYP3A5*1/*1, CYP3A5*1/*3 patients exhibit a lower metabolic ratio of amlodipine in vivo and hypertensive patients with CYP3A5*1/*3 genotypes show a higher decrease in blood pressure.14 These findings suggest that CYP3A5 genetic variants are really involved in amlodipine response, having an impact on treatment effectivity of amlodipine.
Although studies have identified that CYP3A5 polymorphisms do play a role in blood pressure control and treatment responses of amlodipine, the effect of a particular genotype in amlodipine-induced peripheral edema remains unclear. Here, we aim to test whether any genetic variants associate with amlodipine-induced peripheral edema. For this, we used the targeted region sequencing method to genotype the SNPs in all known haplotypes of CYP3A5, and evaluated their effect on amlodipine-induced peripheral edema in Chinese Han hypertensive patients.
Materials and Methods
Participants
We recruited a total of 240 patients with essential hypertension who have ever been treated with amlodipine or L-amlodipine for more than 4 weeks. The participants were outpatients from Beijing Chaoyang Hospital of Capital Medical University and the Second Hospital of Jilin University. The demographic characteristics and baseline data were collected, including gender, age, body mass index (BMI), smoking and drinking status, blood pressure, heart rate and use of medications. Participants were also questioned about peripheral edema symptoms (leg or ankle edema). Two trained physicians evaluated the adverse drug reactions caused by the amlodipine or L-amlodipine based on the WHO-UMC scale.29 We classified the participant as a peripheral edema case if WHO-UMC causality categories were certain, probable, or possible. The others were grouped as controls.
Sample Collection and DNA Sequencing
The DNA Sampling Swabs (Taitong Gene Testing Equipments Co., Ltd., Suzhou, China) were used for collecting oral buccal mucosa cells. Genomic DNA was isolated from the swabs, using Hi-Swab DNA kit (TIANGEN, Beijing, China) according to the manufacturer’s instructions. Quantification of the DNA concentration was performed using the manufacturer’s protocol with the Qubit® dsDNA HS Assay Kit (Yeasen, Shanghai, China). Methods for targeted region capture and NGS have been reported previously.30 Illumina HiSeq X was used to sequence the captured DNA with paired-end reads of 150 bp length. Table S1 lists the targeted regions of sequencing and the 24 SNPs of CYP3A5.
SNP Calling and Genotyping
High-quality sequencing reads were obtained from the raw data by removing reads that contained adapters, were with unknown bases, or had a low-quality using the Trimmomatic (v0.36)31 program. The high-quality reads were aligned to human reference genome hg19 using the Burrows-Wheeler Aligner (BWA, v0.7.15)32 using the default parameters. The Genome Analysis ToolKit (GATK, v3.8)33 was used for indels realignment, quality score recalibration, variant calling, and genotyping (using Haplotype Caller).
Statistical Analysis
Demographic and clinical characteristics of different groups were compared by t-test or Chi-square (χ2) test according to the data category. The associations between gene polymorphisms and the risk of peripheral edema were assessed by codominant model, dominant model, recessive model and allele model by calculating the odds ratios (ORs) and 95% confidence intervals (CIs) using logistic regression with or without adjustment by gender and alcohol status. Stratification was done by gender. Analyses above were carried out on R-4.03. PLINK 1.934 was used to calculate the minor allele frequency and assess Hardy–Weinberg equilibrium (HWE) for each SNP. Additionally, linkage disequilibrium (LD) block and haplotype were assessed by Haploview35 software. The D’ and r2 values for all pairs of SNPs were calculated. P value<0.05 was considered as the significant level.
Results
General Characteristics
Two hundred and forty enrolled patients were separated into 64 cases and 176 controls. The general characteristics of the study population are summarized in Table 1. In agreement with previous reports, a higher incidence of CCB-induced peripheral edema was observed in women. A significant difference was found in the sex ratio between cases and controls (P=0.00048). In addition, the frequency of drinkers was 29.69% in the case group and 46.02% in the control group (P=0.034). There was no significant difference between the two groups regarding other characteristics.
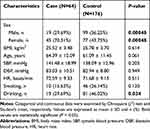 |
Table 1 Characteristics of the Study Population |
Association of CYP3A5 Gene Polymorphisms with the Risk of Amlodipine-Induced Peripheral Edema
All the observed SNPs and the minor allele frequencies (MAF) in two groups are listed in Table 2. Except rs15524, rs4646453 and rs776746, the other SNPs were rare or not detected in the studied population. Therefore, we focused on these three SNPs (MAF>0.05) for further studies. Distributions of genotype frequencies of the SNPs did not show any deviation in Hardy–Weinberg equilibrium (P>0.05).
 |
Table 2 Observed CYP3A5 Variations and Frequencies |
The genotype and allele allocations of the test polymorphisms differed considerably between cases and controls (Table 3). In more detail, the frequencies of alleles rs15524 G, rs4646453 A, and rs776746 T were significantly lower in cases than those in the control group (G vs A: OR=0.53, P=0.011; A vs C: OR=0.54, P=0.019; T vs C: OR=0.58, P=0.03; respectively). Furthermore, there was a statistically significant difference in genotype of the rs15524 and rs4646453 between the two groups in dominant model with or without adjustment by gender and alcohol status (GG+AA vs AA: OR=0.5, P=0.021; AA+AC vs CC: OR=0.54, P=0.04). As for rs776746, the presence of TT+CT genotype demonstrated a significantly lower risk after gender and alcohol status adjustment (TT+CT vs CC: OR=0.57, adjusted P=0.044).
 |
Table 3 Genotype Frequencies of Study SNPs in Case and Control Groups |
Stratified analyses were performed to access the differential effect of gender on the association between amlodipine-induced edema and the polymorphisms. No significant differences were identified, but the relevant genotypes still showed lower risk in all subgroups. The details are presented in Table 4.
 |
Table 4 Stratified Analyses Between SNPs and Risk of Amlodipine-Induced Peripheral Edema |
Linkage Disequilibrium and Haplotype Analysis of Three CYP3A5 Gene Polymorphisms
We employed Haploview program to assess the Linkage disequilibrium (LD) block and haplotype of the three SNPs of CYP3A5. The LD analysis indicated that these SNPs were in strong LD with each other (Figure 1, rs15524 and rs4646453 D’=0.965, r2=0.821; rs15524 and rs776746 D’=0.979, r2=0.919; rs776746 and rs4646453 D’=0.989, r2=0.898). Frequencies of four haplotypes were found to be more than 1% in the haplotype analysis (Table 5). The most represented haplotype in the whole cohort of controls and cases was ACC, followed by GAT, GCC and GCT. Two haplotypes (ACC and GAT) were significantly associated with the risk of amlodipine-induced peripheral edema. The frequency of the ACC haplotype was higher in the cases than controls (79.7% vs 67.9%, P=0.012), whereas the frequency of the GAT haplotype was lower in the cases (17.2% vs 28.1%, P=0.015).
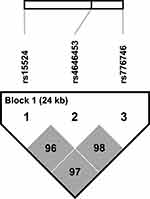 |
Figure 1 Linkage disequilibrium coefficients (|D’|) and LD block among the three polymorphisms of CYP3A5. |
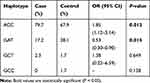 |
Table 5 Haplotype Frequencies of CYP3A5 Gene and the Association with the Amlodipine-Induced Peripheral Edema Risk |
Discussion
Studies have demonstrated that genetic polymorphisms may influence the gene function, thus causing alterations in the pharmacokinetics (PK) and pharmacodynamics (PD) of the gene-associated drugs. Moreover, genetic variants have been confirmed for their association with appearance of adverse reactions to drugs, such as ACE inhibitors-induced cough36 and rosuvastatin-induced myotoxicity.37 To our knowledge, this is the first study to investigate the relationships of CYP3A5 polymorphisms and amlodipine-induced peripheral edema by using a case–control retrospective study in the Chinese population. Our study reveals that amlodipine-induced peripheral edema may associate with genetic polymorphisms in CYP3A5 gene. We find that the distribution of allele and genotype frequencies of the three SNPs are significantly different between cases and controls. Specifically, the alleles rs15524 G, rs4646453 A, and rs776746 T reduce the risk of amlodipine-induced peripheral edema. On the other hand, A, C, and C increase the risk and haplotype analysis also confirms this. These findings support the hypothesis that genetic variation in CYP3A5 is involved in the development of amlodipine-induced peripheral edema.
CYP3A5 is highly polymorphic with significant inter-individual variation in the enzyme activity contributing to the absorption, metabolism and tissue distribution of drugs.24,38 Genetic polymorphisms of CYP3A5 may potentially alter its protein’s expression and function, and subsequently influence the clearance of any drugs metabolized by CYP3A5.38 The most studied SNP related to CYP3A5 functional variation is rs776746 (also known as CYP3A5*3 or CYP3A5*3C). Homozygous carriers of this SNP (*3/*3 or CC) lack functional CYP3A5 protein because of the frameshift mutation and truncation of the translated protein.39 Previous studies have confirmed that CYP3A5*3 is associated with drug metabolism, and CYP3A5*3/*3 carriers have decreased metabolism of nifedipine40 and tacrolimus,41 compared to CYP3A5*1/*1 and CYP3A5*1/*3 carriers. Patients with CYP3A5*3/*3 who are treated with tacrolimus may have an increased risk of nephrotoxicity as compared to patients without it.25,26 These findings suggest that the CYP3A5 *3/*3 carriers have low activity of CYP3A5 enzyme and decreased metabolism for certain drugs dependent on it, resulting in the appearance of an adverse reaction to the drug. Conforming to these findings, our study also indicates that there is an association between CYP3A5*3 and the occurrence of amlodipine-induced peripheral edema and that *3/*3 (CC) carriers have a higher risk of peripheral edema.
CYP3A5*1D (rs15524) is another frequently studied SNP in CYP3A5, which is in the 3ʹ-untranslated region (UTR). CYP3A5*1D is differentially distributed among populations ranging from 77.5% in Americans to 71.4% in East Asians, 40.6% in Africans, and even rarer in Europeans (less than 8%) (GnomAD).21 SNPs in UTRs may influence the stability of mRNA, thus affecting the expression and activity of the enzyme.42,43 According to a study identifying the potential SNPs related to miRNA, rs15524 may influence the hsa-miR-500a-5p that targets CYP3A5, affecting its expression.44 In addition, studies have found that CYP3A5*1D influences the pharmacokinetics of many drugs, including tacrolimus45 and carbamazepine.46 Therefore, rs15524 may act as a genetic marker and should be considered while studying or prescribing drugs metabolized by CYP3A5. Here, we find that the frequencies of rs15524 G allele and GA+GG genotypes in cases are lower than controls, which indicates that this SNP is associated with reduced risk of incidence of amlodipine-induced peripheral edema. In other words, CYP3A5*1D/*1D carriers may have an increased risk than others.
As for CYP3A5*1E (rs4646453), it is associated with a decreasing risk of amlodipine-induced peripheral edema. However, there is litter information about the function of the rs4646453. To some extent, this is possible because the SNP locates in the intron region. Studies have revealed that CYP3A5 rs4646453 is in LD with rs776746,47 and there is a strong LD between rs15524 and rs776746.39,46 Our findings are consistent with these studies and further promote that the presence of LDs with rs776746 may partly explain the role of rs15524 and rs4646453 in amlodipine-induced peripheral edema.
Given all these findings, there are some limitations in our present study that we would like to acknowledge. First, the population in our study is Chinese Han, and it is known that the frequency of mutations differs among ethnic groups. Hence, our results may apply well to Chinese population but are probably not suitable for populations of other ethnic groups. Second, we have performed a retrospective study and not a prospective one. The study is also limited by the smaller number of samples. The third limitation is not involving other genes contributing to amlodipine metabolism, such as CYP3A4. As we mentioned before, polymorphisms of CYP3A4 are rare in Chinese, and the evidence so far that CYP3A4 polymorphisms influence amlodipine metabolism is scarce. So, we took no account of them currently. But it would be better for future studies to include these related genes to better explain the connection between genetic factors and amlodipine-induced peripheral edema. Hence, more case–control studies with large number of polyethnic samples and involvement of diversified factors are necessary.
None the less, our study does investigate the novel relationship between the genetic variants and amlodipine-induced peripheral edema. In conclusion, we provide evidence that CYP3A5 polymorphisms are involved in the occurrence of amlodipine-induced peripheral edema, and the three genetic variants of CYP3A5 have the potential to serve as novel biomarkers for amlodipine-induced adverse reactions. Our findings thus provide new insights into amlodipine-induced peripheral edema and are of importance in developing and prescribing personalized and precise medicine for hypertension.
Data Sharing Statement
The raw data are available on reasonable request to the correspondence author Songnian Hu.
Ethics and Consent Statement
All the participants have signed the written informed consent, and approval was obtained from the Research Ethics Board of Beijing Chaoyang Hospital of Capital Medical University and the Research Ethics Board of the Second Hospital of Jilin University. This study has been conducted in accordance with the World Medical Association Declaration of Helsinki.
Acknowledgments
We thank all the participants included in this study.
Disclosure
All authors declare that they have no conflicts of interest for this work.
References
1. Flynn JT, Pasko DA. Calcium channel blockers: pharmacology and place in therapy of pediatric hypertension. Pediatr Nephrol. 2000;15(3):302–316. doi:10.1007/s004670000480
2. Sica DA. Pharmacotherapy review: calcium channel blockers. J Clin Hypertens (Greenwich). 2006;8(1):53–56. doi:10.1111/j.1524-6175.2005.04140.x
3. Fares H, DiNicolantonio JJ, O’Keefe JH, Lavie CJ. Amlodipine in hypertension: a first-line agent with efficacy for improving blood pressure and patient outcomes. Open Heart. 2016;3(2):e000473. doi:10.1136/openhrt-2016-000473
4. Kala N, Babu S, Manjeu J, Aadivalavan A, Khan R. Allele-specific polymerase chain reaction for the detection of single nucleotide polymorphism in amlodipine-induced gingival enlargement. J Clin Pharm Ther. 2018;43(1):110–113. doi:10.1111/jcpt.12587
5. Makani H, Bangalore S, Romero J, et al. Peripheral edema associated with calcium channel blockers: incidence and withdrawal rate–a meta-analysis of randomized trials. J Hypertens. 2011;29(7):1270–1280. doi:10.1097/HJH.0b013e3283472643
6. Kes S, Caglar N, Canberk A, et al. Treatment of mild-to-moderate hypertension with calcium channel blockers: a multicentre comparison of once-daily nifedipine GITS with once-daily amlodipine. Curr Med Res Opin. 2003;19(3):226–237. doi:10.1185/030079903125001677
7. The ClinCalc DrugStats Database. Amlodipine drug usage statistics, United States; 2008–2018. Available from: https://clincalc.com/DrugStats/Drugs/Amlodipine.
8. Epstein BJ, Roberts ME. Managing peripheral edema in patients with arterial hypertension. Am J Ther. 2009;16(6):543–553. doi:10.1097/MJT.0b013e3181afbf9f
9. Messerli FH, Grossman E. Pedal edema–not all dihydropyridine calcium antagonists are created equal. Am J Hypertens. 2002;15(11):1019–1020. doi:10.1016/S0895-7061(02)03087-X
10. Bhatnagar V, Garcia EP, O’Connor DT, et al. CYP3A4 and CYP3A5 polymorphisms and blood pressure response to amlodipine among African-American men and women with early hypertensive renal disease. Am J Nephrol. 2010;31(2):95–103. doi:10.1159/000258688
11. Irvin MR, Lynch AI, Kabagambe EK, et al. Pharmacogenetic association of hypertension candidate genes with fasting glucose in the GenHAT study. J Hypertens. 2010;28(10):2076–2083. doi:10.1097/HJH.0b013e32833c7a4d
12. Sorensen IF, Vazquez AI, Irvin MR, et al. Pharmacogenetic effects of ‘candidate gene complexes’ on stroke in the GenHAT study. Pharmacogenet Genomics. 2014;24(11):556–563. doi:10.1097/FPC.0000000000000088
13. Kim KA, Park PW, Lee OJ, et al. Effect of CYP3A5*3 genotype on the pharmacokinetics and pharmacodynamics of amlodipine in healthy Korean subjects. Clin Pharmacol Ther. 2006;80(6):646–656. doi:10.1016/j.clpt.2006.09.009
14. Lu Y, Zhong H, Tang Q, et al. Construction and verification of CYP3A5 gene polymorphisms using a Saccharomyces cerevisiae expression system to predict drug metabolism. Mol Med Rep. 2017;15(4):1593–1600. doi:10.3892/mmr.2017.6214
15. Eichelbaum M, Burk O. CYP3A genetics in drug metabolism. Nat Med. 2001;7(3):285–287. doi:10.1038/85417
16. Zhu Y, Wang F, Li Q, et al. Amlodipine metabolism in human liver microsomes and roles of CYP3A4/5 in the dihydropyridine dehydrogenation. Drug Metab Dispos. 2014;42(2):245–249. doi:10.1124/dmd.113.055400
17. Katoh M, Nakajima M, Shimada N, Yamazaki H, Yokoi T. Inhibition of human cytochrome P450 enzymes by 1,4-dihydropyridine calcium antagonists: prediction of in vivo drug-drug interactions. Eur J Clin Pharmacol. 2000;55(11–12):843–852. doi:10.1007/s002280050706
18. Dorofeeva MN, Shikh EV, Sizova ZM, et al. Antihypertensive effect of amlodipine in co-administration with omeprazole in patients with hypertension and acid-related disorders: cytochrome P450-associated aspects. Pharmgenomics Pers Med. 2019;12:329–339. doi:10.2147/PGPM.S217725
19. Whirl-Carrillo M, McDonagh EM, Hebert JM, et al. Pharmacogenomics knowledge for personalized medicine. Clin Pharmacol Ther. 2012;92(4):414–417. doi:10.1038/clpt.2012.96
20. Gaedigk A, Ingelman-Sundberg M, Miller NA, et al. The pharmacogene variation (PharmVar) consortium: incorporation of the human cytochrome P450 (CYP) allele nomenclature database. Clin Pharmacol Ther. 2018;103(3):399–401. doi:10.1002/cpt.910
21. Lek M, Karczewski KJ, Minikel EV, et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature. 2016;536(7616):285–291. doi:10.1038/nature19057
22. Saiz-Rodriguez M, Almenara S, Navares-Gomez M, et al. Effect of the most relevant CYP3A4 and CYP3A5 polymorphisms on the pharmacokinetic parameters of 10 CYP3A substrates. Biomedicines. 2020;8(4):94. doi:10.3390/biomedicines8040094
23. Wojnowski L. Genetics of the variable expression of CYP3A in humans. Ther Drug Monit. 2004;26(2):192–199. doi:10.1097/00007691-200404000-00019
24. Kuehl P, Zhang J, Lin Y, et al. Sequence diversity in CYP3A promoters and characterization of the genetic basis of polymorphic CYP3A5 expression. Nat Genet. 2001;27(4):383–391. doi:10.1038/86882
25. Min SI, Kim SY, Ahn SH, et al. CYP3A5 *1 allele: impacts on early acute rejection and graft function in tacrolimus-based renal transplant recipients. Transplantation. 2010;90(12):1394–1400. doi:10.1097/TP.0b013e3181fa93a4
26. Satoh S, Saito M, Inoue T, et al. CYP3A5 *1 allele associated with tacrolimus trough concentrations but not subclinical acute rejection or chronic allograft nephropathy in Japanese renal transplant recipients. Eur J Clin Pharmacol. 2009;65(5):473–481. doi:10.1007/s00228-008-0606-3
27. Zhang YP, Zuo XC, Huang ZJ, et al. CYP3A5 polymorphism, amlodipine and hypertension. J Hum Hypertens. 2014;28(3):145–149. doi:10.1038/jhh.2013.67
28. Huang Y, Wen G, Lu Y, et al. CYP3A4*1G and CYP3A5*3 genetic polymorphisms alter the antihypertensive efficacy of amlodipine in patients with hypertension following renal transplantation. Int J Clin Pharmacol Ther. 2017;55(2):109–118. doi:10.5414/CP202559
29. Uppsala Monitoring Centre. The use of the WHO-UMC system for standardised case causality assessment; 2018. Available from: https://www.who-umc.org/media/164200/who-umc-causality-assessment_new-logo.pdf.
30. Guo Z, Wang H, Tao J, et al. Development of multiple SNP markers panels affordable to breeders through genotyping by target sequencing (GBTS) in maize. Mol Breeding. 2019;39. doi:10.1007/s11032-019-0940-4
31. Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30(15):2114–2120. doi:10.1093/bioinformatics/btu170
32. Li H, Durbin R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics. 2010;26(5):589–595. doi:10.1093/bioinformatics/btp698
33. DePristo MA, Banks E, Poplin R, et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet. 2011;43(5):491–498. doi:10.1038/ng.806
34. Purcell S, Neale B, Todd-Brown K, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007;81(3):559–575. doi:10.1086/519795
35. Barrett JC, Fry B, Maller J, Daly MJ. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics. 2005;21(2):263–265. doi:10.1093/bioinformatics/bth457
36. Mas S, Gasso P, Alvarez S, et al. Pharmacogenetic predictors of angiotensin-converting enzyme inhibitor-induced cough: the role of ACE, ABO, and BDKRB2 genes. Pharmacogenet Genomics. 2011;21(9):531–538. doi:10.1097/FPC.0b013e328348c6db
37. Liu JE, Liu XY, Chen S, et al. SLCO1B1 521T > C polymorphism associated with rosuvastatin-induced myotoxicity in Chinese coronary artery disease patients: a nested case-control study. Eur J Clin Pharmacol. 2017;73(11):1409–1416. doi:10.1007/s00228-017-2318-z
38. Lamba JK, Lin YS, Schuetz EG, Thummel KE. Genetic contribution to variable human CYP3A-mediated metabolism. Adv Drug Deliv Rev. 2002;54(10):1271–1294. doi:10.1016/S0169-409X(02)00066-2
39. Xie HG, Wood AJ, Kim RB, Stein CM, Wilkinson GR. Genetic variability in CYP3A5 and its possible consequences. Pharmacogenomics. 2004;5(3):243–272. doi:10.1517/phgs.5.3.243.29833
40. Haas DM, Quinney SK, Clay JM, et al. Nifedipine pharmacokinetics are influenced by CYP3A5 genotype when used as a preterm labor tocolytic. Am J Perinatol. 2013;30(4):275–281. doi:10.1055/s-0032-1323590
41. Niioka T, Satoh S, Kagaya H, et al. Comparison of pharmacokinetics and pharmacogenetics of once- and twice-daily tacrolimus in the early stage after renal transplantation. Transplantation. 2012;94(10):1013–1019. doi:10.1097/TP.0b013e31826bc400
42. Mei J, Yan T, Huang Y, et al. A DAAM1 3ʹ-UTR SNP mutation regulates breast cancer metastasis through affecting miR-208a-5p-DAAM1-RhoA axis. Cancer Cell Int. 2019;19:55. doi:10.1186/s12935-019-0747-8
43. Saunders MA, Liang H, Li WH. Human polymorphism at microRNAs and microRNA target sites. Proc Natl Acad Sci U S A. 2007;104(9):3300–3305. doi:10.1073/pnas.0611347104
44. Wei R, Yang F, Urban TJ, et al. Impact of the interaction between 3ʹ-UTR SNPs and microRNA on the expression of human xenobiotic metabolism enzyme and transporter genes. Front Genet. 2012;3:248. doi:10.3389/fgene.2012.00248
45. Liu J, Ouyang Y, Chen D, et al. Donor and recipient P450 gene polymorphisms influence individual pharmacological effects of tacrolimus in Chinese liver transplantation patients. Int Immunopharmacol. 2018;57:18–24. doi:10.1016/j.intimp.2018.02.005
46. Wang P, Yin T, Ma HY, et al. Effects of CYP3A4/5 and ABCB1 genetic polymorphisms on carbamazepine metabolism and transport in Chinese patients with epilepsy treated with carbamazepine in monotherapy and bitherapy. Epilepsy Res. 2015;117:52–57. doi:10.1016/j.eplepsyres.2015.09.001
47. Hyland PL, Freedman ND, Hu N, et al. Genetic variants in sex hormone metabolic pathway genes and risk of esophageal squamous cell carcinoma. Carcinogenesis. 2013;34(5):1062–1068. doi:10.1093/carcin/bgt030
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
