Back to Journals » International Journal of General Medicine » Volume 14
Application of Single Nucleotide Polymorphism Microarray in Prenatal Diagnosis of Fetuses with Central Nervous System Abnormalities
Authors Xie X , Wu X , Su L , Cai M, Li Y, Huang H, Xu L
Received 8 June 2021
Accepted for publication 28 July 2021
Published 6 August 2021 Volume 2021:14 Pages 4239—4246
DOI https://doi.org/10.2147/IJGM.S323899
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Scott Fraser
Xiaorui Xie, Xiaoqing Wu, Linjuan Su, Meiying Cai, Ying Li, Hailong Huang, Liangpu Xu
Medical Genetic Diagnosis and Therapy Center, Fujian Key Laboratory for Prenatal Diagnosis and Birth Defect, Fujian Provincial Maternity and Children’s Hospital, Affiliated Hospital of Fujian Medical University, Fuzhou, People’s Republic of China
Correspondence: Liangpu Xu
Medical Genetic Diagnosis and Therapy Center, Fujian Key Laboratory for Prenatal Diagnosis and Birth Defect, Fujian Provincial Maternity and Children’s Hospital, Affiliated Hospital of Fujian Medical University, Fuzhou, People’s Republic of China
Tel +86 59187554929
Email [email protected]
Background: The current gold standard of karyotype analysis for prenatal diagnosis of fetuses with central nervous system (CNS) abnormalities has some limitations. Here, we assessed the value of single nucleotide polymorphism (SNP) arrays as a diagnostic tool.
Methods: The results of prenatal diagnosis of 344 fetuses with CNS abnormalities as determined by ultrasonographic screening were retrospectively analyzed. All fetuses underwent chromosomal karyotype analysis and genome-wide SNP array analysis simultaneously. The resultant rates and frequencies of genomic abnormalities were compared.
Results: Karyotype analysis found 45 (13.2%) abnormal CNS cases, while SNP array found 60 (17.4%) cases. SNP array detected 23 (6.7%) cases of submicroscopic abnormalities that karyotype analysis did not find. The detection rate of karyotype analysis was 8.1% in the group with isolated CNS anomalies, but 16.5% in the group with CNS abnormalities plus extra ultrasound anomalies. Detection rates of SNP array were 12.4% and 20.8% in these two groups, respectively. Statistical analysis showed that the detection rates of both methods were significantly higher in the group with CNS malformations and other ultrasound anomalies than in the group with isolated CNS anomalies. Abnormal chromosomes were detected most frequently in fetuses with holoprosencephaly.
Conclusion: Genome-wide SNP array technology can significantly improve the positive detection rate of fetuses with CNS abnormalities. Combining karyotype analysis and SNP array technology is recommended for detecting the development of fetuses with abnormal CNS.
Keywords: central nervous system abnormalities, chromosome, karyotype analysis, single nucleotide polymorphism microarray, prenatal diagnosis
Introduction
Fetal central nervous system (CNS) defects are caused by abnormal CNS development during the embryonic period. Common CNS defects include anencephaly, neural tube defects, choroid plexus cyst, ventriculomegaly, hydrocephalus, abnormalities of the corpus callosum and cavum septum pellucidum, holoprosencephaly, lissencephaly, cerebellar and posterior fossa abnormalities, as well as some neurological syndromes, such as Dandy-Walker malformation and Joubert syndrome. Collectively, fetal CNS malformations are the most common and serious congenital malformations.1 The prevalence of CNS abnormalities in live births is 0.14%–0.16%, reaching as high as 3%–6% in stillbirths.2–4 Depending on the mechanism of formation, fetal CNS defects may be the intracranial manifestation of malformations in other systems, or they may indicate a fetal chromosomal abnormality, which is usually considered the primary cause.4,5
Although karyotype analysis is considered the gold standard for prenatal cytogenetic detection,6 it has some limitations. The technique cannot consistently detect microdeletions and microduplications less than 5 Mb in size. In addition, karyotype analysis requires cell culture, has a long reporting period, and has a higher risk of sample contamination that leads to experiment failure.7 Duplication/deletion of chromosomal segments less than 5–10 Mb can cause microdeletion and microduplication syndromes that play an important role in multiple malformations and mental retardation.8–10 Indeed, about 6%–15% of genetic diseases are associated with microduplication/microdeletion of genomic segments (also known as copy number variations, CNVs).11,12 Conventional karyotype analysis cannot detect CNVs, which are only detectable through high-resolution techniques.
Chromosomal microarray analysis (CMA) has become a first-line technique for prenatal diagnosis of fetuses with structural abnormalities identified through ultrasound.13–15 CMA can be separated into comparative genomic hybridization (CGH) chips and single nucleotide polymorphism (SNP) chips. Both techniques can detect microdeletions and microduplications across the whole genome, as small as 50–100 kb. The SNP array can detect regions of homozygosity, triploidy, and maternal cell contamination, which the CGH array cannot. In addition, since CMA does not require cell culture, the results can be reported within 3–4 days; therefore, it is increasingly widely used in clinical diagnosis.
In this study, chromosome karyotype and SNP array analyses were performed on 344 fetuses with CNS abnormalities. The results were compared to evaluate the application value of SNP arrays in prenatal diagnosis, which should provide guidance for genetic counseling.
Methods
Study Population
This retrospective study included 344 pregnant women admitted to Fujian Provincial Maternity and Children’s Hospital, China, from January 2016 to July 2020, and excluded those with twin pregnancies and those who did not undergo both chromosome karyotype and SNP array analyses. All 344 fetuses were diagnosed with CNS abnormalities via ultrasonographic screening. Written informed consent for participation was received for all patients. The study was approved by the Protection of Human Ethics Committee of Fujian Maternity and Child Health Hospital and complied with the Declaration of Helsinki.
Interventional Surgery
Interventional surgeries were performed by obstetricians using standard clinical procedures under ultrasound guidance.16,17 Chorionic villi (~10 mg) were extracted using choriocentesis at 9–13 weeks, amniotic fluid (~20–40 mL) was extracted via amniocentesis at 18–24 weeks, and cord blood (~4 mL) was extracted via umbilical vein puncture after 24 weeks.
Chromosome Karyotype Analysis
Fetal chromosome karyotype analysis was performed in accordance with our laboratory’s routine operation at the 320–400 band level resolution. Giemsa banding (G-banding) karyotypes were analyzed and diagnosed according to the International System for Human Cytogenomic Nomenclature (ISCN2016).18
SNP Array and Data Interpretation
Fetal DNA was extracted using the QIAGEN DNA Mini Kit (Qiagen, Valencia, CA, USA) and analyzed using the CytoScan 750K gene chip detection platform (Affymetrix Inc., Santa Clara, CA, USA). CNV thresholds were set to report deletions >200 kb or duplications >500 kb. Data were analyzed using the Chromosome Analysis Suite (ChAS; Affymetrix). SNP array results were assessed with reference to the following databases: Database of Genomic Variants (DGV, http://projects.tcag.ca/variation), Database of Chromosomal Imbalance and Phenotype in Humans Using Ensembl Resources (DECIPHER, htts://decinher.sanger.ac.uk/), International Standards for Cytogenomic Arrays (ISCA, https://www.iscaconsortium.org/) Consortium, and Online Mendelian Inheritance in Man (OMIM, http://www.omim.org). Peripheral blood from parents was extracted for SNP array detection when appropriate. These databases, scientific literature, and ultrasonography findings were used to systematically evaluate the clinical significance of CNVs, which were then categorized as pathogenic, variants of unknown significance (VUSs), or benign, following the American College of Medical Genetics and Genomics standards and guidelines for interpretation and reporting of postnatal constitutional CNVs.19 VUSs were further subdivided into likely pathogenic, VUSs with no subclassification, and likely benign.
Statistical Analyses
Data are expressed as frequencies and rates. The detection rate was compared in groups using a chi-square test, with a P value < 0.05 indicative of statistical significance.
Results
Patient Clinical Characteristics
The average age of the 344 pregnant women was 29.9 years (range: 18–45); gestational age ranged from 12–37 weeks. Two cases were diagnosed with choriocentesis, 178 cases with amniocentesis, and 164 cases with umbilical vein puncture. Among the 344 cases, 137 (39.8%) had simple nervous system malformations and 207 (60.2%) had combined extrinsic nervous system malformations.
Karyotype Analysis
Fetal amniotic fluid cell cultures failed in 2 out of 344 cases; the overall success rate was 99.4%. Of the 342 successful cultures, 297 (86.8%) had a normal karyotype (including two chromosome polymorphisms). Of the 45 (13.2%) cases with an abnormal karyotype, 28 had abnormal chromosomal numbers, including 6 cases of trisomy 21, 11 cases of trisomy 18, 5 cases of trisomy 13, and 6 cases of sexual and other chromosome abnormalities (including three chimeras). Among the 17 chromosomal structural abnormalities detected, 7 cases had balanced chromosomal structural abnormalities, and the remaining 10 had large-fragment duplication/deletion (Table 1).
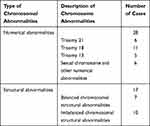 |
Table 1 Karyotype Analysis Results Showing Chromosomal Abnormalities |
SNP Array
The SNP array revealed 284 cases with normal results and 60 cases with abnormal results, with an abnormality rate of 17.4% (60/344). The detection rate appeared higher than traditional chromosome karyotype analysis, but this difference was not statistically significant (χ2 = 2.43, P > 0.05). Among the 60 abnormal SNP array results, 25 cases had numerical abnormalities (including one chimera), and 12 cases had structural abnormalities in large segments. In addition, the SNP array detected 23 (6.7%) cases of submicroscopic abnormalities that karyotype analysis did not find. In one case of structural abnormalities in large segments, SNP array identified two additional microdeletions. After searching the database and literature, we concluded that 4 of the 24 (one case had two microdeletions) submicroscopic abnormalities were clearly pathogenic mutations, 5 were likely pathogenic, 8 were mutations with unknown clinical significance, and 7 were likely benign mutations. Table 2 presents details of the 23 cases with submicroscopic abnormalities.
 |
Table 2 Copy Number Variations Detected in Fetuses with CNS Anomalies Using SNP Array |
Detection Rates of Anomalies in Fetuses with Isolated CNS Anomalies and Fetuses with CNS Anomalies Plus Extra Ultrasound Anomalies
The detection rates of karyotype analysis were 8.1% (11/136) in the group with isolated CNS anomalies and 16.5% (34/206) in the group with CNS anomalies plus extra ultrasound anomalies; the respective detection rates of SNP array were 12.4% (17/137) and 20.8% (43/207) (Table 3). For both analyses, detection rates were significantly higher in the group with CNS malformations and other ultrasound anomalies than in the group with isolated CNS anomalies (karyotype: χ2 = 5.08, P < 0.05; SNP array: χ2 = 4.01, P < 0.05).
 |
Table 3 Detection Rates of Abnormalities in Fetuses with CNS Anomalies Linked to Different Ultrasound Findings |
Incidence of Chromosomal Abnormalities in Different Types of Fetal CNS Anomalies
Both karyotype and SNP array analyses detected abnormal chromosomes most frequently in fetuses with holoprosencephaly. Table 4 depicts the relationship between different types of CNS anomalies and the incidence of chromosomal abnormalities.
 |
Table 4 Types of Fetal CNS Anomalies and Abnormal Chromosome Incidence |
Discussion
The CNS is one of the most complex systems in the human body, and CNS malformations are the most common of all fetal defects.20 Common teratogenic factors include chromosomal abnormalities, intrauterine infections, and the use of certain drugs. Understanding the etiology of CNS anomalies is very important for evaluating fetal prognosis and recurrence risk. Numerous studies have shown that fetal CNS anomalies and numerical chromosomal abnormalities are closely linked.21–23 Advancements in molecular biology have gradually revealed genomic mutations that cause microdeletion or microduplication syndromes associated with fetal CNS malformations.24 In recent years, CMA has become increasingly popular for prenatal diagnosis because of its short detection cycle, high throughput, and high resolution. In our study, karyotype analysis and SNP array analysis detected 45 (13.2%) and 60 (17.4%) abnormal cases, respectively; the different detection rates were not statistically significant. Nevertheless, SNP array identified an additional 23 cases (6.7%) carrying submicroscopic abnormalities that escaped karyotype analysis, and 39.1% (9/23) of these cases were pathogenic or likely pathogenic, consistent with previous studies.25,26
Moreover, some CNVs were clearly related to CNS abnormalities. For example, duplication of the 7q36 region in case 3 (Table 2) involves the SHH gene, which is linked to holoprosencephaly.27 Likewise, in case 9, the microdeletion in the 14q21.2q21.3 is related to the MDHA2 gene, which is involved in the development of the nervous system.28 The microdeletion of 16p11.2 in cases 14 and 15 is an important genetic risk factor for neurodevelopmental disorders, and it also affects the development of brain structure.29–31
These findings indicate that chromosome microarray is necessary when an ultrasound suggests fetal CNS abnormalities but the karyotype analysis suggests the opposite. An SNP array provides more information for assessing fetal prognosis. The clinical significance of genetic information is conducive to genetic counseling and assessment of recurrence risk. The detection rate of karyotype analysis was 8.1% in fetuses with isolated CNS anomalies and 16.5% in fetuses with CNS anomalies plus extra ultrasound anomalies, while detection rates of SNP array were 12.4% and 20.8% in these two groups, respectively. Detection rates were significantly higher in the group with CNS malformations plus other ultrasound anomalies than in the group with isolated CNS anomalies. Thus, fetuses with CNS anomalies (especially if combined with extra system malformations) likely have more chromosomal abnormalities. These findings are similar to previous reports, although some of the studies only showed a higher detection rate of non-isolated CNS abnormalities and no significant difference between the two groups.3,32
In this study, detection of holoprosencephaly (66.7%, 4/6) had the highest positive rate among the different types of CNS abnormalities. All abnormal cases were trisomy 18 syndromes. Studies show that 39% of reported trisomy 13 cases in fetuses had neurological malformations, such as holoprosencephaly and dilatation of the cisterna magna.
Despite several advantages, SNP array cannot completely replace karyotype analysis because it cannot detect balanced chromosomal structural abnormalities, such as balanced translocations and inversions. Although most of these anomalies are inherited from parents and do not always affect fetal phenotypes, they can cause spontaneous abortion during reproduction or generate unbalanced chromosomal structures in the fetus. In addition, SNP arrays may ignore a low proportion of mosaic chromosomal abnormalities. Therefore, combining SNP arrays with karyotype analysis will provide more genetic information for clinical guidance and the two methods cannot be replaced by each other.
This study had some possible limitations. First, the study focused on fetuses with CNS abnormalities, whether structural malformations or soft index abnormalities, and we did not analyze the correlation between the two and chromosomal abnormalities. Second, the two methods used in this study cannot identify pathogenic mutations at the gene level, which can be addressed in future research by the application of Next Generation Sequencing (NGS) technology.
Conclusion
In summary, fetal CNS abnormalities, which are related to chromosomal aneuploidy and CNVs, are important indicators for prenatal diagnosis. Chromosomal abnormality rate increases significantly when malformations occur across different systems. SNP array can improve the detection rate of genetic causes in fetuses with CNS abnormalities during prenatal diagnosis. Therefore, when such fetuses are identified through ultrasound, a prenatal diagnosis that includes both karyotype analysis and SNP array should be recommended. The integrated application of karyotype analysis and SNP array can provide us with more genetic information to avoid a missed diagnosis, provide more basis for genetic counseling, and better understand the genetic etiology of CNS abnormalities.
Acknowledgments
We thank all the patients and research workers for their participation.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Van den Veyver IB. Prenatally diagnosed developmental abnormalities of the central nervous system and genetic syndromes: a practical review. Prenat Diagn. 2019;39(9):666–678. doi:10.1002/pd.5520
2. Onkar D, Onkar P, Mitra K. Evaluation of fetal central nervous system anomalies by ultrasound and its anatomical co-relation. J Clin Diagn Res. 2014;8(6):AC05–07.
3. Sun L, Wu Q, Jiang SW, et al. Prenatal diagnosis of central nervous system anomalies by high-resolution chromosomal microarray analysis. Biomed Res Int. 2015;2015:426379. doi:10.1155/2015/426379
4. Huang J, Wah IY, Pooh RK, Choy KW. Molecular genetics in fetal neurology. Semin Fetal Neonatal Med. 2012;17(6):341–346. doi:10.1016/j.siny.2012.07.007
5. Hof MCVD, Wilson RD. Fetal soft markers in obstetric ultrasound. J Obstet Gynaecol Ca. 2005;27(6):592–636.
6. Schaaf CP, Wiszniewska J, Beaudet AL. Copy number and SNP arrays in clinical diagnostics. Annu Rev Genomics Hum Genet. 2011;12:25–51. doi:10.1146/annurev-genom-092010-110715
7. Reddy UM, Page GP, Saade GR, et al. Karyotype versus microarray testing for genetic abnormalities after stillbirth. N Engl J Med. 2012;367(23):2185–2193. doi:10.1056/NEJMoa1201569
8. Engels H, Brockschmidt A, Hoischen A, et al. DNA microarray analysis identifies candidate regions and genes in unexplained mental retardation. Neurology. 2007;68(10):743–750. doi:10.1212/01.wnl.0000256367.70365.e0
9. Stankiewicz P, Beaudet AL. Use of array CGH in the evaluation of dysmorphology, malformations, developmental delay, and idiopathic mental retardation. Curr Opin Genet Dev. 2007;17(3):182–192. doi:10.1016/j.gde.2007.04.009
10. Vissers LE, de Vries BB, Veltman JA. Genomic microarrays in mental retardation: from copy number variation to gene, from research to diagnosis. J Med Genet. 2010;47(5):289–297. doi:10.1136/jmg.2009.072942
11. Vissers LE, Veltman JA, van Kessel AG, Brunner HG. Identification of disease genes by whole genome CGH arrays. Hum Mol Genet. 2005;14(2):R215–223. doi:10.1093/hmg/ddi268
12. Wapner RJ, Martin CL, Levy B, et al. Chromosomal microarray versus karyotyping for prenatal diagnosis. N Engl J Med. 2012;367(23):2175–2184. doi:10.1056/NEJMoa1203382
13. Liao C, Fu F, Li R, et al. Implementation of high-resolution SNP arrays in the investigation of fetuses with ultrasound malformations: 5 years of clinical experience. Clin Genet. 2014;86(3):264–269. doi:10.1111/cge.12271
14. Charan P, Woodrow N, Walker SP, Ganesamoorthy D, McGillivray G, Palma-Dias R. High-resolution microarray in the assessment of fetal anomalies detected by ultrasound. Aust N Z J Obstet Gynaecol. 2014;54(1):46–52. doi:10.1111/ajo.12170
15. Miller DT, Adam MP, Aradhya S, et al. Consensus statement: chromosomal microarray is a first-tier clinical diagnostic test for individuals with developmental disabilities or congenital anomalies. Am J Hum Genet. 2010;86(5):749–764. doi:10.1016/j.ajhg.2010.04.006
16. American College of O, Gynecologists. ACOG Practice Bulletin No. 88, December 2007. Invasive prenatal testing for aneuploidy. Obstet Gynecol. 2007;110(6):1459–1467. doi:10.1097/01.AOG.0000291570.63450.44
17. Bian X. Practical Prenatal Diagnosis. Beijing: People’s Military Medical Press; 2008.
18. International Standing Committee on Human Cytogenomic Nomenclature. McGowan-Jordan J, Simons A, Schmid M. ISCN: An International System for Human Cytogenomic Nomenclature (2016). Basel, New York: Karger; 2016.
19. Kearney HM, Thorland EC, Brown KK, Quintero-Rivera F, South ST; Working Group of the American College of Medical Genetics Laboratory Quality Assurance C. American College of Medical Genetics standards and guidelines for interpretation and reporting of postnatal constitutional copy number variants. Genet Med. 2011;13(7):680–685. doi:10.1097/GIM.0b013e3182217a3a
20. Petracchi F, Crespo L, Michia C, Igarzabal L, Gadow E. Holoprosencephaly at prenatal diagnosis: analysis of 28 cases regarding etiopathogenic diagnoses. Prenat Diagn. 2011;31(9):887–891.
21. Lehman CD, Nyberg DA, Winter TC, Kapur RP, Resta RG, Luthy DA. Trisomy 13 syndrome: prenatal US findings in a review of 33 cases. Radiology. 1995;194(1):217–222. doi:10.1148/radiology.194.1.7997556
22. Gezer C, Ekin A, Ozeren M, et al. Chromosome abnormality incidence in fetuses with cerebral ventriculomegaly. J Obstet Gynaecol. 2014;34(5):387–391. doi:10.3109/01443615.2014.896885
23. Hayashi Y, Suzumori N, Sugiura T, Sugiura-Ogasawara M. Prenatal findings of holoprosencephaly. Congenit Anom (Kyoto). 2015;55(3):161–163. doi:10.1111/cga.12103
24. Tang XH, Yang BC, Zhu S, et al. Prenatal diagnosis of chromosome abnormalities and nine microdeletion syndromes using both traditional karyotyping and BoBs. Chin J Obstet Gynecol. 2016;51(5):325–330.
25. D’Amours G, Kibar Z, Mathonnet G, et al. Whole-genome array CGH identifies pathogenic copy number variations in fetuses with major malformations and a normal karyotype. Clin Genet. 2012;81(2):128–141. doi:10.1111/j.1399-0004.2011.01687.x
26. Hillman SC, Pretlove S, Coomarasamy A, et al. Additional information from array comparative genomic hybridization technology over conventional karyotyping in prenatal diagnosis: a systematic review and meta-analysis. Ultrasound Obst Gyn. 2011;37(1):6–14. doi:10.1002/uog.7754
27. Paulussen AD, Schrander-Stumpel CT, Tserpelis DC, et al. The unfolding clinical spectrum of holoprosencephaly due to mutations in SHH, ZIC2, SIX3 and TGIF genes. Eur J Hum Genet. 2010;18(9):999–1005. doi:10.1038/ejhg.2010.70
28. Litwack ED, Babey R, Buser R, Gesemann M, O’Leary DD. Identification and characterization of two novel brain-derived immunoglobulin superfamily members with a unique structural organization. Mol Cell Neurosci. 2004;25(2):263–274. doi:10.1016/j.mcn.2003.10.016
29. Hanson E, Bernier R, Porche K, et al. The cognitive and behavioral phenotype of the 16p11.2 deletion in a clinically ascertained population. Biol Psychiatry. 2015;77(9):785–793. doi:10.1016/j.biopsych.2014.04.021
30. Rosenfeld JA, Coppinger J, Bejjani BA, et al. Speech delays and behavioral problems are the predominant features in individuals with developmental delays and 16p11.2 microdeletions and microduplications. J Neurodev Disord. 2010;2(1):26–38. doi:10.1007/s11689-009-9037-4
31. Martin-Brevet S, Rodriguez-Herreros B, Nielsen JA, et al. Quantifying the effects of 16p11.2 copy number variants on brain structure: a multisite genetic-first study. Biol Psychiatry. 2018;84(4):253–264. doi:10.1016/j.biopsych.2018.02.1176
32. Song T, Xu Y, Li Y, et al. Detection of submicroscopic chromosomal aberrations by chromosomal microarray analysis for the prenatal diagnosis of central nervous system abnormalities. J Clin Lab Anal. 2020;34(10):e23434. doi:10.1002/jcla.23434
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
