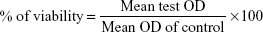Back to Journals » International Journal of Nanomedicine » Volume 10 » Issue 1
Anticancer activity of biostabilized selenium nanorods synthesized by Streptomyces bikiniensis strain Ess_amA-1
Authors Ahmad M, Yasser M, Sholkamy E , Ali AM, Mehanni M
Received 12 February 2015
Accepted for publication 18 March 2015
Published 6 May 2015 Volume 2015:10(1) Pages 3389—3401
DOI https://doi.org/10.2147/IJN.S82707
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Thomas Webster
Maged Sayed Ahmad,1 Manal Mohamed Yasser,1 Essam Nageh Sholkamy,1,2 Ali Mohamed Ali,3,4 Magda Mohamed Mehanni3
1Department of Botany, Faculty of Science, University of Beni-Suef, Beni-Suef, Egypt; 2Department of Botany and Microbiology, College of Science, King Saud University, Riyadh, Saudi Arabia; 3Department of Botany and Microbiology, College of Science, Minia University, El-Minia, Egypt; 4Department of Biological Sciences, College of Science, King Faisal University, Saudi Arabia
Abstract: Selenium is an important component of human diet and a number of studies have declared its chemopreventive and therapeutic properties against cancer. However, very limited studies have been conducted about the properties of selenium nanostructured materials in comparison to other well-studied selenospecies. Here, we have shown that the anticancer property of biostabilized selenium nanorods (SeNrs) synthesized by applying a novel strain Ess_amA-1 of Streptomyces bikiniensis. The strain was grown aerobically with selenium dioxide and produced stable SeNrs with average particle size of 17 nm. The optical, structural, morphological, elemental, and functional characterizations of the SeNrs were carried out using techniques such as UV-vis spectrophotometry, transmission electron microscopy, energy dispersive X-ray spectrometry, and Fourier transform infrared spectrophotometry, respectively. The MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide) assay revealed that the biosynthesized SeNrs induces cell death of Hep-G2 and MCF-7 human cancer cells. The lethal dose (LD50%) of SeNrs on Hep-G2 and MCF-7 cells was recorded at 75.96 µg/mL and 61.86 µg/mL, respectively. It can be concluded that S. bikiniensis strain Ess_amA-1 could be used as renewable bioresources of biosynthesis of anticancer SeNrs. A hypothetical mechanism for anticancer activity of SeNrs is also proposed.
Keywords: biosynthesis, selenium nanorods, Streptomyces, anticancer activity
Introduction
Selenium (Se) is an important trace element that plays a crucial role in human health and regulates many crucial cellular functions mediated through its incorporation into selenoproteins.1,2 This inorganic element also occurs in trigonal and monoclinic phases of crystalline microstructures. The monoclinic phase is less stable and occurs in α, β, and γ forms, which differ only in the way that the crystals are packed.3 The antioxidant function of Se is conferred by some of these selenoproteins that directly or indirectly protect against oxidative stress. The extensive experimental evidence indicates based on the in vitro, animal, geographic, and prospective studies that Se supplementation reduces the incidence of various types of cancers. Since as early as the 1960s, geographical studies have proved a consistent trend for populations with low Se compounds intake to have higher cancer mortality rates.2,4 However, at elevated doses, Se compounds usually turn into a prooxidant with well-established cell growth inhibiting/killing properties.1 Thus, the uses of Se compounds for anticancer therapy have been greatly explored during the last decade and results of studies have shown that Se compounds reduce the risk of various types of cancers, such as mammary, prostate, lung, colon, and liver cancers.1,4–7 The research findings also suggest that the concentration, chemical species, and redox potential of Se compounds are critical for their anticancer activity.1,5 These Se compounds have more promising anticancer activity at high dosage; however, high doses of Se compounds give rise to greater concerns about its toxicity.1,5 In this regard, selenium nanostructured materials (SNMs) could reduce the risk of Se toxicity and be widely used in cancer biology due to its promising anticancer activity and less toxicity, compared to Se compounds (inorganic and organic).6–9 SNMs also exhibit unique physical, chemical, and biological properties compared to that of Se compounds.10 Various types of SNMs, which are stabilized and modified with different kinds of biological macromolecules, are reported to possess excellent anticancer activities.1,11–13 Based on this, some researchers suggest that biological macromolecules stabilized and modified may have potential applications as anticancer agents for the killing of human cancer cells. For these reasons, the study of SNMs has gained considerable importance in recent years and therefore, various types SNMs have been obtained by employing physico-chemical methods, ie, amorphous,3,11 trigonal, nanorods,12,13 nanoribbons,14 hexagonal prism,13 nanoplates,15 nanotubes,16 and spheres.13 Therefore, SNMs are being widely used in basic and applied areas of chemistry, physics, environmental science, material science, and biomedicine.7,13,17,18 However, concern is now growing regarding the environmental impact of the SNM synthesis process based on physico-chemical methods that require high pressures, temperatures, and toxic chemicals. These methods have some drawbacks: (i) production of stable SNMs dispersions only at lower concentrations and unsuitability for large-scale production; (ii) requirement of additional stabilizing agents; (iii) production of hazardous by-products, and (iv) increased pollution in the environment.
Consequently, significant efforts are ongoing toward the development of novel nontoxic methods for the synthesis and surface modification/stabilization of SNMs.19–21 Biogenic methods are a renewable, clean, nontoxic, and environmentally friendly procedure for the synthesis of these types of nanomaterials.7,22,23 Recently, biogenic methods have been utilized for the synthesis of a variety of SNMs at ambient conditions.6,19,24–26 It is well established that biosynthesized SNMs have several important characteristics, including more stability and higher biological activity, due to the surface functionalization by biological macromolecules secreted by fungi and bacteria.7,9,17,23,27 Among the different microorganisms, Actinomycetes are less explored for the synthesis of SNMs. Actinomycetes are a diverse group of filamentous true bacteria found in a variety of habitats in terrestrial and aquatic environments.28 Hence, there are reports that have shown that actinomycetes are efficient bioagents for the intracellular and extracellular synthesis of metal nanostructured materials (MNMs).22 Most of the studies have been done on species of Streptomyces genus due to its inherent capability for the production of redox active macromolecules/secondary metabolites.29,30 The capability of Streptomyces genus for the biosynthesis of MNMs has previously been reported.18,22,31–34 However, the synthesis of biostabilized selenium nanorods (SeNrs) using any strain of actinomycetes has not been reported yet. In the present study, we reported: (i) a simple, environmentally friendly, and renewable biogenic method for synthesizing disperse and stable SeNrs by Streptomyces bikiniensis strain Ess_amA-1 and (ii) presented evidence that biosynthesized SeNrs have the anticancer activity. To the best of our knowledge, this is the first report on the synthesis of SeNrs by S. bikiniensis as a novel renewable bioresource and opens up the possibility of commercially viable biogenic production of SeNrs for novel anticancer nanostructured materials.
Materials and methods
Isolation of S. bikiniensis
An insect Tapinoma simrothi was collected from Eldrieh, Riyadh, Saudi Arabia (24.7 N latitude and 46.7 E longitude), and used for the isolation of S. bikiniensis. The suspension of T. simrothi was prepared in normal saline solution (NSS) for the isolation of saccharolytic actinomycetes and an appropriate dilution was spread on the starch casein agar (SCA) medium (pH 7.2±0.2), supplemented with antibiotics (cycloheximide [40 mg/L], nystatin [30 mg/L], and nalidixic acid [10 mg/L]).35,36 The inoculated Petri plates were incubated aerobically at 30°C until the appearance of powdery texture colonies with branching filaments and aerial mycelia. The selected colonies were subcultured and further purified by streaking and among them the strain Ess_amA-1was selected and maintained on International Streptomyces Project 2 (ISP-2) agar medium by periodical subculturing.
Morphological and physiological characterization of S. bikiniensis strain Ess_amA-1
The color of aerial mycelium was determined from mature and sporulating aerial mycelia of the actinomycete colonies on different media such as ISP-2, ISP-4, ISP-6, ISP-7, Czapek Dox, and SCA. The color was determined using color names lists.37 The color of the soluble pigments was determined visually by observing the color changes in the medium due to the diffusing pigments produced by strain Ess_amA-1.38 Carbohydrates and physiological tests were performed using the specific media and methods.39–41 All the cultures were incubated at 30°C for 7 days. The assay for enzymatic activity was performed according to Bibb et al.42
Genomic DNA extraction and purification
Total genomic DNA of strain Ess_amA-1 was isolated from the mycelium biomass (0.1 g), which was harvested from the freshly grown culture in ISP-2 medium as described. Briefly, the collected mycelium biomass was crushed with liquid nitrogen and the powder was mixed with 500 μL lysis buffer (containing 50 mM Tris-HCl, pH 8.0; 5 mM EDTA, pH 8.0; 50 mM NaCl; and 20 μL lysozyme, 10 mg/mL). The cells were lysed by vigorous vortexing and lysate was incubated at 37°C for 30 minutes. Subsequently, 20 μL SDS (10% w/v) and 20 μL of proteinase K (10 mg/mL) were added into the Eppendorf tube and incubated at 55°C for 30 minutes. The cell lysate was cooled down and extracted once with an equal volume of phenol and chloroform (1:1 v/v). The aqueous phase was collected by centrifugation at 10,000 rpm for 5 minutes. Total genomic DNA was precipitated from the obtained aqueous phase by the addition of two volumes of chilled isopropanol. The precipitated genomic DNA was palletized by centrifugation at 13,000 rpm for 30 minutes and the pellet was washed with 70% ethanol. The washed pellet was air dried under laminar flow and dissolved in 50 μL TE buffer (containing 50 mM Tris-HCl and 1 mM EDTA; pH 7.2).
Polymerase chain reaction amplification of 16S ribosomal RNA (16S rRNA) gene and nucleotide sequencing
For molecular characterization, 16S ribosomal RNA (16S rRNA) gene was amplified using primer set Star-F (5′-GAGTTTGATCMTGGCTCAG-3′) and 1387-R (5′-CGGGCGGTGTGTACAAGG-3′). Briefly, ~50 ng genomic DNA was used as the template in 25 μL of polymerase chain reaction (PCR) mixture containing 0.20 mM dNTPs (Deoxyribonucleotide triphosphates), 25 pmole of each primer, 5× Taq polymerase buffer (5 μL), 2.0 mM MgCl2, and 0.2 U Taq DNA polymerase. The PCR parameters consisted of 35 cycles of denaturation at 94°C for 1 minute, annealing at 51°C for 1 minute, extension at 72°C for 1 minute with an initial denaturation, and the final extension was performed for 3 minutes and 7 minutes at 94°C and 72°C, respectively. Amplification was performed on the PCR machine (Sens Quest co., D-37085 Goettingen, Germany). The amplicon was analyzed and sequenced by GATC Biotech, European Custom Sequencing Centre (Cologne, Germany).
Multiple sequence alignments and phylogenetic analysis
The obtained 16S rRNA gene sequence was compared with the homologous sequences retrieved from GenBank using the Blastn tool.43 Multiple sequence analysis with the sequences of different actinomycetes groups was performed using CLUSTALW with default parameters.44 A phylogenetic tree was constructed by the neighbor-joining method with nucleotide pair-wise genetic distances corrected by the Kimura two-parameter method45 using the TreeCon tool. The reliability of the tree topology was subjected to a bootstrap test and numbers at the nodes indicate bootstrap support values as a percentage of 1,000 replications. All branches with <50% bootstrap support were judged as inconclusive and were collapsed and branch lengths for all trees were normalized to 0.02% divergence. Based on biochemical and molecular characterization, the characterized strain was designated as S. bikiniensis strain Ess_amA-1.
Biosynthesis of SeNrs
Briefly, sterile 100 mL ISP-2 medium containing 1 mM selenium dioxide (SeO2) was inoculated with 1 mL of the fresh inoculums (OD600, 0.5) of strain Ess_amA-1 and incubated in an orbital shaker incubator (150 rpm) at 30°C for 48 hours. A control flask containing ISP-2 without SeO2 was inoculated with a test strain and incubated under the same conditions. The reduction of SeO2 into the elemental selenium (S0) and the nucleation/growth of SeNrs were monitored by sampling an aliquot of the medium at different time periods of incubation (6 hours, 12 hours, and 48 hours). The cells were then removed by filtration and the resulting cell-free filtrate was then centrifuged at 14,000 rpm for 15 minutes to obtain the biosynthesized SeNrs.
Characterization of biosynthesized SeNrs
The optical, structural, morphological, elemental, and functional characterizations of the SeNrs were carried out using UV–Vis spectrophotometer, transmission electron microscope (TEM), energy dispersive X-ray (EDAX) spectrometer, and Fourier transform infrared (FTIR) spectrophotometer, respectively. In order to ascertain the optical characteristics of synthesized SNTs, the absorption spectrum was recorded by Lambda 35 double beam UV–Vis spectrophotometer (Hitachi, Japan) in the wavelength range of A200–A800 nm using a quartz cuvette. The size and structure of the biosynthesized SeNrs were analyzed by JEM-1010 TEM (JEOL, Tokyo, Japan) at an accelerating voltage of 80 kV. For this analysis, the sample was prepared by placing drops of SeNrs aqueous solution on carbon coated copper grids and air dried under dark conditions. The elemental analysis of SeNrs was done using the energy dispersive X-ray spectroscopy (EDS) equipped with JSM-6380 LA scanning electron microscope (SEM) (JEOL, Tokyo, Japan). For functional characterization of SeNrs, the FTIR spectrum was recorded in the range of 400–4,000 wave number (cm−1) on Nicolet 6700 FTIR spectrometer in the transmittance mode at a 4 cm−1 resolution.
3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide assay
The anticancer activity of biosynthesized SeNrs was tested on human breast adenocarcinoma cell line (MCF-7) and human liver carcinoma cell line (Hep-G2) cells (ATCC, Manassas, VA, USA) using the 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) dye reduction assay. This assay is based on the reduction of MTT dye to a blue colored formazan product by mitochondrial dehydrogenase. The cells were cultured in a humid environment at 37°C and 5% CO2 in a cell culture minimum essential medium (Thermo Fisher Scientific, Waltham, MA, USA) supplemented with 15% fetal bovine serum and 1% penicillin/streptomycin (Thermo Fisher Scientific). At 85%–90% confluence, cells were harvested using 0.25% trypsin/EDTA solution and subcultured into a 96-well plate. The MTT colorimetric assay developed by Mosmann46 with modification was used to screen the cytotoxic activity of SeNrs. Briefly, the MCF-7 and Hep-G2 cells (1×104 cells/well) were grown overnight in 96-well flat bottom culture plates, and then exposed to seven different concentrations (1.0, 2.0, 5.0, 10, 25, 50, and 100 μg/mL) of SeNrs for 24 hours. In addition, negative/vehicle control and positive control (doxorubicin) were also used for comparison. After the completion of the desired treatment, 10 μL of MTT reagent (Thermo Fisher Scientific) was added to each well and further incubated for 3 hours at 37°C. Finally, the medium with MTT solution was removed, and 100 μL of DMSO (Sigma-Aldrich Co., St Louis, MO, USA) was added to each well and further incubated for 20 minutes. The optical density (OD) of each well was measured at 570 nm using a microplate reader (Synergy; BioTek, Winooski, VT, USA). The percentage of cytotoxicity compared to the untreated cells was determined. Triplicates were maintained for each treatment. Lethal concentration (LC50) was determined by calculating the cell viability:
|
Statistical analysis
The triplicate sets of data for the various parameters evaluated were subjected to analysis of variance in accordance with the experimental design (completely randomized design) using SAS statistical packages (Cary, version 6.12; SAS Institute Inc., NC, USA) to quantify and evaluate LSD. The values were calculated at P level of 0.05%.
Results and discussion
Characterization of strain Ess_amA-1
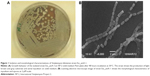
In the present study, the actinomycete strain Ess_amA-1 was isolated from an insect T. simrothi with the aim of exploiting its SeNrs synthesizing potential for anticancer therapy. The presumptive taxonomic identification of the strain was done by its morphological and biochemical characteristics. For the identification of morphological characteristics, a specimen of the strain was examined under the bright-field microscope and analysis revealed that the strain Ess_amA-1 produces a light brown and gray substrate (Figure 1A). These morphological characteristics of the strain were precisely confirmed by the SEM. The image showed that the smooth-surfaced spores are held in straight chains (rectiflexibiles) (Figure 1B).47,48 The morphological characteristics of the strain Ess_amA-1 were further substantiated by physiological and biochemical tests. The results of the physiological and biochemical characteristics are summarized in Tables 1 and 2. The strain showed fast growth behavior on media (ISP-2, 4, 6, and 7) and moderate growth behavior on ISP-5 and Czapek-Dox agar medium; melanin pigment production was determined on ISP-6 medium. Thus, based on the morphological and biochemical characteristics of strain Ess_amA-1 was identified as a member of Streptomyces genus.38 For the authentic taxonomic characterization of the strain, 16S rRNA gene sequencing was performed and obtained data were analyzed carefully. The strain was identified as a member of S. bikiniensis by 16S rRNA gene sequencing and in silico analysis that has been deposited in NCBI GenBank (Accession Number: KF588366). The total length of the 16S rRNA gene sequenced in the present study has 1,206 base pairs, and NCBI Blastn analysis revealed ~99% sequence homology with the strain of S. bikiniensis clone ZD Hu (GenBank, Accession No: AY946043.1). The related sequence data sets were retrieved from the Blastn result and phylogenetic analysis indicated a close genetic relatedness of strain Ess_amA-1 with S. bikiniensis and we therefore designated the strain as S. bikiniensis strain Ess_amA-1 (Figure 2). The members of the Streptomyces genus have been largely exploited for the production of bioactive secondary metabolites (ie, antimicrobials, antitumorals, antihypertensives, and immunosuppressants) with wide uses in medicine and agriculture.30,49,50 Thus, the species of the genus Streptomyces are well established bioresources for the production of valuable nanostructured materials.18,22,31–34,51–57
Biosynthesis of SeNrs
The bioreductive capability of the strain was utilized for the synthesis of SeNrs. The strain when challenged with 1 mM SeO2 exhibited a time-dependent change in the color of the ISP-2 liquid culture medium from light gray to red after 6 hours incubation period. The intensity of the red color of culture medium increased upon further incubation up to 48 hours (Figure 3). The emergence of a red-brick color in the culture medium after incubation of 48 hours was a clear indication that the strain biogenically easily reduces selenite ions to insoluble elemental Se (Se0) form.6,8,25,27,58 The yield of SeNrs was determined and was approximately 7.74 mg/100 mL. The reaction mixture of SeNrs biosynthesis can be optimized to increase the yield and purity by altering the physico-chemical and cultural conditions in the used medium: i) precursor salt, ii) carbon and nitrogen source, iii) pH and oxygen supply, and iv) addition of electron donor, etc.59
Characterizations (optical, morphological, elemental, and functional) of SeNrs
The synthesis of SeNrs in liquid culture medium was monitored by the UV–Vis spectroscopy that showed a strong and broad surface plasmon resonance (SPR) peak at ~620 nm, which is a characteristic of SeNrs (Figure 4A). However, no absorption peak corresponding to the SeNrs in the control flask (without SeO2) was observed. It is well known that due to Mie scattering, SeNrs exhibit absorption at the wavelength ~620 nm. As evident from previous reports, the presence of a single SPR peak evokes biogenic synthesis of SeNrs by S. bikiniensis strain Ess_amA-1, and this was further confirmed by TEM and EDS techniques.58 The time-dependent intensity of red color increase of the culture medium indicated the gradual growth in the size and shape of SeNrs during the incubation period. Hence, the SeNrs were analyzed at three different incubation times, 6 hours, 24 hours, and 48 hours, employing the TEM. The time-dependent change in the shape and morphology of SeNrs was noticed. During the incubation, the shape of reduced elemental selenium (Se0) gradually changed from the spherical structure to a rod-like structure. Figure 4Ba shows the spherical shape of Se0 irregular nanospheres, which possess an average diameter of 50–100 nm after 6 hours of incubation. However, these spherical shape structures start to lose their integrity as relatively low aspect-ratio anisotropic structures (rods). After 12 hours, aggregates of higher aspect-ratio rods emerging from a few growth centers were observed (Figure 4Bb). After 48 hours, the length of the structures increased in one dimension and they were converted into the rod-like structures (Figure 4Bc). The SeNrs with an average length of 600 nm, average diameter of 17 nm, and aspect ratio of 35:1 were observed. Nevertheless, most of the biosynthesis methods have reported the production of spherical shape selenium nanoparticles (SeNPs).24,26,27,60 However, few recent studies reported the biosynthesis of Se nanorods via spherical SeNPs that acted as seeds for growth.6,17 The high free energy of SeNPs evoked the Ostwald ripening process, which may be responsible for the growth of spherical SeNPs into SeNrs.17,61 The TEM data also revealed the nucleation/growth of the SeNrs (6 hours and 12 hours incubation) that is probably due to the presence of aromatic amino acids produced by the strain Ess_amA-1, thus indicating possible adhesion of biological macromolecules on the surface of SeNrs. These data are consistent with the previously documented occurrence of biological macromolecules associated to SNMs of microbial origin.7,9,22 The EDX analysis revealed the presence of Se peak at 1.37 keV, confirming that the SeNrs was successfully synthesized.23 However, the peaks of carbon (C) and oxygen (O) were believed to be derived from biological macromolecules present on the surface of SeNrs (Table 3). These biological macromolecules may be responsible for the reduction, growth, and stabilization of biosynthesized SeNrs.25 FTIR spectroscopy was performed to identify the functional groups of the biological macromolecules responsible for the reduction of SeO2 into the elemental selenium (S0) and the nucleation/growth of SeNrs. The FTIR spectrum shows the characteristic stretching vibration bands of proteins on the surface of biosynthesized SeNrs (Figure 4C). The band positions at 3,430 cm−1 were assigned to the stretching vibrations of –N–H and C=O from the amide A and amide I bands of proteins/enzymes, respectively.62 The free amine or cysteine groups of proteins have a strong ability to bind to metal NPs.63,64 FTIR data suggest that the proteins/enzymes produced by the strain Ess_amA-1 are primarily responsible for the synthesis of the SeNrs. The proteins/enzymes present on the surface of SeNrs were acting as natural capping agents, preventing agglomeration and promising anticancer activity. Taken together, the data obtained from EDX and FTIR analyses revealed the purity of SeNrs on the basis of absence of signature peaks of other species of SNMs such as SeO2 NPs.
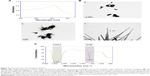  | Figure 4 Characterization of biosynthesized SeNrs. |
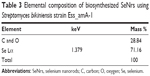  | Table 3 Elemental composition of biosynthesized SeNrs using Streptomyces bikiniensis strain Ess_amA-1 |
Assessment of anticancer activity of SeNrs
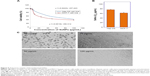
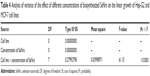
Nanostructured selenium materials have attracted substantial attention due to its excellent biological activity and low toxicity.17 With promising applications in cancer nanotechnology, SNMs are being touted as the new anticancer and chemopreventive agents.65 Very recently, the cytotoxic effect of the inorganic and organic Se compounds on the MCF-7 cell line was assessed.1,9 Various types of SNMs stabilized and surface modified with different types of biological macromolecules/functional groups are reported to possess excellent anticancer activities via the including induction of reactive oxygen species (ROS) production, cell cycle arrest, mitochondrial dysfunction, DNA fragmentation, and cell apoptosis.1,66 These SNMs have also been shown to augment the anticancer properties of chemotherapeutic drugs like adriamycin and doxorubicin.67,68 Therefore, the SeNrs were evaluated for their anticancer properties against Hep-G2 and MCF-7 cell lines. The biosynthesized SeNrs showed growth inhibition of Hep-G2 and MCF-7 cells in a dose-dependent manner (Table 4). The inhibitory effect of the SeNrs was significantly higher on the MCF-7 cells than on the Hep-G2 cells. For instance, SeNrs at 10 μg/mL, 25 μg/mL, 50 μg/mL, and 100 μg/mL reduced MCF-7 cell viability to 69.1%, 54.4%, 44.3%, and 37.5%, respectively. All of those values were significantly lower than those of the Hep-G2 cells: 86.9%, 72.5%, 56.4%, and 42.3%, respectively, at LSD (0.05) =5.7% (Figure 5A). The lethal dose (ID50%) of SeNrs on Hep-G2 and MCF-7 cells was obtained at 75.96 μg/mL and 61.86 μg/mL, respectively, as shown in Figure 5B. The data indicated that the effect of SeNrs on MCF-7 cells was significantly more than on Hep-G2 cells. Moreover, microscopic cell morphological observation after MTT staining showed that Hep-G2 and MCF-7 cells treated with SeNrs showed a dose-dependent reduction in cell numbers, loss of cell-to-cell contact, cell shrinkage, and formation of apoptotic bodies (Figure 5C; MCF-7 cells data not shown). These results collectively suggested that biosynthesized SeNrs have the anticancer property against cancer cells and can serve as potential anticancer agents. However, a few recent studies have reported lower toxicity and selectivity of SNMs toward normal cells.1,20 The mechanism involved in the selectivity of the SNMs remains unexplained. Therefore, we tried to explain the plausible mechanism of SNMs for selective killing of cancer cells.
Plausible anticancer mechanism of SeNrs
The inorganic and organic selenium compounds play an essential role in human life and a number of them are considered to possess chemopreventive and therapeutic properties against cancer.66 In situ surface functionalized SNMs via the biosynthesis procedure have recently gained much attention as potential anticancer agents due to their excellent anticancer activity, biocompatibility, and low toxicity, when compared to inorganic and organic Se compounds.1,9,69 Conjugation with functional ligands/groups, indeed, can not only prevent the aggregation of SNMs via plus-to-minus charge interactions, but also enhance the anticancer efficacy.1 These SNMs are established as promising antioxidants (redox modulating) but can also act as prooxidants and thereby exhibit potential anticancer properties in the presence of transition metal ions (Cu). It is well established that tissue, cellular, and serum copper levels are considerably elevated in various malignancies.70,71 These SNMs are able to bind cell chromatin materials (both DNA and Cu[II]) forming a ternary complex. A redox reaction of the Se compound and Cu(II) in the ternary complex may occur, leading to the reduction of Cu(II) to Cu(I), whose reoxidation generates a variety of ROS. Therefore, cancer cells may be more subject to electron shuttle between copper ions and SeNrs to generate ROS, thereby exhibiting the killing effect on Hep-G2 and MCF-7 cells (Figure 6). Thus, our hypothesis is that the anticancer mechanism of SeNrs involves the mobilization of endogenous copper, possibly chromatin-bound copper, and the consequent prooxidant action.
Conclusion
The isolated S. bikiniensis strain Ess_amA-1 has the inherent potential to produce more stabilized, bioefficacious, and ecofriendly SeNrs than physico-chemically synthesized SNMs, and can be exploited for mass-scale production. The biosynthesized SeNrs showed anticancer activity against Hep-G2 and MCF-7 cells under in vitro conditions. SNMs are potent anticancer agents, with a modest effect on normal cells. The exact mechanism by which this anticancer activity is mediated remains unclear to the scientific community. In this paper, we suggest a hypothesis that the anticancer mechanism of SeNrs involves mobilization of elevated endogenous copper of cancer cells and consequent prooxidant action. Nevertheless, in-depth studies should be conducted to investigate the anticancer action mechanism of SNMs.
Acknowledgment
This work was supported by King Saud University, Deanship of Scientific Research, College of Science, Research Center.
Disclosure
The authors report no conflicts of interest in this work.
References
Aboul-Fadl, T. Selenium derivatives as cancer preventive agents. Current Medicinal Chemistry – Anti-Cancer Agents. 2005;5(6):637–652. | ||
Rayman MP. Selenium and human health. Lancet. 2012;379(9822):1256–1268. | ||
Shikuo L, Yuhua S, Anjian X, et al. Rapid, room-temperature synthesis of amorphous selenium/protein composites using Capsicum annuum L extract. Nanotechnology. 2007;18(40):405101. | ||
Rayman MP. Selenium in cancer prevention: a review of the evidence and mechanism of action. Proc Nutr Soc. 2005;64(4):527–542. | ||
Gao F, Yuan Q, Gao L, et al. Cytotoxicity and therapeutic effect of irinotecan combined with selenium nanoparticles. Biomaterials. 2014;35(31):8854–8866. | ||
Srivastava N, Mukhopadhyay M. Biosynthesis and structural characterization of selenium nanoparticles mediated by Zooglea ramigera. Powder Technol. 2013;244:26–29. | ||
Beheshti N, Soflaei S, Shakibaie M, et al. Efficacy of biogenic selenium nanoparticles against Leishmania major: in vitro and in vivo studies. J Trace Elem Med Biol. 2013;27(3):203–207. | ||
Torres SK, Campos VL, León CG, et al. Biosynthesis of selenium nanoparticles by Pantoea agglomerans and their antioxidant activity. J Nanopart Res. 2012;14(11):1236. | ||
Forootanfar H, Adeli-Sardou M, Nikkhoo M, et al. Antioxidant and cytotoxic effect of biologically synthesized selenium nanoparticles in comparison to selenium dioxide. J Trace Elem Med Biol. 2014;28(1):75–79. | ||
Lampis S, Zonaro E, Bertolini C, Bernardi P, Butler CS, Vallini G. Delayed formation of zero-valent selenium nanoparticles by Bacillus mycoides SeITE01 as a consequence of selenite reduction under aerobic conditions. Microb Cell Fact. 2014;13(1):35. | ||
Johnson JA, Saboungi ML, Thiyagarajan P, Csencsits R, Meisel D. Selenium nanoparticles: a small-angle neutron scattering study. J Phys Chem B. 1999;103(1):59–63. | ||
Zhang B, Ye X, Dai W, Hou W, Zuo F, Xie Y. Biomolecule-assisted synthesis of single-crystalline selenium nanowires and nanoribbons via a novel flake-cracking mechanism. Nanotechnology. 2006;17(2):385–390. | ||
Kumar A, Sevonkaev I, Goia DV. Synthesis of selenium particles with various morphologies. J Colloid Interface Sci. 2014;416(0):119–123. | ||
Cao X, Xie Y, Zhang S, Li F. Ultra-thin trigonal selenium nanoribbons developed from series-wound beads. Adv Mater. 2004;16(7):649–653, 578. | ||
Yin H, Xu Z, Bao H, Bai J, Zheng Y. Single crystal trigonal selenium nanoplates converted from selenium nanoparticles. Chem Lett. 2005;34(1):122–123. | ||
Zhang H, Yang D, Ji Y, Ma X, Xu J, Que D. Selenium nanotubes synthesized by a novel solution phase approach. J Phys Chem B. 2004;108(4):1179–1182. | ||
Srivastava P, Braganca JM, Kowshik M. In vivo synthesis of selenium nanoparticles by Halococcus salifodinae BK18 and their anti-proliferative properties against HeLa cell line. Biotechnol Prog. 2014;30(6):1480–1487. | ||
Subbaiya R, Selvam M. Synthesis and characterization of silver nanoparticles from Streptomyces olivaceus sp-1392 and its anticancerous activity against non-small cell lung carcinoma cell line (NCI-H460). Curr Nanosci. 2014;10(2):243–249. | ||
Sarkar J, Saha S, Dey P, Acharya K. Production of selenium nanorods by phytopathogen, Alternaria alternata. Adv Sci Lett. 2012;10(1):111–114. | ||
Feng Y, Su J, Zhao Z, et al. Differential effects of amino acid surface decoration on the anticancer efficacy of selenium nanoparticles. Dalton Trans. 2014;43(4):1854–1861. | ||
Yang F, Tang Q, Zhong X, et al. Surface decoration by Spirulina polysaccharide enhances the cellular uptake and anticancer efficacy of selenium nanoparticles. Int J Nanomedicine. 2012;7:835–844. | ||
Golinska P, Wypij M, Ingle AP, Gupta I, Dahm H, Rai M. Biogenic synthesis of metal nanoparticles from actinomycetes: biomedical applications and cytotoxicity. Appl Microbiol Biotechnol. 2014;98(19):8083–8097. | ||
Shakibaie M, Forootanfar H, Golkari Y, Mohammadi-Khorsand T, Shakibaie MR. Anti-biofilm activity of biogenic selenium nanoparticles and selenium dioxide against clinical isolates of Staphylococcus aureus, Pseudomonas aeruginosa, and Proteus mirabilis. J Trace Elem Med Biol. 2015;29(0):235–241. | ||
Mishra RR, Prajapati S, Das J, Dangar TK, Das N, Thatoi H. Reduction of selenite to red elemental selenium by moderately halotolerant Bacillus megaterium strains isolated from Bhitarkanika mangrove soil and characterization of reduced product. Chemosphere. 2011;84(9):1231–1237. | ||
Srivastava N, Mukhopadhyay M. Biosynthesis and structural characterization of selenium nanoparticles using Gliocladium roseum. J Cluster Sci. 2015. | ||
Oremland RS, Herbel MJ, Blum JS, et al. Structural and spectral features of selenium nanospheres produced by Se-Respiring bacteria. Appl Environ Microbiol. 2004;70(1):52–60. | ||
Mahmoudvand H, Fasihi Harandi M, Shakibaie M, et al. Scolicidal effects of biogenic selenium nanoparticles against protoscolices of hydatid cysts. Int J Surg. 2014;12(5):399–403. | ||
Tiwari K, Gupta RK. Diversity and isolation of rare actinomycetes: an overview. Crit Rev Microbiol. 2013;39(3):256–294. | ||
Dela Cruz R, Gao Y, Penumetcha S, Sheplock R, Weng K, Chander M. Expression of the Streptomyces coelicolor SoxR regulon is intimately linked with Actinorhodin production. J Bacteriol. 2010;192(24):6428–6438. | ||
Hwang KS, Kim HU, Charusanti P, Palsson BT, Lee SY. Systems biology and biotechnology of Streptomyces species for the production of secondary metabolites. Biotechnol Adv. 2014;32(2):255–268. | ||
Zonooz NF, Salouti M. Biological synthesis of gold nanoparticles by cell free supernatant of Streptomyces sp. ERI-3. Asian J Chem. 2012;24(10):4647–4649. | ||
Thenmozhi M, Kannabiran K, Kumar R, Gopiesh Khanna V. Antifungal activity of Streptomyces sp. VITSTK7 and its synthesized Ag2O/Ag nanoparticles against medically important Aspergillus pathogens. J Mycol Med. 2013;23(2):97–103. | ||
Subashini J, Gopiesh Khanna V, Kannabiran K. Anti-ESBL activity of silver nanoparticles biosynthesized using soil Streptomyces species. Bioprocess Biosyst Eng. 2014;37(6):999–1006. | ||
Sadhasivam S, Shanmugam P, Yun K. Biosynthesis of silver nanoparticles by Streptomyces hygroscopicus and antimicrobial activity against medically important pathogenic microorganisms. Colloids Surf B Biointerfaces. 2010;81(1):358–362. | ||
Srinivasan MC, Laxman RS, Deshpande MV. Physiology and nutritional aspects of actinomycetes: an overview. World J Microbiol Biotechnol. 1991;7(2):171–184. | ||
Cochrane VW. Physiology of actinomycetes. Annu Rev Microbiol. 1961;15(1):1–24. | ||
Pridham TG. Color and Streptomycetes: report of an international workshop on determination of color of streptomycetes. Appl Microbiol. 1965;13(1):43–61. | ||
Shirling EB, Gottlieb D. Methods for characterization of Streptomyces species. Int J Syst Bacteriol. 1966;16(3):313–340. | ||
Pridham TG, Gottlieb D. The utilization of carbon compounds by some Actinomycetales as an aid for species determination. J Bacteriol. 1948;56(1):107–114. | ||
Gorajana A, Venkatesan M, Vinjamuri S, et al. Resistoflavine, cytotoxic compound from a marine actinomycete, Streptomyces chibaensis AUBN1/7. Microbiol Res. 2007;162(4):322–327. | ||
Luedemann GM, Brodsky BC. Taxonomy of gentamicin-producing micromonospora. Antimicrob Agents Chemother (Bethesda). 1963;161:116–124. | ||
Bibb M, Freeman R, Hopwood D. Physical and genetical characterisation of a second sex factor, SCP2, for Streptomyces coelicolor A3(2). Mol Gen Genet. 1977;154(2):155–166. | ||
Altschul SF, Madden TL, Schäffer AA, et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997;25(17):3389–3402. | ||
Thompson JD, Higgins DG, Gibson TJ. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994;22(22):4673–4680. | ||
Kimura M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol. 1980;16(2):111–120. | ||
Mosmann T. Rapid colorimetric assay for cellular growth and survival: application to proliferation and cytotoxicity assays. J Immunol Methods. 1983;65(1–2):55–63. | ||
Ohnishi Y, Ishikawa J, Hara H, et al. Genome sequence of the streptomycin-producing microorganism Streptomyces griseus IFO 13350. J Bacteriol. 2008;190(11):4050–4060. | ||
de Lima Procópio RE, da Silva IR, Martins MK, de Azevedo JL, de Araújo JM. Antibiotics produced by Streptomyces. Braz J Infect Dis. 2012;16(5):466–471. | ||
Aigle B, Lautru S, Spiteller D, et al. Genome mining of Streptomyces ambofaciens. J Ind Microbiol Biotechnol. 2014;41(2):251–263. | ||
Van Wezel GP, McDowall KJ. The regulation of the secondary metabolism of Streptomyces: new links and experimental advances. Nat Prod Rep. 2011;28(7):1311–1333. | ||
Manivasagan P, Venkatesan J, Kang KH, Sivakumar K, Park SJ, Kim SK. Production of α-amylase for the biosynthesis of gold nanoparticles using Streptomyces sp. MBRC-82. Int J Biol Macromol. 2015;72:71–78. | ||
Soni N, Prakash S. Antimicrobial and mosquitocidal activity of microbial synthesized silver nanoparticles. Parasitol Res. 2014;114(3):1023–1030. | ||
Manikprabhu D, Lingappa K. Synthesis of silver nanoparticles using the Streptomyces coelicolor klmp33 pigment: an antimicrobial agent against extended-spectrum beta-lactamase (ESBL) producing. Escherichia coli. Mater Sci Eng C Mater Biol Appl. 2014;45:434–437. | ||
Sanjenbam P, Gopal JV, Kannabiran K. Anticandidal activity of silver nanoparticles synthesized using Streptomyces sp.VITPK1. J Mycol Med. 2014. | ||
Karthik L, Kumar G, Kirthi AV, Rahuman AA, Bhaskara Rao KV. Streptomyces sp. LK3 mediated synthesis of silver nanoparticles and its biomedical application. Bioprocess Biosyst Eng. 2014;37(2):261–267. | ||
El-Naggar NEA, Abdelwahed NA, Darwesh OM. Fabrication of biogenic antimicrobial silver nanoparticles by Streptomyces aegyptia NEAE 102 as eco-friendly nanofactory. J Microbiol Biotechnol. 2014;24(4):453–464. | ||
Sivalingam P, Antony JJ, Siva D, Achiraman S, Anbarasu K. Mangrove Streptomyces sp. BDUKAS10 as nanofactory for fabrication of bactericidal silver nanoparticles. Colloids Surf B Biointerfaces. 2012;98:12–17. | ||
Mary Jacob J, Balakrishnan RM, Kumar UB. Biosynthesis of lead selenide quantum rods in marine Aspergillus terreus. Mater Lett. 2014;124(0):279–281. | ||
Korbekandi H, Ashari Z, Iravani S, Abbasi S. Optimization of biological synthesis of silver nanoparticles using Fusarium oxysporum. Iran J Pharm Res. 2013;12(3):289–298. | ||
Dhanjal S, Cameotra SS. Aerobic biogenesis of selenium nanospheres by Bacillus cereus isolated from coalmine soil. Microb Cell Fact. 2010;9:52. | ||
Jeong U, Xia Y. Synthesis and crystallization of monodisperse spherical colloids of amorphous selenium. Adv Mater Deerfield. 2005;17(1):102–106. | ||
Barth A. Infrared spectroscopy of proteins. Biochim Biophys Acta. 2007;1767(9):1073–1101. | ||
Saravana Kumar P, Balachandran C, Duraipandiyan V, Ramasamy D, Ignacimuthu S, Al-Dhabi N. Extracellular biosynthesis of silver nanoparticle using Streptomyces sp. 09 PBT 005 and its antibacterial and cytotoxic properties. Appl Nanosci. 2015;5(2):169–180. | ||
Mandal S, Phadtare S, Sastry M. Interfacing biology with nanoparticles. Curr Appl Phys. 2005;5(2):118–127. | ||
Estevez H, Garcia-Lidon JC, Luque-Garcia JL, Camara C. Effects of chitosan-stabilized selenium nanoparticles on cell proliferation, apoptosis and cell cycle pattern in HepG2 cells: comparison with other selenospecies. Colloids Surf B Biointerfaces. 2014;122:184–193. | ||
Pi J, Yang F, Jin H, et al. Selenium nanoparticles induced membrane bio-mechanical property changes in MCF-7 cells by disturbing membrane molecules and F-actin. Bioorg Med Chem Lett. 2013;23(23):6296–6303. | ||
Tan L, Jia X, Jiang X, et al. In vitro study on the individual and synergistic cytotoxicity of adriamycin and selenium nanoparticles against Bel7402 cells with a quartz crystal microbalance. Biosens Bioelectron. 2009;24(7):2268–2272. | ||
Ramamurthy CH, Sampath KS, Arunkumar P, et al. Green synthesis and characterization of selenium nanoparticles and its augmented cytotoxicity with doxorubicin on cancer cells. Bioprocess Biosyst Eng. 2013;36(8):1131–1139. | ||
Prasad KS, Selvaraj K. Biogenic synthesis of selenium nanoparticles and their effect on As(III)-induced toxicity on human lymphocytes. Biol Trace Elem Res. 2014;157(3):275–283. | ||
Hadi SM, Bhat SH, Azmi AS, Hanif S, Shamim U, Ullah MF. Oxidative breakage of cellular DNA by plant polyphenols: a putative mechanism for anticancer properties. Semin Cancer Biol. 2007;17(5):370–376. | ||
Azmi AS, Bhat SH, Hanif S, Hadi SM. Plant polyphenols mobilize endogenous copper in human peripheral lymphocytes leading to oxidative DNA breakage: a putative mechanism for anticancer properties. FEBS Lett. 2006;580(2):533–538. |
 © 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms.php
and incorporate the Creative Commons Attribution
- Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms.php
and incorporate the Creative Commons Attribution
- Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.