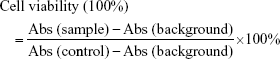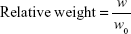Back to Journals » International Journal of Nanomedicine » Volume 13
Anti-EGFR-iRGD recombinant protein modified biomimetic nanoparticles loaded with gambogic acid to enhance targeting and antitumor ability in colorectal cancer treatment
Authors Zhang Z, Qian H , Huang J, Sha H, Zhang H, Yu L, Liu B, Hua D, Qian XP
Received 4 April 2018
Accepted for publication 25 May 2018
Published 31 August 2018 Volume 2018:13 Pages 4961—4975
DOI https://doi.org/10.2147/IJN.S170148
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Linlin Sun
Zhen Zhang,1,2 Hanqing Qian,1 Jie Huang,1 Huizi Sha,1 Hang Zhang,1 Lixia Yu,1 Baorui Liu,1 Dong Hua,3 Xiaoping Qian1
1Comprehensive Cancer Center, Nanjing Drum Tower Hospital, Clinical Cancer Institute of Nanjing University, Nanjing, Jiangsu, People’s Republic of China; 2Department of Integrated Traditional Chinese Medicine and Western Medicine Oncology, Affiliated Hospital of Jiangnan University, Wuxi, Jiangsu, People’s Republic of China; 3Department of Medical Oncology, Affiliated Hospital of Jiangnan University, Wuxi, Jiangsu, People’s Republic of China
Background: Red blood cell membrane-coated nanoparticle (RBCm-NP) platform, which consist of natural RBCm and synthetic polymeric core, can extend circulation time in vivo with an improved biocompatibility and stability of this biomimetic nanocarrier. To achieve better bioavailability of antitumor drugs that were loaded in RBCm-NPs, the functionalization of coated RBCm with specific targeting ability is essential. Bispecific recombinant protein anti-EGFR-iRGD, containing both tumor penetrating peptide (internalizing RGD peptide) and EGFR single-domain antibody (sdAb), seems to be an optimal targeting ligand for RBCm-NPs in the treatment of multiple tumors, especially colorectal cancer with high EGFR expression.
Materials and methods: We modified the anti-EGFR-iRGD recombinant protein on the surface of RBCm-NPs by lipid insertion method to construct iE–RBCm–PLGA NPs and confirmed the presentation of active tumor-targeting ability in colorectal cancer models with high EGFR expression when compared with RBCm–PLGA NPs. In addition, potential anti- tumor drug gambogic acid (GA) was loaded into the NPs to endow the antitumor efficiency of iE–RBCm–GA/PLGA NPs. It was simultaneously evaluated whether GA can reach better biocompatibility benefiting from the improved antitumor efficiency of iE–RBCm–GA/PLGA NPs in colorectal cancer models.
Results: We successfully modified anti-EGFR-iRGD proteins on the surface of biomimetic NPs with integrated and stable “shell–core” structure. iE–RBCm–PLGA NPs showed its improved targeting ability in vitro (multicellular spheroids [MCS]) and in vivo (nude mice bearing tumors). Besides, no matter on short-term cell apoptosis at tumor site (terminal deoxyribonucleotidyl transferase-mediated dUTP nick end labeling [TUNEL]) and long-term tumor inhibition, iE–RBCm–GA/PLGA NPs achieved better antitumor efficacy than free GA in spite of the similar effects of cytotoxicity and apoptosis to GA in vitro.
Conclusion: We expect that the bispecific biomimetic nanocarrier can extend the clinical application of many other potential antitumor drugs similar to GA and become a novel drug carrier in the colorectal cancer treatment.
Keywords: biomimetic nanocarrier, anti-EGFR-iRGD recombinant protein, gambogic acid, targeting ability, antitumor efficiency, colorectal cancer
Introduction
Nanoparticle (NP)-based drug delivery platforms are often used for various drugs to achieve good bioavailability in complex biological environments.1–3 With the development of multi-type prepared materials for NPs, it is not limited to the biocompatible and biodegradable synthetic materials,4,5 such as polylactic acid (PLA), poly(lactic-co-glycolic acid) (PLGA),6 polycaprolactone (PCL), and polyethylene glycol (PEG). Natural liposome7 cell membrane with inimitable structure and function attracts more attention for another new material of cell membrane-camouflaged NPs.8–11 Different from conventional NPs, the biomimetic nanocarriers served as the new class of NPs integrated two distinct materials – synthetic copolymer with customizability and flexibility and natural biomembrane with functionality and complexity. Biomembrane-cloaked NPs can indicate immune evasion to prolong circulation time in vivo, and NPs are able to accumulate at tumor sites subsequently via enhanced permeability and retention (EPR) effect. Hence, the application of biomimetic nanocarriers will be of great importance in antitumor therapy.12–15
Many kinds of cell membranes have been used to build the biomimetic NPs, and red blood cell membrane (RBCm) was the earliest and most widely explored,16–24 benefiting from the enormous quantity in vivo as well as simple structure and properties. However, compared with passive target through EPR effect, active target can optimize the bioavailability of antitumor drugs and minimize off-target side effects in vivo. The surface of RBCm25 lacks specific proteins or molecules that can induce active targeting ability. Tumor-targeting ligands can make it possible by modified on the surface of RBCm-coated polymeric nanoparticles (RBCm-NPs). To retain as much biological components with immune evasion capabilities as possible on the RBCm, physical lipid insertion approach instead of chemical conjugation technique was selected to modify ligands onto the bilayered RBC lipid membranes26,27 instead of chemical conjugation technique28,29 was selected to modify ligands onto the bilayered RBC lipid membranes. Except modification of targeting ligands, other specific cell membranes30–32 such as cancer cell membranes have been used to achieve active targeting ability for NPs in recent research. Due to adhesion proteins on the surface, cancer cell membrane-coated polymeric NPs (CCNPs) seem to have self-targeting ability to homotypic tumor site in vivo.33–36 Nevertheless, process to capture cancer cells from blood circulation is complicated and the tumor heterogeneity between cancer cells is non-negligible as well. Therefore, available RBCm with active targeting ligands is likely to be a better option for biomimetic NPs in antitumor therapy.
Due to special microenvironment of solid tumor, which shows abnormal homeostatic regulation, hypoxic cancer cells, increased interstitial fluid pressure, and existence of extracellular matrix (ECM), antitumor drugs can accumulate preferentially at tumor site via targeting nanocarriers, but the poor penetration of drugs into solid tumor tissues may further prevent the antitumor efficiency. For this reason, the selection of targeting ligands is not only to ensure active targeting ability but also to promote drugs to penetrate into solid tumor tissues efficiently. As tumor penetrating and cell internalizing peptide, iRGD peptide37–39 with the sequence of CRGDK/RGPD/EC can target αvβ3/αvβ5 integrin receptor and neuropilin-1 (NRP-1)40,41 that highly expressed in tumor tissues. It has been reported that iRGD peptide promoted co-administered drugs to penetrate into extravascular tumor tissue by increasing vascular and tissue permeability.38,39,42–46 In addition, epidermal growth factor receptor (EGFR) is overexpressed in multiple human solid tumors including colorectal cancer and showed 27.3% high expression of colorectal cancer in the previous report.47 Thus, EGFR targeting-based therapy has always been of important significance for colorectal cancer treatment. Therefore, the combination of iRGD peptide and EGFR antibody seems to be desired for targeting ligands in colorectal cancer. In our previous research, we have constructed a new recombinant protein named anti-EGFR–iRGD,48,49 containing both iRGD peptide and EGFR single-domain antibody (sdAb).50–53 Besides, we also confirmed the dual targeting ability and improved synergistic antitumor effect with antitumor drugs for anti-EGFR-iRGD recombinant protein.48,49,54,55 Thus, it can be seen that anti-EGFR-iRGD recombinant protein is expected to an optimal targeting ligand for RBCm NPs to achieve better tumor-targeting ability and antitumor efficiency.
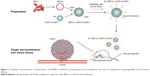
Gambogic acid (GA) is served as the major active ingredient of gamboge and can be potentially developed as a new antitumor drug.56–58 Recently, the incredible antitumor effect of GA on colorectal cancer has been reported in various researches.59,60 Therefore, GA is seemed to be an appropriate drug loaded in the bispecific targeting biomimetic NPs. Here, we lipidated anti-EGFR-iRGD proteins with palmitate chemically61,62 by detergent dialysis method63–68 and, then, the lipid chain of palmitate–anti-EGFR-iRGD proteins could insert into bilayered RBC lipid membranes to obtain RBCm NPs modified with anti-EGFR-iRGD proteins (iE–RBCm–PLGA NPs). We evaluated the targeting ability of iE–RBCm–PLGA NPs and antitumor effect of iE–RBCm–GA/PLGA NPs loaded with potential antitumor drug GA in colorectal cancer (Figure 1).
Materials and methods
Materials and reagents
GA (purity ≥99%), PLGA (molecular weight =40,000), N-N-succinimidyl palmitate, dichloromethane (DCM), and polyvinyl alcohol (PVA) were purchased from Aladdin Industrial Corporation (Shanghai, People’s Republic of China).
Anti-EGFR-iRGD recombinant proteins
Escherichia coli BL21 (DE3), which express recombinant proteins anti-EGFR-iRGD, was preserved by our laboratory.48,49 The E. coli massively proliferated when cultured in Lysogeny broth (LB) after resuscitation. Proteins anti-EGFR-iRGD were then expressed in E. coli BL21 (DE3) after induction by isopropyl β-D-1-thiogalactopyranoside (IPTG) at 37°C for 4 h. The cells were collected and suspended in 5 mM imidazole buffer solution and subsequently disrupted using probe sonicator (Sonics, Newtown, CT, USA). The supernatant of cell lysate was purified using the protein liquid chromatography system (ÄKTA™ Pure; GE Healthcare UK Ltd, Little Chalfont, UK), and the purified proteins were obtained after dialysis to remove midazole.
Cells and cell culture
Human colon cancer cell lines Caco-2 (ATCC® HTB37™), HT-29 (ATCC® HTB-38™), and SW480 (ATCC® CCL228™) were preserved in our laboratory and purchased from American Type Culture Collection (ATCC) (Manassas, VA, USA). The cells were cultured in Roswell Park Memorial Institute (RPMI-1640) medium (MultiCell, Calgary, AB, Canada) with 10% calf serum (Every Green, Hangzhou, People’s Republic of China) at 37°C in a humidified atmosphere with 5% CO2.
Animals
Experimental Animal Center of Nanjing Drum Tower Hospital supplied male BALB/c mice (nu/nu, weight: 18−25 g, age: 5–8 weeks). All animal-handling procedures were performed in compliance with guidelines set by the Animal Care Committee, Nanjing Drum Tower Hospital. All animal experiments were approved by the Animal Care Committee, Drum Tower Hospital, Nanjing, People’s Republic of China.
Preparation of iE–RBCm–GA/PLGA NPs
Preparation of RBCm–GA/PLGA NPs
The membranes of RBCs were collected in low osmotic pressure from broken RBCs, which separated from whole blood of BALB/c mice. Then, RBCm vesicles were prepared using sonication approach. Blank PLGA and GA-loaded PLGA NPs’ core were prepared using O/W emulsion method with the optimal ratio of GA and PLGA in weight. The fusion of RBCm-derived vesicles with GA/PLGA NPs’ core was conducted via mini-extruder (Avanti Polar Lipids, Alabaster, AL, USA). All the preparation of RBCm–GA/PLGA NPs was described in detail in our previous report.60
Modification of proteins anti-EGFR-iRGD on the surface of RBCm–GA/PLGA NPs
Recombinant proteins anti-EGFR-iRGD were dissolved at 1.5 mg/mL in phosphate-buffered saline (1× PBS), containing 0.3% sodium deoxycholate (DOC), 0.1% sodium azide, and 0.1% sodium bicarbonate. N-succinimidyl palmitate was added to the protein solution, which was prewarmed to 37°C to yield a final concentration of 0.1 mg/mL. This mixture was then incubated at room temperature for 12–18 hours. The lipid-derivatized proteins (palmitate–anti-EGFR-iRGD proteins) were purified on a 10 mL sephadex G-25 column equilibrated with PBS containing 0.1% DOC.
The resulted palmitate–anti-EGFR-iRGD proteins were incubated with RBCm–GA/PLGA NPs at 37°C for >30 min and centrifuged to remove excess proteins. Finally, the targeting moieties (anti-EGFR-iRGD proteins) were incorporated into RBCm through the aid of lipid tethers to build iE–RBCm–GA/PLGA NPs.
Characterization of iE–RBCm–GA/PLGA NPs
To confirm that the recombinant proteins anti-EGFR-iRGD were modified on the surface of RBCm–GA/PLGA NPs successfully, the chemical states of N of iE–RBCm–GA/PLGA NPs and RBCm–GA/PLGA NPs were characterized by X-ray photoelectron spectroscopy (XPS; Kratos XSAM 800; Kratos Analytical, Manchester, UK). The samples were frozen dried before examination.
The composite structure of the iE–RBCm–GA/PLGA NPs was examined using a transmission electron microscopy (TEM; JEM-100S, Tokyo, Japan). A drop of the NPs suspension was deposited onto the cupper grid and, then, air-dried at room temperature. The sample of iE–RBCm–GA/PLGA NPs was stained with 1% uranyl acetate before observation.
To measure the diameter and polydispersity of iE–RBCm–GA/PLGA NPs, the dynamic light scattering (DLS) method was conducted by using the Brookhaven BI-900AT instrument/zeta potential analyzer (Brookhaven Instruments Corporation, Holtsville, NY, USA). To evaluate the stability of the NPs, the iE–RBCm–GA/PLGA NPs in 1× PBS buffer (pH =7.4) were kept at 25°C and the diameter was determined by DLS everyday for 7 days.
To study drug release, 1 mL of iE–RBCm–GA/PLGA NPs in dialysis bag with a molecular weight cutoff of 10 kDa was immersed in 10 mL of 1× PBS buffer (pH =7.4). And the dialyzing sample was kept in thermostated shaking water bath (Taichang Medical Apparatus Co., Jiangsu, People’s Republic of China) at 37°C for 14 days. Every day, the release buffer was removed for analysis, and 10 mL of fresh PBS buffer was added for continuing incubation. Then, the GA content in the release buffer was detected using a HPLC system (Agilent Technologies, Santa Clara, CA, USA).
Targeting ability in colorectal cancer
EGFR expression levels of colorectal cancer cell lines
Western blot was used to confirm EGFR expression levels of colorectal cancer cell lines (Caco-2, HT-29, and SW480). The proteins of three cell lines were collected from each supernatant fraction after lysed on ice for 10 min. Samples containing equivalent amounts of proteins were then subjected to sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and blotted onto nitrocellulose membranes (EMD Millipore, Billerica, MA, USA). Membranes were then blocked with 5% skim milk and subsequently incubated with the EGFR-specific polyclonal antibody (Cell Signaling Technology, Danvers, MA, USA) in Tris-Buffered Saline and Tween 20 (Beyotime, Haimen, People’s Republic of China) overnight at 4°C. After several washes with TBST buffer, the membranes were incubated for 1 h with secondary antibody diluted 1:5,000. The membranes were then processed with enhanced chemiluminescence (ECL) western blot detection reagents. Signals were digitally captured using a MicroChemi chemiluminescent image analysis system (DNR Bio-imaging Systems, Jerusalem, Israel).
Formation of HT-29 MCS, uptake, and penetration in MCS
To form multicellular spheroids (MCS), colorectal cancer cells HT-29 were seeded in 96-well ultra-low attachment plates (Corning Incorporated, Corning, NY, USA) at a density of 2,000 cells/well and incubated at 37°C with 5% CO2 for 3–4 days (MCS is ~200 μm in diameter).
PLGA NPs were fluorescence labeled with green 3,3′-dioctadecyloxacarbocyanine perchlorate (DiO; Beyotime) dye or red 1,1′-dioctadecyl-3,3,3′,3′-tetramethylindocarbocyanine perchlorate (DiI; Beyotime). The fluorescent PLGA NPs were coated with RBCm and modified with proteins anti-EGFR-iRGD. NPs were added into the cell culture medium for 4 h of incubation. Then, the HT-29 MCS were collected, washed with 1× PBS buffer, and placed in confocal dishes (In Vitro Scientific, Gerasdorf bei Wien Austria). The cellular uptake of NPs and penetration in MCS were observed using inverted fluorescence microscope (Olympus Corporation, Tokyo, Japan) and confocal laser scanning microscopy (CLSM; Olympus Corporation).
Target and penetration into tumor tissues
A total of 1×107 Caco-2 cells were implanted subcutaneously into the right flank of BALB/c mice. The PLGA NPs were fluorescence labeled with green DiO dye and divided into four equal groups. The four groups acted as PLGA NPs (P group), RBCm–PLGA NPs (RP group), RBCm–PLGA NPs with free anti-EGFR-iRGD proteins (iE + RP group), and RBCm–PLGA NPs modified with anti-EGFR-iRGD proteins (iE–RP group). The samples were injected to Caco-2 tumor-bearing nude mice via the tail vein. Tumor-bearing nude mice were sacrificed after 24 h, and subcutaneous tumors were collected. Immunofluorescence analysis was carried out with frozen tumor tissues. For immunostaining, slides were first blocked with 3% bull serum albumin (BSA) for 1 h at room temperature and then exposed to the primary antibody against CD31 (1:300 dilution; GeneTech, Shanghai, People’s Republic of China) at 4°C overnight and the second antibody (1:300 dilution) at 37°C for 30 min. Finally, nuclei were stained with 2-(4-amidinophenyl)-6-indolecarbamidine dihydrochloride (DAPI; Beyotime) and slides were visualized using CLSM.
Fluorescence images of real-time tumor targeting in vivo
To investigate the tumor targeting and accumulation of NPs in vivo, 1,1′-dioctadecyl-3,3,3′,3′-tetramethylindotricarbocyanine iodide (DiR; Bridgen, Beijing, People’s Republic of China) dye was used to label the different NPs as abovementioned. The four different groups (P group, RP group, iE + RP group, and iE–RP group) were injected to Caco-2 tumor-bearing nude mice via the tail vein. At different time points (24, 72, and 168 h), the mice were anesthetized and imaged using the Maestro EX in vivo fluorescence imaging system (CRi, Inc., Hopkinton, MA, USA). At the end of test, the tumor-bearing mice were sacrificed to collect tumors and main organs including heart, liver, spleen, lungs, and kidneys for images of biodistribution in vivo.
Antitumor efficiency of iE–RBCm–GA/PLGA NPs in colorectal cancer
Cytotoxicity
MTT assay was carried out to evaluate the cytotoxity of iE–RBCm–GA/PLGA NPs against colorectal cancer cells HT-29 in vitro. Cells were seeded in 96-well plates at the density of 5,000 cells/well and then incubated at 37°C with 5% CO2 for attachment. Each well was added with 20 μL of 5 mg/mL MTT solution after cells were exposed to various concentrations of NPs and free GA for 12, 24, and 48 h and then incubated for another 4 h. The supernatant was discarded, and 150 μL of dimethylsulphoxide (DMSO) was added to each well to dissolve the formazan crystals. The absorption at 490 nm was measured by Multiskan Spectrum Microplate Reader (BioTek, Winooski, VT, USA), and the cell viabilities were calculated by the following formula:
|
Apoptosis assay in vitro and in vivo
Apoptosis in vitro was detected using Annexin-V-FLUOS Staining Kit (Hoffman-La Roche Ltd., Basel, Switzerland) according to the instruction. A total of 1×106 HT-29 cells were exposed to certain various concentrations of iE–RBCm–GA/PLGA NPs and free GA for 12, 24, 48, and 72 h, and then, cells in each wall were collected and washed with 1× PBS buffer. Washed cells were resuspended in 100 μL of Annexin-V-FLUOS-labeling solution containing 100 μL of 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid; PI: propidium iodide (HEPES) buffer, 2 μL of propidium iodide (PI), and 2 μL of Annexin-V-Fluorescein, and then, samples were incubated in the dark for 15 min at room temperature. The apoptosis of HT-29 was analyzed on flow cytometer (EMD Millipore).
For in vivo experiment, iE–RBCm–GA/PLGA NPs (iE–RGP group), RBCm–GA/PLGA NPs (RGP group), GA, iE–RBCm–PLGA NPs (iE–RP group), and normal saline (NS) were injected to Caco-2 tumor-bearing nude mice via tail vein. After 48 h, mice were sacrificed. Subcutaneous tumors were collected and fixed with paraformaldehyde. Terminal deoxyribonucleotidyl transferase-mediated dUTP nick end labeling (TUNEL; Hoffman-La Roche Ltd.) method was used to detect the apoptotic cells of tumor tissues in vivo. Then, nuclei were stained with DAPI, and slides were visualized using CLSM.
In vivo antitumor effect and safety
Caco-2 tumor-bearing nude mice were randomly divided into five groups (n=6 per group) and were injected with iE–RBCm–GA/PLGA NPs (6 mg/kg, iE–RGP group), RBCm–GA/PLGA NPs (6 mg/kg, RGP group), GA solution (6 mg/kg, GA group), iE–RBCm–PLGA NPs (iE–RP group), and normal saline (NS group) via the tail vein once a week, a total of four times. The tumor size and weight of mice were measured on day 7 postimplantation and every other day after injection. Tumor volume was calculated as V=D×d2/2, where D and d are the longest and the shortest diameters of the tumor in millimeters, respectively. Survival was predefined as tumor size <2,000 mm3 prior to the initiation of the study. In order to reduce the impact of initial weight differences among different groups, relative weight was calculated by the following formula:
|
w represents the absolute weight, and w0 represents the average weight of the group on day 7.
When mice were sacrificed, the tumor tissues and main organs including hearts, lungs, livers, spleens, and kidneys of mice in each group were collected for histology observation. The Ki67 immunohistochemistry was used to evaluate the proliferation of the cells in the tumors. And systemic toxicity was assessed via organ slices that stained by hematoxylin and eosin (H&E).
Statistical analysis
Statistical analysis was performed by the SPSS V.23.0 software (IBM Corporation, Armonk, NY, USA). One-way analysis of variance (ANOVA) or Independent Student’s t-test was used to determine the differences between various treatments. Data are presented as the mean ± SD. Statistically significant results were subjected to Tukey’s post hoc test (GraphPad Prism 6.0; GraphPad Software, Inc., La Jolla, CA, USA). All quantitative data reported represent the mean of at least three independent experiments. The resultant P-value was expressed as ***P≤0.001, **P≤0.01, and *P<0.05. Differences were considered to be statistically significant if P-value was <0.05.
Results
Preparation and physicochemical characterization of iE–RBCm–GA/PLGA NPs
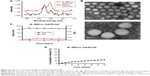
The lipid terminus of palmitate–anti-EGFR-iRGD proteins can be inserted into the coating RBCm of GA/PLGA NPs to modify the bispecific targeting. To confirm that the proteins were modified on the surface of NPs, the surface chemistry of the iE–RBCm–GA/PLGA NPs and RBCm–GA/PLGA NPs was characterized using XPS and the spectra of nitrogen (N) were depicted in Figure 2A. Compared with RBCm–GA/PLGA NPs, the peak of elemental N of iE–RBCm–GA/PLGA NPs moved upward obviously,69 demonstrating that the proteins were modified on the surface of NPs successfully.
The structure of the iE–RBCm–GA/PLGA NPs with uranyl acetate negative staining was visualized by TEM and the shell–core structure was revealed in the image (Figure 2B). The hydrodynamic diameter of iE–RBCm–GA/PLGA NPs was about 153±3.83 nm, and little change in the particle size and polydispersity index (PDI) of iE–RBCm–GA/PLGA NPs could be observed in a week (Figure 2C). The NPs suspended in 1× PBS buffer (pH =7.4) showed the long-term stability in vitro. Besides, the iE–RBCm–GA/PLGA NPs also displayed a slow and sustained release in 14 days (Figure 2D). Approximately 30% of GA was released from iE–RBCm–GA/PLGA NPs in neutral PBS buffer (pH =7.4) over 14 days.
Targeting ability in colorectal cancer
To select suitable cell line, the EGFR expression of three different colorectal cancer cells (Caco-2, HT-29, and SW480) was measured. The result showed that EGFR was highly expressed in Caco-2 and HT-29 (Figure 3A). And compared with Caco-2 and HT-29, the EGFR levels in SW480 cells could not be detected. Besides, due to that cells HT-29 could form MCS with symmetrical and spherical shapes and cells Caco-2 could be implanted to form subcutaneous tumors, HT-29 was used in following experiments in vitro, while Caco-2 was used in in vivo.
The penetration ability of the three groups of NPs (iE–RP, iE + RP, and RP) were analyzed with MCS. The result showed that green fluorescence intensity of iE–RP group was higher than the other two groups under the fluorescence microscope (Figure 3B), which reflected that more iE–RBCm–PLGA NPs were taken up by MCS. We also further evaluated the penetration of the three groups of NPs in MCS under CLSM. As shown in Figure 3C, the fluorescence signal of iE–RP group was localized in both interior and periphery of MCS, while RP group in the vicinity of periphery. The image of iE + RP group also showed a little increased penetration when compared with RP group. It may benefit from tumor-penetrating peptide iRGD, allowing co-administered NPs to penetrate into MCS. However, the penetration of iE–RBCm–PLGA NPs was deeper than RBCm–PLGA NPs without anti-EGFR-iRGD proteins modified on the surface. Therefore, RBCm–PLGA NPs modified with anti-EGFR-iRGD proteins on the surface could obtain positive penetration ability, while the increased penetration ability from free anti-EGFR-iRGD proteins was limited.
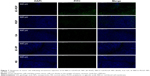
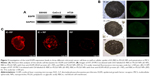
Based on the result that iE–RBCm–PLGA NPs could get deep penetration into MCS in vitro, target and penetration into tumor tissues in vivo were also observed and evaluated by both immunostaining tumor tissue slices and real-time near-infrared (NIR) fluorescence imaging. In the image of tumor tissue slice (Figure 4), green fluorescence signal represents NPs visualized with DiO dye, red fluorescence signal represents blood vessels of tumor issue, and blue fluorescence signal represents tumor cells. Except P group, green fluorescent NPs could be seen in the images of other three groups (iE–RP, iE + RP, and RP). Besides, considerable amounts of green fluorescence NPs were around blood vessels and tumor cells of tumor issue in the iE–RP group, which was much more than that in the iE + RP group and the RP group. It meant that iE–RBCm–PLGA NPs could accumulate efficiently at tumor site with active targeting ability. In addition, a certain number of RBCm–PLGA NPs accumulated at tumor site benefiting from acceptable biocompatibility and EPR effect. Only free anti-EGFR-iRGD proteins instead of being modified on the surface could not obtain obvious active targeting delivery function for RBCm–PLGA NPs.
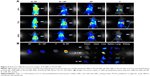
The biodistribution of four groups NPs (iE–RP, iE + RP, RP, and P) were monitored via real-time NIR fluorescence imaging (Figure 5A). At 24 hours postinjection, different fluorescence signals in intensity appeared at the tumor site of iE–RP, iE + RP, and RP groups except P group and iE–RP group showed the strongest fluorescence signal among them. The result demonstrated that iE–RBCm–PLGA NPs have enhanced tumor-targeting ability than RBCm–PLGA NPs in vivo. At 72 h postinjection, the intensity of the fluorescent signal continued to increase, which reflected the “long circulating” advantage of biomimetic nanocarriers. During the process, the fluorescence intensity of iE + RP and RP were similar and weaker than that of iE–RP group. It meant that free anti-EGFR-iRGD proteins cannot guide RBCm–PLGA NPs to accumulate efficiently at tumor site. At 168 h postinjection, the fluorescence intensity at tumor site decreased compared with the previous time point. Then, the mice of all groups were sacrificed, and ex vivo fluorescence intensity images were obtained for tumor tissues as well as other major organs including hearts, lungs, livers, spleens, and kidneys. As shown in Figure 5B, the fluorescence signal intensity of subcutaneous tumor in iE–RP group was significantly higher than the rest of the three groups. Moreover, compared with the high intensity of fluorescence signal of liver and spleen, the fluorescence signal of subcutaneous tumor in the iE–RP group was also visible. This matched with above results in vivo.
Antitumor efficiency in colorectal cancer
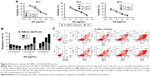
MTT assay was used to evaluate the cytotoxicity of iE–RBCm–GA/PLGA NPs against HT-29 cells in vitro. For 12, 24, and 48 h incubation, both iE–RBCm–GA/PLGA NPs and GA showed a dose-dependent cytotoxicity against HT-29 cells (Figure 6A). In addition, compared with respective cell viability, iE–RBCm–GA/PLGA NPs and free GA almost had no difference against HT-29 at all the time points.
To illustrate the possible mechanism of cytotoxicity in vitro, the further analysis of apoptosis that may lead to cytotoxicity was also conducted. When HT-29 cells were exposed to different concentrations of iE–RBCm–GA/PLGA NPs from 12 to 72 h, obvious increase in apoptotic rate (Q2 + Q4) was observed (Figure 6B). And when the concentration of iE–RBCm–GA/PLGA NPs and GA was >1.5 μg/mL, the cell apoptosis of HT-29 cells was apparent. Therefore, flow cytometric analysis was conducted to assess apoptosis induced by iE–RBCm–GA/PLGA NPs and GA at certain concentration (1.5 μg/mL) for 12, 24, 48, and 72 h (Figure 6C). The result reflected that the rate of both early (Q4) stage and late (Q2) stage apoptotic HT-29 cells were increasing with the time. At all the time points, both two groups showed similar apoptosis rate of HT-29 cells. It matched with the results of MTT assay and suggested that both iE–RBCm–GA/PLGA NPs and free GA have the same efficient antitumor effect in vitro.
In vivo, we sequentially investigated whether the antitumor activity of iE–RBCm–GA/PLGA NPs could act as effective therapy against tumor growth. BALB/c nude mice-bearing subcutaneous inoculated Caco-2 tumors, acted as colorectal cancer model, were used to observe both the apoptosis at tumor site and the long-term tumor burden in vivo. The treatments were administrated via tail vein with the solution of iE–RBCm–GA/PLGA NPs, RBCm–GA/PLGA NPs, GA, iE–RBCm–PLGA NPs, and NS. Apoptotic cells were visualized as green fluorescence signal, and all the tumor cells were visualized as blue fluorescence signal in the image of subcutaneous tumor tissue slice (Figure 7). The result showed a large amount of apoptotic cells and a small number of tumor cells in the visual field of slice in the iE–RGP group, which indicated that iE–RBCm–GA/PLGA NPs with active targeting ability accumulated and penetrated efficiently at tumor site within a short time to lead significant cell apoptosis in vivo. The presence of fluorescence signal in the RGP group suggested that RBCm–GA/PLGA NPs also could passively target and accumulate at tumor site to lead apoptosis due to the stability, biocompatibility and “long circulating” characteristic in vivo. However, it still was slightly inferior than iE–RBCm–GA/PLGA NPs. In addition, the green fluorescence signal that represents apoptotic cells was not obvious in the GA group compared with the control group (iE–RP and NS) within 48 h. It reflected that GA-loaded NPs achieved better bioavailability than free GA in vivo, and the NPs with active targeting ability could expand this advantage.
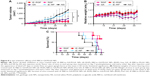
Antitumor efficacy of iE–RBCm–GA/PLGA NPs in vivo was evaluated via the long-term subcutaneous tumor burden in the colorectal cancer model of mice. As shown in Figure 8A, compared with the control group (iE–RP and NS), the tumor volumes of iE–RGP treatment group grew quite slowly (***P≤0.001, n=6), which showed an expected result of iE–RBCm–GA/PLGA NPs in terms of tumor growth inhibition. RGP treatment group also showed satisfactory antitumor efficacy at GA dose of 6 mg/kg, reflecting the advantage of biomimetic nanocarriers for tumor therapy. Moreover, the fact that the weights of mice in GA-loaded NPs groups were increasing gradually rather than remained as in control groups, indicating that mice of these groups benefited from antitumor efficiency and did not suffer from the increasing tumor burden (Figure 8B). GA group showed certain tumor inhibition effect, but the weight of mice in GA group was decreased, which indicated that mice suffered from the toxicity of free GA. It also reflected in the result of survival (Figure 8C); mice in control group with increasing tumor burden began to die first (from day 40). In contrast, mice in the iE–RGP group began to die after 38 days (from day 66) after the last administration and extended the median survival time at least 27 days more compared with the control group. Simultaneously, mice in the GA group died within 17 days (from days 64 to 80) due to cumulative toxicity during the treatment.
In Ki67 staining images, the nuclei of positive cells were stained brown, indicating high proliferation ability. The result showed that there were less positive cells in the tumor tissue of the iE–RGP group compared with the control group and so was of RGP group (Figure 9A). Moreover, H&E-stained section images of major organs in each group showed no noticeable signal of organ damage (Figure 9B). It suggested that iE–RBCm–GA/PLGA NPs had optimal antitumor efficiency and safety in vivo.
Discussion
On the basis of the RBC-NP platform with good stability and biocompatibility, the modification of anti-EGFR-iRGD recombinant protein with two target antigens endowed the active bispecific targeting ability for this synthetic biomimetic nanocarrier. The physical insertion into the RBCm avoided damaging the existing surface proteins and remained the integrality and stability of structure. Moreover, potential antitumor drug GA that loaded into iE–RBCm–PLGA NPs brought the synthetic nanocarrier antitumor efficacy. The resulting iE–RBCm–GA/PLGA NPs not only lifted restrictions on the application of GA with poor water solubility but also delivered GA to tumor site efficiently to improve its bioavailability in vivo.
The targeting ability of iE–RBCm–PLGA NPs and antitumor efficiency of GA loaded iE–RBCm–GA/PLGA NPs were taken as the object to investigate the effect on the colorectal cancer in vitro and in vivo. In terms of the targeting and penetrating efficiency of iE–RBCm–PLGA NPs, we evaluated it at the level of MCS and nude mice bearing subcutaneous inoculated tumors. Compared with RBCm–PLGA NPs, more iE–RBCm–PLGA NPs could be taken up by MCS and showed better penetration into MCS in vitro. iE–RBCm–PLGA NPs also showed better targeting efficiency in nude mice model of colorectal cancer with obvious accumulation at tumor site in a certain time. It indirectly reflected that anti-EGFR-iRGD protein successfully modified on the surface of NPs via lipid insertion, and iE–RBCm–PLGA NPs with the modification of anti-EGFR-iRGD protein achieved satisfactory targeting and penetrating efficiency in vitro and in vivo. Besides, RBCm–PLGA NPs with free anti-EGFR-iRGD protein showed a little advantage in targeting and penetrating ability when compared with RBCm–PLGA NPs, suggesting that the targeting and penetrating ability of free anti-EGFR-iRGD to guide more RBCm–PLGA NPs to tumor site were limited. In consideration of anti-EGFR-iRGD recombinant protein with dual targeting of αvβ3 and EGFR, we emphatically studied specific EGFR target in colorectal cancer in future work. Colorectal cancer models with the high expression of EGFR and low or no expression of EGFR were established to explore the target differentiation of iE–RBCm–PLGA NPs between different colorectal cancer models.
In terms of the antitumor efficiency of iE–RBCm–GA/PLGA NPs, we also evaluated it in vitro and in vivo, which contained cytotoxicity, apoptosis, and tumor inhibition. iE–RBCm–GA/PLGA NPs and free GA were taken up via different mechanisms – endocytosis and pure diffusion – and free GA could diffuse into the cancer cells more easily compared with NPs. However, iE–RBCm–GA/PLGA NPs modified with active targeting ligands enhanced the transport capacity across cellular membrane. As a result, iE–RBCm–GA/PLGA NPs showed similar cytotoxicity compared with free GA at the same dose in vitro. Simultaneously, the cell apoptosis of both iE–RBCm–GA/PLGA NPs and GA were almost the same in vitro, but the apoptosis of cancer cells at subcutaneous tumor site in vivo seemed to be different. Considering complex and changeable biological environment in vivo, free GA did not induce obvious cell apoptosis as shown in in vitro. Nevertheless, iE–RBCm–GA/PLGA NPs showed significant cell apoptosis within 48 h benefiting from the active targeting ability. Difference was also observed in subsequent experiment of tumor growth inhibition. Compared with control group, iE–RBCm-GA/PLGA NPs inhibited the growth of subcutaneous tumor efficiently with safety and nontoxicity. Although GA exerted certain effect on tumor inhibition, it accompanied with serious toxicity, which appeared as decreasing weight and death within a short time. In addition, RBCm–GA/PLGA NPs with good stability and biocompatibility achieved a significant inhibition of tumor growth at a dose of 6 mg/kg, indicating the superiority of biomimetic nanocarrier in cancer therapy. However, it did not objectively show that iE–RBCm-GA/PLGA NPs possess the ability of active targeting delivery. Subsequently, we repeated and adjusted this experiment with the dose of GA intended to be reduced accordingly to observe whether iE–RBCm–GA/PLGA NPs with active and efficient tumor targeting ability could improve the bioavailability of GA in vivo at a dose that is lower than effective dose for free GA.
Conclusion
In our study, recombinant protein anti-EGFR-iRGD was modified on the surface of RBCm-coated NPs to build bispecific targeting biomimetic NPs, combining biocompatibility, stability, and active target. In regard to targeting ability, when compared with RBCm–PLGA NPs, more iE–RBCm–PLGA NPs could be taken up by MCS and penetrate into MCS in vitro and more iE–RBCm–PLGA NPs targeted and accumulated at tumor site in vivo. It was confirmed that the modification of anti-EGFR-iRGD protein significantly improved the targeting ability of iE–RBCm–PLGA NPs. In regard to antitumor efficacy, despite the similar antitumor effect of iE–RBCm–GA/PLGA NPs and free GA that shown in vitro, the antitumor efficacy of GA weakened in vivo, along with considerable toxicity. In contrast, iE–RBCm–GA/PLGA NPs still retained antitumor efficiency via short-term cell apoptosis at tumor site and long-term tumor inhibition in vivo. Moreover, iE–RBCm–GA/PLGA NPs reduced the side effects of GA and ensured safety in vivo. Therefore, the bispecific targeting biomimetic NPs reinforce the advantage of drug delivery. We expected that the new nanocarrier could extend the clinical application of many other potential antitumor drugs similar to GA in the future.
Acknowledgment
This study was supported by grants from Provincial Natural Science Foundation of Jiangsu (no BK20161107), National Natural Science Foundation (no 81602106 and 81502037), Jiangsu Health and Family Planning Commission development program (no H2817042) and Nanjing Health and Family Planning Commission key program (no ZKX17012).
Disclosure
The authors report no conflicts of interest in this work.
References
Shi D, Bedford NM, Cho HS. Engineered multifunctional nanocarriers for cancer diagnosis and therapeutics. Small. 2011;7(18):2549–2567. | ||
Mahmoodi NO, Ghavidast A, Amirmahani N. A comparative study on the nanoparticles for improved drug delivery systems. J Photochem Photobiol B. 2016;162:681–693. | ||
Cheng Z, Al ZA, Hui JZ, Muzykantov VR, Tsourkas A. Multifunctional nanoparticles: cost versus benefit of adding targeting and imaging capabilities. Science. 2012;338(6109):903–910. | ||
Gunatillake PA, Adhikari R. Biodegradable synthetic polymers for tissue engineering. Eur Cell Mater. 2003;5:1–16; discussion 16. | ||
Aryal S, Hu CM, Zhang L. Polymeric nanoparticles with precise ratiometric control over drug loading for combination therapy. Mol Pharm. 2011;8(4):1401–1407. | ||
Acharya S, Sahoo SK. PLGA nanoparticles containing various anticancer agents and tumour delivery by EPR effect. Adv Drug Deliv Rev. 2011;63(3):170–183. | ||
Gao W, Hu CM, Fang RH, Zhang L. Liposome-like nanostructures for drug delivery. J Mater Chem B. 2013;1(48):1–8. | ||
Luk BT, Zhang L. Cell membrane-camouflaged nanoparticles for drug delivery. J Control Release. 2015;220(pt B):600–607. | ||
Fang RH, Luk BT, Hu CM, Zhang L. Engineered nanoparticles mimicking cell membranes for toxin neutralization. Adv Drug Deliv Rev. 2015;90:69–80. | ||
Zhang J, Gao W, Fang RH, Dong A, Zhang L. Synthesis of nanogels via cell membrane-templated polymerization. Small. 2015;11(34):4309–4313. | ||
Copp JA, Fang RH, Luk BT, et al. Clearance of pathological antibodies using biomimetic nanoparticles. Proc Natl Acad Sci U S A. 2014;111(37):13481–13486. | ||
Wicki A, Witzigmann D, Balasubramanian V, Huwyler J. Nanomedicine in cancer therapy: challenges, opportunities, and clinical applications. J Control Release. 2015;200:138–157. | ||
Fang RH, Kroll AV, Zhang L. Nanoparticle-based manipulation of antigen-presenting cells for cancer immunotherapy. Small. 2015;11(41):5483–5496. | ||
Voon SH, Kiew LV, Lee HB, et al. In vivo studies of nanostructure-based photosensitizers for photodynamic cancer therapy. Small. 2014;10(24):4993–5013. | ||
Minelli C, Lowe SB, Stevens MM. Engineering nanocomposite materials for cancer therapy. Small. 2010;6(21):2336–2357. | ||
Hu CM, Zhang L, Aryal S, Cheung C, Fang RH, Zhang L. Erythrocyte membrane-camouflaged polymeric nanoparticles as a biomimetic delivery platform. Proc Natl Acad Sci U S A. 2011;108(27):10980–10985. | ||
Aryal S, Hu CM, Fang RH, et al. Erythrocyte membrane-cloaked polymeric nanoparticles for controlled drug loading and release. Nanomedicine (Lond). 2013;8(8):1271–1280. | ||
Piao JG, Wang L, Gao F, You YZ, Xiong Y, Yang L. Erythrocyte membrane is an alternative coating to polyethylene glycol for prolonging the circulation lifetime of gold nanocages for photothermal therapy. ACS Nano. 2014;8(10):10414–10425. | ||
Guo Y, Wang D, Song Q, et al. Erythrocyte membrane-enveloped polymeric nanoparticles as nanovaccine for induction of antitumor immunity against melanoma. ACS Nano. 2015;9(7):6918–6933. | ||
Fang RH, Hu CM, Zhang L. Nanoparticles disguised as red blood cells to evade the immune system. Expert Opin Biol Ther. 2012;12(4):385–389. | ||
Rao L, Bu LL, Xu JH, et al. Red blood cell membrane as a biomimetic nanocoating for prolonged circulation time and reduced accelerated blood clearance. Small. 2015;11(46):6225–6236. | ||
Luk BT, Fang RH, Hu CM, et al. Safe and immunocompatible nanocarriers cloaked in RBC membranes for drug delivery to treat solid tumors. Theranostics. 2016;6(7):1004–1011. | ||
Rao L, Xu JH, Cai B, et al. Synthetic nanoparticles camouflaged with biomimetic erythrocyte membranes for reduced reticuloendothelial system uptake. Nanotechnology. 2016;27(8):085106. | ||
Luk BT, Hu CM, Fang RH, et al. Interfacial interactions between natural RBC membranes and synthetic polymeric nanoparticles. Nanoscale. 2014;6(5):2730–2737. | ||
Tsai RK, Rodriguez PL, Discher DE. Self inhibition of phagocytosis: the affinity of ‘marker of self’ CD47 for SIRPalpha dictates potency of inhibition but only at low expression levels. Blood Cells Mol Dis. 2010;45(1):67–74. | ||
Fang RH, Hu CM, Chen KN, et al. Lipid-insertion enables targeting functionalization of erythrocyte membrane-cloaked nanoparticles. Nanoscale. 2013;5(19):8884–8888. | ||
Fu Q, Lv P, Chen Z, et al. Programmed co-delivery of paclitaxel and doxorubicin boosted by camouflaging with erythrocyte membrane. Nanoscale. 2015;7(9):4020–4030. | ||
Zhou H, Fan Z, Lemons PK, Cheng H. A facile approach to functionalize cell membrane-coated nanoparticles. Theranostics. 2016;6(7):1012–1022. | ||
Bell GI. Models for the specific adhesion of cells to cells. Science. 1978;200(4342):618–627. | ||
Hu CM, Fang RH, Wang KC, et al. Nanoparticle biointerfacing by platelet membrane cloaking. Nature. 2015;526(7571):118–121. | ||
Dehaini D, Wei X, Fang RH, et al. Erythrocyte-platelet hybrid membrane coating for enhanced nanoparticle functionalization. Adv Mater. 2017;29(16):1606209. | ||
Parodi A, Quattrocchi N, van de Ven AL, et al. Synthetic nanoparticles functionalized with biomimetic leukocyte membranes possess cell-like functions. Nat Nanotechnol. 2013;8(1):61–68. | ||
Sun H, Su J, Meng Q, et al. Cancer-cell-biomimetic nanoparticles for targeted therapy of homotypic tumors. Adv Mater. 2016;28(43):9581–9588. | ||
Fang RH, Hu CM, Luk BT, et al. Cancer cell membrane-coated nanoparticles for anticancer vaccination and drug delivery. Nano Lett. 2014;14(4):2181–2188. | ||
Rao L, Bu LL, Cai B, et al. Cancer cell membrane-coated upconversion nanoprobes for highly specific tumor imaging. Adv Mater. 2016;28(18):3460–3466. | ||
Zhu JY, Zheng DW, Zhang MK, et al. Preferential cancer cell self-recognition and tumor self-targeting by coating nanoparticles with homotypic cancer cell membranes. Nano Lett. 2016;16:5895–5901. | ||
Sugahara KN, Teesalu T, Karmali PP, et al. Tissue-penetrating delivery of compounds and nanoparticles into tumors. Cancer Cell. 2009;16(6):510–520. | ||
Sugahara KN, Teesalu T, Karmali PP, et al. Coadministration of a tumor-penetrating peptide enhances the efficacy of cancer drugs. Science. 2010;328(5981):1031–1035. | ||
Mantis C, Kandela I, Aird F. Reproducibility Project: Cancer Biology. Replication study: coadministration of a tumor-penetrating peptide enhances the efficacy of cancer drugs. Elife. 2017;6:e17584. | ||
Teesalu T, Sugahara KN, Kotamraju VR, Ruoslahti E. C-end rule peptides mediate neuropilin-1-dependent cell, vascular, and tissue penetration. Proc Natl Acad Sci U S A. 2009;106(38):16157–16162. | ||
Haspel N, Zanuy D, Nussinov R, Teesalu T, Ruoslahti E, Aleman C. Binding of a C-end rule peptide to the neuropilin-1 receptor: a molecular modeling approach. Biochemistry. 2011;50(10):1755–1762. | ||
Chen R, Braun GB, Luo X, Sugahara KN, Teesalu T, Ruoslahti E. Application of a proapoptotic peptide to intratumorally spreading cancer therapy. Cancer Res. 2013;73(4):1352–1361. | ||
Zhang J, Wang L, Fai CH, et al. Co-delivery of paclitaxel and tetrandrine via iRGD peptide conjugated lipid-polymer hybrid nanoparticles overcome multidrug resistance in cancer cells. Sci Rep. 2017;7:46057. | ||
Gao F, Zhang J, Fu C, et al. iRGD-modified lipid-polymer hybrid nanoparticles loaded with isoliquiritigenin to enhance anti-breast cancer effect and tumor-targeting ability. Int J Nanomedicine. 2017;12:4147–4162. | ||
Huang Y, Li X, Sha H, et al. sTRAIL-iRGD is a promising therapeutic agent for gastric cancer treatment. Sci Rep. 2017;7(1):579. | ||
Zhu Z, Xie C, Liu Q, et al. The effect of hydrophilic chain length and iRGD on drug delivery from poly(ε-caprolactone)-poly(N-vinylpyrrolidone) nanoparticles. Biomaterials. 2011;32(35):9525–9535. | ||
Ceccarelli C, Piazzi G, Paterini P, et al. Concurrent EGFr and Cox-2 expression in colorectal cancer: proliferation impact and tumour spreading. Ann Oncol. 2005;16(suppl 4):iv74–iv79. | ||
Sha H, Li R, Bian X, et al. A tumor-penetrating recombinant protein anti-EGFR-iRGD enhance efficacy of paclitaxel in 3D multicellular spheroids and gastric cancer in vivo. Eur J Pharm Sci. 2015;77:60–72. | ||
Sha H, Zou Z, Xin K, et al. Tumor-penetrating peptide fused EGFR single-domain antibody enhances cancer drug penetration into 3D multicellular spheroids and facilitates effective gastric cancer therapy. J Control Release. 2015;200:188–200. | ||
Ward ES, Güssow D, Griffiths AD, Jones PT, Winter G. Binding activities of a repertoire of single immunoglobulin variable domains secreted from Escherichia coli. Nature. 1989;341(6242):544–546. | ||
Teillaud JL. From whole monoclonal antibodies to single domain antibodies: think small. Methods Mol Biol. 2012;911:3–13. | ||
Muyldermans S. Nanobodies: natural single-domain antibodies. Annu Rev Biochem. 2013;82:775–797. | ||
Harmsen MM, De Haard HJ. Properties, production, and applications of camelid single-domain antibody fragments. Appl Microbiol Biotechnol. 2007;77(1):13–22. | ||
Bian X, Wu P, Sha H, et al. Anti-EGFR-iRGD recombinant protein conjugated silk fibroin nanoparticles for enhanced tumor targeting and antitumor efficiency. Onco Targets Ther. 2016;9:3153–3162. | ||
Xin X, Sha H, Shen J, Zhang B, Zhu B, Liu B. Coupling Gd-DTPA with a bispecific, recombinant protein anti-EGFR-iRGD complex improves tumor targeting in MRI. Oncol Rep. 2016;35(6):3227–3235. | ||
Wang T, Wei J, Qian X, Ding Y, Yu L, Liu B. Gambogic acid, a potent inhibitor of survivin, reverses docetaxel resistance in gastric cancer cells. Cancer Lett. 2008;262(2):214–222. | ||
Wang LH, Li Y, Yang SN, et al. Gambogic acid synergistically potentiates cisplatin-induced apoptosis in non-small-cell lung cancer through suppressing NF-κB and MAPK/HO-1 signalling. Br J Cancer. 2014;110(2):341–352. | ||
Yin D, Yang Y, Cai H, Wang F, Peng D, He L. Gambogic acid-loaded electrosprayed particles for site-specific treatment of hepatocellular carcinoma. Mol Pharm. 2014;11(11):4107–4117. | ||
Wen C, Huang L, Chen J, et al. Gambogic acid inhibits growth, induces apoptosis, and overcomes drug resistance in human colorectal cancer cells. Int J Oncol. 2015;47(5):1663–1671. | ||
Zhang Z, Qian H, Yang M, et al. Gambogic acid-loaded biomimetic nanoparticles in colorectal cancer treatment. Int J Nanomedicine. 2017;12:1593–1605. | ||
Chou SH, Shetty AV, Geng Y, et al. Palmitate-derivatized human IL-2: a potential anticancer immunotherapeutic of low systemic toxicity. Cancer Immunol Immunother. 2013;62(3):597–603. | ||
Kim SA, Peacock JS. The use of palmitate-conjugated protein A for coating cells with artificial receptors which facilitate intercellular interactions. J Immunol Methods. 1993;158(1):57–65. | ||
Huang A, Huang L, Kennel SJ. Monoclonal antibody covalently coupled with fatty acid. A reagent for in vitro liposome targeting. J Biol Chem. 1980;255(17):8015–8018. | ||
Chen A, Zheng G, Tykocinski ML. Hierarchical costimulator thresholds for distinct immune responses: application of a novel two-step Fc fusion protein transfer method. J Immunol. 2000;164(2):705–711. | ||
Liu S, Breiter DR, Zheng G, Chen A. Enhanced antitumor responses elicited by combinatorial protein transfer of chemotactic and costimulatory molecules. J Immunol. 2007;178(5):3301–3306. | ||
Zheng G, Chen A, Sterner RE, et al. Induction of antitumor immunity via intratumoral tetra-costimulator protein transfer. Cancer Res. 2001;61(22):8127–8134. | ||
Liu S, Foster BA, Chen T, Zheng G, Chen A. Modifying dendritic cells via protein transfer for antitumor therapeutics. Clin Cancer Res. 2007;13(1):283–291. | ||
Chen A, Zheng G, Tykocinski ML. Quantitative interplay between activating and pro-apoptotic signals dictates T cell responses. Cell Immunol. 2003;221(2):128–137. | ||
Zhu J, Tang C, Kottke-Marchant K, Marchant RE. Design and synthesis of biomimetic hydrogel scaffolds with controlled organization of cyclic RGD peptides. Bioconjug Chem. 2009;20(2):333–339. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.