Back to Journals » Journal of Blood Medicine » Volume 10
Analysis of the reannealing- instead of melting-curve in the detection of JAK2 V617F mutation by HRM method
Authors Moradabadi A, Fatemi A, Noroozi-Aghideh A
Received 5 February 2019
Accepted for publication 10 July 2019
Published 22 July 2019 Volume 2019:10 Pages 235—241
DOI https://doi.org/10.2147/JBM.S204222
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Martin Bluth
Alireza Moradabadi,1 Ahmad Fatemi,2 Ali Noroozi-Aghideh1
1Department of Hematology, Faculty of Paramedicine, Aja University of Medical Sciences, Tehran, Iran; 2Department of Hematology, School of Allied Medical Sciences, Kerman University of Medical Sciences, Kerman, Iran
Background: The Janus kinase 2 (JAK2) has an important role in the intracellular signaling in normal and neoplastic cells. JAK2 mutation, called JAK2 V617F, is frequently found in Philadelphia chromosome-negative myeloproliferative neoplasms. We aimed to assess the analytical efficiency of high-resolution melting (HRM) method using reannealing-curve analysis in comparison with routine melting-curve analysis for JAK2 V617F mutation detection.
Method: Twenty-three samples including one negative synthetic standard DNA, two 50% and 75% positive synthetic standard DNA samples, five wild-type samples and 15 samples positive for JAK2 V617F were examined by HRM. Melting and reannealing stages were performed, and then, raw and normalized curves were compared between the two stages.
Results: In melting-curve analysis, the wild-type and mutant samples had different melting temperatures (75/53°C and 75/10°C, respectively). In normalized curves corresponding to reannealing method, mutant samples were better separated from the baseline than in melting method as well as for samples with different mutant DNA burden from each other. Furthermore, wild-type samples were more homogenous in the normalized curves corresponding to reannealing than in melting method. This means that patients with a low allelic burden may be wrongly interpreted as normal in the common melting method.
Conclusion: We suggest the use of reannealing instead of the melting-curve analysis for the detection of sequence variations, especially for large-scale mutation and allele burden measurement in clinical settings. However, more evaluations with more sample size will better improve the benefits of reannealing-curve analysis in research and clinic.
Keywords: HRM, melting, reannealing
Introduction
The Janus kinase 2 (JAK2) plays an important role in the intracellular signaling in normal and neoplastic cells.1 A point mutation (c.1849GNT) results in a mutant protein (GTC→TTC), called JAK2 V617F in which valine is substituted by phenylalanine at position 617.2 JAK2 V617F has been detected frequently in Philadelphia chromosome-negative myeloproliferative neoplasms, such as polycythemia vera (65–97%), essential thrombocythemia (23–57%) and primary myelofibrosis (35–57%). Accordingly, detection of this mutation has been included within the WHO diagnostic criteria for polycythemia vera (PV), essential thrombocytosis (ET) and primary myelofibrosis (PMF).3–6 In recent years, several methods have been developed to detect the JAK2 V617F mutation.7,8 However, each of them has its own limitations and disadvantages, such as low sensitivity and high costs, particularly in large-scale diagnosis.9 A more recently established technique, high-resolution melting (HRM) curve analysis represents a new mutation scanning technology without the need for time and cost consuming post-PCR processing.9 HRM-curve analysis is a simple, economic and sensitive technique consisting of PCR that distinguishes DNA samples in the presence of a DNA-binding dye, based on their melt temperatures (Tm) as they are denatured from a double-stranded to a single-stranded form, by precise increasing the temperature.10 Eva Green is a sensitive, stable and cost-effective DNA-binding dye, and therefore suitable for DNA melt curve analysis and several other PCR-based detection methods.11 The melting pattern of a PCR product depends on its length, sequence, GC content or strand complementarity. Homozygous mutations will result in the Tm shift compared with the wild-type sample. In contrast, heterozygous mutations present different shape of the melting curve.12,13 Recently, many researchers focused on the most efficient methods for JAK2 V617F mutation detection.14–17 Table 1 summarizes some of the previous studies on JAK2V617F mutation detection, which focused on HRM method and compared this method with other routine molecular methods.
 |
Table 1 Some of the previous studies on JAK2V617F mutation detection |
In this work, we aimed to describe the basis of the melting- and reannealing-curve analysis in real-time instruments and to assess the analytical efficiency of HRM method using reannealing-curve analysis in comparison with routine melting-curve analysis for JAK2 V617F mutation detection. Also, we tried to introduce a new approach to improve the sensitivity and specificity of the routine HRM method.
Materials and methods
Sample
A total of 23 samples including one negative synthetic standard DNA, two positive synthetic standard DNA samples (positivity of 50% and 75%), five wild-type samples (negative mutation for JAK2 V617F) and 15 mutant samples (positive for JAK2 V617F). Patients with hematocrit higher than 52% for males and 48% for females were recruited in this study. About 3 mL of venous blood were collected in EDTA for complete blood count and DNA extraction. Demographic characteristics and hematologic parameters are summarized in Table 2.
 |
Table 2 Demographic characteristics and hematologic parameters |
Ethical considerations
This study was approved by ethical committee of Kerman University of Medical Sciences. The ethics approval code is IR.KMU.REC.1395.812. This code obtained before the study starts and the patients provided written informed consent conducted in accordance with the Declaration of Helsinki.
DNA extraction
DNA was extracted using the QIAamp DNA Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s protocol and was kept at 4°C before use. DNA purity was determined by calculating the ratio of absorbance at 260 and 280 nm (A260/A280) using Nano Drop ND-1000 Spectrophotometer (Nano Drop Technologies, Thermo Scientific, Wilmington, DE, USA).
HRM-curve analysis
The specific primers for using in the amplification stage of HRM were designed using Vector NTI version11 as follows:
JAK2, F: 5ʹ-TTGAAGCAGCAAGTATGATG-3ʹ;
JAK2, R: 5ʹ-CTTACTCTCGTCTCCACAG-3ʹ.
The target region was 86 bp in length and located in the 1783–1869 nucleotide position. These primers were synthesized by Eurofins MWG Operon (Ebensburg, Germany) (Figure 1).
 |
Figure 1 Schematic view of amplification target for HRM method and JAK2 V617F (C.1849G>T) point mutation. |
Concentration of DNA samples was set as 100 pmol/µL to make minimum interference in melting analysis and better interpretations and comparison of the results.
HRM method was conducted by Rotor-Gene Q real-time thermal cycler and results were confirmed by sequencing and TaqMan Probe methods. The HRM reaction mix without DNA template was used as non-template control. To check the integrity and overall quality, PCR products were run on 2% agarose gel and visualized by staining with ethidium bromide. The final reaction volume of 20 µL consisted of 10 µL of Master Mix (typit HRM; Qiagen), 1.4 µl primer pair and 2 µL DNA in 6.6 µL distilled water. The thermal cycling conditions consisted of initial melting stage at 95°C for 5 mins followed by 35 cycles of 5 s at 94°C, annealing at 60°C for 10 s and then a final extension stage of 5 mins at 72°C.
After that, HRM analysis was continued in the two stages: first, melting stage by precise increasing the temperature and second, reannealing stage by precise decreasing the temperature. Melting stage consisted of temperature increase from 60°C to 94°C with a ramp rate of 0.02%. This procedure starts with double-strand DNA (dsDNA) molecules with 100% fluorescence emission due to the binding interactions between Eva Green dye and dsDNA molecules. By the temperature increase, the dsDNA began melting at which all ssDNA molecules dissociate gradually and fluorescence starts to reduce. Tm is the temperature at which one-half of dsDNA is separated to ssDNA. On the other hand, fluorescence emission is about 50%. In contrast, reannealing stage starts with a gradual temperature decrease from 94°C to 60°C with a ramp rate of 0.02% that all of ssDNA molecules associate together gradually to form ssDNA, and therefore, fluorescent emission increases to 100%. In both methods, there are some variations between samples because of the different DNA copy number and fluorescence emission status (depending on the pre-HRM PCR reaction). Therefore, the normalized region should be used to overcome this defect. The first normalized region for the reannealing stage is considered as 0% fluorescent emission and the second normalized region is considered as 100%. Conversely, the first and second normalized regions for melting stage are considered as 100% and 0% fluorescent emission, respectively. These regions are the same for all samples and melting analysis is performed to find the Tm or 50% fluorescent emission. The amplification plot for each sample was generated by the Rotor-Gene Q- Pure Detection software linked to the instrument. The generated data were subsequently imported to the HRM software for melting-curve analysis. The fluorescence change of each sample was plotted against temperature and normalized to produce the aligned melt curves. Finally, the results of melting and reannealing stages were compared.
Results
Quality and concentration of DNA samples were also quickly estimated by agarose gel electrophoresis (Figure 2).
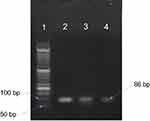 |
Figure 2 Electrophoresis of PCR products on 1% agarose gel. Amplified product is 86 bp. 1: Size marker (50 bp); 2, 3: samples, 4: positive control. |
HRM analysis by melting and reannealing methods
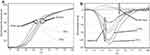
According to the results of melting-curve analysis, the wild-type and mutant DNA samples had different melting temperatures (75/53°C and 75/10°C, respectively) (Figure 3). First and second normalization regions were defined in the raw curves (Figures 4 and 5). Then, normalized curves were obtained for both melting and reannealing methods in which the difference in melting temperatures was apparent (Figures 6A and 7A). For the better discrimination between wild-type and mutant samples, one wild-type sample was defined as baseline (normalized minus G). As shown in Figures 6B and 7B, mutant sample was completely separated from the baseline. In this context, samples with more mutant DNA burden were in more distance from the baseline (50% and 75% mutant samples compared to the baseline). Interestingly, in normalized minus G curves corresponding to reannealing method, mutant samples were better separated from the baseline than in melting method as well as for samples with different mutant DNA burden from each other (for example, 50% from 75% mutant samples). Furthermore, wild-type samples were more homogenous in the normalized minus G curves corresponding to reannealing than in melting method. This means that patients with a low allelic burden may be wrongly interpreted as normal in melting method.
 |
Figure 3 Melting curve of the wild-type and mutant DNA samples. |
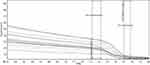 |
Figure 4 Raw melt curves with normalized region. |
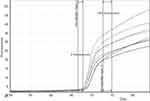 |
Figure 5 Raw reannealing curves with normalized region. |
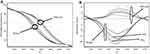 |
Figure 6 (A) Normalized curves for melting method; (B) normalized minus G curves for melting method: wild type, mutant and, 50% and 75% synthetic positive samples are shifted from the baseline. |
Discussion
HRM analysis is an efficient and sensitive PCR-based approach for determining the gene mutation with capability to differentiate heterozygous and homozygous mutations. Furthermore, this method provides the possibility of allele burden measurement in clinical evaluations.19,20 Previous studies using the dilution series of wild-type and mutant DNA samples have shown an analytical sensitivity of about 5% for HRM method in the detection of mutations.19 In a study by Chien-Yu Lin et al, the limit of detection for HRM method was 6%, and in comparison, with AS-PCR it was 2.5%; this finding shows that the HRM method is a sensitive method, can also be performed in closed tube, and is cost-effective. Due to the sensitivity of this method (with 6% limit of detection),16 it can be useful in following up the patients’ response to the treatment and determining MRD for them. So the analysis of HRM is essential for detection. Here for the first time, we compared the reannealing- and melting-curve analysis methods following the HRM analysis with the aim to introduce the more efficient one for JAK2 V617F mutation detection and better differentiation of the samples with different sequence alterations. To the best of our knowledge, there is no previous study that focused on the reannealing step following the HRM-curve analysis. Therefore, there was no previous report for discussion and result comparison. In a study by Serge Carillo et al,15 nested PCR before the common HRM analysis was introduced as Nested HRM which improved the efficacy of the HRM-curve analysis. They reported that Nested HRM had higher sensitivity than common HRM analysis in clinical applications in comparison with other currently used methods. According to the results, samples with mutation allele burden were better differentiated in reannealing than melting step, as well as better separation of the wild type from the mutant samples. Since the length of the PCR products is similar, the Tm shift in curves is directly related to the sequence alterations. Therefore, the Tm shift in the sample with 70% mutant allele burden was more than in sample with 30% mutant allele burden. Complete separation of the mutant from wild-type samples is an essential step in mutation detection and allele burden measurement. In conclusion, we suggest the use of reannealing instead of the melting-curve analysis for the detection of sequence variations, especially for large-scale mutation and allele burden measurement in clinical settings. However, more evaluations with more sample size will better improve the benefits of reannealing-curve analysis in research and clinics.
Acknowledgment
The authors would like to thank all friends and colleagues of Pathology and Research Center of Aja University of Medical Sciences for their relentless hard work and efforts.
Disclosure
The authors declare that they have no conflict of interests in this work.
References
1. Socoro-Yuste N, de Peredo AG, Dagher M, et al. . pH-negative MPN red blood cells display deregulation of IQGAP1-rho GTPase signaling depending on CALR/JAK2 status. Haematologica. Pavia (Italy): Ferrata Storti Foundation; 2015:95–96.
2. Feenstra JDM, Nivarthi H, Gisslinger H, et al. Whole-exome sequencing identifies novel MPL and JAK2 mutations in triple-negative myeloproliferative neoplasms. Blood. 2016;127:325–332. doi:10.1182/blood-2015-07-661835
3. Baxter EJ, Scott LM, Campbell PJ, et al. Acquired mutation of the tyrosine kinase JAK2 in human myeloproliferative disorders. Lancet. 2005;365:1054–1061. doi:10.1016/S0140-6736(05)71142-9
4. Zhao R, Xing S, Li Z, et al. Identification of an acquired JAK2 mutation in polycythemia vera. J Biol Chem. 2005;280:22788–22792. doi:10.1074/jbc.C500138200
5. Levine RL, Wadleigh M, Cools J, et al. Activating mutation in the tyrosine kinase JAK2 in polycythemia vera, essential thrombocythemia, and myeloid metaplasia with myelofibrosis. Cancer Cell. 2005;7:387–397. doi:10.1016/j.ccr.2005.03.023
6. Kralovics R, Passamonti F, Buser AS, et al. A gain-of-function mutation of JAK2 in myeloproliferative disorders. N Engl J Med. 2005;352:1779–1790. doi:10.1056/NEJMoa051113
7. Er T-K, Lin S-F, Chang J-G, et al. Detection of the JAK2 V617F missense mutation by high resolution melting analysis and its validation. Clinica Chimica Acta. 2009;408:39–44. doi:10.1016/j.cca.2009.07.002
8. Moradabadi A, Farsinejad A, Khansarinejad B, Fatemi A. Development of a high resolution melting analysis assay for rapid identification of JAK2 V617F missense mutation and its validation. Experimental Hematology & Oncology. 2019;8(1):10.
9. Moradabadi AR, Farsinejad AR, Fatemi A. Modified PCR-RFLP for detection of JAK2V617F mutation in patients with myeloproliferative neoplasm. Sci J Iran Blood Transfus Org. 2017;14:289–294.
10. Federici MTT, Branda Sica A, Artigas R, Briano C, Dutra F, Llambí S. High-resolution melting (HRM) curve analysis: new approach used to detect blad and dumps in Uruguayan Holstein breed. Arch Vet Sci. 2018;23. doi:10.5380/avs.v23i4
11. Tahmasebi H, Dehbashi S, Arabestani MR. High resolution melting curve analysis method for detecting of carbapenemases producing pseudomonas aeruginosa. JKIMSU. 2018;7:70–77.
12. Ritort F. Open questions about DNA melting: comment on” DNA melting and energetics of the double helix” by Maxim Frank-Kamenetskii et al. Phys Life Rev. 2018;25:34–36. doi:10.1016/j.plrev.2018.03.009
13. Vologodskii A, Frank-Kamenetskii MD. DNA melting and energetics of the double helix. Phys Life Rev. 2018;25:1–21. doi:10.1016/j.plrev.2017.11.012
14. Cao H-C, Lin J, Qian J, et al. Detection of the JAK2 mutation in myeloproliferative neoplasms by asymmetric PCR with unlabeled probe and high‐resolution melt analysis. J Clin Lab Anal. 2011;25:300–304. doi:10.1002/jcla.20474
15. Carillo S, Henry L, Lippert E, et al. Nested high-resolution melting curve analysis: a highly sensitive, reliable, and simple method for detection of JAK2 Exon 12 mutations – clinical relevance in the monitoring of polycythemia. J Mol Diagn. 2011;13:263–270. doi:10.1016/j.jmoldx.2010.12.002
16. Lin C-Y, Ho C-M, Tamamyan G, Yang S-F, Peng C-T, Chang J-G. Validating the sensitivity of high-resolution melting analysis for JAK2 V617F mutation in the clinical setting. J Clin Lab Anal. 2016;30:838–844. doi:10.1002/jcla.21945
17. Matsumoto N, Mori S, Hasegawa H, et al. Simultaneous screening for JAK2 and calreticulin gene mutations in myeloproliferative neoplasms with high resolution melting. Clinica Chimica Acta. 2016;462:166–173. doi:10.1016/j.cca.2016.09.023
18. Wang J, Ren X, Bai X, et al. Identification of gene mutation in patients with osteogenesis imperfect using high resolution melting analysis. Sci Rep. 2015;5:13468. doi:10.1038/srep13468
19. Heideman D, Thunnissen FB, Doeleman M, et al. A panel of high resolution melting (HRM) technology-based assays with direct sequencing possibility for effective mutation screening of EGFR and K-ras genes. Anal Cell Pathol. 2009;31:329–333.
20. Milbury CA, Li J, Makrigiorgos GM. COLD-PCR-enhanced high-resolution melting enables rapid and selective identification of low-level unknown mutations. Clin Chem. 2009;55:2130–2143. doi:10.1373/clinchem.2009.131029
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.