Back to Journals » International Journal of Nanomedicine » Volume 15
Affibody-Modified Gd@C-Dots with Efficient Renal Clearance for Enhanced MRI of EGFR Expression in Non-Small-Cell Lung Cancer
Authors Wu Y, Li H, Yan Y, Wang K, Cheng Y, Li Y, Zhu X, Xie J , Sun X
Received 30 December 2019
Accepted for publication 25 May 2020
Published 30 June 2020 Volume 2020:15 Pages 4691—4703
DOI https://doi.org/10.2147/IJN.S244172
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Mian Wang
Yongyi Wu,1,2,* Haoxiang Li,1,2,* Yuling Yan,1,2,* Kai Wang,1,2 Yongna Cheng,1,2 Yangyang Li,1,2 Xinyuan Zhu,3 Jin Xie,4 Xilin Sun1,2
1NHC and CAMS Key Laboratory of Molecular Probe and Targeted Theranostics, Molecular Imaging Research Center (MIRC), Harbin Medical University, Harbin, Heilongjiang, 150028, People’s Republic of China; 2TOF-PET/CT/MR Center, The Fourth Hospital of Harbin Medical University, Harbin, Heilongjiang 150028, People’s Republic of China; 3School of Chemistry and Chemical Engineering, Shanghai Key Laboratory for Molecular Engineering of Chiral Drugs, Shanghai Jiao Tong University, Shanghai 200240, People’s Republic of China; 4Department of Chemistry and Bio-Imaging Research Center, The University of Georgia, Athens, Georgia 30602, USA
*These authors contributed equally to this work
Correspondence: Xilin Sun
NHC and CAMS Key Laboratory of Molecular Probe and Targeted Theranostics, Molecular Imaging Research Center, Harbin Medical University, 766 Xiangan N Street, Songbei District, Harbin, Heilongjiang 150028, People’s Republic of China
Fax +86-451-82576509
Email [email protected]
Jin Xie
Department of Chemistry, University of Georgia, Athens, GA 30602, USA
Email [email protected]
Purpose: Gd-encapsulated carbonaceous dots (Gd@C-dots) have excellent stability and magnetic properties without free Gd leakage, therefore they can be considered as a safe alternative T1 contrast agent to commonly used Gd complexes. To improve their potential for cancer diagnosis and treatment, affibody-modified Gd@C-dots targeting non-small-cell lung cancer (NSCLC) EGFR-positive tumors with enhanced renal clearance were developed and synthesized.
Materials and Methods: Gd@C-dots were developed and modified with Ac-Cys-ZEGFR:1907 through EDC/NHS. The size, morphology, and optical properties of the Gd@C-dots and Gd@C-dots-Cys-ZEGFR:1907 were characterized. Targeting ability was evaluated by in vitro and in vivo experiments, respectively. Residual gadolinium concentration in major organs was detected with confocal imaging and inductively coupled plasma mass spectrometry (ICP-MS) ex vivo. H&E staining was used to assess the morphology of these organs.
Results: Gd@C-dots with nearly 20 nm in diameter were developed and modified with Ac-Cys-ZEGFR:1907. EGFR expression in HCC827 cells was higher than NCI-H520. In cell uptake assays, EGFR-expressing HCC827 cells exhibited significant MR T1WI signal enhancement when compared to NCI-H520 cells. Cellular uptake of Gd@C-dots-Cys-ZEGFR:1907 was reduced, when Ac-Cys-ZEGFR:1907 was added. In vivo targeting experiments showed that the probe signal was significantly higher in HCC827 than NCI-H520 xenografts at 1 h after injection. In contrast to Gd@C-dots, Gd@C-dots-Cys-ZEGFR:1907 nanoparticles can be efficiently excreted through renal clearance. No morphological changes were observed by H&E staining in the major organs after injection of Gd@C-dots-Cys-ZEGFR:1907.
Conclusion: Gd@C-dots-Cys-ZEGFR:1907 is a high-affinity EGFR-targeting probe with efficient renal clearance and is therefore a promising contrast agent for clinical applications such as diagnosis and treatment of NSCLC EGFR-positive malignant tumors.
Keywords: Gd@C-dots, EGFR, MRI, nanoparticles, efficient renal clearance
Introduction
Nanomedicine is a promising approach that may provide significant breakthroughs in medicine and healthcare.1 Indeed, nanotechnology has allowed to overcome important limitations of traditional magnetic resonance imaging (MRI) contrast agents due to the properties and surface modifications of nanoparticles, ultimately leading to the improvement of diagnosis efficacy.2 MRI contrast agents increase the sensitivity of MR scans and thereby improve diagnostic accuracy, particularly in malignant disease. To date, paramagnetic Gd-chelates, such as Gd-DTPA, Gd-DTPA-BMA, Gd-DOTA, are more widely used in clinical diagnosis among the MRI contrast agents. However, these chelates are non-specific contrast agents and therefore cannot differentiate molecular expression in tumors, which is necessary to guide targeted therapy. Moreover, gadolinium linear complexes can release free gadolinium ions in the body, which may cause severe nephrogenic systemic fibrosis (NSF) due to their toxicity, particularly in patients with renal diseases or poor renal functions.3,4
Recently, nanoparticle-based MRI contrast agents have received increasing attention because of their many advantages. For instance, Gd nanoparticles exhibit reduced toxicity profiles by preventing Gd3+ leakage from traditional chelates. Thus, several Gd nanoparticles have been developed over the past decade, including Gd2O3,5 Gd-based silica nanoparticles,6 and Gd-Si oxide nanoparticles.7 However, due to their surface properties and relatively large size, these nanoparticles accumulate heavily in the reticuloendothelial (RES) organs after systemic injection, particularly in the liver and spleen. To overcome this limitation, we recently developed Gd-based nanoparticles with efficient renal clearance.8 The Gd@C-dots have shown relatively high T1 relaxivity with steerable size and low toxicity, and can be eliminated by the renal system quickly without leakage of free Gd(III) to the plasma or urine. And this efficient renal clearance makes Gd@C-dots a safer MRI agent.8
Although nanoparticles modify with specific targeting peptides provide accurate cancer diagnosis, imaging the lung with MRI remains challenging.9 Most of nuclear magnetic resonance (NMR) signals generated from MRI based on the hydrogen nuclei of water molecules. However, there’s a lot of air in the lungs which leads to an inherently low signal-to-noise ratio. Moreover, the high difference between the magnetic susceptibility of the air cavity and the tissue in the lung brings the chemical displacement artifact of the air. Finally, because of the long scanning time, the artifact of respiratory movement becomes a problem to solve. Thus, the use of MRI contrast agents can make up for the deficiencies and improve diagnostic accuracy. There is therefore currently a need to develop effective contrast agents to improve the potential of MRI in detecting malignant lung cancer.
Epidermal growth factor receptor (EGFR) is a well-established tumor biomarker, which is overexpressed in a wide range of human tumors, such as breast cancer, brain tumor, and particularly lung cancer, where it is associated with tumor proliferation, metastasis and angiogenesis.10,11 Besides, anti-EGFR therapy is one of the most advanced and effective treatments for EGFR-overexpressing tumors.12,13 Thus, distinguishing EGFR expression in tumors is crucial for precision anti-EGFR therapy. Affibodies have shown great promise as tumor-targeting agents.14–16 Several anti-EGFR affibodies, including Ac-Cys-ZEGFR: 1907, exhibit high affinities in the nM ranges, and have been used for tumor imaging. To date, Ac-Cys-ZEGFR: 1907 shows the best in vivo tumor-targeting properties.16–19
In this study, we developed Gd@C-dots nanoparticles with surface carboxyl groups modified with Ac-Cys-ZEGFR: 1907 by following a synthesis method reported in.20 We assessed the applicability of this nanoprobe in diagnosing EGFR-positive tumors with MRI in non-small-cell lung cancer (NSCLC) (Figure 1).
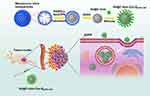 |
Figure 1 Schematic of preparation for Gd@C-dots-Cys-ZEGFR:1907 and its mechanism targeting to EGFR. |
Materials and Methods
Synthesis of Gd@C-Dots-Cys-ZEGFR: 1907
The preparation was optimized to obtain proper sized Gd@C-dots. Briefly, we first synthesized mesoporous silica nanoparticles (MSNs) with an average diameter of 160 nm and a pore size of 20 nm (hereafter designated as MSN-20). For the synthesis of MSNs, according to previous reports.20 Briefly, dissolve 0.6 g cetyl trimethyl ammonium bromide (CTAB) in 300 mL water and add 2.1 mL 2 M NaOH, stirring magnetically and heating to 70 °C, add 3 mL TEOS, 18 mL ethyl acetate and 1.8 mL APS to the mixture and stir at 70 °C for 3 h. After centrifugation, wash with ethanol for 3 times, and then disperse into ethanol. Add 50 mg NH4NO3 and stir for 3 h at 60 °C. Wash with ethanol for 2 times. Dry at 60 °C overnight. MSN-20 was obtained by incubation in Na2CO3 solution (pH 12) at 50 °C. For the synthesis of Gd@C-dots, MSN-20 were first incubated in a solution containing 100 mM Gd(NO3)3 and 10 mM Gd-DTPA. Then, the Gd-loaded MSNs were left to dry on a crucible and were calcined at 300 °C for 2 h in air. The raw products were dispersed in water and purified with centrifugal filter units (MWCO = 100–3K removed aggregations of nanoparticles and unreacted precursors, respectively). The yielded Gd@C-dots were spherical, with an average diameter of ~20 nm and relatively narrow size distribution. The affibody analogue, Ac-Cys-ZEGFR:1907 Ac-CVDNKFNKEMWAAWEEIRNLPNLNGWQMTAFIASL
VDDPSQSANLLAEAKKLNDAQAPK-NH2, was prepared by solid-phase peptide synthesis as previously described.21–23 The peptide was purified by RP-HPLC on a C-4 column and the purity of the product was confirmed by analytical HPLC.21,24 There was a notable amination condensation reaction in Gd@C-dots and Ac-Cys-ZEGFR: 1907. Gd@C-dots were dispersed in borate buffer, and carbodiimide (EDC) and N-hydroxysuccinimide (NHS) (10X) were added (pH=5.5). The mixture was magnetically stirred for 30 min and centrifuged with centrifugal filter units (MWCO = 3K). Then the reaction products were dispersed in borate buffer and Ac-Cys-ZEGFR: 1907 was added. The mixture (pH=8) was incubated for 2 h with gentle agitation and the product was collected with a centrifugal filtration unit (Millipore filter unit: MWCO 10K) and re-dispersed in PBS (pH = 7.4).
Characterization of Gd@C-Dots-Cys-ZEGFR:1907
The size, morphology, and optical properties of the Gd@C-dots and Gd@C-dots-Cys-ZEGFR:1907 were characterized by dynamic light scattering (DLS), transmission electron microscopy (TEM), SpectraMax M2. The result of DLS measurement is presented at number-based distribution. Absorption and photoluminescence spectra were used to confirm the attachment of affibody onto the surface of Gd@C-dots. We assessed the T1 relaxivity of Gd@C-dots-Cys-ZEGFR:1907 with Gd concentrations ranging from 0.0625 to 0.5 mM and suspended in 1% agarose gel in 500 μL PCR tubes. T1 weighted MR images of the samples were acquired on a 3.0T MR system with a four-channel small animal coil. The parameters used were as follows: T1WI:TR/TE = 600/9.4ms; 128×128 matrices; and repetition times = 4. To measure the longitudinal relaxation time of each sample, multi-TR FSE sequence with TE (11.8ms), multi-TR (60, 100, 200, 500, 800, 1000, 1200, 1500, 2000, 2500ms, respectively) was performed.
Cell Culture and Tumor Xenograft Model
HCC827 human lung adenocarcinoma cell line (EGFR expression-positive cell line) and NCI-H520 human squamous cell carcinoma cell line (EGFR expression negative cell line) were purchased from ATCC. They were cultured routinely in RPMI 1640 (Hyclone USA) supplemented with 10% fetal calf serum (Hyclone USA) under 5% CO2, at 37 °C. All cultures were passaged 3–4 times per week to maintain logarithmic growth. All the animal experiments performed in this study were approved by the Harbin Medical University animal ethics committee (Harbin Medical University Harbin, China) and the study was carried out in strict accordance with the recommendations in the Guide for the Care and Use of Laboratory Animals of the National Institutes of Health. Athymic nude mice (female, 4–6 weeks) were purchased from Beijing Vital River Laboratory Animal Technology Company. Mice were fed in an SPF room. The tumor model was established by subcutaneous injection of 5 × 106 HCC827 or NCI-H520 cells into the right lower flank of athymic nude mice. When the tumor volumes reached 300–500mm3, MR scans were performed.
Western Blots
To assess EGFR expression, total protein samples were extracted from HCC827 and NCI-H520 cells in lysis buffer (RIPA) containing protease inhibitor (PMSF). The resulting lysates were centrifuged at 13.2×103 rpm for 15 min and the supernatants were collected. The protein concentrations were measured using the BCA protein assay kit (Solebo biotech Ltd). SDS–PAGE and Western blotting were performed using 80 μg of proteins. The lysates were resolved by electrophoresis (70 V for 25 min and 110 V for 1.5 h) and transferred onto NC membranes. After blocking in 5% non-fat milk for 2 h, the blots were incubated with the first antibody: rabbit anti-human EGFR monoclonal (1:1000, Abcam) and mouse anti-human GAPDH (1:1500, Abcam) were used as a loading control overnight at 4 °C. The blots were washed and incubated with the second antibody: goat anti-rabbit IgG HRP-linked antibody (1:5000, Cell Signaling Technology); goat anti-mouse antibody (1:5000, Cell Signaling Technology) at room temperature for 1.5 h. Western blot bands were captured by the ECL Western blotting detection system (BD). GAPDH was used as a loading control. After development, the films were scanned with BIO-RAD Gel Doc XRS+. The images were opened and analyzed by ImageLab (BIO-RAD) software. Three samples of each tumor cell type were prepared for Western blot to obtain semi-quantitative data for statistical analyses.
Immunofluorescence
HCC827 and NCI-H520 cells were cultured on a sterile glass at 37 °C with 5% CO2. After washing with PBS, the cells were fixed with 4% paraformaldehyde (PFA) for 10 minutes. The mice were sacrificed after MRI scanning for harvesting the tumor tissues for immunofluorescence. Frozen tumor tissue slices (5 μm) of HCC827 and NCI-H520 were fixed with 4% paraformaldehyde (PFA) for 10 minutes. After blocking with 5% goat serum incubated for 1 h, the cells and tumor tissue slices were incubated with primary antibody rabbit anti-human EGFR monoclonal (1:50, Abcam) overnight at 4 °C, washed and incubated with secondary antibody goat anti-rabbit (1:250, Invitrogen). The samples were observed with a confocal microscope (Nikon).
In vitro Cell Uptake Studies
HCC827 and NCI-H520 cells were cultured on a sterile glass in petri dishes and then incubated in 1 mL of media containing Gd@C-dots-Cys-ZEGFR:1907 (50μg Gd/mL) for 1h at 37 °C. After incubation, the cells were washed with PBS three times and then fixed with 4% PFA. Similarly, 106 of HCC827 and NCI-H520 cells in 6-well plates were incubated with Gd@C-dots-Cys-ZEGFR:1907 (50μg Gd/mL) for 1h at 37 °C, 5% CO2. For the blocking experiment, HCC827 cells were pretreated with 500 μM Ac-Cys-ZEGFR:1907 for 45 min before the addition of Gd@C-dots-Cys-ZEGFR:1907. After washing with PBS three times, the cells were detached by tyrosine treatment and harvested by centrifugation. They were counted with a hemocytometer and then suspended in 1% agarose in 500ul PCR tubes for MR imaging. T1WI MR images were acquired with the following parameters: TR/TE = 600/9.4ms; matrix: 128×128; and NEX: 4. HCC827 cells after uptake also observed by transmission electron microscopy (TEM).
In vivo MR Imaging of Subcutaneous Tumor Xenografts
HCC827 and NCI-H520 tumor xenograft models were intravenously injected with Gd@C-dots-Cys-ZEGFR:1907 (3.2 mg Gd/kg). Gd@C-dots and Gd-DTPA-BMA (3.2 mg Gd/kg) were also injected into HCC827 tumor xenograft models. T1-weighted images were acquired before injection, and 10 min, 30 min, 1 h, 2 h, and 4 h after injection with the following parameters: T1-weighted: Fast Spin Echo, FOV: 8 cm × 4.8 cm; thickness 2.0 mm; TR: 600ms; TE: 9.8 ms; flip angle: 142°; matrix: 128 × 128; NEX: 4. Signal-to-background ratio (SBR) was calculated by finely analyzing regions of interest (ROIs) of the MR images and SBR/SBR0 was calculated to represent the signal changes. Signal intensity (SI) of tumor, liver, kidney and muscle were measured before and after the injection of the contrast agents. The mean SI measurements of three mice per group were used for statistical analysis. For each mouse, 3–5 ROIs were selected to measure SI of tumors, liver, kidney and muscle. The SBR values were calculated according to SBR= SI organ/SI muscle.
Ex vivo Biodistribution of Gd@C-Dots-Cys-ZEGFR:1907
To assess the biodistribution of Gd@C-dots-Cys-ZEGFR:1907, the mice were sacrificed 4 h after injection and the organs and tumors were collected for fluorescence and ICP-MS analyses. In ICP-MS experiments, tissue was dispersed in 5 mL of aqua regia (67% HNO3/37% hydrochloric acid (w/w)) for 4 h at 80 °C and, subsequently, the samples were diluted with HNO3 2% (w/w) matrix to adjust the volume to 5 mL, filtered (0.22 μm) and analyzed by ICP-MS. For calibration of the inductively coupled plasma mass spectrometry (ICP-MS), single-element standard solutions were prepared by successive dilution in a HNO3 2% (w/w) matrix from a 1000-ppm Gd standard acquired from SCP Science. Based on the fluorescence characteristics of Gd@C-dots, the liver, spleen, and kidney were collected before injection and 4 h after injection. Slices of 5 μm thickness were prepared and nuclei were stained with propidium iodide (PI). The distribution of Gd@C-dots-Cys-ZEGFR:1907 in major organs was analyzed with a confocal fluorescence microscope.
Toxicity of Gd@C-Cys-ZEGFR:1907 in Mice
In vivo toxicity of Gd@C-dots-Cys-ZEGFR:1907 was determined in mice (18−25 g, n=3). Typically, Gd@C-dots-Cys-ZEGFR:1907 (3.2 mg Gd/kg) were injected into the tail vein. As control group, an equal volume of saline solution was injected into the tail vein (18–25 g, n=3). The kidney, liver, spleen, and lung were harvested at 4 h after injection. Hematoxylin and eosin (H&E) staining were performed to investigate any morphology changes after injection with Gd@C-dots-Cys-ZEGFR:1907.
Statistical Analysis
Statistics were performed using the data analysis package within GraphPad Prism 6.0 for Windows (GraphPad Software, San Diego, CA). Unless otherwise stated, tests compare two means with two-tail unpaired Student’s t-tests, with equal variance assumed. Error bars indicate standard error of the mean (SEM) unless otherwise stated. All data are expressed as mean ± standard deviation (SD). The in vivo targeting ability of Gd@C-dots-Cys-ZEGFR:1907 at different time points, and with different contrast agents, were analyzed by one-way ANOVA. Comparisons of biodistribution of Gd@C-dots-Cys-ZEGFR:1907 in liver, spleen, kidney, lung, and tumor were analyzed by two-way ANOVA. P<0.05 was considered statistically significant. All MR images were analyzed by ImageJ software.
Results
Characterization of Gd@C-Dots-Cys-ZEGFR:1907
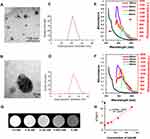
We prepared Gd@C-dots-Cys-ZEGFR:1907 by conjugation of the affibody Ac-Cys-ZEGFR:1907 coupled to the carboxyl group onto the surface of Gd@C-dots nanoparticles. High-resolution TEM imaging of the nanoparticles (Figure 2A and B) showed that it had regular shape and relatively uniform size. DLS revealed that the hydrodynamic diameters of Gd@C-dots and Gd@C-dots-Cys-ZEGFR:1907 were 21.5±2.2 nm (Figure 2C) and 25.1±3.0 nm (Figure 2D), respectively. Gd@C-dots showed a broad absorption band between 200 nm and 500 nm, with a shoulder appearing at 280 nm which had changed after the attachment of affibody. A red shift emerged in the photoluminescence spectrum of Gd@C-dots-Cys-ZEGFR:1907 meaning that the molecular structure of the probe changed. Gd@C-dots and Gd@C-dots-Cys-ZEGFR:1907 were also highly fluorescent, and both can be excited by light of a wide range of wavelengths to emit photoluminescence (Figure 2E and F). Figure 2G showed a representative T1WI image of different Gd concentrations of Gd@C-dots-Cys-ZEGFR:1907. T1 relaxivity analyzed with 3.0 T MR on agarose samples of Gd@C-dots-Cys-ZEGFR:1907 revealed a relativity r1 of 2.55 s−1mM−1 (Figure 2H). This enhanced r1 was mainly attributed to the increase in the rotational correlation time (τR) as a result of binding Gd to a nanoparticle.
EGFR Expression in HCC827 and NCI-H520 Cells
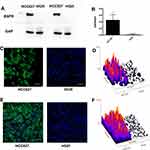
To measure total EGFR expression levels in HCC827 and NCI-H520 cells, we performed Western blots and immunofluorescence analyses. Total EGFR in HCC827 cells detected by Western blots was significantly higher than in NCI-H520 cells (Figure 3A and B, P<0.05). The results were consistent with the immunofluorescence analysis (Figure 3C and D), which also showed that total EGFR expression in HCC827 cells was higher than in NCI-H520 cells.
EGFR Expression in HCC827 and NCI-H520 Tumor Tissue
Based on the results above, HCC827 and NCI-H520 xenografts models were generated to present lung tumors with high- and no-EGFR expression, respectively, for subsequent imaging studies. Subcutaneous tumor tissues (n=3 for each tumor model) were harvested for assessment of EGFR expression by immunofluorescence which indicated HCC827 tumor tissue showed high EGFR expression, whereas NCI-H520 xenografts had no detectable EGFR expression (Figure 3E and F).
In vitro Gd@C-Dots-Cys-ZEGFR:1907 Cell Uptake
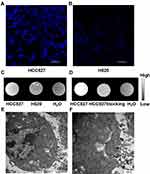
Gd@C-dots-Cys-ZEGFR:1907 was incubated with HCC827 and NCI-H520 respectively. After 1 h of incubation, we could detect a significantly higher fluorescence signal (ex/em: 360/460 nm) in HCC827 cells when compared to NCI-H520 cells (Figure 4A and B), that was in agreement with our T1-weighted MRI results. Moreover, when compared to the negative control, the T1-weighted MR signal in HCC827 cells was notably enhanced (61.8%), whereas the signal in NCI-H520 cells only increased slightly (19.8%) (Figure 4C). These results show that EGFR-positive cells have a high uptake of Gd@C-dots-Cys-ZEGFR:1907. Also, cellular uptake in HCC827 cells was dramatically decreased upon pretreatment of the Gd@C-dots with Ac-Cys-ZEGFR:1907 (Figure 4D), indicating that it was mostly mediated by specific binding of the nanoparticles to EGFR. These differences in cellular uptake can also be discerned by MRI. TEM revealed that most of nanoprobes were targeted to the plasma membrane and a smaller proportion was transported into the cytoplasm by endocytosis (Figure 4E and F). Together these results demonstrate that Gd@C-dots-Cys-ZEGFR:1907 can target EGFR specifically in vitro.
Gd@C-Dots-Cys-ZEGFR:1907 Imaging in Subcutaneous Tumor Xenografts
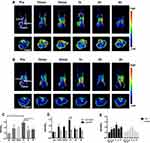
To assess the EGFR-targeting specificity of Gd@C-dots-Cys-ZEGFR:1907 in vivo, the particles were intravenously (i.v.) injected (3.2 mg Gd/kg) into HCC827 and NCI-H520 tumor xenografts nude mice models (n=3). T1-weighted images were acquired before and 10 min, 30 min, 1 h, 2 h, and 4 h after injection. The T1WI tumor signal was significantly enhanced in EGFR-positive HCC827 models, when compared to NCI-H520 tumor models (Figure 5A and B). We detected an initial increase in the signal throughout the body. At 1 h after injection, the signal started decaying and after 4 h, it had subsided to pre-injection levels. In HCC827 tumor models, SBR/SBR0 at 1 h after injection (137.4 ± 4.6%) was higher than that before and 2 h (109.95 ± 9.5%) or 4 h (112.5 ± 4.3%) after injection (Figure 5C, P<0.05). Moreover, the signal enhancement in HCC827 tumors (137.4 ± 4.6%) was higher than that in NCI-H520 tumors (105.3 ± 2.5%) at 1 h after injection (Figure 5D, P<0.001). Finally, the signal enhancement of the non-targeting contrast agents Gd@C-dots and Gd-DTPA-BMA were significantly lower than the Gd@C-dots-Cys-ZEGFR:1907 in the HCC827 tumor model at 1 h after injection (Figure S1, P<0.0001). Although Gd-DTPA-BMA had strong signal enhancement after injection, there was no tumor-specific.
Biodistribution of Gd@C-Dots-Cys-ZEGFR:1907
Biodistribution experiments showed that Gd@C-dots-Cys-ZEGFR:1907 signal intensity increased in the kidney but started decaying after 1 h, and reached pre-injection levels after 2 h. The signal in the liver increased and remained stable throughout the experiment (Figure 5E). Following a significant increase of the signal in the bladder from 10 min to 2 h after injection, it decreased to pre-injection levels at 4 h (Figure 6A). In contrast, there was no signal enhancement with Gd@C-dots (Figure 6B). These results demonstrated that Gd@C-dots modified with Ac-Cys-ZEGFR:1907 owned effective renal clearance.
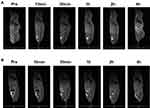 |
Figure 6 MR imaging of renal clearance. (A) Sagittal T1-weighted MR images of Gd@C-dots-Cys-ZEGFR:1907 in animal. (B) Sagittal T1-weighted MR images of Gd@C-dots in animal. |
Ex vivo Biodistribution of Gd@C-Dots-Cys-ZEGFR:1907
Based on fluorescence properties of Gd@C-dots-Cys-ZEGFR:1907, the biodistribution of Gd@C-dots-Cys-ZEGFR:1907 in liver, kidney, and spleen tissues was examined. No detectable probe fluorescence signal was observed in the kidney, whereas nanoparticle accumulation in the liver and spleen tissues were detected by fluorescence (Figure S2a). Moreover, Gd concentration measured by ICP-MS revealed that more than 60% of particles had been eliminated from the body after 4 hours. Most of the residual Gd@C-dots-Cys-ZEGFR:1907 accumulated in the liver and spleen (Figure S2b, S2c).
Toxicity of Gd@C-Dots-Cys-ZEGFR:1907 in Mice
Low biotoxicity is a crucial property of MRI contract probes used in clinical applications. To determine the biotoxicity of Gd@C-dots-Cys-ZEGFR:1907, we harvested the major organs of mice 4 h after injection of Gd@C-dots-Cys-ZEGFR:1907 or saline solution. H&E stainings on these tissues showed that the lung, liver, spleen, and kidney in mice injected with Gd@C-dots-Cys-ZEGFR:1907 had no morphological differences from those in mice injected with saline solution (Figure S3).
Discussion
Gd(III) chelates are the most commonly used MRI contrast agents in clinical settings. However, although these probes offer detailed anatomical images, Gd(III) chelates cannot provide molecular expression information and may cause toxicity, particularly in kidney failure patients.25 In this study, we developed EGFR-targeting Gd@C-dots that can be efficiently eliminated by the renal system. Moreover, this study shows that Gd@C-dots-Cys-ZEGFR:1907 does not cause biotoxicity in mice models. These results are consistent with our previous research claiming that the gadolinium in Gd@C-dots maintains an optimal T1 relaxivity without free Gd(III) release in the body. Moreover, a high concentration of Gd@C-dots results in low cellular toxicity when compared to Gd-DTPA.8 Other studies also indicate that carbon dots nanoparticles do not exert any significant toxic effect in vivo and in vitro. Biocompatibility, low cost, chemical inertness, and fluorescence are also recognized properties of carbon dots.26,27 Thus, Gd@C-dots modified with a targeting ligand may represent a potential safe molecular probe for clinical applications.
An ideal imaging probe can home efficiently to the diseased area after systemic injection, with the unbound being rapidly excreted from the host. However, because of their relatively large size, most nanoparticles tend to accumulate in major organs, such as liver, spleen, and lung.28–30 It is well established that nanoparticles measuring up to 5 nm can be cleared via the kidney, and our previous research showed that Gd@C-dots, which have diameters ~12 nm, can also be efficiently cleared by the renal system.8 In the current study, we demonstrate that Gd@C-dots-Cys-ZEGFR:1907 are efficiently excreted via urine, even though they measure ~20 nm, which exceeds the commonly assumed threshold for renal clearance. In contrast, Gd@C-dots on their own appear to be excreted less efficiently. Although the exact mechanism is unknown, the unique surface of Gd@C-dots-Cys-ZEGFR:1907 may likely facilitate clearance. Ac-Cys-ZEGFR:1907 is quite hydrophilic and can be cleared via the kidney.17,21 On the other hand, Gd@C-dots have a shell of amorphous carbon decorated with carboxyl groups. Thus, Gd@C-dots-Cys-ZEGFR:1907 has a hydrophilic surface and a hydrophobic core, thereby affording the particles with good colloidal stability and the capacity to cross certain types of biological barriers. Indeed, it has previously been suggested that, in addition to particle size and shape, surface properties may facilitate renal clearance.31,32 Mass balance of Gd@C-dots-Cys-ZEGFR:1907 can be revealed by in vivo imaging and ex-vivo biodistribution that significant increase of Gd-related signal in the bladder from 10 min to 2 h post injection indicating that Gd@C-dots-Cys-ZEGFR:1907 was mainly cleared by urinary system, from which 60% of Gd@C-dots-Cys-ZEGFR:1907 can be excreted out at 4 h post injection as measured by ICP-MS in ex-vivo biodistribution study. In addition, like most nanoparticles, Gd@C-dots-Cys-ZEGFR:1907 can accumulate non-specifically in live and spleen as confirmed in both images and biodistribution results, for the reason of phagocytes in the reticuloendothelial. Future research should address these questions to improve the design of nanoparticles used for MRI.
Despite recent advances in the development of contrast agents and MR scanning technology, lung cancer diagnosis with MRI remains challenging, causes include limited spatial resolution, mismatched susceptibility between the air cavities and tissues in the lung, and respiratory and cardiac motion artifacts. Nevertheless, MRI provides excellent tissue differentiation, and high temporal and spatial resolution, which are fundamental for accurate cancer diagnosis. On the other hand, EGFR plays a curial role in NSCLC diagnosis and treatment.33 Thus, MRI with EGFR-targeting probes represents a promising approach for lung cancer diagnosis, and research efforts should be invested in developing safe and sensitive EGFR molecular targeting MR contrast agents. In this study, Gd@C-dots modified with Ac-Cys-ZEGFR:1907 shows highly specific EGFR-targeting ability in cell uptake assays and in vivo experiments with EGFR-expressing NSCLC subcutaneous xenograft mice models. Interestingly, we found that nanoparticle size affects the sensitivity of targeting. Indeed, 3 nm Gd@C-dots-Cys-ZEGFR:1907 shows lower signal enhancement in EGFR-expressing HCC827 xenografts (Figure S4a). This may be explained by poor binding to the targeted EGFR peptide, due to the reduced surface area of the nanoparticle. When compared with larger particles (~100 nm), which allow many ligand-receptor interactions per particle, a small (~5 nm) nanoparticle coated with 50-kDa proteins may interact with only one or two cell receptors. Thus, a particle size that allows interactions with enough receptor peptide is necessary for targeted molecular imaging.34 Moreover, small particles are unsuitable for passive accumulation in the tumor compartment, due to a high rate of interstitial space permeation.35 Finally, small nanoparticles have short circulation time and rapid clearance from the kidney, which significantly reduces the probability of reaching and interacting with the surface of tumor cells. Indeed, after only 4 hours, the 3 nm Gd@C-dots-Cys-ZEGFR:1907 had been eliminated from the body (Figure S4b). Thus, our results suggest that a size of ~20 nm optimizes the affinity and targeting ability of Gd@C-dots-Cys-ZEGFR:1907 to EGFR-expressing tumors in vivo, while allowing efficient renal clearance.
Conclusion
We have developed a new contrast agent for MRI, Gd@C-dots-Cys-ZEGFR:1907, with a high affinity to EGFR and efficient renal clearance. Gd@C-dots-Cys-ZEGFR:1907 shows EGFR-specific binding in EGFR-expressing tumor cells in vitro and in vivo in NSCLC subcutaneous xenograft mice models. Moreover, Gd@C-dots-Cys-ZEGFR:1907 are efficiently eliminated from the kidney, liver, and spleen, and without accumulation in the kidney. Finally, these nanoparticles do not cause biotoxicity. These results suggest that Gd@C-dots-Cys-ZEGFR:1907 is a robust and highly specific MR contrast agent with clinical translation potential for the diagnosis of NSCLC.
Abbreviations
Gd@C-dots, Gd-encapsulated carbonaceous dots; NSCLC, non-small-cell lung cancer; NSF, nephrogenic systemic fibrosis; EGFR, epidermal growth factor receptor; DLS, dynamic light scattering; TEM, transmission electron microscopy; ICP-MS, inductively coupled plasma mass spectrometry.
Statement
The abstract of this paper was presented at the World Molecular Imaging Conference 2018 as a poster presentation with interim findings. The poster’s abstract was published in “Poster Abstracts” in Molecular Imaging and Biology: Volume 20, Issue 1 Supplement, December 2018 https://www.proxydgb.buap.mx:2067/journal/11307/20/1/suppl (Pages 585-787 ID27).
Ethical Approval
All animal experiments were approved by the Harbin Medical University Animal Ethics Committee in accordance with Chinese legislation and followed relevant guidelines. The institutional review board approved this study on mice with NSCLC tumor xenografts.
Funding
This work was supported by the National Natural Science Foundation of China (81627901, 81471724), the Tou-Yan Innovation Team Program of the Heilongjiang Province (2019-15), National Basic Research Program of China (2015CB931800), and the Key Laboratory of Molecular Imaging Foundation (College of Heilongjiang Province).
Disclosure
The authors have declared that no competing interest exists and that they have no conflicts of interest in this work.
References
1. Chang EH, Harford JB, Eaton MA, et al. Nanomedicine: past, present and future a global perspective. Biochem Biophys Res Commun. 2015;468(3):511–517. doi:10.1016/j.bbrc.2015.10.136
2. Le Duc G, Roux S, Paruta-Tuarez A, et al. Advantages of gadolinium based ultrasmall nanoparticles vs molecular gadolinium chelates for radiotherapy guided by MRI for glioma treatment. Cancer Nanotechnol. 2014;5(1):4. doi:10.1186/s12645-014-0004-8
3. Strijkers GJ, Mulder WJ, van Tilborg GA, Nicolay K. MRI contrast agents: current status and future perspectives. Anticancer Agents Med Chem. 2007;7(3):291–305. doi:10.2174/187152007780618135
4. Yang L, Krefting I, Gorovets A, et al. Nephrogenic systemic fibrosis and class labeling of gadolinium-based contrast agents by the Food and Drug Administration. Radiology. 2012;265(1):248–253. doi:10.1148/radiol.12112783
5. Shen ZT, Zheng S, Gounis MJ, Sigalov AB. Diagnostic magnetic resonance imaging of atherosclerosis in apolipoprotein e knockout mouse model using macrophage-targeted gadolinium-containing synthetic lipopeptide nanoparticles. PLoS One. 2015;10(11):e0143453. doi:10.1371/journal.pone.0143453
6. Plissonneau M, Pansieri J, Heinrich-Balard L, et al. Gd-nanoparticles functionalization with specific peptides for ss-amyloid plaques targeting. J Nanobiotechnol. 2016;14(1):60. doi:10.1186/s12951-016-0212-y
7. Cabrera-Garcia A, Vidal-Moya A, Bernabeu A, et al. Gd-Si Oxide nanoparticles as contrast agents in magnetic resonance imaging. Nanomaterials. 2016;6:6. doi:10.3390/nano6060109
8. Chen H, Wang GD, Tang W, et al. Gd-encapsulated carbonaceous dots with efficient renal clearance for magnetic resonance imaging. Adv Mater. 2014;26(39):6761–6766. doi:10.1002/adma.201402964
9. Bianchi A, Dufort S, Lux F, et al. Targeting and in vivo imaging of non-small-cell lung cancer using nebulized multimodal contrast agents. Proc Natl Acad Sci U S A. 2014;111(25):9247–9252. doi:10.1073/pnas.1402196111
10. Mendelsohn J, Baselga J. Status of epidermal growth factor receptor antagonists in the biology and treatment of cancer. J Clin Oncol. 2003;21(14):2787–2799. doi:10.1200/JCO.2003.01.504
11. Mitsudomi T, Yatabe Y. Epidermal growth factor receptor in relation to tumor development: EGFR gene and cancer. FEBS J. 2010;277(2):301–308. doi:10.1111/j.1742-4658.2009.07448.x
12. Moroni M, Veronese S, Benvenuti S, et al. Gene copy number for epidermal growth factor receptor (EGFR) and clinical response to antiEGFR treatment in colorectal cancer: a cohort study. Lancet Oncol. 2005;6(5):279–286. doi:10.1016/S1470-2045(05)70102-9
13. Zhou W, Ercan D, Chen L, et al. Novel mutant-selective EGFR kinase inhibitors against EGFR T790M. Nature. 2009;462(7276):1070–1074. doi:10.1038/nature08622
14. Nilsson FY, Tolmachev V. Affibody molecules: new protein domains for molecular imaging and targeted tumor therapy. Curr Opin Drug Discov Devel. 2007;10(2):167–175.
15. Tolmachev V, Orlova A, Nilsson FY, Feldwisch J, Wennborg A, Abrahmsen L. Affibody molecules: potential for in vivo imaging of molecular targets for cancer therapy. Expert Opin Biol Ther. 2007;7(4):555–568. doi:10.1517/14712598.7.4.555
16. Qi S, Miao Z, Liu H, Xu Y, Feng Y, Cheng Z. Evaluation of four affibody-based near-infrared fluorescent probes for optical imaging of epidermal growth factor receptor positive tumors. Bioconjug Chem. 2012;23(6):1149–1156. doi:10.1021/bc200596a
17. Friedman M, Orlova A, Johansson E, et al. Directed evolution to low nanomolar affinity of a tumor-targeting epidermal growth factor receptor-binding affibody molecule. J Mol Biol. 2008;376(5):1388–1402. doi:10.1016/j.jmb.2007.12.060
18. Zhao P, Yang X, Qi S, et al. Molecular imaging of hepatocellular carcinoma xenografts with epidermal growth factor receptor targeted affibody probes. Biomed Res Int. 2013;2013:759057. doi:10.1155/2013/759057
19. Su X, Cheng K, Jeon J, et al. Comparison of two site-specifically (18) F-labeled affibodies for PET imaging of EGFR positive tumors. Mol Pharm. 2014;11(11):3947–3956. doi:10.1021/mp5003043
20. Chen H, Wang GD, Sun X, et al. Mesoporous silica as nanoreactors to prepare Gd‐encapsulated carbon dots of controllable sizes and magnetic properties. Adv Funct Mater. 2016;26(22):3973–3982. doi:10.1002/adfm.201504177
21. Miao Z, Ren G, Liu H, Jiang L, Cheng Z. Small-animal PET imaging of human epidermal growth factor receptor positive tumor with a 64Cu labeled affibody protein. Bioconjug Chem. 2010;21(5):947–954. doi:10.1021/bc900515p
22. Gao J, Chen K, Miao Z, et al. Affibody-based nanoprobes for HER2-expressing cell and tumor imaging. Biomaterials. 2011;32(8):2141–2148. doi:10.1016/j.biomaterials.2010.11.053
23. Hoppmann S, Miao Z, Liu S, et al. Radiolabeled affibody-albumin bioconjugates for HER2-positive cancer targeting. Bioconjug Chem. 2011;22(3):413–421. doi:10.1021/bc100432h
24. Troy T, Jekic-McMullen D, Sambucetti L, Rice B. Quantitative comparison of the sensitivity of detection of fluorescent and bioluminescent reporters in animal models. Mol Imaging. 2004;3(1):9–23. doi:10.1162/153535004773861688
25. Thomsen HS, Marckmann P, Logager VB. Nephrogenic systemic fibrosis (NSF): a late adverse reaction to some of the gadolinium based contrast agents. Cancer Imaging. 2007;7:130–137. doi:10.1102/1470-7330.2007.0019
26. Wang K, Gao Z, Gao G, et al. Systematic safety evaluation on photoluminescent carbon dots. Nanoscale Res Lett. 2013;8(1):122. doi:10.1186/1556-276X-8-122
27. Himaja AL, Karthik PS, Singh SP. Carbon dots: the newest member of the carbon nanomaterials family. Chem Rec. 2015;15(3):595–615. doi:10.1002/tcr.201402090
28. Roggers R, Kanvinde S, Boonsith S, Oupicky D. The practicality of mesoporous silica nanoparticles as drug delivery devices and progress toward this goal. AAPS Pharm Sci Tech. 2014;15(5):1163–1171. doi:10.1208/s12249-014-0142-7
29. Wang Q, Zhuang X, Mu J, et al. Delivery of therapeutic agents by nanoparticles made of grapefruit-derived lipids. Nat Commun. 2013;4:1867. doi:10.1038/ncomms2886
30. Li J, Pant A, Chin CF, et al. In vivo biodistribution of platinum-based drugs encapsulated into multi-walled carbon nanotubes. Nanomedicine. 2014;10(7):1465–1475. doi:10.1016/j.nano.2014.01.004
31. Liu J, Yu M, Zhou C, Zheng J. Renal clearable inorganic nanoparticles: a new frontier of bionanotechnology. Materials Today. 2013;16(12):477–486. doi:10.1016/j.mattod.2013.11.003
32. Ehlerding EB, Chen F, Cai W. Biodegradable and renal clearable inorganic nanoparticles. Adv Sci. 2016;3:2. doi:10.1002/advs.201600165
33. Chariou PL, Lee KL, Wen AM, Gulati NM, Stewart PL, Steinmetz NF. Detection and imaging of aggressive cancer cells using an epidermal growth factor receptor (EGFR)-targeted filamentous plant virus-based nanoparticle. Bioconjug Chem. 2015;26(2):262–269. doi:10.1021/bc500545z
34. Albanese A, Tang PS, Chan WC. The effect of nanoparticle size, shape, and surface chemistry on biological systems. Annu Rev Biomed Eng. 2012;14:1–16. doi:10.1146/annurev-bioeng-071811-150124
35. Perrault SD, Walkey C, Jennings T, Fischer HC, Chan WC. Mediating tumor targeting efficiency of nanoparticles through design. Nano Lett. 2009;9(5):1909–1915. doi:10.1021/nl900031y
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.