Back to Journals » Infection and Drug Resistance » Volume 13
Absence of Integrase Inhibitor-Associated Resistance Among Antiretroviral Therapy-Naïve HIV-1-Infected Adults in Guangdong Province, China, in 2018
Authors Lan Y, Li L, Chen W, Deng X, Li J, Fan Q, Cai X, Cai W, Hu F
Received 2 October 2020
Accepted for publication 25 November 2020
Published 8 December 2020 Volume 2020:13 Pages 4389—4394
DOI https://doi.org/10.2147/IDR.S284917
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Prof. Dr. Héctor Mora-Montes
Yun Lan,1,* Linghua Li,2,* Weilie Chen,1,* Xizi Deng,1 Junbin Li,2 Qinghong Fan,1 Xiaoli Cai,1 Weiping Cai,2 Fengyu Hu1
1Institute of Infectious Disease, Guangzhou Eighth People’s Hospital, Guangzhou Medical University, Guangzhou, Guangdong, People’s Republic of China; 2Infectious Disease Center, Guangzhou Eighth People’s Hospital, Guangzhou Medical University, Guangzhou, Guangdong, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Weiping Cai
Infectious Disease Center, Guangzhou Eighth People’s Hospital, Guangzhou Medical University, 627 Dongfeng East Road, Yuexiu District, Guangzhou 510030, People’s Republic of China
Tel +86-20-83710825
Email [email protected]
Fengyu Hu
Institute of Infectious Disease, Guangzhou Eighth People’s Hospital, Guangzhou Medical University, 627 Dongfeng East Road, Yuexiu District, Guangzhou 510030, People’s Republic of China
Tel +86-20-83710851
Email [email protected]
Background: Antiretroviral therapy (ART) containing an integrase strand transfer inhibitor (INSTI) plus two nucleoside reverse-transcriptase inhibitors has been recommended as a first-line regimen for ART-naïve HIV-1-infected patients in the latest Chinese Guidelines for Diagnosis and Treatment of HIV/AIDS.
Objective: To determine the prevalence of INSTI-related mutations among ART-naïve HIV-1-infected adults in Guangdong, China, in 2018.
Methods: The entire integrase gene was amplified from blood plasma. Demographic and epidemiological information was collected. INSTI mutations and antiretroviral susceptibility were interpreted using the Stanford University HIV Drug Resistance Database HIVdb program.
Results: Of 927 samples, 827 integrase sequences were successfully obtained. Among them, no major resistance mutations to INSTIs were identified, and four accessory mutations, including T97A (0.12%, 1/827), A128T (0.24%, 2/827), E157Q (0.85%, 7/827), and G163R (0.24%, 2/827), were found in twelve individuals. Two patient samples contained the G163R mutation conferring low-level resistance to elvitegravir and raltegravir.
Conclusion: The overall prevalence of INSTI mutations remains low. Drug resistance mutation testing for the detection of INSTI drug resistance mutations in HIV treatment-naïve patients should be considered due to the circulation of polymorphisms contributing to INSTI resistance and the expected increasing use of this class of drugs.
Keywords: HIV-1, integrase inhibitor, drug resistance mutation, Guangdong
Introduction
Human immunodeficiency virus-1 (HIV-1) remains a global public health problem of unprecedented proportions. Antiretroviral therapy (ART) is a good choice for the prevention and treatment of HIV infection and has reduced the mortality and morbidity associated with HIV infection.1 However, the long-term success of ART is now threatened by HIV drug resistance.1,2 The World Health Organization (WHO) recommends monitoring and periodic surveillance for the appearance of HIV resistance to all antiretroviral treatment programmes.3
HIV-1 integrase plays an important role in HIV-1 replication by catalysing two distinct reactions, 3ʹ-end processing and strand transfer.4 Integrase strand transfer inhibitors (INSTIs) have become an important part of ART because INSTIs have high efficiency, good tolerability, low toxicity, and reduced concern about CYP3A4-associated drug–drug interactions for most patients.5
INSTI-containing regimens are recommended as first-line treatment in the latest Chinese Guidelines for Diagnosis and Treatment of HIV/AIDS after the efficacy of INSTIs was demonstrated in randomized clinical trials.6–9 In Guangdong, the following regimens which include INSTIs are recommended as first-line regimens for approximately 2 years: two nucleotide reverse transcriptase inhibitors backbone with raltegravir (RAL) or Dolutegravir (DTG). Although INSTIs have a higher genetic barrier to the development of HIV-1 resistance,9–11 INSTI inevitably shows drug resistance similar to other anti-HIV drugs.12 Some data show that rare major INSTI mutations were found among primary infected HIV-1 patients in Europe, Venezuela, Canada, sub-Saharan Africa, and the US.13–17 In China, few data are available on the prevalence of HIV-1 INSTI resistance among ART-naïve patients: no major INSTI drug resistance mutations and rare accessory INSTI drug resistance mutations (0.52%, 2/385) were found among 385 ART-naïve individuals in Beijing; a total of 10 (1.95%) major INSTI drug resistance mutations and 8 (1.56%) accessory INSTI drug resistance mutations were found among 513 ART-naïve individuals in Yunnan.18,19 Given that INSTIs are currently used in HIV-1-infected individuals in Guangdong, a developed province on the southern coast of China, it is essential to obtain the resistance status to this class of antiretrovirals, but there is no information about mutations related to INSTIs among HIV-1-infected individuals in Guangdong. Here, we characterized HIV-1 INSTI-associated resistance among ART-naïve HIV-1-infected adults in Guangdong Province, China.
Materials and Methods
Study Subjects and Sample Collection
Guangdong Province has established a system of sentinel hospitals in which patients wishing to receive HIV treatment are cared for by highly trained Infectious Disease specialists. The Guangzhou Eighth People’s Hospital was the leader of all these sentinel hospitals. The enrolment of the subjects included 927 ART-naïve patients who were over 16 years old and who had initiated their therapy at the eleven sentinel tertiary care hospital of Guangdong from January to December 2018, randomly. Five millilitres of blood with EDTA were drawn prior to ART, and the samples were centrifuged immediately, aliquoted, and stored at −80°C. The demographic and epidemiological information of the patients, including risk group, sex, age, ethnicity, and marital status, was collected. The baseline plasma viral load plus CD4+ T cell count were quantified and drug resistance was monitoring at Guangzhou Eighth People’s Hospital. Patients were previously genotyped based on partial pol regions (HXB2 2253–3318) and were excluded if they had ever used antiretroviral drugs.
RNA Extraction and RT-PCR Amplification
HIV-1 RNA was isolated from 200 µL plasma samples using the magnetic-bead-based Viral RNA Extraction Kit (Daan, China) according to the manufacturer’s instructions. The amplification of the entire Integrase gene (864 bp, HXB2 4230–5093) was performed using one-step reverse transcription PCR (RT-PCR) using the PrimeScript One Step RT-PCR Kit (Taraka, China), followed by nested PCR using PrimeSTAR HS DNA Polymerase (Taraka, China). The outer and inner primers were synthesized as previously described.20
The cycling conditions for cDNA synthesis and the first round of PCR amplification were as follows: 45 minutes at 50°C (cDNA synthesis) and initial denaturation for 3 minutes at 94°C, 30 cycles of 94°C for 30 seconds, 55°C for 30 seconds, and 72°C for 90 seconds, followed by a final extension at 72°C for 5 minutes. The cycling conditions for the second round of PCR amplification were as follows: initial denaturation for 3 minutes at 94°C, 40 cycles of 94°C for 30 seconds, 55°C for 45 seconds, and 72°C for 70 seconds, followed by a final extension at 72°C for 5 minutes. Negative controls were utilized to detect possible contamination.
The amplified PCR products were electrophoresed on a 1% agarose gel with negative and positive controls and a DNA size marker (Takara, China). After gel electrophoresis, the positive products were purified using the QIAquick Gel Extraction Kit (QIAGEN, Germany) according to the manufacturer’s protocol and sequenced by a genomics company (Tianyi Huiyuan, China). The primers used for Sanger dideoxy sequencing were the inner primers used for the second round of PCR amplification.
Drug Resistance Testing and Interpretation
The obtained sequences were assembled and edited using Sequencher DNA sequence analysis softwareV5.4.6. All sequences were manually edited and then submitted to the Stanford University HIV Drug Resistance Database (https://hivdb.stanford.edu/) to identify INSTI mutations. Sequences with a low-level category of resistance or greater were defined as having Integrase drug resistance.
Results
A total of 927 HIV-1 patients were tested using the protocol described above, and 100 of them were excluded for cannot obtain the positive PCR products, so the integrase region in 827 patients was successfully amplified, purified and sequenced. The demographic characteristics of these patients (n=827) are shown in Table 1. The participants were aged between 17 and 79 years, with a median age of 35 years. Of these patients, 86.22% (713/827) were male, with a median age of 34 years (ranging from 17 to 79), and 13.78% (114/827) were female, with a median age of 43 years (ranging from 17 to 69). Most of them (91.17%, 754/827) were diagnosed in the last two years (2017–2018). The dominant transmission route was men who have sex with men (MSM; 49.46%, 409/827), followed by heterosexual contact (HET; 39.66%, 328/827). The median (range) HIV-1 viral load was 520,801 (1035–6765,100) IU/mL. The median (range) CD4+ T cell count was 234 (1–1425) cells/mm3, and 42.32% (350/827) of the participants had a CD4+ T cell count of <200 cells/mm3. Based on partial pol sequences, HIV-1 genotypes were identified: CRF01_AE (37.97%), CRF07_BC (32.41%), CRF55_01B (12.33%), B (3.39%), CRF59_01B (2.66%), CRF08_BC (2.18%), Other (0.97%), and URF (8.10%).
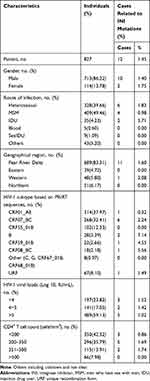 |
Table 1 Demographic and Viral Characteristics of 827 Study Population |
Of 827 successfully amplified and sequenced integrase sequences, no individual with major INSTI mutations was identified. Twelve individuals had INSTI accessory mutations. The overall prevalence of INSTI drug resistance mutations was 1.45% (12/827). The mutations were T97A (0.12%, 1/827), A128T (0.24%, 2/827), E157Q (0.85%, 7/827), and G163R (0.24%, 2/827). No sample with dual or triple INSTI drug resistance mutations was found in the study.
The characteristics of the twelve individuals harbouring INSTI mutations are shown in Table 2. Unequal distributions of mutations were found in the transmission routes, geographical region and genotype. The rates of INSTI mutations associated with the transmission routes were 5.71%, 1.83%, and 0.98% for IDU, HET, and MSM, respectively. The rates of INSTI mutations associated with the geographical region were 2.08% and 1.60% for the Western and Pear River Deltas, respectively. No INSTI mutations were detected in eastern and northern Guangdong. The rate of INSTI mutations among Subtype B was the highest (7.14%, 2/28), followed by CRF08_BC (5.56%, 1/18), CRF59_01B (4.55%, 1/22), CRF07_BC (2.24%, 6/268), URF (1.49%, 1/67), and CRF01_AE (0.32%, 1/314).
 |
Table 2 Characteristics of Patients Harboring HIV-1 Integrase Inhibitor Mutations |
Discussion
Two patient samples contained the G163R mutation, which is polymorphic in subtype F viruses from ARV-naive patients but is otherwise non-polymorphic,21,22 and confers low-level resistance to elvitegravir (EVG) and RAL. One patient sample contained the T97A mutation, which is a polymorphic INSTI-selected mutation that, depending on subtype, occurs in 1% to 5% of viruses from untreated persons,23 has minimal effects on INSTI susceptibility but in combination with other major resistance mutations, conferring potential low-level resistance to EVG and RAL. Seven patient samples contained the E157Q mutation, which is a polymorphic mutation that confers potential low-level resistance, and it generally only occurs in combination with other high- or intermediate-level resistance mutations.24 Two patient samples contained the A128T mutation, which is a relatively nonpolymorphic possibly INSTI-selected mutation that does not appear to reduce INSTI susceptibility.
Updated guidelines containing INSTI regimens have been recommended for approximately 2 years in Guangdong, China. Drug resistance testing is not routinely performed for ART-naïve HIV-1-infected individuals. HIV-1 is characterized by a high degree of natural variability due to its high replication rate and lack of a proofreading mechanism in RNA viral polymerase reverse transcription and genetic recombination.12,25,26 Small genetic errors or mutations can lead to the development of drug resistance. In the present study, we found that INSTI drug resistance mutations remain uncommon in Guangdong, China. No mutations related to high-level or medial-level resistance to four INSTIs were found. G163R (0.24%, 2/827) confers low-level resistance to EVG and RAL. G163R was also detected in one patient (0.2%, 1/513) in Yunnan Province.19 Other INSTI accessory mutations found in this study do not reduce susceptibility to INSTIs. The low rates of INSTI drug resistance mutations may be due to natural polymorphisms in the integrase region of HIV-1 virus, and surveillance is also needed for the further use of INSTIs.
In this study, different rates of INSTI mutations among different genotypes were found (Table 2). It has been reported that different genotypes may develop different mutational pathways leading to varying levels of drug resistance. Natural polymorphisms may influence the development of resistance against INSTIs in different HIV-1 subtypes.27,28 Here, we found that the mutation rate of Subtype B was higher than that of other genotypes, but the total number and types of drug-resistant sites were limited. Further study will consider grouping the patients according to genotype and following them by using INSTI-containing regimens.
A limitation of this study is that we used Sanger dideoxy sequencing to identify established clinically significant drug resistance mutations. Although Sanger sequencing is the gold standard method in HIV drug resistance testing, it fails to identify drug resistance minority variants that are below 20% of the virus population.1,29,30 The method probably underestimates the real prevalence of resistance mutations among treatment-naïve patients.
Conclusion
In conclusion, our results demonstrate that drug resistance mutations associated with INSTIs, including T97A, A128T, E157Q, and G163R, were detected among ART-naïve HIV-1-infected adults in Guangdong, China, in 2018. The overall prevalence of INSTI mutations remains low. Drug resistance mutation testing for the detection of INSTI drug resistance mutations in HIV treatment-naïve patients should be considered due to the circulation of polymorphisms contributing to INSTI resistance and the expected increasing use of this class of drugs.
Ethical Approval
This study was approved by the Institutional Review Board of the Guangzhou Eighth People’s Hospital (20171491), and written informed consent of the patient was obtained before collecting serum samples. A parent or legal guardian provided informed consent for any patients under the age of 18 years, and that this study was conducted in accordance with the Declaration of Helsinki.
Funding
This study was funded by the Guangzhou Science and Technology Plan Project (202002030028), and the National Major Scientific and Technological Project (2018ZX10302103-002, 2017ZX10202101-003 and 2018ZX10732101-001-014).
Disclosure
The authors report no conflicts of interest in this work.
References
1. Clutter DS, Jordan MR, Bertagnolio S, et al. HIV-1 drug resistance and resistance testing. Infect Genet Evol. 2016;46:292–307. doi:10.1016/j.meegid.2016.08.031
2. Rossouw TM, Nieuwoudt M, Manasa J, et al. HIV drug resistance levels in adults failing first-line antiretroviral therapy in an urban and a rural setting in South Africa. HIV Med. 2017;18(2):104–114. doi:10.1111/hiv.12400
3. World Health Organization: HIV drug resistance report; 2019 Available from: https://www.who.int/hiv/pub/drugresistance/hivdr-report-2019/en/.
4. Pommier Y, Johnson AA, Marchand C. Integrase inhibitors to treat HIV/AIDS. Nat Rev Drug Discov. 2005;4(3):236–248. doi:10.1038/nrd1660
5. Chen Q, Cheng X, Wei D, et al. Molecular dynamics simulation studies of the wild type and E92Q/N155H mutant of elvitegravir-resistance HIV-1 integrase. Interdiscip Sci. 2015;7(1):36–42. doi:10.1007/s12539-014-0235-8
6. AIDS and Hepatitis C Professional Group, Society of Infectious Diseases, Chinese Medical Association; Chinese Center for Disease Control and Prevention. Chinese guidelines for diagnosis and treatment of HIV/AIDS (Version 2018). Zhonghua Nei Ke Za Zhi. 2018;57(12):867–884. doi:10.3760/cma.j.issn.0578-1426.2018.12.002.
7. DeJesus E, Rockstroh JK, Lennox JL, et al. Efficacy of raltegravir versus efavirenz when combined with tenofovir/emtricitabine in treatment-naïve HIV-1-infected patients: week-192 overall and subgroup analyses from STARTMRK. HIV Clin Trials. 2012;13(4):228–232. doi:10.1310/hct1304-228
8. Raffi F, Jaeger H, Quiros-Roldan E, et al. Once-daily dolutegravir versus twice-daily raltegravir in antiretroviral-naïve adults with HIV-1 infection (SPRING-2 study): 96 week results from a randomised, double-blind, non-inferiority trial. Lancet Infect Dis. 2013;13(11):927–935. doi:10.1016/S1473-3099(13)70257-3
9. Gallant J, Lazzarin A, Mills A, et al. Bictegravir, emtricitabine, and tenofovir alafenamide versus dolutegravir, abacavir, and lamivudine for initial treatment of HIV-1 infection (GS-US-380-1489): a double-blind, multicentre, Phase 3, randomised controlled non-inferiority trial. Lancet. 2017;390(10107):2063–2072. doi:10.1016/S0140-6736(17)32299-7
10. Eron JJ, Cooper DA, Steigbigel RT, et al. Efficacy and safety of raltegravir for treatment of HIV for 5 years in the BENCHMRK studies: final results of two randomised, placebo-controlled trials. Lancet Infect Dis. 2013;13(7):587–596. doi:10.1016/S1473-3099(13)70093-8
11. Rockstroh JK, Lennox JL, Dejesus E, et al. Long-term treatment with raltegravir or efavirenz combined with tenofovir/emtricitabine for treatment-naïve human immunodeficiency virus-1-infected patients: 156-week results from STARTMRK. Clin Infect Dis. 2011;53(8):807–816. doi:10.1093/cid/cir510
12. Choi E, Mallareddy JR, Lu D, et al. Recent advances in the discovery of small-molecule inhibitors of HIV-1 integrase. Future Sci OA. 2018;4(9):FSO338. doi:10.4155/fsoa-2018-0060
13. Casadellà M, van Ham PM, Noguera-Julian M, et al. Primary resistance to integrase strand-transfer inhibitors in Europe. J Antimicrob Chemother. 2015;70(10):2885–2888. doi:10.1093/jac/dkv202
14. Rangel HR, Garzaro D, Fabbro R, et al. Absence of primary integrase resistance mutations in HIV type 1-infected patients in Venezuela. AIDS Res Hum Retroviruses. 2010;26(8):923–926. doi:10.1089/aid.2010.0039
15. Ji H, Patterson A, Taylor T, et al. Prevalence of primary drug resistance against HIV-1 integrase inhibitors in Canada. J Acquir Immune Defic Syndr. 2018;78(1):e1–e3. doi:10.1097/QAI.0000000000001649
16. Inzaule SC, Hamers RL, Noguera-Julian M, et al. Primary resistance to integrase strand transfer inhibitors in patients infected with diverse HIV-1 subtypes in sub-Saharan Africa. J Antimicrob Chemother. 2018;73(5):1167–1172. doi:10.1093/jac/dky005
17. Stekler JD, McKernan J, Milne R, et al. Lack of resistance to integrase inhibitors among antiretroviral-naïve subjects with primary HIV-1 infection, 2007–2013. Antivir Ther. 2014;20(1):77–80. doi:10.3851/IMP2780
18. Liu L, Dai L, Yao J, et al. Lack of HIV-1 integrase inhibitor resistance among 392 antiretroviral-naïve individuals in a tertiary care hospital in Beijing, China. AIDS. 2019;33(12):1945–1947. doi:10.1097/QAD.0000000000002282
19. Deng XM, Liu JF, Zhang M, et al. Mutations of primary integrase gene resistance of HIV/AIDS patients in Yunnan province. Clin J AIDS STD. 2019;25(4):327–341. doi:10.13419/j.cnki.aids.2019.04.01
20. Van Laethem K, Schrooten Y, Covens K, et al. A genotypic assay for the amplification and sequencing of integrase from diverse HIV-1 group M subtypes. J Virol Methods. 2008;153(2):176–181. doi:10.1016/j.jviromet.2008.07.008
21. Blanco JL, Varghese V, Rhee SY, Gatell JM, Shafer RW. HIV-1 integrase inhibitor resistance and its clinical implications. J Infect Dis. 2011;203(9):1204–1214. doi:10.1093/infdis/jir025
22. Boyd SD, Maldarelli F, Sereti I, et al. Transmitted raltegravir resistance in an HIV-1 CRF_AG-infected patient. Antivir Ther. 2011;16(2):257–261. doi:10.3851/IMP1749
23. Rhee SY, Gonzales MJ, Kantor R, et al. Human immunodeficiency virus reverse transcriptase and protease sequence database. Nucleic Acids Res. 2003;31(1):298–303. doi:10.1093/nar/gkg100
24. Bradley-Stewart A, Urcia C, MacLean A, Aitken C, Gunson R. HIV-1 integrase inhibitor resistance among treatment naïve patients in the West of Scotland. J Clin Virol. 2017;92:7–10. doi:10.1016/j.jcv.2017.04.012
25. Korber B, Gaschen B, Yusim K, Thakallapally R, Kesmir C, Detours V. Evolutionary and immunological implications of contemporary HIV-1 variation. Br Med Bull. 2001;58(1):19–42. doi:10.1093/bmb/58.1.19
26. Neher RA, Leitner T, Fraser C. Recombination rate and selection strength in HIV intra-patient evolution. PLoS Comput Biol. 2010;6(1):e1000660. doi:10.1371/journal.pcbi.1000660.
27. Han YS, Mesplède T, Wainberg MA. Differences among HIV-1 subtypes in drug resistance against integrase inhibitors. Infect Genet Evol. 2016;46:286–291. doi:10.1016/j.meegid.2016.06.047
28. Depatureaux A, Mesplède T, Quashie P, et al. HIV-1 Group O resistance against integrase inhibitors. J Acquir Immune Defic Syndr. 2015;70(1):9–15. doi:10.1097/QAI.0000000000000698
29. Van Laethem K, Theys K, Vandamme AM. HIV-1 genotypic drug resistance testing: digging deep, reaching wide? Curr Opin Virol. 2015;14:16–23. doi:10.1016/j.coviro.2015.06.001
30. Schuurman R, Brambilla D, de Groot T, et al. Underestimation of HIV type 1 drug resistance mutations: results from the ENVA-2 genotyping proficiency program. AIDS Res Hum Retroviruses. 2002;18(4):243–248. doi:10.1089/088922202753472801
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
