Back to Journals » Infection and Drug Resistance » Volume 12
A novel plasmid carrying carbapenem-resistant gene blaKPC-2 in Pseudomonas aeruginosa
Authors Hu YY , Wang Q, Sun QL , Chen GX, Zhang R
Received 29 November 2018
Accepted for publication 26 February 2019
Published 14 May 2019 Volume 2019:12 Pages 1285—1288
DOI https://doi.org/10.2147/IDR.S196390
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Joachim Wink
Yan-Yan Hu, Qi Wang, Qiao-Ling Sun, Gong-Xiang Chen, Rong Zhang
2nd Affiliated Hospital of Zhejiang University, School of Medicine, Zhejiang University, Hangzhou, People’s Republic of China
Abstract: A carbapenem-resistant Pseudomonas aeruginosa strain PA1011 (ST463) was isolated from a patient in a surgical intensive care unit. PCR detection showed that PA1011 carried the blaKPC-2 gene. A plasmid was isolated and sequenced using the Illumina NextSeq 500 and PacBio RSII sequencing platforms. The plasmid was named pPA1011 and carried the carbapenem-resistant gene blaKPC-2. pPA1011 was a 62,793 bp in length with an average G+C content of 58.8%. It was identified as a novel plasmid and encoded a novel genetic environment of blaKPC-2 gene (ΔIS6-Tn3-ISKpn8-blaKPC-2-ISKpn6-IS26).
Keywords: plasmid, genetic environment, KPC, ST463, Pseudomonas aeruginosa
Pseudomonas aeruginosa, one of the most common and important nosocomial bacteria, often causes infections associated with blood, urinary tract, skin and soft tissue. The morbidity and mortality due to P. aeruginosa infections were very high among the severe patients, especially those in the intensive care unit wards.1 Carbapenems have been the most efficient antibiotics for treating multidrug-resistant P. aeruginosa. However, with the extensive use of carbapenem, the incidence of carbapenem-resistant P. aeruginosa increased significantly. In Zhejiang, China, the imipenem and meropenem resistance rates of P. aeruginosa had climbed up to 39.0% and 27%, respectively.2 Plasmid-mediated carbapenemase gene is most risky due to its potential to disseminate rapidly among different strains and species. blaKPC-2 gene was first discovered and also most prevalent in Hangzhou, China.3,4 Our previous study has reported that a KPC-2-producing P. aeruginosa clone (ST463) spread in Hangzhou, China.5 In this study, a blaKPC-2-bearing plasmid was isolated from P. aeruginosa ST463 strain PA1011 and sequenced. And then ,the genetic structure of the plasmid was analyzed.
Strain PA1011 was isolated from the sputum of a 58-year-old male inpatient with gastroesophageal variceal hemorrhage caused by liver cirrhosis, and was identified to be P. aeruginosa by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (Bruker Daltonik GmbH, Bremen, Germany). The patient was admitted to digestive department of the Second Affiliated Hospital of Zhejiang University, School of Medicine (Hangzhou, China) on October 2010, and then transferred to surgical intensive care unit. He had a surgery, followed by antimicrobial treatment and mechanical ventilation during hospitalization, and died after 20 days in hospital. Levofloxacin, imipenem and teicoplanin were used before strain PA1011 was isolated. The detailed antimicrobial treatment and bacterial isolation are described in Figure 1. Antimicrobial susceptibility test of PA1011 was performed by the micro-broth dilution method, and it was resistant to carbapenems, monobactams, fluoroquinolones, β-lactam combination agents, cephems, and only susceptible to lipopeptides and aminoglycosides.
Plasmid DNA of PA1011 was extracted using the QIAGEN Plasmid Midi Prep kit according to the manufacturer’s instructions (Valencia, CA, USA). Its sequence was determined using the Illumina NextSeq 500 (San Diego, CA, USA) and PacBio RSII (Pacific Biosciences) platforms. Raw PacBio reads were de novo assembled with SMRT analysis version 2.3 (HGAP3 algorithm) with default settings. The assembled sequence was corrected and polished with Illumina reads by Quiver v1, and closed with Circlator v. 1.5.1.6 The plasmid was annotated with the RAST tool (version 2.0)7 and Prokka (version 1.12.21).8 The plasmid incompatibility type was searched using the online tool PlasmidFinder (
Plasmid pPA1011 was a 62,793 bp in length, encoding 185 predicted open reading frames (ORFs) with an average G+C content of 58.8%. pPA1011 contains a typical backbone structure, including regions involved in replication initiation (repA), partitioning (parA), plasmid maintenance, and type IV secretion (vir gene cluster), but it could not be assigned into any of known incompatibility groups. BLAST analysis showed that pPA1011 was overall similar to p14057 (accession number: KY296095)9 with 99% identity and 82% query coverage. p14057 was a 51.66 kb plasmid isolated from P. aeruginosa of human in Nanjing, China, which also carries blaKPC-2 gene (Figure 2). The major differences between p14057 and pPA1011 were a region encoding phage-related proteins and a region encoding hypothetical proteins on pPA1011. In addition, pPA1011 encodes multiple mobile genetic elements including an ISKpn27, ISKpn6, a copy of Tn3-like transposon and two copies of IS26.
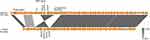
KPC enzymes were detected most frequently in K. pneumoniae and other Enterobacteriaceae species in Hangzhou, China.10,11 Our previous study has first detected KPC enzymes in P. aeruginosa ST463. ST463 was a novel sequence type of KPC-producing P. aeruginosa, and only reported in Hangzhou, China. The genetic environment surrounding blaKPC-2 in pPA1011 was identical to pKP048 (accession number: FJ628167)12 from K. pneumoniae and the previously reported plasmid pE1 from Escherichia coli,13 with the ORFs organized as follows: Tn3-transposase, Tn3-resolvase, ISKpn8, the blaKPC-2 gene, and ISKpn6 (Figure 3). It appeared to be a combination of a Tn3-based transposon and a partial Tn4401 transposon. This genetic content (Tn3-ISKpn8-blaKPC-2-ISKpn6) of blaKPC-2 has been found among different enterobacterial species in the hospital.14 Notably, the blaKPC-2-bearing fragment was flanked by DIS6 upstream and IS26 downstream, which may have facilitated the spread of blaKPC-2 gene cassette in P. aeruginosa in Zhejiang, China.
 | Figure 3 Schematic representation of the genetic environment surrounding the blaKPC-2 gene on pPA1011. Genes and their corresponding transcription orientations are indicated by colored arrows. Shading denotes regions of homology (100% nucleotide identity). The accession numbers of pKP048, p14057 and Tn4401 for reference are KT362706, KY296095 and FJ628167, respectively. The sequence of pE1 from Copyright©2014. American Society for Microbiology. Adapted from Cai JC, Zhang R, Hu YY, Zhou HW, Chen GX. Emergence of Escherichia coli sequence type 131 isolates producing KPC-2 carbapenemase in China. Antimicrob Agents Chemother. 2014;58(2):1146–1152.13 |
Conclusion
This study presented the complete sequence of a novel plasmid pPA1011 in KPC-producing ST463 P. aeruginosa, an important nosocomial pathogen. The findings provided a deep insight into the mechanism of horizontal genetic transfer of blaKPC-2 in P. aeruginosa. pPA1011 carried a novel genetic environment of blaKPC-2 gene, which should be closely monitored to control the rapid spread of blaKPC-2 gene in P. aeruginosa.
Acknowledgment
This work was funded by the National Natural Science Foundation of China (No. 81501805 and No. 81871705) and Natural Science Foundation of Zhejiang Province (No. LQ16H200002).
Ethics approval
Not applicable.
Consent for publication
Not applicable.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Lister PD, Wolter DJ, Hanson ND. Antibacterial-resistant Pseudomonas aeruginosa: clinical impact and complex regulation of chromosomally encoded resistance mechanisms. Clin Microbiol Rev. 2009;22(4):582–610. doi:10.1128/CMR.00040-09
2. Xie XY, Yu YS, Zhang R. Annual Review of Hospital Infection Resistance Survey in Zhejiang Province.
3. Wei ZQ, Du XX, Yu YS, Shen P, Chen YG, Li LJ. Plasmid-mediated KPC-2 in a Klebsiella pneumoniae isolate from China. Antimicrob Agents Chemother. 2007;51(2):763–765. doi:10.1128/AAC.01053-06
4. Zhang R, Zhou HW, Cai JC, Chen GX. Plasmid-mediated carbapenem-hydrolysing beta-lactamase KPC-2 in carbapenem-resistant Serratia marcescens isolates from Hangzhou, China. J Antimicrob Chemother. 2007;59(3):574–576. doi:10.1093/jac/dkl541
5. Hu YY, Gu DX, Cai JC, Zhou HW, Zhang R. Emergence of KPC-2-producing Pseudomonas aeruginosa sequence type 463 isolates in Hangzhou, China. Antimicrob Agents Chemother. 2015;59(5):2914–2917. doi:10.1128/AAC.04903-14
6. Hunt M, Silva ND, Otto TD, Parkhill J, Keane JA, Harris SR. Circlator: automated circularization of genome assemblies using long sequencing reads. Genome Biol. 2015;16:294. doi:10.1186/s13059-015-0667-4
7. Overbeek R, Olson R, Pusch GD, et al. The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic Acids Res. 2014;42(Database issue):D206–D214. doi:10.1093/nar/gkt1226
8. Seemann T. Prokka: rapid prokaryotic genome annotation. Bioinformatics. 2014;30(14):2068–2069. doi:10.1093/bioinformatics/btu153
9. Shi L, Liang Q, Feng J, et al. Coexistence of two novel resistance plasmids, blaKPC-2-carrying p14057A and tetA(A) -carrying p14057B, in Pseudomonas aeruginosa. Virulence. 2018;9(1):306–311. doi: 10.1080/21505594.2017.1372082
10. Nordmann P, Cuzon G, Naas T. The real threat of Klebsiella pneumoniae carbapenemase-producing bacteria. Lancet Infect Dis. 2009;9(4):228–236. doi:10.1016/S1473-3099(09)70054-4
11. Nordmann P, Naas T, Poirel L. Global spread of Carbapenemase-producing Enterobacteriaceae. Emerg Infect Dis. 2011;17(10):1791–1798. doi:10.3201/eid1710.110655
12. Jiang Y, Yu D, Wei Z, Shen P, Zhou Z, Yu Y. Complete nucleotide sequence of Klebsiella pneumoniae multidrug resistance plasmid pKP048, carrying blaKPC-2, blaDHA-1, qnrB4, and armA. Antimicrob Agents Chemother. 2010;54(9):3967–3969. doi:10.1128/AAC.00137-10
13. Cai JC, Zhang R, Hu YY, Zhou HW, Chen GX. Emergence of Escherichia coli sequence type 131 isolates producing KPC-2 carbapenemase in China. Antimicrob Agents Chemother. 2014;58(2):1146–1152. doi:10.1128/AAC.00912-13
14. Cai JC, Zhou HW, Zhang R, Chen GX. Emergence of Serratia marcescens, Klebsiella pneumoniae, and Escherichia coli Isolates possessing the plasmid-mediated carbapenem-hydrolyzing beta-lactamase KPC-2 in intensive care units of a Chinese hospital. Antimicrob Agents Chemother. 2008;52(6):2014–2018. doi:10.1128/AAC.01539-07
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.