Back to Journals » OncoTargets and Therapy » Volume 13
A Combination of circRNAs as a Diagnostic Tool for Discrimination of Papillary Thyroid Cancer
Authors Shi E, Ye J, Zhang R, Ye S, Zhang S, Wang Y , Cao Y, Dai W
Received 30 January 2020
Accepted for publication 27 April 2020
Published 19 May 2020 Volume 2020:13 Pages 4365—4372
DOI https://doi.org/10.2147/OTT.S247796
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Federico Perche
Erlan Shi, Jun Ye, Rong Zhang, Shuai Ye, Suwan Zhang, Yunsheng Wang, Yonghong Cao, Wu Dai
Department of Endocrinology, Hefei Hospital Affiliated to Anhui Medical University, The Second People’s Hospital of Hefei, Hefei 230011, People’s Republic of China
Correspondence: Wu Dai; Yonghong Cao
Department of Endocrinology, Hefei Hospital Affiliated to Anhui Medical University, The Intersection of Guangde Road and Leshui Road, Hefei 230011, People’s Republic of China
Email [email protected]; [email protected]
Background and Aim: Circular RNAs (circRNAs) have been highlighted to exert essential biological functions in papillary thyroid cancer (PTC). The purpose of this study was explore diagnostic utility of circRNAs in PTC patients.
Patients and Methods: The distinctive expression profile of serum circRNAs was determined by individual quantitative real-time PCR (qRT-PCR) in two independent cohorts of 113 PTC patients, 80 thyroid nodules, and 111 healthy controls (HCs). A combination of circRNAs (circRNA-based combination index) was constructed by logistic regression.
Results: Individual qRT-PCR identification showed that two circRNAs (circRAPGEF5 and hsa_circ_0058124) were significantly up-regulated in PTC patients compared with HCs and thyroid nodules. Receiver-operating characteristic (ROC) curve analysis suggested that a combination of circRNAs was superior to individual circRNA in distinguishing PTC patients from HCs and thyroid nodules with area under ROC curve of more than 0.80. Furthermore, the combination of circRNAs increased significantly after systematic treatment, suggesting that it could monitor PTC dynamics. Additionally, the combination of circRNAs was independently correlated with PTC presence.
Conclusion: The combination of these altered circRNAs was correlated with PTC and may serve as a novel diagnostic approach.
Keywords: circular RNA (circRNA), papillary thyroid cancer (PTC), thyroid nodule, diagnosis, circRAPGEF5, hsa_circ_0058124
Introduction
Thyroid cancer (TC) is one of the common endocrine malignancies worldwide, and its incidence has increased rapidly over recent decades, especially papillary TC (PTC).1,2 PTC is the most common histological type of TC and accounts for about 85% of all TCs. Despite synchronous lymph node metastases in around 30% of PTCs, its 5-year overall survival (OS) rate is almost 98% (Howlader et al, SEER Cancer Statistics Review, 1975–2012. National Cancer Institute, Bethesda, 2015). However, only 59% of patients with progressed TC have a 5-year OS rate.3 Although fine-needle aspiration (FNA) presently guides therapeutic strategies of patients with a thyroid nodule, the prevalence of indeterminate and non-diagnostic cytology from FNA is still high at around 30%.3 Therefore, more efforts are focused on trying to identify a new approach for the clinical assessment of advanced cancer.
Amounting evidence shows that epigenetic molecular markers may improve clinical assessment of many diseases.4,5 Various circular RNAs (circRNAs) have been verified by genome-wide analyses,6 which are involved in many pathological processes through transcriptional and post-transcriptional regulatory mechanisms.7 However, the system characteristics of circRNAs in PTC patients remain unclear. Previous reports have demonstrated that circRAPGEF5, hsa_circ_0058124, circRNA_102171 and circBACH2 contribute to PTC proliferation and metastasis though a similar mechanism of competing endogenous RNA.8–10 However, clinical significance of these circRNAs remains unknown.
In this study, we firstly detected the differential expression of four circRNAs in the training cohort of 52 PTC patients, 38 patients with thyroid nodules and 50 healthy controls (HCs). We then confirmed these findings in the test cohort of 61 PTC patients, 42 patients with thyroid nodules and 61 HCs, and showed their temporal profiles at pre- and post-treatment. Furthermore, we investigated the combined diagnostic and predictive power of circRAPGEF5 and hsa_circ_0058124 in PTC patients.
Patients and Methods
Patients and Samples
The study design was approved by the Medical Ethics Committee of Hefei Hospital affiliated to Anhui Medical University (Hefei, China). We obtained the written informed consent from all enrolled participants. Tissue and blood samples were processed at Hefei Hospital affiliated to Anhui Medical University and followed with ethical and legal standards. This study was also conducted in accordance with the Declaration of Helsinki.
A total of 113 PTC patients, 80 patients with thyroid nodules and 111 HCs were collected in this study and randomly divided into the training and test cohorts. The training cohort included 52 PTC patients, 38 patients with thyroid nodules and 50 HCs to screen nominated circRNAs. The test cohort enrolled 61 PTC patients, 42 thyroid nodules and 61 HCs to confirm the expression of screened circRNAs and their potential association with the risk of PTC. In brief, a total of 113 serum samples were collected from all participants with PTC receiving surgery at Hefei Hospital affiliated to Anhui Medical University and The People’s Hospital of Chizhou with a definite pathological diagnosis of PTC during surgery. Patients with PTC did not undergo radiotherapy, chemotherapy, and blood transfusion before surgery. Clinical features of all enrolled patients were recorded including sex, age, and tumor staging in Table 1.
 |
Table 1 Clinical Characteristics of Patients Enrolled in the Training and Test Cohorts |
Blood Samples
Blood samples were obtained from 113 PTC patients at 2 h before surgery and 7 days after operation, respectively. Furthermore, we collected 63 samples from PTC patients after treatment. A total of 113 samples were collected from newly diagnosed PTC patients, and 80 patients with thyroid nodules were also collected. Additionally, 111 healthy volunteers were enrolled as the referred group who were from medical examination without evidence of malignancies. Serum samples were isolated from blood samples after being centrifuged at 3000 rpm for 10 min and were stored at −80 °Cuntil RNA extraction.
RNA Preparation and Quantitative Real-Time PCR
Total RNA was extracted from serum samples using TRIzol reagent (Invitrogen, MA, USA) as described previously.4 RNA integrity was determined by agarose gel electrophoresis (1% gel). cDNA was synthesized using the RNA-to-cDNA kit (Takara Vio, Dalian, China) according to the manufacturer’s instruction. CircRNA relative expression was determined using the ABI 7500 System (Applied Biosystems, USA) and a SYBR Green kit (Takara, Dalian, China) with the following qPCR cycling program: 45 cycles including denaturation at 95°C for 5 sec, annealing at 60 for 30 sec and extension at 72°C for 30 sec. GAPDH was considered as an internal control and RNA expression was calculated using the ΔCt method. All primers were synthesized by Sangon Biotech Co. Ltd. (Shanghai, China).
Statistical Analysis
All statistical analyses were conducted using GraphPad Prism software (version 4.0; Inc., CA, USA) and SPSS software (version 20.0; IBM Corp., NY, USA). Continuous variables are expressed as the median (interquartile range, IQR) or mean (S.D.) and compared by Student’s t test or the Mann–Whitney U-test (n=2 groups) as well as one-way ANOVA or the Mann–Whitney U-test (n>2 groups) as appropriate. Comparisons of categorical variables were conducted by the Chi-squared test or Fisher’s exact test. Discriminative capacity of circRNAs was assessed by receiver-operating characteristic (ROC) curves and their area under the ROC curve (AUC). Optimal cut-off values of circRNAs were determined by Youden’s index. Logistic regression model was used to determine circle RNAs’ regression coefficient (0.392 for circRAPGEF5 and 0.792 for hsa_circ_0058124), and a combination of circRNAs was calculated on this equation (0.392×ΔCtcircRAPGEF5+0.792×ΔCthsa_circ_0058124). The predictive ability of circlRNAs was further analyzed by logistic regression analyses. P<0.05 was considered statistically significant.
Results
Clinical Characteristics of the Study Participants
A total of 52 patients with PTC, 38 patients with thyroid nodules and 50 HCs were enrolled in the training cohort to screen the nominated circRNAs. To further verify the expression of screened circRNAs and their potential association with the risk of PTC, we conducted a case-control study including 61 PTC patients, 42 thyroid nodules and 61 HCs in the test cohort. The clinical characteristics of two cohorts are summarized in Table 1.
Biomarker Screening and Validation
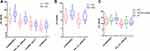
To investigate clinical significance of the four candidate circRNAs, their differential expression was firstly measured by qRT-PCR in the training cohort of 52 PTC patients and 50 HCs (Figure 1A). The results showed that circRAPGEF5 and hsa_circ_0058124 were differentially expressed in the serum samples of 52 PTC patients (P<0.05). We further analyzed the expression of circRAPGEF5 and hsa_circ_0058124 by qRT-PCR in 50 pre- and post-treatment serum samples from PTC patients (Figure 1B). These two circRNAs (circRAPGEF5 and hsa_circ_0058124) were obviously decreased after systematic treatment (P<0.05). Additionally, we also found that circRAPGEF5 and hsa_circ_0058124 exhibited increased expression in PTC patients compared with thyroid nodules (P<0.05, Figure 1C).
Stability of circRNAs Under Extreme Conditions
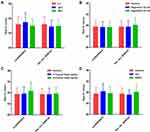
To explore the stability of two circRNAs as diagnostic biomarkers in PTC, serum samples were processed under different conditions, such as incubation at 4, −20, and −80°C for 4 h, digestion with RNase A, freeze thaw cycles repeatedly and acidic or alkaline conditions. Our results showed that the raw Ct values of circRAPGEF5 and hsa_circ_0058124 were not significantly fluctuated (Figure 2).
Training Cohort: Diagnostic Power of circRNAs in PTC
To study diagnostic power of circRAPGEF5 and hsa_circ_0058124 for PTC patients, we performed a ROC curve analysis in the training cohort. We found that the AUC was 0.711 for circRAPGEF5 and 0.790 for hsa_circ_0058124. Their sensitivity and specificity are detailed in Table 2. Furthermore, we observed that circRAPGEF5 presented the best diagnostic power with a cutoff value of 8.78 (Table 2). To assess the diagnostic ability of the combined circRNAs (circRAPGEF5 and hsa_circ_0058124) for PTC patients, we used the regression coefficient of circRAPGEF5 and hsa_circ_0058124 from a logistic regression model as weight values to construct a combination of circRNAs. The findings showed that the combination of circRNAs significantly decreased in PTC patients compared with HCs (P<0.05, Figure 3A), and it was superior to the individual circRNAs in PTC identification (Figure 3B and Table 2), suggesting that it had the greatest diagnostic ability with an AUC value of 0.860 (95% CI: 0.784–0.937). Taken together, a combination of circRNAs could precisely distinguish PTC patients from HCs. Additionally, we used 50 serum samples from 52 PTC patients after systematic treatment in the training cohort to investigate whether the combination of circRNAs fluctuated after systematic treatment. We observed that the combination of circRNAs significantly increased at post-treatment (P<0.001, Figure 3C), suggesting its capacity to supervise PTC dynamics.
 |
Table 2 Capacity of Two circRNAs Discriminating PTC from HC |
Test Cohort: Diagnostic Power of circRNAs in PTC
To further independently confirm the results from the training cohort, we enrolled a test cohort of 61 PTC patients and 61 HCs to investigate the levels of serum circRNAs. Similar to the findings in the training cohort, the AUC was 0.692 for circRAPGEF5 and 0.724 for hsa_circ_0058124 in the test cohort, and their sensitivity and specificity were ascertained by cutoff values in the training cohort (Table 2). Furthermore, the combination of circRNAs significantly decreased in PTC patients (P<0.001, Figure 4A), which was superior to that of circRNAs alone (P<0.05, Figure 4B). Additionally, a total of 30 serum samples were obtained after systematic treatment. The combination of circRNAs was significantly increased after treatment (P<0.001, Figure 4C), suggesting that it could monitor PTC dynamics.
To assess the correlation of combination of circRNAs with clinical characteristics, we re-analyzed PTC patients from the training and test cohorts. All PTC patients were divided into low and high groups according to the median of the combination of circRNAs. The findings showed that a low value combination of circRNAs was significantly correlated with TNM staging (P<0.001), lymph node metastasis (P<0.001) and distant metastasis (P=0.035, Table 3).
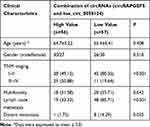 |
Table 3 Correlation of Combination of circRNAs with Clinical Characteristics in Two Cohorts |
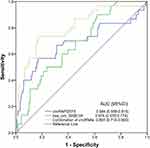
A diagnostic marker should not distinguish PTC patients from HC, but should also confirm PTC from other similar symptoms. To explore the diagnostic power of the two circRNAs for PTC patients compared with other benign diseases, we enrolled 80 patients with thyroid nodules to study their discriminatory power. The results showed that the combination of circRNAs could distinguish PTC from thyroid nodules with an AUC value of 0.807 (Figure 5), and the corresponding sensitivity and specificity were 0.821 and 0.640, respectively.
Predictive Capacity of circRNAs in the Training and Test Cohorts
To evaluate the predictive capacity of the two identified circRNAs for patients with PTC and thyroid nodules, we combined the two cohorts of patients and conducted a logistic regression analysis. Firstly, the HCs were defined as a reference group. The findings showed that increased circRAPGEF5, hsa_circ_0058124 and low combination of circRNAs were independently correlated with PTC presence after adjusting for age and sex (Table 4). Subsequently, patients with thyroid nodules were defined as a reference group. Increased circRAPGEF5, hsa_circ_0058124 and low combination of circRNAs were also independently correlated with PTC presence after adjustment (Table 4).
 |
Table 4 Logistic Regression Analysis for Predictive Ability of Two Identified circRNAs |
Discussion
Accumulating reports have suggested that control of the transcriptome is an essential mechanism responsible for tumorigenesis and progression of PTC.11 Among them, circRNAs play a critical role in both transcriptional and post-transcriptional regulation in the tumorigenesis and development of various malignancies.12 As a new focus of the regulation of transcriptome, circRNAs have been deeply investigated as diagnostic biomarkers with their molecular stability, substantial diversity and tissue specificity. circRNAs deepened our understanding of the complex regulated network of noncoding RNAs. To the best of our knowledge, the present study is the first attempt to explore diagnostic significance of combined multi-circRNAs in PTC. A panel of two dysregulated circRNAs was confirmed in this study.
The circRAPGEF5 is from 5 exons of RAPGEF5 gene and increased in PTC tissues, which functions as tumor promoter through miR-198/FGFR1 axis.8 Furthermore, hsa_circ_0058124 is also upregulated in PTC tissues and correlated with a poor prognosis for PTC patients, which functions as a miRNA sponge to regulate miRNA-218-5p/NOTCH3/GATAD2A axis.9 These reports explored their molecular mechanisms and prognostic significance in PTC. However, clinical significance of circRAPGEF5 and hsa_circ_0058124 in PTC patients remains unclear. The current study aimed to investigate their diagnostic value in PTC.
Our study successfully established a panel of two circRNAs (circRAPGEF5 and hsa_circ_0058124), which were differentially expressed in PTC patients by individual qRT-PCR validation. We also confirmed their distinct expression profiles in PTC, thyroid nodules, and healthy volunteers. Our findings showed that these differentiated circRNAs may predict and discriminate the presence of PTC from thyroid nodules and healthy volunteers. The signatures of circRAPGEF5 and hsa_circ_0058124 were obviously associated with tumor progression and reflected dynamic changes in PTC. Therefore, we revealed that these circRNAs may function as diagnostic biomarkers for PTC.
Our report may provide a novel approach into the diagnosis of PTC. However, some limitations should be acknowledged in the present study. First, as the number of patients to estimate the clinical significance of these circRNAs was limited, it is urgent to validate the correlation of circRAPGEF5 and hsa_circ_0058124 expression and PTC in large samples. Second, some heterogeneity are exhibited in the treatment used for PTC patients after surgical resection leading to different clinical outcomes.
Conclusions
In conclusion, this study applied a systematic strategy to verify a differential circRNAs expression profile in the serum samples of PTC patients. Two differentially expressed circRNAs manifested discriminative potential as novel markers for PTC, allowing tumor progression of PTC to be assessed. Further investigations are required to confirm clinical significance of serum circRNAs for PTC.
Abbreviations
PTC, papillary thyroid cancer; circRNA, circular RNA; qRT-PCR; quantitative real-time PCR; HC, healthy control.
Ethics and Consent Statement
The study design was approved by the Medical Ethics Committee of Hefei Hospital affiliated to Anhui Medical University (Hefei, China). We obtained the written informed consent from all enrolled participants. Tissue and blood samples were processed at Hefei Hospital affiliated to Anhui Medical University and followed with ethical and legal standards.
Author Contributions
WD, YHC, ELS, JY, RZ, SY, SWZ, YSW contributed to experimental design, procedure and data acquisition. DW, YHC, ELS contributed to data interpretation as well as manuscript drafting or revision. All authors contributed to data analysis, drafting or revising the article, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Miller KD, Siegel RL, Lin CC, et al. Cancer treatment and survivorship statistics, 2016. CA Cancer J Clin. 2016;66(4):271–289. doi:10.3322/caac.21349
2. Kitahara CM, Sosa JA. The changing incidence of thyroid cancer. Nat Rev Endocrinol. 2016;12(11):646–653. doi:10.1038/nrendo.2016.110
3. Kunavisarut T. Diagnostic biomarkers of differentiated thyroid cancer. Endocrine. 2013;44(3):616–622. doi:10.1007/s12020-013-9974-2
4. Deng QW, Li S, Wang H, et al. Differential long noncoding RNA expressions in peripheral blood mononuclear cells for detection of acute ischemic stroke. Clin Sci. 2018;132(14):1597–1614. doi:10.1042/CS20180411
5. Deng Q, He B, Gao T, et al. Up-regulation of 91H promotes tumor metastasis and predicts poor prognosis for patients with colorectal cancer. PLoS One. 2014;9(7):e103022. doi:10.1371/journal.pone.0103022
6. Yu T, Wang Y, Fan Y, et al. CircRNAs in cancer metabolism: a review. J Hematol Oncol. 2019;12(1):90. doi:10.1186/s13045-019-0776-8
7. Yin Y, Long J, He Q, et al. Emerging roles of circRNA in formation and progression of cancer. J Cancer. 2019;10(21):5015–5021. doi:10.7150/jca.30828
8. Liu W, Zhao J, Jin M, Zhou M. circRAPGEF5 contributes to papillary thyroid proliferation and metastatis by regulation miR-198/FGFR1. Mol Ther Nucleic Acids. 2019;14:609–616. doi:10.1016/j.omtn.2019.01.003
9. Yao Y, Chen X, Yang H, et al. Hsa_circ_0058124 promotes papillary thyroid cancer tumorigenesis and invasiveness through the NOTCH3/GATAD2A axis. J Exp Clin Cancer Res. 2019;38(1):318. doi:10.1186/s13046-019-1321-x
10. Bi W, Huang J, Nie C, et al. CircRNA circRNA_102171 promotes papillary thyroid cancer progression through modulating CTNNBIP1-dependent activation of beta-catenin pathway. J Exp Clin Cancer Res. 2018;37(1):275. doi:10.1186/s13046-018-0936-7
11. Abdullah MI, Junit SM, Ng KL, Jayapalan JJ, Karikalan B, Hashim OH. Papillary thyroid cancer: genetic alterations and molecular biomarker investigations. Int J Med Sci. 2019;16(3):450–460. doi:10.7150/ijms.29935
12. Chen Y, Li C, Tan C, Liu X. Circular RNAs: a new frontier in the study of human diseases. J Med Genet. 2016;53(6):359–365. doi:10.1136/jmedgenet-2016-103758
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.