Back to Journals » Clinical, Cosmetic and Investigational Dermatology » Volume 11
Transforming growth factor-β1 gene polymorphism in psoriasis vulgaris
Authors El-hadidi HH, Hassan AS, El-hanafy G , Amr KS, Abdelmesih SF , Abdelhamid MF
Received 17 April 2018
Accepted for publication 14 June 2018
Published 22 August 2018 Volume 2018:11 Pages 415—419
DOI https://doi.org/10.2147/CCID.S171403
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Jeffrey Weinberg
Heba H El-hadidi,1 Akmal S Hassan,1 Ghada El-hanafy,1 Khalda S Amr,2 Sameh F Abdelmesih,3 Mahmoud F Abdelhamid3
1Department of Dermatology, Kasr Al-Ainy University Hospitals, Cairo University, Giza, Egypt; 2Department of Molecular Genetics, National Research Center, Cairo, Egypt; 3Department of Dermatology, National Research Center, Cairo, Egypt
Background: Increased transforming growth factor beta 1 (TGF-β1) in the epidermis and serum has been found in psoriatic patients. The mechanism for this increase remains unclear.
Objective: To study the TGF-β1 gene polymorphism at codon 10 and its relation to psoriasis susceptibility in a sample of Egyptian patients.
Materials and methods: This cross-sectional study involved 70 patients with psoriasis vulgaris and 100 age- and sex- comparable healthy volunteers as a control group. Genomic DNA was prepared from peripheral blood lymphocytes from all subjects using QIAamp DNA mini kit (QIAGEN Inc., Germany). The TGF-β1 polymorphism was genotyped by PCR-based restricted fragment length polymorphism (PCR-RFLP) analysis. Amplification of codon 10, located in exon 1 of TGFβ1 gene was done through PCR reaction using gene-specific primers.
Results: Statistically significant difference was found between psoriasis patient and controls as regards TGF-β1 (T869C) polymorphism (P=0.045). The presence of TT genotype was associated with a 3-fold risk of psoriasis compared to CC genotype (P=0.016, OR: 3.13 95% CI: 1.24–7.88). T allele was significantly more frequent in psoriasis patients (P=0.017). TGF-β1 gene mutation was significantly higher among psoriasis patients with positive family history (P=0.007).
Conclusion: TGF-β1 gene polymorphism at codon 10 (T869C) is significantly associated with susceptibility to psoriasis in Egyptian patients. This polymorphism is more common in patients with a positive family history of psoriasis.
Keywords: psoriasis, TGF-β1, gene polymorphism
Introduction
Psoriasis is an immune-mediated inflammatory disease with a genetic basis. Inflammatory cells and their secreted products such as cytokines, chemokines and growth factors ultimately lead to keratinocyte hyperproliferation, epidermal thickening, and angiogenesis with marked ectasia of blood vessels.1
Transforming growth factor-β (TGF-β) is a multipotent cytokine that regulates cell growth and differentiation. Three isoforms of TGF-β (TGF-β1, 2 and 3) have been recognized in human tissues. TGF-β1 is the predominant isoform in the majority of tissues including the skin.2 The skin has been shown to be an important target tissue of TGF-β1, and its receptors are detected in epidermal keratinocytes.3 TGF-β1 has a contradicting role in the pathogenesis of psoriasis. It has been demonstrated to inhibit the growth of keratinocytes, but occasionally it stimulates keratinocyte proliferation due to a secondary effect of the increased inflammatory cytokines and chemokines in psoriasis IL-1, IL-6, IL-8. TGF-β1 also stimulates the growth of fibroblasts and induces angiogenesis and vasodilatation observed in early psoriasis.4–6 Strong evidence was provided that psoriasis-like skin inflammation is closely related to overexpression of latent TGF-β1 in the epidermis.3 Moreover, it mediates psoriasis-like lesions in mice.7
Increased TGF-β1 in the epidermis and the serum has been found in psoriatic patients, which was closely correlated with disease severity.8–10 However, the mechanism of increased serum levels of TGF-β1 in psoriasis remains unclear. In other diseases, TGF-β1 polymorphism significantly affects serum levels of TGF-β1.11,12
Regulation of cytokine levels has been shown to be under genetic control. Genetic polymorphisms in the coding and promoter sequences of their genes affect the rate of cytokine expression. Association of allelic variations in specific cytokine genes has been reported in several diseases including T-cell-mediated diseases of the skin.13
The human TGF-β1 gene is located on the long arm of chromosome 19. TGF-β1gene has some polymorphisms; two in the promoter region at positions 800 G/A and 509 C/T and three in the coding sequence at positions 869 T/C, 915 G/C and 1628 C/A. TGF-β1 gene polymorphism has been detected in many immune-mediated diseases such as systemic lupus erythematosus,14 systemic sclerosis,15 rheumatoid arthritis,16,17 Crohn’s disease18 and asthma,19 yet it remains to be determined whether these polymorphisms are linked with psoriasis.
This study aimed to assess the possible role TGF-β1 gene polymorphism has in psoriasis and its relation to psoriasis susceptibility in a group of Egyptian patients.
Materials and methods
This cross-sectional study involved 70 patients with psoriasis, and 100 healthy volunteers served as controls. All participants were recruited from the Dermatology Outpatient Clinics of Kasr-Al-Aini Hospital and National Research Center. The study was approved by the Dermatology Research Ethical Committee (DermaREC) of the Faculty of Medicine, Cairo University. A written informed consent was signed by each patient and control. The parents of the four patients under the age of 18 provided the written informed consent for participation of their children. Inclusion criteria were patients with psoriasis vulgaris who did not receive any treatment for at least 2 months before the recruitment to allow proper grading. Patients with other types of psoriasis and controls with a positive family history of psoriasis or other immune or inflammatory disorders were excluded.
All patients were subjected to careful history taking and clinical examination. The extent of psoriasis was determined using the Psoriasis Area and Severity Index (PASI) score.20 Accordingly, the patients were classified into mild (PASI<7), moderate (PASI 7–12) and severe psoriasis (PASI>12).
Detection of TGF-β1 gene polymorphism at codon 10
Three ml of venous blood were withdrawn under complete aseptic conditions from every patient and control subject. Blood was then collected in sterile ethylene diamine tetra-acetic acid (EDTA) vacationer tubes. Samples were either stored in the same vacationer at –20°C or used directly within 24 hours for detection of TGF-β1 gene polymorphism by PCR.
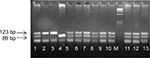
Genomic DNA was prepared from peripheral blood lymphocytes using QIAamp DNA mini kit (QIAGEN Inc., Germany). The TGF-β1 polymorphism was genotyped by PCR-based restricted fragment length polymorphism (PCR-RFLP) analysis. Amplification of codon 10 located in exon 1 of TGF-β1 gene was performed through PCR reaction using gene-specific primers.21 The presence of TGF-β1 polymorphism yielded specific products of 123 bp. The following primers were used: 5′-ACCACACCAGCCCTGTTCGC-3′ and the reverse primer sequence was 5′- AGTAGCCACAGCAGCGGTAGCAGCTGC-3′. PCR reaction conditions of 25 µL were: initial denaturation 94°C for 5 minutes; 35 cycles of 94°C for 30 seconds, 66°C for 30 seconds, and 72°C for 30 seconds; with final extension 72°C for 7 minutes. Amplified PCR product was analyzed for successful amplification on 2% agarose gel electrophoresis stained with ethidium bromide. RFLP analysis of succeeded PCR product was digested with 10 U of PstI-fast restriction enzyme (Fermentas, Waltham, MA, USA) at 37°C for 15 minutes. Digested PCR products were analyzed in a 3% agarose gel. The genotypes were recorded as Leu/Leu (TT), Leu/Pro (TC) and Pro/Pro (CC); CC genotype is the wild type.
Statistical analysis
Statistical analysis was done using IBM SPSS Statistics version 22 (IBM Corporation, Armonk, NY, USA). Numerical variables were described using mean and SD or median avnd range. Categorical variables were expressed as frequency and percentage. Comparison between two means was made using Student’s t-test and or Mann–Whitney test as appropriate. Chi-square or Fisher’s exact test was used to examine the relationship between categorical variables. Logistic regression was used for risk estimation expressed as odds ratio (OR) and its 95% CI. Hardy–Weinberg equilibrium (HWE) was tested for patients, controls and their combination using Chi-squared test. P-values <0.05 were determined as significant.
Results
The psoriasis group included 41 males (58.6%) and 29 females (41.4%) with a mean age of 42.5±13.4 years ranging between 16 and 70 years. The control group included 55 males (55%) and 45 females (45%) with a mean age of 30.6±6.5 years ranging between 18 and 40 years. Early-onset type one psoriasis (onset before 40 years) was found in 42 patients (60%), while 28 (40%) had late-onset type two disease. The disease duration ranged between 6 months and 50 years, with a median of 10 years. According to PASI score, 38 patients (54.3%) had mild, 19 (27.1%) had moderate, and 13 (18.6%) had severe disease. Ten patients (14.3%) had psoriatic arthritis and 48 (68.6%) of patients had a positive family history of psoriasis.
Genotyping of TGF-β1 codon 10 gene (Figure 1) is shown in Table 1. Psoriasis patients had a significantly lower proportion of the wild CC genotype (P=0.045). In a logistic regression model, the presence of TT genotype was associated with a 3-fold risk of psoriasis compared to CC genotype (P=0.016, OR: 3.13 95% CI: 1.24–7.88). On the other hand, TC genotype was not associated with a significant risk of disease (P=0.239). T allele was significantly more frequent in psoriasis patients (P=0.017).
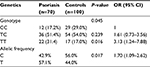  | Table 1 Genotyping and allelic frequency of TGF-β1 gene in psoriasis patients and control group |
TGF-β1 gene mutation was significantly higher among psoriasis patients with positive family history (P=0.007). Otherwise, TGF-β1 gene mutation was not associated with age, sex, onset, duration, and severity of disease, and psoriatic arthritis (Table 2). Genotype distribution was consistent with HWE (χ2=0.175, P=0.676 for patients), (χ2=0.917, P=0.338 for controls), (χ2=0.591, P=0.442 for combined group).
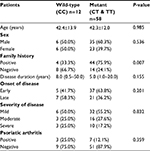  | Table 2 Relation between of TGF-β1 genotypes of psoriasis patients and the characteristics of patients and disease Notes: Data are presented as mean ± SD, median (range), or No. (%). |
Discussion
This study showed that T869C polymorphism of TGF-β1 was significantly associated with susceptibility to psoriasis (P=0.045). Homozygous mutation (TT) was associated with 3-fold risk of psoriasis, but heterozygous mutation (TC) was not. TGF-β1 gene mutation was significantly more common among patients with positive family history (P=0.007). The point mutation in position 869 at codon 10 involves alleles T (leucine) and C (proline). The latter denote that the frequency in genotypes among psoriatic patients and control groups are not similar, with the frequency of T allele, which represents high producer genotype (meaning increased liability to induce polymorphism), is more than C allele which represents low producer genotyping (meaning a decreased liability of polymorphism induction).
Our findings are different from a Polish study of TGF-β1 gene codon 10 (T/C) and codon 25 (G/C). The authors did not find a significant difference between psoriasis patients and controls in the two codons, ie, TGF-β1 gene polymorphism in codon 10 and 25 are not associated with susceptibility to psoriasis vulgaris.22 However, they reported a significant difference in TGF-β1 polymorphism between early and late onset psoriasis. In the current study, there was no significant difference between early and late onset psoriasis (P=0.201). We believe that this contradiction regarding genotypic differences between patients and controls and between early and late onset of the disease could be due to different genetic factors that can present between different ethnicities. Moreover, the etiopathogenesis of psoriasis is multifactorial.23
In the present study, we did not find a significant association between TGF-β1 gene codon 10 polymorphism and the presence of psoriatic arthritis (P=0.359). Interestingly, previous studies16,17 found an association between rheumatoid arthritis and polymorphism of TGFβ1 gene at nucleotide T869C.
Settin et al24 assessed cytokine gene polymorphisms in psoriatic Egyptian patients from the Nile delta. The authors found significantly higher frequencies of certain genotypes of different cytokines such as IL-6, IL-10 and TNF-α. They concluded that the genetic polymorphisms related to those cytokines showed a particular pattern of association with psoriasis. These results together with that of the current study may indicate that the presence of polymorphic genotypes of different cytokines in Egyptian psoriatic patients may play a role in the etiology of the disease.
In conclusion, the present study demonstrated that the TGF-β1 gene polymorphism of codon 10 at T869C nucleotide is associated with susceptibility to psoriasis in Egyptian patients. This polymorphism is more common in patients with a positive family history of psoriasis. Whether this polymorphism is associated with increased production of TGF-β1 detected in psoriasis with a subsequent increase in keratinocyte proliferation, angiogenesis, and asodilatation observed in psoriasis, is yet to be determined. Further studies are warranted to investigate the role of TGF-β1 in psoriatic skin and whether it can be considered as the basis of future gene therapy in the treatment of psoriasis.
Acknowledgments
The molecular sequence used deposited in GenBank (accession number: BC000125.1).
Disclosure
The authors report no conflicts of interest in this work.
References
Sabat R, Philipp S, Höflich C, et al. Immunopathogenesis of psoriasis. Exp Dermatol. 2007;16(10):779–798. | ||
Lawrence DA. Identification and activation of latent transforming growth factor beta. Methods Enzymol. 1991;198:327–336. | ||
Han SW, Kim TY, Hwang PG, et al. Predictive and prognostic impact of epidermal growth factor receptor mutation in non-small-cell lung cancer patients treated with gefitinib. J Clin Oncol. 2005;23(11):2493–2501. | ||
Bonifati C, Ameglio F. Cytokines in psoriasis. Int J Dermatol. 1999;38(4):241–251. | ||
Cutroneo KR. TGF-beta-induced fibrosis and SMAD signaling: oligo decoys as natural therapeutics for inhibition of tissue fibrosis and scarring. Wound Repair Regen. 2007;15(Suppl. 1):S54–S60. | ||
Han G, Williams CA, Salter K, Garl PJ, Li AG, Wang XJ. A role for TGFbeta signaling in the pathogenesis of psoriasis. J Invest Dermatol. 2010;130(2):371–377. | ||
Zhang Y, Meng XM, Huang XR, Wang XJ, Yang L, Lan HY. Transforming growth factor-β1 mediates psoriasis-like lesions via a Smad3-dependent mechanism in mice. Clin Exp Pharmacol Physiol. 2014;41(11):921–932. | ||
Flisiak I, Chodynicka B, Porebski P, Flisiak R. Association between psoriasis severity and transforming growth factor beta(1) and beta(2) in plasma and scales from psoriatic lesions. Cytokine. 2002;19(3):121–125. | ||
Flisiak I, Zaniewski P, Chodynicka B. Plasma TGF-beta1, TIMP-1, MMP-1 and IL-18 as a combined biomarker of psoriasis activity. Biomarkers. 2008;13(5):549–556. | ||
Meki AR, Al-Shobaili H. Serum vascular endothelial growth factor, transforming growth factor β1, and nitric oxide levels in patients with psoriasis vulgaris: their correlation to disease severity. J Clin Lab Anal. 2014;28(6):496–501. | ||
Akhurst RJ. TGF beta signaling in health and disease. Nat Genet. 2004;36(8):790–792. | ||
Mao JH, Saunier EF, de Koning JP, et al. Genetic variants of TGF-β1 act as context-dependent modifiers of mouse skin tumor susceptibility. Proc Natl Acad Sci USA. 2006;103(21):8125–8130. | ||
Arkwright PD, Pravica V, Geraghty PJ, et al. End-organ dysfunction in cystic fibrosis: association with angiotensin I converting enzyme and cytokine gene polymorphisms. Am J Respir Crit Care Med. 2003;167(3):384–389. | ||
Lu LY, Cheng HH, Sung PK, Yeh JJ, Shiue YL, Chen A. Single-nucleotide polymorphisms of transforming growth factor-beta1 gene in Taiwanese patients with systemic lupus erythematosus. J Microbiol Immunol Infect. 2004;37(3):145–152. | ||
Ohtsuka T, Yamakage A, Yamazaki S. The polymorphism of transforming growth factor-beta1 gene in Japanese patients with systemic sclerosis. Br J Dermatol. 2002;147(3):458–463. | ||
Sugiura Y, Niimi T, Sato S, et al. Transforming growth factor beta1 gene polymorphism in rheumatoid arthritis. Ann Rheum Dis. 2002;61(9):826–828. | ||
Zhou TB, Zhao HL, Fang SL, Drummen GP. Association of transforming growth factor-β1 T869C, G915C, and C509T gene polymorphisms with rheumatoid arthritis risk. J Recept Signal Transduct Res. 2014;34(6):469–475. | ||
Cantor MJ, Nickerson P, Bernstein CN. The role of cytokine gene polymorphisms in determining disease susceptibility and phenotype in inflammatory bowel disease. Am J Gastroenterol. 2005;100(5):1134–1142. | ||
Chiang CH, Chuang CH, Liu SL, Shen HD. Genetic polymorphism of transforming growth factor β1 and tumor necrosis factor α is associated with asthma and modulates the severity of asthma. Respir Care. 2013;58(8):1343–1350. | ||
Fredriksson T, Pettersson U. Severe psoriasis – oral therapy with a new retinoid. Dermatologica. 1978;157(4):238–244. | ||
Wu L, Chau J, Young RP, et al. Transforming growth factor-beta1 genotype and susceptibility to chronic obstructive pulmonary disease. Thorax. 2004;59(2):126–129. | ||
Baran W, Szepietowski JC, Mazur G, Baran E. TGF-beta(1) gene polymorphism in psoriasis vulgaris. Cytokine. 2007;38(1):8–11. | ||
Mir AM, Christiano AM, Rapini RP. Basics of genetics. Chapter 53. Bolognia Textbook of Dermatology. 2nd ed. Bolognia JL, Jorrizo JL, Rapini RP, editors. Spain: Mosby Elsevier; 2008:693–704. | ||
Settin AA, Hassan HA, El-Baz RA, Hassan TA. Association of cytokine gene polymorphisms with psoriasis in cases from the Nile delta of Egypt. Indian J Dermatol. 2011;56(3):272–277. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.