Back to Journals » Pharmacogenomics and Personalized Medicine » Volume 12
The influence of an IL-4 variable number tandem repeat (VNTR) polymorphism on breast cancer susceptibility
Authors AL-Eitan LN , Rababa'h DM, Alghamdi MA , Khasawneh RH
Received 24 June 2019
Accepted for publication 6 August 2019
Published 26 August 2019 Volume 2019:12 Pages 201—207
DOI https://doi.org/10.2147/PGPM.S220571
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Martin Bluth
Laith N AL-Eitan,1,2 Doaa M Rababa’h,1 Mansour A Alghamdi,3 Rame H Khasawneh4
1Department of Applied Biological Sciences, Jordan University of Science and Technology, Irbid 22110, Jordan; 2Department of Biotechnology and Genetic Engineering, Jordan University of Science and Technology, Irbid 22110, Jordan; 3Anatomy Department, Faculty of Medicine, College of Medicine, King Khalid University, Abha, Saudi Arabia; 4Department of Hematopathology, King Hussein Medical Center (KHMC), Jordanian Royal Medical Services (RMS), Amman 11118, Jordan
Correspondence: Laith N AL-Eitan
Jordan University of Science and Technology, P.O. Box 3030, Irbid 22110, Jordan
Tel + 962 2 720 1000 Ext. 23464
Fax + 962 2 720 1071
Email [email protected]
Backgrounds: Breast cancer (BC) is one of the most widespread cancers globally. Understanding the etiology of BC may help in determining the various risk factors involved in its malignancy. Certain genetic mutations are considered to play a key role in increasing the risk of BC.
Objectives: In this study, we explored the correlation between a variable number tandem repeat (VNTR) polymorphism in the IL-4 gene and BC.
Methods: PCR and subsequent gel electrophoresis were used to genotype this variant in 360 Jordanian women (180 BC patients and 180 controls). In addition, phenotype–genotype analysis was carried out.
Results: Our findings illustrate that there is no significant relationship between the variant genotypes in the IL-4 gene and BC among Jordanian females. Other than body mass index and tumor differentiation (p< 0.05), none of the clinical and pathological parameters of BC patients exhibited any association with the variant genotypes.
Conclusions: From this study, we propose that the IL-4 genetic variant does not impact BC development and progression but that it could influence the disease prognosis.
Keywords: breast, cancer, IL-4, genetic variation, prognosis
Introduction
According to the World Health Organization, breast cancer (BC) constitutes a major burden on worldwide female health and is the leading cause of death among women.1 However, survival rates among BC patients in the developed world are much higher than those in less developed countries.2 In 2012, the estimated number of women with BC was 651,000, accounting for more than a third of worldwide cancer cases.3,4 In Jordan, the mean age of BC diagnosis is 51 years, and it is the most common fatal cancer among Jordanian females.5
BC is a complex disease which is influenced by a range of factors, including the environment, sociobiology, family history, hormones, obesity, and the immune system of the individual.6,7 Despite the fact that only 5–6% of BC cases are inherited, genetic mutations are one of the most predictable risk factors that contribute to BC development, with many studies reporting different genetic variants in critical genes that may induce BC onset.8,9
One of the most frequently investigated genetic variants is the variable number tandem repeat (VNTR), a region in the genome with interindividual differences in length consisting of repeated nucleotides that lie adjacent to one another.10 Different studies have screened the VNTR in the IL-4 gene in order to detect if it is related to the progression of different cancers, including BC.11 IL-4 is a cytokine produced by T lymphocytes and is known for its antitumor ability, the latter of which is hypothesized to occur by inducing apoptosis in BC cells and regulating estrogen synthesis.12–14
Together with the published data on the subject, the aforementioned functions of IL-4 make it ideal for candidate gene association analysis.15–17 The IL-4 gene is located on chromosome 5 and contains a 70-bp VNTR polymorphism within intron 3 (Figure 1) that includes two common alleles: allele 1 (one repeat) and allele 2 (two repeats). These alleles are more frequent among the population than allele 3 (3 repeats), which is considered a rare variant.18,19
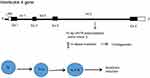 |
Figure 1 VNTR polymorphism locus within the IL-4 gene.Abbreviation: VNTR, variable number tandem repeat. |
In this case–control study, we aim to explore the association between the VNTR polymorphism in the IL-4 gene and the development of BC among Jordanian women. This study also investigates the relationship between clinicopathological parameters and disease progression among BC patients.
Materials and methods
Study design
This study involved 180 healthy Jordanian women with no family history of BC and 180 Jordanian BC patients, all of whom were randomly chosen and matched for gender and age. Written informed consent was obtained from the participants in this study. Ethical approval was obtained from the Institutional Review Board (IRB) at Jordan University of Science and Technology (ethical approval code number 32/104/2017). This study was also conducted in accordance with the Declaration of Helsinki. Furthermore, clinical and demographic information was obtained from BC patients. Blood samples were collected in cooperation with Jordanian Royal Medical Services (JRMS).
Molecular analysis
Each participant gave 5 mL of peripheral blood, and DNA was extracted using the QIAamp DNA Blood Mini Kit (Qiagen, Valencia, CA, USA). The quantity and purity of the extracted DNA were verified using the NanoDrop (Thermo Fisher Scientific Inc., DE, USA). PCR was used to genotype the IL-4 VNTR using a specific forward primer (5′AGGCTGAAAGGGGGAAAGC3′) and reverse primer (5′CTGTTCACCTCAACTGCTCC3′).20 The researchers added 50 ng of DNA to a 25 µL PCR tube containing 12.5 µL Master Mix, 10 µL deionized distilled water, and 2 µL of each of the forward and reverse primers. The PCR program consisted of an initial denaturation phase at 94°C for 5 mins followed by 30 cycles of denaturation at 94°C for 50 s, annealing at 61°C for 30 s, and extension at 72°C for 45 s. PCR was concluded by a final extension step at 72°C for 5 mins.21 The PCR product was separated and visualized using 2% agarose, and a 100 bp ladder was used to identify their sizes.
Statistical analysis
Frequency distribution for genotypes and alleles was statistically calculated. Additionally, Pearson’s Chi-squared test was used to calculate the differences between cases and controls.22 The genetic association and phenotype-genotype analyses were conducted using the Statistical Package for the Social Sciences (SPSS), version 25.0 (SPSS, Inc., Chicago, IL, USA). Statistical significance was set at p< 0.05.
Results
VNTR of the IL-4 gene
The genotypes of the VNTR within the IL-4 gene were analyzed using PCR and subsequent gel electrophoresis. Figure 2 shows the visualization of the resulting genotypes on the agarose gel. Three genotypes were detected in terms of their band sizes: allele 1 (1R) was present at a size of 183 bp and allele 2 (2R) at 253 bp, while the presence of the heterozygous alleles (2R\1R) was indicated by the appearance of both bands.
However, the genotypic and allelic frequency distribution of both the control and patient groups was analyzed and tested using Pearson’s Chi-squared test (Table 1). As Table 1 illustrates, 79.4% of patients had 2R on both copies of the IL-4 gene, which was not markedly different from the control group. Our results did not estimate the homozygous 1R genotype among patients, while only two healthy individuals carried the (1R\1R) genotype. The frequency of the heterozygous genotype (2R\1R) among patients was 26.7% compared to 19.5% in the control group; however, this was found to be nonsignificant (p= 0.303). Moreover, the allelic distribution of 1R and 2R genotypes was not significantly different between controls and patients (Table 1). Moreover, genetic association analysis using genetic models was performed. 2R/1R vs 2R/2R was the only applicable model and revealed no association between IL-4 VNTR and BC (OR: 0.67, 95% CI: 0.41–1.11, χ2: 2.47).
 |
Table 1 Genotypic and allelic frequency distribution of the VNTR in the IL-4 gene for cases and controls |
In addition to genetic association analysis, in this study, the correlation between the clinicopathological parameters and the VNTR of IL-4 was investigated. Table 2 demonstrates the relationship between a number of clinical features and the VNTR genotypes of BC patients. We found that 66% of BC patients were older than 45 years and had the 2R\2R genotype, while only 33% of the cases were under the age of 45 and had the 2R\2R genotype. However, this difference between the two groups was not statistically significant (p= 0.8).
 |
Table 2 Association of IL-4 polymorphism with clinical characteristics of breast cancer patients |
Other clinical features, such as age at first pregnancy and age at menarche, were also explored in terms of the relationship between exposure to endogenous estrogen and different VNTR genotypes among BC patients. In the present study, we found that the only critical clinical parameter in the context of the VNTR genotypes was body mass index (BMI). Most patients with a BMI of more than 25 and double 2R genotypes (79%) were far more frequent than patients who had only one 2R and one 1R (p= 0.035). Additionally, the relationship between the pathological characteristics exhibited by BC patients and VNTR genotypes was studied (Table 3). In this study, we found none of the features to be critical for BC risk among the different VNTR genotypes, except for tumor differentiation (p= 0.028).
 |
Table 3 Association of IL-4 polymorphism with pathological characteristics of breast cancer patients |
Discussion
BC is a serious problem that affects both genders, but it is particularly common among females.1,4 While the cause of BC has yet to be definitively determined, it is believed to be triggered by a combination of many risk factors, including environment, physiology, and individual genetics.2,23 With regard to the genetic component of the disease, several studies have investigated the relationship between different genetic polymorphisms and the increased risk of developing BC in different populations.24–27 For example, many studies have been conducted on the Jordanian population to investigate the genetic predisposition to BC. A range of genetic variants within significant genes have been explored among Jordanian BC patients, such as the IL-10 haplotype,28 ERα PvuII and XbaI polymorphisms,29 BRCA1/230,31 and RAD51.32
In the present study, we screened 180 healthy Jordanian women and 180 Jordanian BC patients to determine the relationship between the VNTR of a 70-bp sequence located in intron 322,23 within the IL-4 gene and BC. Two different alleles were detected within the VNTR region of the IL-4 gene: 2 repeats and 3 repeats. IL-4 is a pleiotropic cytokine secreted by a wide range of epithelial cells, including BC cells. This cytokine has been implicated in the progression of BC and resistance to common therapeutic regimens driven by tumor-initiating cells (TICs).33 The distribution frequency of the IL-4 VNTR alleles varies among different populations, for example, the estimated frequencies for (1R, 2R) in China,34 Malaysia,11 North India35, and Taiwan36 were: (76.9%, 23.1%), (62.81%, 37.19%), (21.7%, 78.8%), respectively. In the current study, we found that 1R frequency among healthy Jordanian was 13.3%, while the 2R frequency was 86.7%.
The data related to IL-4 VNTR and BC are very scarce. However, the 1R allele of the IL-4 VNTR has been assigned as a risk factor for malignancy development, and in Taiwan 1R allele of IL-4 VNTR was associated with bladder36 and oral cancers in China.37 In addition, the 1R/2R genotype of IL-4VNTR increased the risk of developing cervical cancer among North Indian women.38
In a recent study, Ibrahimi et al suggested that RP2\RP2 genotypes might be a potent protective factor against BC.39 In addition, Jia et al40 examined in a comprehensive meta-analysis the correlation between different types of genetic variants within the IL-4 gene, including the VNTR, and cancer risk. In their analysis, only one study among Indian women reported a significant correlation between IL-4 VNTR polymorphism and BC risk. That study stands in contrast to our results, which suggest that the IL-4 VNTR is not associated with BC risk among Jordanian women.
In the present study, we described a group of the clinical and pathological features that might be associated with increased BC risk among Jordanian women. Moreover, we speculated regarding the correlation between these parameters and the VNTR of IL-4. Our results reveal that 33.8% of patients under the age of 45 had the homozygous 2R genotype, which is comparable to the proportion of patients with the heterozygous 2R\1R genotype at the same age (32.6%). Furthermore, age at first pregnancy (<20) and age at first menstruation (<13) were analyzed to detect the effect of endogenous estrogen. Though patients with the 2R\2R genotype were evidently more numerous than patients with the 1R\2R genotype at the critical age, there was no statistically significant association between each of the age at first pregnancy and the age at first menstruation on the one hand, and BC on the other. Moreover, 34% of patients with a family history of BC possessed the 2R\2R genotype, while 26.1% of such patients had the 1R\2R genotype. Despite this, we suggest that family history did not interfere with IL-4 polymorphism in this study.
On the other hand, the relationship between allergy and BC has not been fully elucidated. Hedderson et al41 declared that allergy history may be linked with reduced risk of BC for women who were diagnosed early. Although we did not find any significant relationship between BC and allergy, the estimated frequency of patients with a history of allergy and who possessed the 2R\2R genotype (29.8%) was lower than patients with the same genotype but without any history of allergy (70.2%). Otherwise, we could not assess any correlation between BC and the remaining clinical characteristics of menstrual condition, breastfeeding status, smoking, and comorbidity. Remarkably, the only clinical characteristic to show any significant association with BC was BMI. Of patients with the 2R\2R genotype, 79% had a BMI >25, while 60.9% of patients with a BMI >25 had the 2R\1R genotype. Correspondingly, we can mark BMI as a factor that may influence BC prognosis among patients who carry or are susceptible to the IL-4 variant.
Similarly, we analyzed the pathological factors that may influence BC risk. Molecular receptors such as progesterone receptor (PR) and estrogen receptor (ER) status were examined. (ER) is one of the crucial predictive markers in BC prognosis and in anticipating response to hormone therapies.42 We found that, of the patients carrying the 2R\2R genotype, 54.7% and 79.50% were screened with positive progesterone and ERs, respectively, which is slightly higher than among patients carrying the 1R\2R genotype.
Moreover, patients were placed into two different groups (low differentiation and medium to high differentiation) based on the rate of cancer progression. The variation between patients carrying the 2R\2R and 1R\2R genotypes was striking, as 70.5% of patients with medium and high differentiation had the 2R\1R genotype compared to 66.4% of those with the 2R\2R genotype. Correspondingly, our result revealed an association between tumor differentiation of BC and IL-4 VNTR, suggesting that the 2R allele reduces the risk of BC. Other parameters, such as tumor size, lymph node involvement, and histology classification, did not show any significant differences between the different VNTR genotypes among BC patients. However, in this study, tumor differentiation was the only pathological factor to show a significant relationship with IL-4 VNTR.
In conclusion, there is no significant relationship between the VNTR polymorphism within the IL-4 gene and BC among Jordanian women. Furthermore, BMI and the tumor differentiation parameters of BC were associated with the variant genotypes. To the best of our knowledge, this study is one of only a few in the Arab world to tackle the genetic component of BC in an Arab, and particularly the Jordanian, population. Due to the rapid rise of BC incidence in the developing world, more studies must be carried out in order to enhance survival rates by tailoring treatment regimens to the individual. Together with the established prognostic factors, the selection of BC genetic markers can help facilitate the rise of personalized medicine in the Arab world.
Data availability
The datasets generated and/or analyzed over the course of the study are not publicly available but are available from the corresponding author on reasonable request.
Ethical approval and informed consent
All procedures performed in studies involving human participants were in accordance with the ethical standards of the IRB at Jordan University of Science and Technology with ethical code number 32/104/2017. Informed consent was obtained from all individual participants included in the study.
Acknowledgment
The authors thank the JRMS, Amman, Jordan, for approving this study in the first instance and making the clinical data and samples available for the study. This study was funded by the Deanship of Research (RN: 126/2017), Jordan University of Science and Technology.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Bhikoo R, Srinivasa S, Yu TC, Moss D, Hill AG. Systematic review of breast cancer biology in developing countries (part 2): asian subcontinent and South East Asia. Cancers (Basel). 2011;3(2):2382–2401. doi:10.3390/cancers3022382
2. McPherson K, Steel CM, Dixon JM. ABC of breast diseases. Breastcancer-epidemiology, risk factors, and genetics. BMJ. 2000;321(7261):624–628. doi:10.1136/bmj.321.7261.624
3. Tang J, Zhou Q, Zhao F, et al. Association of glutathione S-transferase T1, M1 and P1 polymorphisms in the breast cancer risk: a meta-analysis in Asian population. Int J Clin Exp. 2015;8(8):12430–12447.
4. Youlden DR, Cramb SM, Yip CH, Baade PD. Incidence and mortality of female breast cancer in the Asia-Pacific region. Cancer Biol Med. 2014;11(2):101–115. doi:10.7497/j.issn.2095-3941.2014.02.005
5. [Internet] Ministry of Health, Jordan Cancer Registry. Cancer Incidence in Jordan; 2012, Available from: http://www.moh.gov.jo/.
6. Verma R, Bowen RL, Slater SE, Mihaimeed F, Jones JL. Pathological and epidemiological factors associated with advanced stage at diagnosis of breast cancer. Br Med Bull. 2012;103(1):129–145.
7. Angahar LT. An overview of breast cancer epidemiology, risk factors, pathophysiology, and cancer risks reduction. MOJ Biol Med. 2017;19(4):1–5.
8. Zhang B, Beeghly-Fadiel A, Long J, Zheng W. Genetic variants associated with breast cancer risk: comprehensive field synopsis, meta-analysis, and epidemiologic evidence. Lancet Oncol. 2011;12(5):477–488.
9. Han MR, Zheng W, Cai Q, et al. Evaluating genetic variants associated with breast cancer risk in high and moderate-penetrance genes in Asians. Carcinogenesis. 2017;38(5):511–518.
10. Leclerc M, Neuhausen SL, Schayek H, Laitman Y, Antonis AC, Friedman E. Are VNTRs co-localizing with breast cancer-associated SNPs? Breast Cancer Res Treat. 2017;168(1):277–281. doi:10.1007/s10549-017-4588-7
11. Vasudevan R, Norhasniza MN, Patimah I. Association of variable number of tandem repeats polymorphism in the IL-4 gene with end-stage renal disease in Malaysian patients. Genet Mol Res. 2011;10(2):943–947. doi:10.4238/vol10-2gmr1066
12. Nagai S, Toi M. Interleukin-4 and breast cancer. Breast Cancer. 2000;7(3):181–186.
13. Nelms K, Keegan AD, Zamorano J, Ryan JJ, Paul WE. The IL-4 receptor: signaling mechanisms and biologic functions. Annu Rev Immunol. 1999;17:701–738. doi:10.1146/annurev.immunol.17.1.701
14. Gyan BA, Goka B, Cvetkovic JT, et al. Allelic polymorphisms in the repeat and promoter regions of the interleukin-4 gene and malaria severity in Ghanaian children. Clin Exp Immunol. 2004;138:145–150. doi:10.1111/j.1365-2249.2004.02590.x
15. Mout R, Willemze R, Landegent JE. Repeat polymorphisms in the interleukin-4 gene (IL4). Nucleic Acids Res. 1991;19(13):3763. doi:10.1093/nar/19.13.3763
16. Hosseini-Farahabadi S, Tavakkol-Afshari J, Rafatpanah H, Farid Hosseini R, Khaje Daluei M. Association between the polymorphisms of IL-4 gene promoter (−590C>T), IL-13 coding region (R130Q) and IL-16 gene promoter (−295T>C) and allergic asthma. Iran J Allergy Asthma Immunol. 2007;6(1):9–14. doi:06.01/ijaai.914
17. Isidoro-García M, Dávila I, Laffond E, Moreno E, Lorente F, González-Sarmiento R. Interleukin-4 (IL4) and interleukin-4 receptor (IL4RA) polymorphisms in asthma: a case control study. Clin Mol Allergy. 2005;29:3–15.
18. Kok Y, Ong H, Say Y. Interleukin-1 receptor antagonist and interleukin-4 genes variable number tandem repeats are associated with adiposity in Malaysian subjects. J Obes. 2017;1–8. doi:10.1155/2017/4104137
19. Tarlow JK, Blakemore AI, Lennard A, et al. Polymorphism in human IL-1 receptor antagonist gene intron 2 is caused by variable numbers of an 86-bp tandem repeat. Hum Genet. 1993;91(4):403–404. doi:10.1007/BF00217368
20. Salimi S, Khorasani M, Yaghmaei M, Mokhtari M, Moossavi M. Possible association of IL-4 VNTR polymorphism with susceptibility to preeclampsia. Biomed Res Int. 2014;1–5.
21. Salimi S, Khorasani M, Namazi L, Moossavi M, Naghavi A, Yaghmaei M. Association between interleukin 4 gene seventy – base – pair variable number tandem repeats polymorphism and uterine leiomyoma. Gene Cell Tissue. 2014;1(2):19462. doi:10.17795/gct-19462
22. Preacher KJ [Internet]. Calculation for the chi-square test: an interactive calculation tool for chi-square tests of goodness of fit and independence. 2001 [Computer software]. Available from: http://quantpsy.org.
23. Waller M, Moss S, Watson J, Møller H. The effect of mammographic screening and hormone replacement therapy use on breast cancer incidence in England and Wales. Cancer Epidemiol Biomarkers Prev. 2017;16:2257–2261. doi:10.1158/1055-9965.EPI-07-0262
24. Ponder BA. Genetic predisposition to cancer. Br J Cancer. 1991;64(2):203–204. doi:10.1038/bjc.1991.275
25. Liotta L, Petricoin E. Molecular profiling of human cancers. Nature Rev Genet. 2000;1:48–56. doi:10.1038/35049567
26. Apostolou P, Fostira F. Hereditary breast cancer: the era of new susceptibility genes. Biomed Res Int. 2013;2013:747318. doi:10.1155/2013/747318
27. Duan Y, Pan C, Shi J, Chen H, Zhang S. Association between interleukin-4 gene intron 3 VNTR polymorphism and cancer risk. Cancer Cell Int. 2014;14(1):131. doi:10.1186/1475-2867-14-67
28. Atoum MF, Tanashat RQ, Mahmoud SA. Negative association of the HLA-DQB1*02 allele with breast cancer development among Jordanians. Asian Pac J Cancer Prev. 2014;15(17):7337–7341. doi:10.7314/APJCP.2014.15.17.7337
29. Atoum MF, Alzoughool F. Reduction in breast cancer susceptibility due to XbaI gene polymorphism of alpha estrogen receptor gene in Jordanians. Breast Cancer (Dove Med Press). 2017;9:45–49. doi:10.2147/BCTT.S125652
30. Abdel-Razeq H, Al-Omari A, Zahran F, Arun B. Germline BRCA1/BRCA2 mutations among high risk breast cancer patients in Jordan. BMC Cancer. 2018;18(1):152. doi:10.1186/s12885-018-4242-8
31. AL-Eitan L, Jamous R, Khasawneh R. Candidate gene analysis of breast cancer in the Jordanian population of Arab descent: a case-control study. Cancer Invest. 2017. doi:10.1080/07357907.2017.1289217
32. Al-Zoubi MS, Mazzanti CM, Zavaglia K, et al. Homozygous T172T and heterozygous G135C variants of homologous recombination repairing protein RAD51 are related to sporadic breast cancer susceptibility. Biochem Genet. 2016;54(1):83–94. doi:10.1007/s10528-015-9703-z
33. Todaro M, Lombardo Y, Francipane MG, et al. Apoptosis resistance in epithelial tumors is mediated by tumor-cell-derived interleukin-4. Cell Death Differ. 2008;15:762–772. doi:10.1038/sj.cdd.4402305
34. Wu MC
35. Mittal RD, Manchanda PK. Association of interleukin (IL)-4 intron-3 and IL-6-174 G/C gene polymorphism with susceptibility to end-stage renal disease. immunogenetics. 2007;59(2):159–165. doi:10.1007/s00251-006-0182-6
36. Tsai FJ, Chang CH, Chen CC, Hsia TC, Chen HY, Chen WC. Interleukin-4 gene intron-3 polymorphism is associated with transitional cell carcinoma of the urinary bladder. BJU Int. 2005;95(3):432–435. doi:10.1111/j.1464-410X.2005.05315.x
37. Tsai MH, Chen WC, Tsai CH, Hang LW, Tsai FJ. Interleukin-4 gene, but not the interleukin-1 beta gene polymorphism, is associated with oral cancer. J Clin Lab Anal. 2005;19(3):93–98. doi:10.1002/jcla.20087
38. Shekari M, Kordi-Tamandani DM, MalekZadeh K, Sobti RC, Karimi S, Suri V. Effect of anti-inflammatory (IL-4, IL-10) cytokine genes in relation to risk of cervical carcinoma. Am J Clin Oncol. 2012;35(6):514–519. doi:10.1097/COC.0b013e31822d9c12
39. Ibrahimi M, Jamalzei B, Akbari ME, et al. Association between interleukin 4 (IL-4) VNTR, gene polymorphism, and breast cancer susceptibility in Iranian population: experimental and web base analysis. Bratisl Lek Listy. 2018;119(10):651–654. doi:10.4149/BLL_2018_116
40. Jia Y, Xie X, Shi X, Li S. Associations of common IL-4 gene polymorphisms with cancer risk: a meta-analysis. Mol Med Rep. 2017;16(2):1927–1945. doi:10.3892/mmr.2017.6822
41. Hedderson MM, Malone KE, Daling JR, White E. Allergy and risk of breast cancer among young women (United States). Cancer Causes Control. 2003;14(7):619–626.
42. Lange CA, Yee D. Progesterone and breast cancer.Womens. Health (Lond Engl). 2008;4(2):151–162. doi:10.2217/17455057.4.2.151
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.

