Back to Archived Journals » Research and Reports in Biochemistry » Volume 5
The autophagosome: current understanding of formation and maturation
Received 8 November 2014
Accepted for publication 21 November 2014
Published 16 February 2015 Volume 2015:5 Pages 39—58
DOI https://doi.org/10.2147/RRBC.S57405
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Professor Nikolay Dokholyan
Lilith VJC Mannack, Jon D Lane
Cell Biology Laboratories, School of Biochemistry, University of Bristol, Bristol, UK
Abstract: Autophagy is an important and highly conserved catabolic process with roles in development, homeostasis, and cellular stress responses. It describes various distinct pathways for the delivery of cytoplasmic materials (including misfolded protein aggregates and some organelles) to the lysosome for degradation and component recycling. The best understood form of autophagy (macroautophagy) describes the de novo assembly, maturation, and trafficking of a unique double membrane-bound organelle – the autophagosomes – that sequesters cytoplasmic materials and ultimately merges with the lysosomal compartment to form a degradative autolysosome. To rapidly assemble such a structure in response to stimuli, cells express a family of dedicated autophagy-related (ATG) gene products that act sequentially to control membrane events leading first to the nucleation of an isolation membrane or phagophore, followed by phagophore expansion, and sealing to form an autophagosome that traffics to – and ultimately fuses with – the lysosome. These molecules are activated in response to upstream signaling pathways (notably, the mechanistic target of rapamycin [mTOR] pathway), and comprise protein and lipid kinases, putative membrane coats, and unique ubiquitin-like conjugation systems. In concert, a barrage of accessory proteins involved in various membrane trafficking pathways focused on the endosomal compartment are co-opted at the assembly site to facilitate autophagosome biogenesis. Understanding the integrated pathways that coordinate autophagosome assembly at the molecular level will be crucial if we are to realize the potential for autophagy manipulation in future disease therapies.
Keywords: autophagy, ATG proteins, lysosome, phagophore, omegasome, autolysosome, membrane trafficking, ULK1, mTOR, PI(3) kinase, PI3P, LIR motif
Introduction: an overview of autophagy
The term autophagy was coined from ancient Greek, meaning to “self-eat”. It describes several distinct catabolic recycling pathways through which cytoplasmic material is degraded in the lysosome, with the resulting macromolecular building blocks liberated for re-use in the cytoplasm.1 The most widely studied autophagy pathways involve the formation of double-membrane bound vesicles (autophagosomes) that encapsulate cytoplasm and organelles for trafficking to the lysosome (accurately described as “macroautophagy”), but there are other routes for delivery of cytoplasmic material into this degradative compartment; these are known as “microautophagy” and “chaperone-mediated autophagy” (CMA). In microautophagy, the lysosomal membrane invaginates to engulf small portions of cytoplasm containing proteins that are often specifically targeted by Hsc70, as well as a small part of the bulk cytoplasm.2,3 CMA describes a process by which the same chaperone, Hsc70, targets proteins directly (via a consensus KFERQ sequence) for degradation by directly importing them through the lysosomal Lamp2A receptor.4
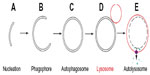
This review focuses on mammalian macroautophagy, which we will hereafter refer to simply as “autophagy”. During this process the canonical double membrane-bound autophagosomes form at nucleation sites known as an isolation membranes or phagophores (Figure 1 for the basic terminology). Importantly, autophagy can be further classified into basal and inducible forms, with basal autophagy serving a housekeeping role, whereas inducible autophagy accounts for the engulfment of bulk cytoplasm in times of stress (eg, during nutrient deprivation). To add to the complexity, both basal and inducible forms can contribute to the sequestration and degradation of selective cellular components. Indeed, there is probably an element of selectivity in all types of autophagy.5 Thus, the terms “selective” and “selective and exclusive” autophagy have recently been introduced to more accurately classify the process.6
Autophagy in homeostasis and disease
Autophagy primarily functions as a stress response pathway that liberates amino acids and other cellular building blocks in times of paucity.7 It was first described in the 1960s,8 but has only been studied beyond a purely descriptive level much more recently,9 leading on from the extensive knowledge gained on the key molecular players through genetic screens in yeast.10,11 Autophagy was long regarded as an alternative cell death pathway (type II cell death), but more recently autophagy has come to be viewed as being much more of a cellular survival mechanism.12 That said there are multiple molecular links between the pathways of apoptosis and autophagy (eg, Marino et al)13 meaning that in some circumstances the regulation of these apparently opposing cellular responses cannot be regarded separately.
Most cells maintain a low level of basal autophagy whose role is probably to clear damaged and damaging proteins that would otherwise accumulate as a consequence of normal cell function.14 Such a role is highlighted by studies on calorie restriction, a trigger that has been shown to extend lifespan in a range of organisms,15 most likely through restricting the buildup of toxic cellular waste. In accordance with these important roles in maintaining normal cellular function, autophagy has been implicated in a wide range of diseases. These include infectious diseases, where autophagy is required to clear invading pathogens and contributes to the acquired immune response, as well as degenerative diseases (eg, neurodegenerative conditions; heart disease; diabetes), and cancer. Jiang and Mizushima16 have provided an excellent general overview of autophagy in disease, while a number of other recent reviews detail the roles of autophagy in immunity and specific types of diseases.16–18 Autophagy impairment affects neurons in particular – presumably because the extended lifespan and poor renewability of this highly specialized cell type makes them especially susceptible to accumulating toxicity, and the links between autophagy and neurodegenerative disease is expertly covered elsewhere.19,20 The role of autophagy in cancer is somewhat more complex.21,22 On the one hand, cells toward the center of an expanding solid tumor are likely to rely on autophagy to maintain a pool of cellular building blocks as nutrients and oxygen become restricted.23 Here, a mechanistic block in autophagy might result in a tumor being unable to sustain itself, thereby making autophagy a potential therapeutic target. On the other hand, as autophagy is important for keeping damaging materials in the cell to a minimum, its downregulation might contribute to tumorigenesis through the buildup of genotoxic stress.22
Selective autophagy
There are several known types of selective autophagy that play roles in cellular homeostasis and responses to infectious challenge. Xenophagy describes the process through which invading microorganisms are specifically targeted for degradation via recognition and engulfment by the autophagy machinery.24 This type of autophagy is important in the immune response to some pathogens.18 Aggrephagy is responsible for the sequestration and degradation of aggregated proteins too large to be accommodated in the proteasome.25 Notably, a number of human diseases have been linked to the inefficient clearance of aggregate-prone proteins, for which treatment could entail the appropriate stimulation of autophagy (eg, Sarkar and Rubinsztein).26 Damaged, surplus, or redundant organelles are also specifically marked for autophagic degradation; an excellent example of this being the mitophagy pathway for mitochondrial degradation.27 Mitophagy is a highly selective process by which only the damaged or redundant parts of the mitochondrial network are specifically exiled before being sequestered into autophagosomes by the actions of a variety of adapter molecules that recruit the autophagy machinery.28 This process is seen during normal cellular homeostasis, and is also upregulated during erythroid terminal differentiation (eg, Betin et al).29 The importance of regulated mitochondrial clearance by mitophagy is evident in Parkinson’s disease, where early onset (hereditary) forms are often associated with loss-of-function mutations in mitophagy-linked proteins, including PINK1 (PTEN-induced putative kinase 1) and Parkin.28 Cell-based in vitro studies have provided a molecular explanation for the cooperative actions of PINK1 and Parkin during mitophagy, in which PINK1 interrogates mitochondria for energetic functionality through a pathway of constitutive import and degradation. This pathway is abrogated in damaged mitochondria, leading to the targeted recruitment of Parkin and subsequent Parkin-mediated ubiquitylation of mitochondrial surface proteins.30 Key to efficient mitophagy is the regulated exile of damaged organelles from the network coordinated by the mitochondrial membrane fission machinery, and in conditions where mitochondrial dynamics are suppressed mitophagy is strongly blocked.31–33 Other organelles can be targeted for autophagic degradation when damaged; for example, parts of the endoplasmic reticulum are sequestered into autophagosomes under endoplasmic reticulum stress conditions (reticulophagy),34 while a pathway for the selective autophagic degradation of peroxisomes (known as pexophagy) also exists.35 Further details on the molecular regulation of selective autophagy can be found in a number of excellent recent reviews (eg, Lamark and Johansen,25 Svenning and Johansen36).
The molecular mechanisms controlling starvation-induced autophagy
There are more than 30 proteins directly involved in autophagy, playing differing roles at the various stages of autophagosome assembly, maturation, and trafficking. Most of these key molecular regulators were first identified in yeast, and the proteins identified in the original screens10,11,37 are now collectively referred to as ATG (AuTophaGy-related) genes.38
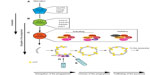
For simplicity, the process of autophagosome biogenesis can be divided into a number of sub-stages: nucleation; elongation of the phagophore; closure of the phagophore to form an autophagosome; trafficking to the lysosome; and fusion of the outer autophagosomal membrane with the lysosome to deliver cargo39 (Figure 1). All of these events involve complex, tightly localized and coordinated molecular signaling, recruitment, and trafficking steps, and the ATG proteins that drive these sequential processes are classed into functional groups corresponding to these substages of autophagy. Correspondingly, the sequential and largely hierarchical recruitment of these autophagy proteins to the site of autophagosome assembly has been well characterized in yeast (where a single assembly site, the pre-autophagosomal structure or “PAS” exists) and in mammals (where multiple assembly sites arise simultaneously and apparently stochastically throughout the cytoplasm). Several recent studies paint a more complex picture of the recruitment and actions of autophagy proteins at assembly sites, with the stability of “early” players positively reinforced by downstream factors whose recruitment critically depends on the former40–43 (Figure 2). The majority of autophagy proteins dissociate from the site of autophagosome assembly just prior to completion of the intact autophagosome (with [green fluorescent protein] GFP-labeled ATG5 structures remaining for no longer than 3–4 minutes during starvation-induced autophagy in human cells in vitro; our unpublished observations;40,42–44), and it is likely that the coordinated removal of certain proteins and protein complexes is equally important for efficient, productive autophagosome assembly.
The nucleation stage: autophagy induction/nutrient sensing
Cellular responses to nutrient availability
In nutrient replete conditions, mTOR (mechanistic target of rapamycin) is held in a constitutively active state, and coordinates cell growth and proliferation, while inhibiting autophagy. Nutrient deprivation, particularly amino acid starvation, inactivates mTOR, which releases the brake on autophagy leading to rapid assembly of autophagosomes.45 Information about nutrient levels in the cell is decoded by mTOR via the Rag (Ras-related small GTP-binding protein) GTPase family,46 which in their GTP bound state (favored in the presence of amino acids) facilitate the translocation of mTOR to cellular locations enriched for the mTOR activator, Rheb (Ras homology enriched brain),47 while simultaneously acting as part of a regulatory platform.48 Energetic stress can be sensed by an increase in adenosine monophosphate (AMP) through AMPK (AMP kinase). One of the consequences of nutrient limitation is a drop in cellular ATP levels (or more accurately, a change in the ATP:AMP ratio). This causes activation of AMPK, which inhibits mTor directly or indirectly by causing downstream inhibition of Rheb. AMPK can also directly activate ULK1 (uncoordinated 51-like kinase 1; the mammalian homologue of yeast ATG1 [see “ULK1 autophagy protein kinase complex” section]) in response to low ATP levels,49,50 further amplifying the autophagy response and providing an additional level of control.
Another way in which mTor can be inactivated in response to environmental change is through the inhibition of upstream regulatory inputs. For example, insulin or growth factor stimulation results in the activation of the PKB (protein kinase B)/Akt pathway, one downstream consequence of which is Rheb activation.51 This means that in the absence of growth factors and/or insulin, a further obstruction to autophagy activation is removed. Furthermore, in glucose rich conditions, the cAMP (cyclic AMP)-dependent protein kinase A (PKA) signaling pathway suppresses autophagy. When glucose levels are limiting, elevated cAMP levels lead to inactivation of the Ras-PKA pathway, which is predicted to trigger ATG1 phosphorylation/activation (at least in yeast), meaning that the Ras-PKA pathway acts in concert with mTor to block autophagy under nutrient rich conditions.51,52 Additionally, LC3 (microtubule-associated protein 1 Light Chain 3), which is a relatively “late” autophagy protein (Figure 2 and following sections), has been shown to be a direct target of PKA in mammalian cells.53 This indicates that the PKA pathway has the potential to regulate autophagy at multiple levels, although it was noted that none of the other mammalian ATG8-related proteins (nor, indeed, yeast ATG8) possesses this PKA consensus sequence.53
The mTor complex 1 (mTorC1)
mTor forms two distinct complexes in mammalian cells – mTorC1 and mTorC2 – with only mTorC1 having nutrient sensing capabilities.54 The mTorC1 complex consists of mTor, mLST8 (mammalian lethal with SEC13 protein 8; also referred to as G protein beta subunit-like [GβL]), and Raptor (regulatory associated protein of mTOR)54 (Figure 2A). Raptor is believed to be the subunit that controls the targeted localization of mTorC1 to signaling hubs, but the function of mLST8 is not known.47 Under basal conditions, mTorC1 is associated with, and suppresses ULK1 and ATG13 activity by phosphorylation.55–57 This means that amino acid restriction results in the activation of ULK1 and ATG13 – two key players acting during early autophagy initiation that will be described in greater detail below.
The ULK1 autophagy protein kinase complex
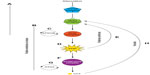
In their activated states – and in concert with FIP200 – ATG101, the autophagy kinase ULK1 and its partner ATG13 form the ULK1-complex55–58 (Figure 2B). The ULK1 complex is the molecular machinery that defines the nucleation stage of autophagic induction (Figure 2B). Its assembly results in the recruitment of the class III phosphoinositol-3 kinase (PI(3)K) complex (see “The autophagy PI(3)K/VPS34 complex” section) – also known as the vacuolar protein sorting 34 (VPS34) complex – and this is required for downstream events in the autophagosome assembly pathway59,60 (Figure 2B). ULK1 complex activity is mechanistically linked to the recruitment and activation of the autophagic VPS34 complex, because Beclin1 (one of its core components [see “The autophagy PI(3)K/VPS34 complex” section ]) is a substrate for ULK1 kinase-mediated phosphorylation, and this regulatory switch initiates the assembly of the VPS34 complex.61 Intriguingly, ULK1 is also capable of inactivating mTor, exemplifying a positive feedback loop in the autophagy pathway62 (Figure 3). ATG13 is thought to be required to support the interaction between ULK1 and FIP200 in mammalian cells,57 as well as the association between ATG101 and ULK1,58 and it may also be directly phosphorylated by mTor. FIP200 is believed to be the mammalian counterpart of yeast ATG17,63 a protein considered to act as a scaffolding factor for the ULK1 complex, as well as potentially inducing local membrane curvature.64 In addition to ULK1 activating downstream autophagy effector proteins, the ULK1 complex serves as a scaffold for ATG9 vesicles,65 indicating that its function may be more important later relative to its time of assembly. Indeed, the yeast ATG1 complex is thought to sense membrane curvature, and thus its ATG17-dependent localization to the phagophore assembly site would be further stabilized by membrane reshaping.64 Interestingly, there is evidence that ATG1 binds directly to ATG8 (LC3), and this has been shown to be required for complete activation of the ATG1 kinase complex66 (Figure 3). Additionally, the ULK1 complex is stabilized by the presence of the phosphoinositide lipid PI3P (see “PI3P recruits effector proteins: WIPI2, ATGS5/12/16” section), which indicates that positive reinforcement between autophagy proteins is critical for optimal autophagy; a lack of PI3P results in a reduction of ULK1 lifespan on the autophagosome initiation site43 (Figure 3).
The ULK1 complex is also regulated by Rab proteins.67 Rabs are small Ras-like GTPases that define membrane compartment identity to aid the specificity of membrane trafficking, and potentially facilitate vesicle docking.68 Rab proteins bind GTP, and usually it is only the GTP bound form that is involved in vesicle/target recognition. Rabs can be activated through specific GEFs (guanine nucleotide exchange factors) and inactivated by GAPs (GTPase activating proteins). Every Rab GAP that has been identified thus far contains a TBC (Tre2/Bub2/Cdc16) domain, which dictates the Rab GAP nomenclature. In a screen for TBC domain containing proteins, Longatti and Tooze67 identified the RabGAP TBC1D14 as a negative regulator of autophagosome formation. The authors went on to validate Rab11 – the Rab regulated by TBC1D14 – as an important mediator of early autophagy events. Rabs have also been found to play distinct roles during later stages of autophagosome biogenesis and maturation as will be detailed later.
It is worth noting that at least five potential mammalian ATG1 homologs have been identified thus far, some of which have been implicated in autophagy under specific settings. ULK1 was the first to be shown to play a role in autophagy, and it remains the most widely studied.69,70 In addition to its regulation by phosphorylation ULK1, may also be posttranslationally regulated by ubiquitination and acetylation,70 adding further complexity to autophagosome assembly control pathways.
Elongation, shaping, and sealing the phagophore
The autophagy PI(3)K/VPS34 complex
Inositol lipids and their phosphorylated derivatives, including PI3P, are important mediators of intracellular signaling, with different phosphorylated PIs marking varied membranous compartments; examples include PI(4)P and PI(4,5)P2 that decorate the Golgi apparatus,71 and PI3P which be found on early endosomes and the nascent autophagosome. The term “omegasome” has been coined to describe the PI3P membrane patch that is generated on or close to the ER (see “PI3P recruits effector proteins: WIPI2, ATGS5/12/16” section) during mammalian autophagy initiation.72 The marking of membranous compartments by differing PIs to some extent defines the compartment, because the presence of a specific PI – in concert with the actions of additional targeting/stabilization factors – dictates the proteins that are recruited, and hence the associated biological activity that is focused on that cytosol-facing patch of membrane.73 As will become clear in the following sections, this scenario certainly holds true for the autophagy system.
In higher eukaryotes, three VPS34 complexes are present: I, II, and III. Complexes I and III regulate autophagy in a negative and positive fashion, respectively, while the class II complex functions in the endosomal network with no direct involvement in autophagy. The class I VPS34 complex is a plasma membrane-associated component of the insulin-signaling pathway that controls mTor activity, and thus indirectly and negatively regulating autophagy.54,74 The core of the autophagic class III VPS34 complex is composed of the catalytic VPS34 subunit, with p150 (VPS15 in yeast), and Beclin1 (VPS30/ATG6 in yeast).75,76 VPS15 is found on all VPS34 complexes in yeast, and is thought to anchor the VPS34 complex to the membrane through N-terminal myristoylation, and positively modulate the activity of VPS34 through an unknown mechanism.70,77 Beclin1 is considered to be the primary regulatory subunit of the class III VPS34 complex, with which both positive and negative modulators bind to determine its context-specific roles during autophagy (Figure 2C). The best-characterized VPS34 class III complex contains ATG14L/Barkor, a peripheral ER protein that concentrates into puncta at early autophagosome initiation sites in response to stimuli.76 It is believed to have the capability to sense and maintain the membrane curvature, and preferentially binds membranes enriched for PI3P.76 However, and as would be predicted from its involvement with the VPS34 complex, ATG14L concentrates at the nascent autophagosome before PI3P is formed,75 implying that, like with ULK1, ATG14L may be stabilized at the nascent autophagosome by the generation of PI3P (Figure 3).
A second VPS34 class III complex that positively regulates autophagy contains UVRAG (UV-irradiation-resistance-associated gene) in the place of ATG14L.78 Endophilin B1 (also known as Bif1 [Bax-interacting factor 1]) associates indirectly with Beclin1 via UVRAG, and in common with ATG14L, Endophilin B1 is thought to sense and induce membrane curvature.79 Unlike the ATG14L-containing complex, the UVRAG class III VPS34 complex also contributes to autophagosome maturation, although this idea remains controversial.70 Furthermore, an inhibitory class III VPS34 complex can assemble from the core elements VPS34, p150, and Beclin1, with UVRAG and “Rubicon” (RUN domain protein as Beclin1 interacting and cysteine-rich containing).80 This complex is thought to prevent both autophagy induction and maturation, and probably plays a role in the endosomal compartment.81
An additional protein that is necessary for the assembly/translocation of the VPS34 complex III is Ambra1 (Activating Molecule in Beclin1-regulated autophagy).82 Under normal conditions, Ambra1 sequesters the VPS34 complex to microtubules via binding to cytoplasmic dynein, but this interaction is disrupted upon starvation allowing Beclin1 to translocate to the ER.60 This putative regulatory system for AMBRA1-mediated autophagy control has been thrown into doubt following the recent demonstration that Ambra1 facilitates the association between ULK1 and TRAF6 (tumor necrosis factor receptor associated factor 6), an E3 ligase, with the ensuing ubiquitination activating ULK1.83 The exact regulation of Ambra1 remains controversial, with different studies also reporting conflicting data with regard to the phosphorylation status of Ambra1.60,83
PI3P generation is vital for autophagy, but in mammalian cells MTMR3 (myotubularin related protein 3; hereafter referred to as Jumpy) – a member of the myotubularin lipid phosphatase family – has been shown to be needed for productive autophagosome assembly by acting as an important limiter of localized PI3P production.84 In support of this theory, yeast Ymr1 (myotubularin-related 1) – the only yeast myotubularin ortholog – has been shown to be needed for the completion of the nascent autophagosome.85 It is possible that PI3P needs to be asymmetrically distributed at the isolation membrane for the purposes of autophagosome assembly. In yeast the inside membrane of the autophagosome is enriched for PI3P,86 which could be achieved via targeted myotubularin-mediated dephosphorylation. Curiously, though, this asymmetric distribution pattern is reversed in mammals.86
PI3P recruits effector proteins: WIPI2, ATG5/12/16
The localized formation of PI3P is an essential step for autophagosome assembly, because when this step is inhibited by drugs such as wortmannin, autophagy is blocked.87,88 One purpose of the generation of PI3P by the VPS34 class III complex is to recruit a number of effector proteins that translate the local PI3P signal into downstream membrane remodeling and autophagosome formation steps; however, until recently, a molecular outline of this process was missing. An essential PI3P effector protein family for autophagy is the WIPI (WD-repeat protein interacting with phosphoinositides) family, which in mammals comprises WIPI1–4 – the mammalian functional homologs of yeast ATG18.89–91 Importantly, in mammalian cells, WIPI2b has recently been shown to recruit ATG16L1,92 which in turn recruits the ATG5-12 conjugate to form the ATG5/12/16L1 complex.93
The ATG5/12/16 complex is needed for ATG8 lipidation (see “Roles and regulation of ATG8 family members in autophagy” section), and is thought to act as a scaffold for the expanding autophagosome, which, in conjunction with ATG8, probably facilitates its growth.94,95 ATG5 is required for targeting ATG5-12 to the autophagosome formation site, and ATG5 can indeed arrive independently of ATG12; however, the presence of ATG12 is required for subsequent steps.96 In mammalian cells, the ATG5/12/16L1 complex localizes to the isolation membrane via ATG16L1. Indeed, if ATG16L1 is mislocalized to the plasma membrane for example, LC3 is lipidated at and incorporates into this compartment.93 ATG16L1 binds directly to FIP200 and this interaction has been shown to be required for ATG16L1 localization to the phagophore.97 Additionally, ATG16 shows a preference for PI3P containing liposomes in vitro.95 In combination with the study of Dooley et al,92 this may indicate that there are several parallel processes by which ATG16L1 localizes to the nascent autophagosome.
ATG16L1 is found in pre-autophagosomal vesicles that derive from the plasma membrane, traffic through the endocytic compartment, and undergo homotypic fusion prior to their integration into the expanding phagophore.98–100 Furthermore, ATG16L1 is believed to be a Rab effector for Rab33A/B, possibly explaining how the autophagy proteins are initiated to arrive at the nascent autophagosome.101 Interestingly, the Rab33 GAP, RUTBC1 (RUN And TBC1 Domain Containing 1), has also been identified as an autophagy modulator.102
Ubiquitin-like conjugation systems in autophagy: ATG5-12 and LC3-PE
In addition to the aforementioned complexes that regulate distinct, temporal membrane-associated events during early autophagosome assembly, there are two ubiquitin-like conjugation systems that are essential for autophagy (Figure 2D and E). The first conjugation reaction involves the covalent attachment of ATG5 to ATG12, and the second reaction consists of ATG8 family members being conjugated to PE (phosphatidylethanolamine)103–105 – a reversible membrane anchor that is required for function of ATG8 in autophagy.104,106–109 Neither ATG8 nor ATG12 display sequence homology with ubiquitin, yet each of their crystal structures reveals obvious ubiquitin-like folding.110 The respective conjugation reactions employ separate E2-like enzymes – ATG10 for ATG5 conjugation to ATG12, ATG3 for the lipidation of ATG8 – but ATG12 and ATG8 share ATG7 as their common E1-like enzyme. In fact, ATG7 binds to either ATG8 or ATG12 in a mutually exclusive manner.111 One interesting difference between these two conjugation systems is that ATG5-12 appears to be constitutively conjugated, whereas the covalent attachment of ATG8 to PE is upregulated following autophagy stimulation.105,112 Importantly, the product of the “upstream” conjugation reaction – the ATG5-12 complex – acts as an E3-like ligase (in association with ATG16) to aid the covalent attachment of ATG8 to PE, which means that the ATG5/12/16 complex acts to speed up the transfer of ATG8 from ATG3 to PE without conveying any additional specificity.113
Membrane vesicle trafficking: ATG9 and ATG16
ATG9 acts immediately downstream of FIP200 in the autophagosome assembly pathway. It localizes to poorly characterized membrane reserves near the trans-Golgi network (TGN) or the mitochondria in nonstarvation conditions, upon which it translocates to the autophagosome formation site during starvation.114–116 This transport step requires actin in yeast,117 and is dependent on myosin II in Drosophila;118,119however it is unclear how transport of ATG9 is regulated in mammalian cells. At the yeast phagophore assembly site, an estimated number of three roughly 30 nm ATG9-containing vesicles fuse to form the phagophore,114 in a process that is dependent on ATG1 and ATG13 for retrieval of ATG9 from the phagophore assembly site.120 ATG9 is not found on the autophagosome itself – most likely ATG9 interacts with the phagophore only briefly in a “kiss and run” fusion event, perhaps to donate membrane.121 Thus ATG9 cycling forms a separate membrane trafficking event – both the anterograde and retrograde movement of ATG9 are required for efficient autophagy.120 Intriguingly, ATG9 has also been found to be a direct target for ATG1 in yeast, with ATG9 phosphorylation critical for the recruitment of ATG2 and ATG18 (the yeast equivalents of mammalian WIPI proteins) to the assembling autophagosome.122 This again shows that there is positive reinforcement within the recruitment hierarchy of autophagy proteins. Interestingly, ATG9 feeds into the MAPK pathway via p38 interacting protein (p38IP), which is regulated by p38α MAPK: p38IP binds to ATG9 and is required for starvation-induced ATG9 trafficking.123
Autophagosome biogenesis requires the actions of SNARE proteins (soluble N-ethylmaleimide-sensitive fusion protein attachment receptors). SNAREs function to aid the docking and ultimately the fusion of transport vesicles with target organelles. In order to do this, SNAREs assemble specific binding pairs – one on each of the compartments that are to fuse – which means that only vesicles with the correct cognate SNARE will dock with the desired target compartment. One of the SNARE-requiring fusion steps is thought to be the homotypic fusion of ATG9 vesicles to create a tubule that turns into the phagophore, another being during ATG9 cycling.99,124 In addition to ATG9 trafficking to the forming autophagosome in membranous vesicles, ATG16L1 has more recently been identified to arrive at the forming autophagosome from the plasma membrane on vesicular carriers.125 In fact, even though ATG16L1 and ATG9 display distinct trafficking pathways, they coincide in the recycling endosome from where they are proposed to traffic together to autophagosomes.125 Interestingly, autophagosome precursor maturation has also been shown to require ATG16L1 vesicle fusion, which is also demonstrably dependent on SNAREs.99
Roles and regulation of ATG8 family members in autophagy
Yeast have a single ATG8 gene, whereas the situation in mammalian cells is more complicated, with human cells expressing at least eight ATG8 paralogs separated into LC3 and GABARAP (gamma aminobutyric acid receptor associated protein) subfamilies. In the LC3 family there are LC3A, LC3B, LC3B2, and LC3C, while the GABARAP family comprises GABARAP, GABARAP-L1/GEC1/ATG8L, GABARAP-L2/GATE-16, and GABARA-L3. All ATG8 family proteins share structural similarities with ubiquitin,126 and all are cleaved shortly after synthesis to reveal a conserved C-terminal glycine (G120 for LC3B and G116 for all the GABARAP family members).104,106,127,128 This exposed glycine is later covalently linked to PE (see “Ubiquitin-like conjugate systems in autophagy: ATG5-12 and LC3-PE” section) as an essential step in autophagosome assembly. The ATG8 family – particularly LC3B in mammals – is widely used as a marker for the forming and completed autophagosome.129 It is the only autophagy protein that remains attached to the autophagosome following its completion, and has multiple proposed roles during autophagosome formation and maturation. LC3 was initially identified as a microtubule binding protein, and LC3B and GABARAPs display microtubule-binding capabilities at their N-termini.130–133 Despite this, no direct microtubule binding functions have yet been found to mediate autophagy.
ATG8 has membrane tethering and hemifusion properties in vitro,134 suggesting that it, and possibly also some members of the expanded family of mammalian ATG8s, may directly facilitate the fusion of the phagophore to form the completed autophagosome; however, it has been noted that ATG8-mediated hemifusion does not occur in liposomes containing physiological concentrations of PE.124 The findings of Nakatogawa et al134 have been confirmed to some extent in experiments using certain mammalian ATG8 family members, both in a cell free system and in cells, where the deletion of the N-termini of GATE-16 and LC3 – responsible for the membrane fusion capabilities of the ATG8 family – resulted in the accumulation of immature phagophores and/or a failure in sealing liposomes.135 Interestingly, the amino acids that mediate membrane fusion are not conserved between LC3 and GATE-16, with positive amino acids responsible for fusion in LC3 and neutral amino acids in GATE-16.135 Furthermore, in mammalian cells the fusion activity of LC3 is dispensable for fusion of autophagic vacuoles with early or late endosomes,136 meaning that it is unlikely that this capability of LC3 is required for autophagosome maturation.
In yeast, ATG8 protein concentration is one factor that has been found to influence autophagosome size, but this has no apparent effect on steady state autophagosome numbers or their rate of assembly.137 The density of ATG8 on the convex surface of the yeast phagophore corresponds with the density expected for a coat protein leading to the proposal that ATG8 acts as a coat adaptor in concert with the ATG12-ATG5 complex.138 This finding has been supported by an in vitro study by Kaufmann et al94 suggesting that ATG8 is part of a coat complex during autophagosome formation, which is subsequently disassembled by ATG4 deconjugase activity. ATG8 also serves a further scaffolding role of recruiting the ATG1 (ULK1) complex to the forming autophagosome,139 this both positively reinforces autophagosome formation (Figure 3) and may support ULK1 targeting of proteins at precise locations on the forming autophagosome. LC3 has been shown to be required for autophagosome expansion, whereas GATE16/GABARAP is involved in autophagosome sealing.140 This important study highlights how regulated and sequential employment of different ATG8 family members in mammalian cells has the potential to fine-tune the autophagosome assembly pathway to meet diverse cellular needs.
LC3 binds readily to the cargo adaptor p62 (sequestosome1 or SQSTM1).5 This means that as mentioned above “nonselective” forms of autophagy have an element of specificity, since p62 binds tightly to ubiquitinated protein targets. In mammalian cells, the various ATG8 family members show differing affinities for p62; LC3 efficiently incorporates p62 into autophagosomes, whereas GABARAP lacks this capability.141 Importantly, p62 is not the only autophagy-associated cargo adaptor; neighbor of BRCA1 gene 1 (NBR1), which acts as a cargo receptor during selective autophagy,142 also interacts directly with the main members of the ATG8 family.143 Conversely, Nix – a mitophagy cargo adapter protein – binds preferentially to GABARAP family members.144 Interestingly, an extensive autophagy network analysis identified 67 ATG8 family interactors, and 31 of these were found to be specific for a single subtype of the ATG8 family.143 Once again, this shows how selective utilization of different classes of mammalian ATG8 proteins can have a profound effect on autophagosome cargo, providing further functional flexibility in the autophagic response.
The LIR motif as an important interacting module in autophagy that confers selectivity to the autophagy response
Many cargo adaptor and other proteins that bind to the ATG8 family contain the consensus sequence WXXI/L, which is known as the LIR (LC3-interacting region) motif.36,145 In most cases, LIR motifs facilitate cargo receptor binding to ATG8 family members, and are therefore important for cargo incorporation into the nascent autophagosome – examples include p62 and NBR1; however, LIR motifs have been identified in ULK1 and ATG12, and these facilitate targeting and/or stabilization of the autophagy machinery at the autophagosome assembly site.66,94 Competition for LIR binding between cargo and the autophagosome assembly machinery has been demonstrated in vitro, revealing a plausible molecular mechanism for the different functions of ATG8 on the inside and outside of the autophagosome (in cargo recognition and as a putative coat, respectively).94
Within the known LIR motifs, there are different classes with differential binding affinities for ATG8 family members. In the case of the LIR motif in NBR1, the binding sequence is YXXI, and exchanging the tyrosine at P1 to tryptophan increases the LC3 binding affinity of this motif, implying that additional factors might facilitate/strengthen this particular interaction.146 The atypical LIR motif identified in ATG12 includes residues Phe185 and Ile111 which are well separated in the primary sequence.94 Consequently, this LIR motif is tertiary structure-dependent; that is to say, it is formed in 3D space by precise folding of the polypeptide chain.94 Importantly, the LIR motif described for the cargo adaptor NDP52 is also atypical, in that it does not conform to the WXXI/L consensus, but nevertheless it binds readily to mammalian LC3C due to compensatory alterations in the aromatic pocket of its cognate ATG8 family member.147 Perhaps most significantly, switching the native isoleucine at position 133 in the NDP52 LIR for tryptophan dramatically increased its affinity for LC3C (into the nanomolar range), but simultaneously destroyed its specificity for LC3C.147
Based on these findings, it can be assumed that other noncanonical LIR domain-containing proteins exist in diverse cargo proteins, meaning that sequence analysis approaches that rely on classical properties in the primary structure alone (eg, iLIR; http://repeat.biol.ucy.ac.cy/iLIR/) will fail to identify these. To add to this complexity, LC3 binding to LIRs may be occluded and/or stimulated by posttranslational modifications; for example, the LIR motif of the mitophagy adaptor, FUNDC1 (FUN14 Domain Containing 1), contains a phosphorylation-competent tyrosine, and dephosphorylation at this site during hypoxia is required to activate LC3 binding.148 Interestingly, FUNDC1 mitophagy activity is also regulated by phosphorylation/dephosphorylation at a two further sites, S13 and S17, by CK2/PGAM5 and ULK1 respectively,149,150 meaning that there are likely to be multiple regulatory influences on LIR motif activity.
Further roles for ATG8 family members during autophagosome maturation
LC3 plays key roles on the outside as well as the inside of the autophagosome. By acting as a Rab GAP adaptor, LC3 integrates autophagosomes into the global membrane trafficking network, enabling them to be sorted to and fuse with the correct endomembrane components.151 Here, the Rab GAP TBC1D5 was found to bind competitively to retromer components on the recycling endosome and to LC3. GABARAP is believed to act as a scaffold to mediate signal transduction through the RabGAP, TBC1D25 (also known as ornithine aminotransferase-like 1 or Oatl1),152 and TBC1D25 is required for fusion of the autophagosome with the lysosome.152 It has GAP activity toward Rab33, and active Rab33 is thought to recruit the ATG5/12/16 complex to the phagophore. This could mean that Rab33 is inactivated as soon as GABARAP is recruited to the phagophore. As the ATG8 family members have been shown to play different roles in autophagosome biogenesis, and TBC1D25 preferentially binds GABARAP,152 it is possible that the regulated removal of other ATG8 family members (eg, LC3B) from the outside of the autophagosome is required for optimal TBC1D25 binding to the mature autophagosome. It may also be that Rab33 needs to be deactivated by GATE16 to complete a phagophore, as GATE16 is required for autophagosomal sealing; thus Rab33 inactivation could be a prelude to autophagosome completion. Rab7 has been shown to be required in conjunction with LC3 and FYCO1 (FYVE and coiled-coil domain containing 1) to form an adaptor protein complex, essential for plus end directed, kinesin-mediated microtubule transport.153 Subsequently Rab7 is required for autophagosome-endosome/MVB (multivesicular body) fusion and a depletion of Rab7 leads to an accumulation of autophagosomes.154 This also means that at least some ATG8 is required on the outside of the autophagosome to mediate autophagosome trafficking.
Roles and regulation of the ATG4 endopeptidase family
ATG4 is a cysteine endopeptidase that is essential for controlling ATG8 processing during autophagy. It serves two distinct roles, priming and delipidation.128 Initially, ATG4 cleaves newly synthesized ATG8 – which is produced in an immature pro-form – to reveal a C-terminal glycine (the priming step). The exposed glycine is subsequently conjugated to PE at the expanding isolation membrane, meaning that the priming step is indispensable for autophagy.128,155 The second role of ATG4 is to deconjugate membrane-bound ATG8 in order for ATG8 to disassociate from the membrane (the delipidation step).107,128 ATG8 delipidation has been shown to be required for the completion of autophagosome assembly in yeast,137,156,157 although it is not yet known whether this holds true in mammalian cells.
Akin to the situation regarding mammalian ATG8 divergence, ATG4 is represented by four paralogs in mammalian cells; namely ATG4A, ATG4B, ATG4C, and ATG4D. These are collectively known as the ATG4 cysteine endopeptidase family, or the autophagins.158 The structure of ATG4 resembles that of the papain family of proteases, despite low sequence homology.159,160 Each of these ATG4 family members displays subtly differing substrate affinities for the various mammalian ATG8 family members, thus potentially allow fine-tuning of the autophagy response with respect to autophagosome form and content in mammalian cells. ATG4B displays the broadest substrate specificity,161–164 while ATG4A is selective for GATE16,165 and ATG4D shows activity exclusively against GABARAP-L1 in vitro.161,163
To add an additional level of regulatory complexity, ATG4B (but not other ATG4 family members is downregulated by RNF5 (Ring Finger Protein 5), an E3 ubiquitin protein ligase on membranous compartments.166 Reduced RNF5 levels leads to an increase in ATG4B, and subsequently lipidated LC3-II levels, but does not result in a change in the levels of the GABARAPs. Another example of ATG4 regulation was demonstrated by Scherz-Shouval et al167 who showed that reactive oxygen species inhibit ATG4A and ATG4B, by modifying an essential cysteine located close to the catalytic site triad of these family members. The authors hypothesized that ATG4s are active in a reducing environment; autophagy induction releases a burst of the oxidizing agent hydrogen peroxide, which inhibits ATG4A/B locally until the newly formed autophagosome leaves the oxidizing environment. At this point, ATG4 becomes active again and delipidates the appropriate ATG8 family member before the autophagosome fuses with the lysosome.167 Atg4C and Atg4D both have a caspase cleavage site, and immediately downstream, a mitochondrial-targeting sequence, suggesting that apoptotic cleavage triggers mitochondrial import.168
In vivo studies of ATG4 deletion in mouse
The apparent specificity of ATG4 endopeptidases for ATG8 family members reported in vitro is not supported by genetic deletion studies in mouse, which generally reveal a milder effect on autophagy than might be expected.169–171 This is perhaps due to adaptive redundancy in the developing mouse, nevertheless the ATG4B and ATG4C knockout models have provided clues as to the individual roles of these family members in vivo. The ATG4C knockout mouse is viable and fertile, showing only an increased susceptibility to fibrosarcoma, decreased locomotor activity after prolonged starvation, and a mild decrease in autophagic activity measured by LC3B lipidation in the cells of the diaphragm.170 Thus, ATG4C is dispensable under normal conditions, but under stress conditions such as prolonged starvation or in the presence of carcinogens, a protective role is revealed. The ATG4B knockout mouse, despite showing a pronounced defect in ATG8 family member lipidation throughout its tissues, also has a mild phenotype with a loss of equilibrioception due to the abnormal formation of otoconia (organic calcium carbonate crystals) in the inner ear.169 It is noteworthy that some tissues showed partial lipidation of LC3B, meaning that another factor may functionally compensate for ATG4B under certain conditions.169 When the ATG4B knockout mouse was further studied, abnormal spheroid-like bodies were found in the neuropil of the deep cerebellar nuclei.171 These abnormal structures were interpreted as lesions, and this translated into slightly poorer performance on a rotarod, giving a second physiological explanation to the impaired locomotion of the mutant mice.171 Additionally, homozygous ATG4B-/- mice were born at slightly lower incidence than Mendelian genetics would predict, possibly indicating an issue in fetal development upon ATG4B deletion.171
Many questions remain about the control of autophagosome assembly and function by ATG4 family members. There is clear evidence that ATG8 priming and delipidation are needed for efficient autophagosome formation, maturation, and vacuolar delivery in yeast,10,107,128,156,172,173 but the picture is much less clear in mammalian cells. It has been shown that overexpression of wild-type and dominant-negative mutant (C74A) ATG4B arrests autophagosome formation at the ATG8 lipidation step,174 and apparently at the lysosomal fusion stage in human erythroid cells,29 but more sophisticated tools will be needed to decipher contribution of the delipidation of ATG8 family members during phagophore closure, autophagosome maturation and trafficking, and lysosomal fusion.
Other factors implicated in the control of autophagosome assembly
VMP1 (Vacuole Membrane Protein 1) is a transmembrane, Beclin1-binding protein that has been shown to be required for autophagy.175 VMP1 localizes to the ER and generates punctate structures upstream of the VPS34 complex upon autophagy stimulation,175 although other work has described constitutive ER-associated VMP1 puncta that do not colocalize with early autophagy markers (ULK1).175 VMP1 competes with BCL-2 for binding to Beclin1, suggesting that it may release Beclin1 at local ER-associated sites.175 VMP1 silencing blocks autophagic flux, and causes an accumulation of abnormally large early autophagy structures (positive for ULK1, WIPI1, DFCP1), suggesting that it might act downstream of the assembly site initiation stage.175 In this respect, it is noteworthy that VMP1 silencing did not increase steady state ATG16L1 puncta numbers,175 arguing that its action is restricted to the PI3P generation step, perhaps to coordinate the turnover of PI3P.
As mentioned above, Rab33B/A is required to localize ATG16 to the phagophore,101 but there are additional Rabs that are believed to function in autophagy, at least in yeast. Ypt1 (yeast protein two 1), and its GEF, TRAPP III (transport protein particle III) – also a multi-protein tethering complex – are needed for autophagy, although with the exact stage and mechanisms of action unknown.176,177 Notably, studies of selective autophagy have implied that Ypt1 recruitment occurs downstream of ATG9.178
Autophagosome maturation
Fully formed autophagosomes travel along the microtubule network toward the lysosome, fusing en route with early endosomes and late endosomes/MVBs, to form a hybrid structure termed an amphisome.179–181 The autophagosome/amphisome acidifies as it matures, either due to the pH of the structures it fuses with and/or through the actions of proton pumps. Indeed, the autophagosome is known to attain a proton pump on its outer membrane, as well as lysosomal LAMP2 (lysosomal associated membrane protein 2), well before the formation of a bone fide autolysosome.179,182,183 Despite autophagosomes having been shown to fuse to a similar extent with early and late endosomes in vitro,136 these different steps appear to use distinct fusion machineries.
Autophagosomes fuse with endosomes and/or MVBs to form amphisomes
Rab11 has been shown to be required for the fusion of autophagosomes to MVBs in K562 cells.184 Rab11 has however also been shown to be required for autophagosome formation,102 meaning that Rab11 may serve two separate roles during assembly and maturation. The Rab GAP TBC1D5 has also been potentially implicated during autophagosome maturation. It binds both LC3 and the endosome competitively, and has two distinct LIR motifs.151 This means that TBC1D5 may form an important bridge between the endosome and the autophagosome, potentially aiding the tethering of these two compartments.
A second function for UVRAG – this time independent of Beclin1, and thus the VPS34 complex in general – is in binding to the endosomal tether C-VPS, to mediate fusion of autophagosomes with endosomes and/or lysosomes.185 Interestingly, it is an entirely different subdomain on UVRAG that binds Beclin1 when in conjunction with the VPS34 complex, than when binding C-VPS; thus Beclin1 and C-VPS do not compete for access to UVRAG.185 UVRAG is probably involved in regulating/mediating fusion within the endosomal network, independently of autophagy.185 The C-VPS complex is the core of either the HOPS (homotypic fusion and vacuole protein sorting) or the CORVET (class C core vacuole/endosome tethering) complex involved in endosome-endosome or endosome-lysosome fusion.186 A SNARE implicated in the autophagosome-endosome fusion stage is VAMP3 (vesicle associated membrane protein 3)/cellubrevin. VAMP3 has been shown to be required for the fusion of autophagosomes with MVBs in erythroid maturation.187 Here, it is worth noting that VAMP3 has also been shown to be required for homotypic fusion of ATG9 and ATG16L1 containing pre-autophagic membrane structures during autophagosome initiation.99
Autophagosome fusion with the lysosome
Fusion of autophagosomal structures with the lysosomal compartment depends upon defined membrane identity, as dictated by the phosphoinositide PI(3,5)P2. This phosphoinositide is required for endomembrane acidification, and for autophagosome maturation in mammalian cells,188 but whether it plays a direct role in autophagy has yet to be confirmed.74 Other phosphoinositides and lipids have been identified as having autophagy roles in a variety of studies, and these are summarized well elsewhere.74 Rab24 has been implicated in autophagy because its position changes drastically upon starvation, to colocalize with autophagic vacuoles.189 Rab24 is thought to influence autophagosome-lysosome fusion with the aid of the drs tumor suppressor.190 Rab7 is implicated in autophagosome/lysosome fusion,154 possibly in conjunction with HOPS complex GEF activity.191–193
The ESCRT (endosomal sorting complex required for transport) machinery has also been implicated during late stages of autophagosomal maturation. ESCRT is required for autophagosome-lysosome fusion in human cells and in Drosophila.194–196 The ESCRT complex was originally identified through its involvement in MVB biogenesis and the sorting of ubiquitinated proteins to MVBs and lysosomes.197 It is possible that the ESCRT machinery is directly involved in autophagosome maturation, but the data also support a situation where autophagy requires functional MVBs.
Proteins acting at the lysosomal membrane have also been implicated in autophagosome maturation; LAMP1 and LAMP2 deficient mouse embryonic fibroblasts (MEFs) have been shown to accumulate immature and mature autophagosomes.198,199 Surprisingly, lysosomal protease activity in these cells appeared to be normal, but the lysosomes showed some structural abnormalities.198,199 This led to the hypothesis that LAMPs may be involved in autophagosome-lysosome fusion, although it is very difficult to tell whether this is a specific effect of inhibition of fusion, or merely due to attenuated lysosomal function.198,199 Impaired phagosome fusion with lysosomes in LAMP1 and LAMP2 double knockout MEFs has been shown to be due to an inability of phagosomes to move toward the lysosome.200 This means that the same could hold true for autophagosome-lysosome fusion; an absence of LAMP1/2 would mean a defect in trafficking, and thus a coincidental fusion defect rather than a mechanistic one.
The autophagosome-lysosome fusion event itself is likely to be governed by SNARE proteins. In yeast there are reports of a number of SNAREs being implicated,201 for example syntaxin 17.202 Upon starvation, syntaxin 17 translocates to fully formed autophagosomes, where it mediates fusion of the autophagosome with the lysosome through its partner SNAREs, SNAP-29 (soluble N-ethylmaleimide-sensitive factor attachment protein 29) and VAMP8.202 Indeed, both Vtib1b (vesicle transport through interaction with the t-SNAREs homologue 1B) and VAMP8 have been shown independently to be required for autophagosome-lysosome fusion,203,204 and TI-VAMP/VAMP7 has been implicated in amphisome-lysosome fusion.187 Mechanistically, syntaxin 17 has been shown to directly interact with the HOPS complex, providing HOPS with a second potential role in autophagosome maturation.205 That differing SNAREs are required for different steps in the autophagosome maturation process indicates a high level of regulation. It is worth noting that these same SNAREs have been identified as a requirement for the maturation of the autophagosome precursors.99 Lastly, and perhaps surprisingly, the ATG5-12 conjugate has also been implicated in autophagosome lysosome fusion; TECPR1 (Tectonin Beta-Propeller Repeat Containing 1), which localizes to autolysosomes, is suggested to bind to PI3P and to ATG5-12 in a manner that excludes ATG16, to mediate tethering and fusion.206
Membrane sources for the autophagosome
There are two models for the source of the membrane in the early autophagosome: one is the maturation model and the other is the assembly model. The maturation model proposes that the membranes forming the early autophagosome are derived from the ER.207 The assembly model states that the membranes needed for autophagosome assembly either stem from a separate, dedicated pool, or are synthesized de novo at the assembly site.208 These two models are not mutually exclusive, and most probably autophagy utilizes both. In addition to the two hypotheses regarding membrane origins, there are two theories on the involvement of the ER in autophagosome biogenesis. One of them states that the ER is merely a scaffold on which the autophagosome forms, whereas the other claims that the autophagosome is contiguous with, and therefore an extension of the ER.
The ER as a cradle for autophagosome biogenesis
Using a reporter protein that binds selectively to PI3P (double FYVE-containing protein 1; DFCP1) Axe et al72 demonstrated that PI3P defines a subdomain of the ER that is formed upon autophagy induction. They showed that this domain forms a cup-like structure, the center of which progressively recruits LC3, implying that the autophagosome forms in a cradle of ER membrane.72 The early stages of autophagosome formation have also been described by electron microscopy tomography. Independent studies found that autophagosomes are synthesized upon the ER, and provided evidence that the nascent autophagosome is cradled between two ER cisternae.209,210 These high-resolution studies suggested that there might be a physical connection between the autophagic isolation membrane and the ER. This would imply that the ER is both a scaffold for autophagosome formation, and a possible donor of membranes. Whilst compelling, it must be pointed out that these observations were based on EM tomography reconstructions, which are necessarily subjective. Interestingly, 70% of autophagosomes have been shown to contain a portion of ER membrane, implying that during the formation process the inner ER scaffold might become enclosed by the nascent autophagosome.209 It is also interesting to note that the isolation membrane often appears unusually electron dense when visualized by electron microscopy following aldehyde fixation, implying that the double membranes might be in very close proximity (at least in fixed samples), which hints at a mechanism for lumenal and membrane protein exclusion from the expanding isolation membrane. At present very little is known about how the ER is reorganized to accommodate the nascent phagophore, and a molecular explanation is needed.
Membranes of diverse organelles contribute to the forming autophagosome
Aside from the ER, many other organelles have been implicated in membrane delivery to the expanding autophagosome. Determining the origin of the autophagosomal membrane is problematic, though, because the autophagosomal membrane appears to be devoid of ancillary markers.211 Early autophagy markers (ATG2, ATG18, and ATG9) have often been observed in close proximity to ER exit sites (ERES) in yeast.212 The involvement of ERES in autophagosome formation has been confirmed by a proteomics study, which showed that phagophores and ERES were physically and functionally linked.213 The authors speculated that in yeast, autophagosome biogenesis is linked to the generation of a COPII compartment that is similar to the ERGIC (ER-Golgi intermediate compartment) in mammals, which agrees with a separate study showing involvement of the ERGIC in autophagosome assembly.213,214 Another finding that backs up the requirement for the ERGIC in mammals or the ERES in yeast is that specific COPII components are required for autophagy.215 The aforementioned study showed that the ERGIC compartment is needed for LC3 lipidation.214 Pharmacological disruption of the ERGIC prevented efficient generation of ATG14 puncta,214 suggesting that a functional ERGIC may be required for very early initiation stages of autophagosome biogenesis (specification of the assembly site itself). If membrane contributions to the phagophore from the ERGIC were a limiting factor, then one might have expected to record an accumulation and not a depletion of ATG14 puncta. Finally, ATG9 has been proposed to traffic from, or at least through the Golgi and the ERGIC, leading to the hypothesis that Golgi membranes contribute to the expanding autophagosome.103,216 Golgi-derived membranes have further been proposed to contribute to the phagophore, as Golgi derived lectins localize to the expanding rims of the phagophore.211
Mitochondria have been suggested to contribute lipids directly to the forming autophagosome.217 This could be either through direct lipid exchange or by a more complex autophagy-specific remodeling event. More recently, the MAM (mitochondrial-associated membrane at the ER) has been identified as a possible site of the autophagosome initiation.218 Here, important autophagosome assembly molecules including Beclin1 and ATG14L have been shown to concentrate,218 and the implications for this with respect to autophagy and ER-to-mitochondrial calcium homeostasis are covered elsewhere.219
The endosomal system is increasingly being implicated as a major contributor to the process of autophagosome assembly. A need for a functional early endosomal system for autophagosome progression has been widely shown (eg, (184)). The recycling endosome may contribute membrane to the autophagosome, as ULK1 has been shown to initially relocalize to the recycling endosome upon starvation.67,102 This study is supported in part by evidence that endosomal sorting nexin 18 is required for autophagy through its role in tubulating the endosome as a prelude to membrane delivery to the autophagosome.220 Additionally, both ATG9 and ATG16L1 have been shown to cycle through the recycling endosome, potentially acquiring membrane en route to contribute to the autophagosome expansion.100,125 In a similar fashion, the plasma membrane has been suggested to donate membrane to the autophagosome via ATG16L1 vesicles.98,221 Furthermore, ATG9 has more recently been shown to also localize to the plasma membrane prior to moving to the recycling endosome, which means that ATG9 could also bring part of the plasma membrane with it. Although internalized separately, ATG9 and ATG16L1 coalesce within the same endocytic subcompartment upon starvation.125 Lipid droplets have been shown to donate lipid to the nascent autophagosome by proximity.222 Further to this, the mobilization of lipids with the aid of a lipase was shown to be required for autophagy. Dupont et al222 identified PNPLA5 (patatin-like phospholipase domain containing 5) as such a lipase. Finally, the plasma membrane has been proposed to act as a reservoir for autophagy proteins that become anchored to connexin 43.223 Upon starvation, connexin 43 is degraded releasing ATG16L1 to amplify the autophagy response.223 As autophagy is such a crucial facet of cellular stress response pathways, it is likely that additional regulatory pathways that converge with autophagy will be revealed in the coming years.
Disclosure
The authors report no conflicts of interest in this work.
References
Klionsky DJ, Baehrecke EH, Brumell JH, et al. A comprehensive glossary of autophagy-related molecules and processes (2nd edition). Autophagy. 2011;7(11):1273–1294. PubMed PMID: 21997368. Epub 2011/10/15. eng. | |
Mukaiyama H, Baba M, Osumi M, et al. Modification of a ubiquitin-like protein Paz2 conducted micropexophagy through formation of a novel membrane structure. Mol Biol Cell. 2004;15(1):58–70. PubMed PMID: 13679515. Pubmed Central PMCID: 307527. | |
Santambrogio L, Cuervo AM. Chasing the elusive mammalian microautophagy. Autophagy. 2011;7(6):652–654. PubMed PMID: 21460618. | |
Li W, Yang Q, Mao Z. Chaperone-mediated autophagy: machinery, regulation and biological consequences. Cell Mol Life Sci. 2011;68(5):749–763. PubMed PMID: 20976518. | |
Pankiv S, Clausen TH, Lamark T, et al. p62/SQSTM1 binds directly to Atg8/LC3 to facilitate degradation of ubiquitinated protein aggregates by autophagy. J Biol Chem. 2007;282(33):24131–24145. PubMed PMID: 17580304. | |
Sawa-Makarska J, Abert C, Romanov J, Zens B, Ibiricu I, Martens S. Cargo binding to Atg19 unmasks additional Atg8 binding sites to mediate membrane-cargo apposition during selective autophagy. Nat Cell Biol. 2014;16(5):425–433. PubMed PMID: 24705553. Pubmed Central PMCID: 4009068. | |
Codogno P, Meijer AJ. Autophagy and signaling: their role in cell survival and cell death. Cell Death Differ. 2005;12(Suppl 2):1509–1518. PubMed PMID: 16247498. | |
De Duve C. The lysosome. Sci Am. 1963;208:64–72. PubMed PMID: 14025755. | |
Klionsky DJ, Ohsumi Y. Vacuolar import of proteins and organelles from the cytoplasm. Annu Rev Cell Dev Biol. 1999;15:1–32. PubMed PMID: 10611955. | |
Tsukada M, Ohsumi Y. Isolation and characterization of autophagy-defective mutants of Saccharomyces cerevisiae. FEBS Lett. 1993;333(1–2):169–174. PubMed PMID: 8224160. | |
Thumm M, Egner R, Koch B, et al. Isolation of autophagocytosis mutants of Saccharomyces cerevisiae. FEBS Lett. 1994;349(2):275–280. PubMed PMID: 8050581. | |
Levine B, Yuan J. Autophagy in cell death: an innocent convict? J Clin Invest. 2005;115(10):2679–2688. PubMed PMID: 16200202. Pubmed Central PMCID: 1236698. | |
Marino G, Niso-Santano M, Baehrecke EH, Kroemer G. Self-consumption: the interplay of autophagy and apoptosis. Nat Rev Mol Cell Biol. 2014;15(2):81–94. PubMed PMID: 24401948. Pubmed Central PMCID: 3970201. | |
Musiwaro P, Smith M, Manifava M, Walker SA, Ktistakis NT. Characteristics and requirements of basal autophagy in HEK 293 cells. Autophagy. 2013;9(9):1407–1417. PubMed PMID: 23800949. | |
Morselli E, Maiuri MC, Markaki M, et al. Caloric restriction and resveratrol promote longevity through the Sirtuin-1-dependent induction of autophagy. Cell Death Dis. 2010;1:e10. PubMed PMID: 21364612. Pubmed Central PMCID: 3032517. | |
Jiang P, Mizushima N. Autophagy and human diseases. Cell Res. 2014;24(1):69–79. PubMed PMID: 24323045. Pubmed Central PMCID: 3879707. | |
Deretic V, Saitoh T, Akira S. Autophagy in infection, inflammation and immunity. Nat Rev Immunol. 2013;13(10):722–737. PubMed PMID: 24064518. | |
Wileman T. Autophagy as a defence against intracellular pathogens. Essays Biochem. 2013;55:153–163. PubMed PMID: 24070478. | |
Banerjee R, Beal MF, Thomas B. Autophagy in neurodegenerative disorders: pathogenic roles and therapeutic implications. Trends Neurosci. 2010;33(12):541–549. PubMed PMID: 20947179. Pubmed Central PMCID: 2981680. | |
Nixon RA. The role of autophagy in neurodegenerative disease. Nat Med. 2013;19(8):983–997. PubMed PMID: 23921753. | |
Guo JY, Xia B, White E. Autophagy-mediated tumor promotion. Cell. 20135;155(6):1216–1219. PubMed PMID: 24315093. Pubmed Central PMCID: 3987898. | |
White E. Deconvoluting the context-dependent role for autophagy in cancer. Nat Rev Cancer. 2012;12(6):401–410. PubMed PMID: 22534666. Pubmed Central PMCID: 3664381. | |
Mathew R, Karantza-Wadsworth V, White E. Role of autophagy in cancer. Nat Rev Cancer. 2007;7(12):961–967. PubMed PMID: 17972889. Pubmed Central PMCID: 2866167. Epub 2007/11/02. eng. | |
Knodler LA, Celli J. Eating the strangers within: host control of intracellular bacteria via xenophagy. Cell Microbiol. 2011;13(9):1319–1327. PubMed PMID: 21740500. Pubmed Central PMCID: 3158265. | |
Lamark T, Johansen T. Aggrephagy: selective disposal of protein aggregates by macroautophagy. Int J Cell Biol. 2012;2012:736905. PubMed PMID: 22518139. Pubmed Central PMCID: 3320095. | |
Sarkar S, Rubinsztein DC. Huntington’s disease: degradation of mutant huntingtin by autophagy. FEBS J. 2008;275(17):4263–4270. PubMed PMID: 18637946. | |
Lemasters JJ. Selective mitochondrial autophagy, or mitophagy, as a targeted defense against oxidative stress, mitochondrial dysfunction, and aging. Rejuvenation Res. 2005;8(1):3–5. PubMed PMID: 15798367. Epub 2005/03/31. eng. | |
MacVicar T. Mitophagy. Essays Biochem. 2013;55:93–104. PubMed PMID: 24070474. | |
Betin VM, Singleton BK, Parsons SF, Anstee DJ, Lane JD. Autophagy facilitates organelle clearance during differentiation of human erythroblasts: evidence for a role for ATG4 paralogs during autophagosome maturation. Autophagy. 2013;9(6):881–893. PubMed PMID: 23508006. Epub 2013/03/20. Eng. | |
Yamano K, Youle RJ. PINK1 is degraded through the N-end rule pathway. Autophagy. 2013;9(11):1758–1769. PubMed PMID: 24121706. | |
MacVicar TD, Lane JD. Impaired OMA1-dependent cleavage of OPA1 and reduced DRP1 fission activity combine to prevent mitophagy in cells that are dependent on oxidative phosphorylation. J Cell Sci. 2014;127(Pt 10):2313–2325. PubMed PMID: 24634514. Pubmed Central PMCID: 4021475. | |
Gomes LC, Benedetto GD, Scorrano L. During autophagy mitochondria elongate, are spared from degradation and sustain cell viability. Nat Cell Biol. 2011;13(5):589–598. PubMed PMID: 21478857. Pubmed Central PMCID: 3088644. Epub 2011/04/12. eng. | |
Tondera D, Grandemange S, Jourdain A, et al. SLP-2 is required for stress-induced mitochondrial hyperfusion. EMBO J. 2009;28(11):1589–1600. PubMed PMID: 19360003. Pubmed Central PMCID: 2693158. Epub 2009/04/11. eng. | |
Bernales S, McDonald KL, Walter P. Autophagy counterbalances endoplasmic reticulum expansion during the unfolded protein response. PLoS Biol. 2006;4(12):e423. PubMed PMID: 17132049. Pubmed Central PMCID: 1661684. | |
Sakai Y, Oku M, van der Klei IJ, Kiel JA. Pexophagy: autophagic degradation of peroxisomes. Biochim Biophys Acta. 2006;1763(12):1767–1775. PubMed PMID: 17005271. | |
Svenning S, Johansen T. Selective autophagy. Essays Biochem. 2013;55:79–92. PubMed PMID: 24070473. | |
Harding TM, Morano KA, Scott SV, Klionsky DJ. Isolation and characterization of yeast mutants in the cytoplasm to vacuole protein targeting pathway. J Cell Biol. 1995;131(3):591–602. PubMed PMID: 7593182. Pubmed Central PMCID: 2120622. | |
Klionsky DJ. Look people, “Atg” is an abbreviation for “autophagy-related.” That’s it. Autophagy. 2012;8(9):1281–1282. PubMed PMID: 22889836. Pubmed Central PMCID: 3442874. | |
Simonsen A, Tooze SA. Coordination of membrane events during autophagy by multiple class III PI3-kinase complexes. J Cell Biol. 2009;186(6):773–782. PubMed PMID: 19797076. Pubmed Central PMCID: 2753151. Epub 2009/10/03. eng. | |
Koyama-Honda I, Itakura E, Fujiwara TK, Mizushima N. Temporal analysis of recruitment of mammalian ATG proteins to the autophagosome formation site. Autophagy. 2013;9(10):1491–1499. PubMed PMID: 23884233. | |
Suzuki K, Kubota Y, Sekito T, Ohsumi Y. Hierarchy of Atg proteins in pre-autophagosomal structure organization. Genes Cells. 2007;12(2):209–218. PubMed PMID: 17295840. | |
Itakura E, Mizushima N. Characterization of autophagosome formation site by a hierarchical analysis of mammalian Atg proteins. Autophagy. 2010;6(6):764–776. PubMed PMID: 20639694. Pubmed Central PMCID: 3321844. | |
Karanasios E, Stapleton E, Manifava M, et al. Dynamic association of the ULK1 complex with omegasomes during autophagy induction. J Cell Sci. 2013;126(Pt 22):5224–5238. PubMed PMID: 24013547. | |
Ktistakis NT, Karanasios E, Manifava M. Dynamics of autophagosome formation: a pulse and a sequence of waves. Biochem Soc Trans. 2014;42(5):1389–1395. PubMed PMID: 25233420. | |
Gallagher LE, Chan EY. Early signalling events of autophagy. Essays Biochem. 2013;55:1–15. PubMed PMID: 24070467. | |
Kim E, Goraksha-Hicks P, Li L, Neufeld TP, Guan KL. Regulation of TORC1 by Rag GTPases in nutrient response. Nat Cell Biol. 2008;10(8):935–945. PubMed PMID: 18604198. Pubmed Central PMCID: 2711503. | |
Sancak Y, Peterson TR, Shaul YD, et al. The Rag GTPases bind raptor and mediate amino acid signaling to mTORC1. Science. 2008;320(5882):1496–1501. PubMed PMID: 18497260. Pubmed Central PMCID: 2475333. | |
Bar-Peled L, Schweitzer LD, Zoncu R, Sabatini DM. Ragulator is a GEF for the rag GTPases that signal amino acid levels to mTORC1. Cell. 2012;150(6):1196–1208. PubMed PMID: 22980980. Pubmed Central PMCID: 3517996. | |
Inoki K, Kim J, Guan KL. AMPK and mTOR in cellular energy homeostasis and drug targets. Annu Rev Pharmacol Toxicol. 2012;52:381–400. PubMed PMID: 22017684. | |
Kim J, Kundu M, Viollet B, Guan KL. AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat Cell Biol. 2011;13(2):132–141. PubMed PMID: 21258367. Pubmed Central PMCID: 3987946. | |
He C, Klionsky DJ. Regulation mechanisms and signaling pathways of autophagy. Annu Rev Genet. 2009;43:67–93. PubMed PMID: 19653858. Pubmed Central PMCID: 2831538. | |
Budovskaya YV, Stephan JS, Deminoff SJ, Herman PK. An evolutionary proteomics approach identifies substrates of the cAMP-dependent protein kinase. Proc Natl Acad Sci U S A. 2005;102(39):13933–13938. PubMed PMID: 16172400. Pubmed Central PMCID: 1236527. | |
Cherra SJ 3rd, Kulich SM, Uechi G, et al. Regulation of the autophagy protein LC3 by phosphorylation. J Cell Biol. 2010;190(4):533–539. PubMed PMID: 20713600. Pubmed Central PMCID: 2928022. | |
Jacinto E. What controls TOR? IUBMB Life. 2008;60(8):483–496. PubMed PMID: 18493947. | |
Hosokawa N, Hara T, Kaizuka T, et al. Nutrient-dependent mTORC1 association with the ULK1-Atg13-FIP200 complex required for autophagy. Mol Biol Cell. 2009;20(7):1981–1991. PubMed PMID: 19211835. Pubmed Central PMCID: 2663915. Epub 2009/02/13. eng. | |
Ganley IG, Wong PM, Gammoh N, Jiang X. Distinct autophagosomal-lysosomal fusion mechanism revealed by thapsigargin-induced autophagy arrest. Mol Cell. 2011;42(6):731–743. PubMed PMID: 21700220. Pubmed Central PMCID: 3124681. | |
Jung CH, Jun CB, Ro SH, et al. ULK-Atg13-FIP200 complexes mediate mTOR signaling to the autophagy machinery. Mol Biol Cell. 2009;20(7):1992–2003. PubMed PMID: 19225151. Pubmed Central PMCID: 2663920. | |
Mercer CA, Kaliappan A, Dennis PB. A novel, human Atg13 binding protein, Atg101, interacts with ULK1 and is essential for macroautophagy. Autophagy. 2009;5(5):649–662. PubMed PMID: 19287211. Epub 2009/03/17. eng. | |
Matsunaga K, Morita E, Saitoh T, et al. Autophagy requires endoplasmic reticulum targeting of the PI3-kinase complex via Atg14L. J Cell Biol. 2010;190(4):511–521. PubMed PMID: 20713597. Pubmed Central PMCID: 2928018. | |
Di Bartolomeo S, Corazzari M, Nazio F, et al. The dynamic interaction of AMBRA1 with the dynein motor complex regulates mammalian autophagy. J Cell Biol. 2010;191(1):155–168. PubMed PMID: 20921139. Pubmed Central PMCID: 2953445. | |
Russell RC, Tian Y, Yuan H, et al. ULK1 induces autophagy by phosphorylating Beclin-1 and activating VPS34 lipid kinase. Nat Cell Biol. 2013;15(7):741–750. PubMed PMID: 23685627. Pubmed Central PMCID: 3885611. | |
Jung CH, Seo M, Otto NM, Kim DH. ULK1 inhibits the kinase activity of mTORC1 and cell proliferation. Autophagy. 2011;7(10):1212–1221. PubMed PMID: 21795849. Pubmed Central PMCID: 3210307. | |
Hara T, Takamura A, Kishi C, et al. FIP200, a ULK-interacting protein, is required for autophagosome formation in mammalian cells. J Cell Biol. 2008;181(3):497–510. PubMed PMID: 18443221. Pubmed Central PMCID: 2364687. | |
Ragusa MJ, Stanley RE, Hurley JH. Architecture of the Atg17 complex as a scaffold for autophagosome biogenesis. Cell. 2012;151(7):1501–1512. PubMed PMID: 23219485. Pubmed Central PMCID: 3806636. | |
Cheong H, Nair U, Geng J, Klionsky DJ. The Atg1 kinase complex is involved in the regulation of protein recruitment to initiate sequestering vesicle formation for nonspecific autophagy in Saccharomyces cerevisiae. Mol Biol Cell. 2008;19(2):668–681. PubMed PMID: 18077553. Pubmed Central PMCID: 2230592. | |
Kraft C, Kijanska M, Kalie E, et al. Binding of the Atg1/ULK1 kinase to the ubiquitin-like protein Atg8 regulates autophagy. EMBO J. 2012;31(18):3691–3703. PubMed PMID: 22885598. Pubmed Central PMCID: 3442273. | |
Longatti A, Tooze SA. Recycling endosomes contribute to autophagosome formation. Autophagy. 2012;8(11):1682–1683. PubMed PMID: 22874560. Pubmed Central PMCID: 3494599. | |
Hutagalung AH, Novick PJ. Role of Rab GTPases in membrane traffic and cell physiology. Physiol Rev. 2011;91(1):119–149. PubMed PMID: 21248164. Pubmed Central PMCID: 3710122. | |
Chan EY, Tooze SA. Evolution of Atg1 function and regulation. Autophagy. 2009;5(6):758–765. PubMed PMID: 19411825. | |
Wirth M, Joachim J, Tooze SA. Autophagosome formation – the role of ULK1 and Beclin1-PI3KC3 complexes in setting the stage. Semin Cancer Biol. 2013;23(5):301–309. PubMed PMID: 23727157. | |
Di Paolo G, De Camilli P. Phosphoinositides in cell regulation and membrane dynamics. Nature. 2006;443(7112):651–657. PubMed PMID: 17035995. | |
Axe EL, Walker SA, Manifava M, et al. Autophagosome formation from membrane compartments enriched in phosphatidylinositol 3-phosphate and dynamically connected to the endoplasmic reticulum. J Cell Biol. 2008;182(4):685–701. PubMed PMID: 18725538. | |
Behnia R, Munro S. Organelle identity and the signposts for membrane traffic. Nature. 2005;438(7068):597–604. PubMed PMID: 16319879. | |
Dall’Armi C, Devereaux KA, Di Paolo G. The role of lipids in the control of autophagy. Curr Biol. 2013;23(1):R33–R45. PubMed PMID: 23305670. Pubmed Central PMCID: 3587843. | |
Itakura E, Kishi C, Inoue K, Mizushima N. Beclin 1 forms two distinct phosphatidylinositol 3-kinase complexes with mammalian Atg14 and UVRAG. Mol Biol Cell. 2008;19(12):5360–5372. PubMed PMID: 18843052. Pubmed Central PMCID: 2592660. | |
Fan W, Nassiri A, Zhong Q. Autophagosome targeting and membrane curvature sensing by Barkor/Atg14(L). Proc Natl Acad Sci U S A. 2011;108(19):7769–7774. PubMed PMID: 21518905. Pubmed Central PMCID: 3093500. | |
Backer JM. The regulation and function of Class III PI3Ks: novel roles for Vps34. Biochem J. 2008;410(1):1–17. PubMed PMID: 18215151. | |
Liang C, Feng P, Ku B, et al. Autophagic and tumour suppressor activity of a novel Beclin1-binding protein UVRAG. Nat Cell Biol. 2006;8(7):688–699. PubMed PMID: 16799551. | |
Takahashi Y, Coppola D, Matsushita N, et al. Bif-1 interacts with Beclin 1 through UVRAG and regulates autophagy and tumorigenesis. Nat Cell Biol. 2007;9(10):1142–1151. PubMed PMID: 17891140. Pubmed Central PMCID: 2254521. | |
Matsunaga K, Saitoh T, Tabata K, et al. Two Beclin 1-binding proteins, Atg14L and Rubicon, reciprocally regulate autophagy at different stages. Nat Cell Biol. 2009;11(4):385–396. PubMed PMID: 19270696. | |
Kang R, Zeh HJ, Lotze MT, Tang D. The Beclin 1 network regulates autophagy and apoptosis. Cell Death Differ. 2011;18(4):571–580. PubMed PMID: 21311563. Pubmed Central PMCID: 3131912. | |
Fimia GM, Stoykova A, Romagnoli A, et al. Ambra1 regulates autophagy and development of the nervous system. Nature. 2007; 447(7148):1121–1125. PubMed PMID: 17589504. | |
Nazio F, Strappazzon F, Antonioli M, et al. mTOR inhibits autophagy by controlling ULK1 ubiquitylation, self-association and function through AMBRA1 and TRAF6. Nat Cell Biol. 2013;15(4):406–416. PubMed PMID: 23524951. | |
Vergne I, Roberts E, Elmaoued RA, et al. Control of autophagy initiation by phosphoinositide 3-phosphatase Jumpy. EMBO J. 2009;28(15):2244–2258. PubMed PMID: 19590496. Pubmed Central PMCID: 2726690. | |
Cebollero E, van der Vaart A, Zhao M, et al. Phosphatidylinositol-3-phosphate clearance plays a key role in autophagosome completion. Curr Biol. 2012;22(17):1545–1553. PubMed PMID: 22771041. Pubmed Central PMCID: 3615650. | |
Cheng J, Fujita A, Yamamoto H, et al. Yeast and mammalian autophagosomes exhibit distinct phosphatidylinositol 3-phosphate asymmetries. Nat Commun. 2014;5:3207. PubMed PMID: 24492518. | |
Blommaart EF, Krause U, Schellens JP, Vreeling-Sindelarova H, Meijer AJ. The phosphatidylinositol 3-kinase inhibitors wortmannin and LY294002 inhibit autophagy in isolated rat hepatocytes. Eur J Biochem. 1997;243(1–2):240–246. PubMed PMID: 9030745. | |
Seglen PO, Gordon PB. 3-Methyladenine: specific inhibitor of autophagic/lysosomal protein degradation in isolated rat hepatocytes. Proc Natl Acad Sci U S A. 1982;79(6):1889–1892. PubMed PMID: 6952238. | |
Lu Q, Yang P, Huang X, et al. The WD40 repeat PtdIns(3)P-binding protein EPG-6 regulates progression of omegasomes to autophagosomes. Dev Cell. 2011;21(2):343–357. PubMed PMID: 21802374. | |
Polson HE, de Lartigue J, Rigden DJ, et al. Mammalian Atg18 (WIPI2) localizes to omegasome-anchored phagophores and positively regulates LC3 lipidation. Autophagy. 2010;6(4):506–522. PubMed PMID: 20505359. | |
Proikas-Cezanne T, Waddell S, Gaugel A, Frickey T, Lupas A, Nordheim A. WIPI-1alpha (WIPI49), a member of the novel 7-bladed WIPI protein family, is aberrantly expressed in human cancer and is linked to starvation-induced autophagy. Oncogene. 2004;23(58):9314–9325. PubMed PMID: 15602573. | |
Dooley HC, Razi M, Polson HE, Girardin SE, Wilson MI, Tooze SA. WIPI2 links LC3 conjugation with PI3P, autophagosome formation, and pathogen clearance by recruiting Atg12-5-16L1. Mol Cell. 2014;55(2):238–252. PubMed PMID: 24954904. Pubmed Central PMCID: 4104028. | |
Fujita N, Itoh T, Omori H, Fukuda M, Noda T, Yoshimori T. The Atg16L complex specifies the site of LC3 lipidation for membrane biogenesis in autophagy. Mol Biol Cell. 2008;19(5):2092–2100. PubMed PMID: 18321988. Pubmed Central PMCID: 2366860. | |
Kaufmann A, Beier V, Franquelim HG, Wollert T. Molecular mechanism of autophagic membrane-scaffold assembly and disassembly. Cell. 2014;156(3):469–481. PubMed PMID: 24485455. | |
Romanov J, Walczak M, Ibiricu I, et al. Mechanism and functions of membrane binding by the Atg5-Atg12/Atg16 complex during autophagosome formation. EMBO J. 2012;31(22):4304–4317. PubMed PMID: 23064152. Pubmed Central PMCID: 3501226. | |
Mizushima N, Yamamoto A, Hatano M, et al. Dissection of autophagosome formation using Apg5-deficient mouse embryonic stem cells. J Cell Biol. 2001;152(4):657–668. PubMed PMID: 11266458. | |
Nishimura T, Kaizuka T, Cadwell K, et al. FIP200 regulates targeting of Atg16L1 to the isolation membrane. EMBO Rep. 2013;14(3):284–291. PubMed PMID: 23392225. Pubmed Central PMCID: 3589088. | |
Ravikumar B, Moreau K, Jahreiss L, Puri C, Rubinsztein DC. Plasma membrane contributes to the formation of pre-autophagosomal structures. Nat Cell Biol. 2010;12(8):747–757. PubMed PMID: 20639872. Pubmed Central PMCID: 2923063. | |
Moreau K, Ravikumar B, Renna M, Puri C, Rubinsztein DC. Autophagosome precursor maturation requires homotypic fusion. Cell. 2011;146(2):303–317. PubMed PMID: 21784250. Pubmed Central PMCID: 3171170. | |
Puri C, Renna M, Bento CF, Moreau K, Rubinsztein DC. Diverse autophagosome membrane sources coalesce in recycling endosomes. Cell. 2013;154(6):1285–1299. PubMed PMID: 24034251. Pubmed Central PMCID: 3791395. | |
Itoh T, Fujita N, Kanno E, Yamamoto A, Yoshimori T, Fukuda M. Golgi-resident small GTPase Rab33B interacts with Atg16L and modulates autophagosome formation. Mol Biol Cell. 2008;19(7):2916–2925. PubMed PMID: 18448665. Pubmed Central PMCID: 2441679. | |
Longatti A, Lamb CA, Razi M, Yoshimura S, Barr FA, Tooze SA. TBC1D14 regulates autophagosome formation via Rab11- and ULK1-positive recycling endosomes. J Cell Biol. 2012;197(5):659–675. PubMed PMID: 22613832. Pubmed Central PMCID: 3365497. | |
Geng J, Klionsky DJ. The Atg8 and Atg12 ubiquitin-like conjugation systems in macroautophagy. ‘Protein modifications: beyond the usual suspects’ review series. EMBO Rep. 2008;9(9):859–864. PubMed PMID: 18704115. Pubmed Central PMCID: 2529362. | |
Tanida I, Komatsu M, Ueno T, Kominami E. GATE-16 and GABARAP are authentic modifiers mediated by Apg7 and Apg3. Biochem Biophys Res Commun. 2003;300(3):637–644. PubMed PMID: 12507496. | |
Tanida I, Ueno T, Kominami E. LC3 conjugation system in mammalian autophagy. Int J Biochem Cell Biol. 2004;36(12):2503–2518. PubMed PMID: 15325588. | |
Kabeya Y, Mizushima N, Yamamoto A, Oshitani-Okamoto S, Ohsumi Y, Yoshimori T. LC3, GABARAP and GATE16 localize to autophagosomal membrane depending on form-II formation. J Cell Sci. 2004;117(Pt 13):2805–2812. PubMed PMID: 15169837. | |
Kirisako T, Ichimura Y, Okada H, et al. The reversible modification regulates the membrane-binding state of Apg8/Aut7 essential for autophagy and the cytoplasm to vacuole targeting pathway. J Cell Biol. 2000;151(2):263–276. PubMed PMID: 11038174. Pubmed Central PMCID: 2192639. | |
Tanida I, Tanida-Miyake E, Ueno T, Kominami E. The human homolog of Saccharomyces cerevisiae Apg7p is a Protein-activating enzyme for multiple substrates including human Apg12p, GATE-16, GABARAP, and MAP-LC3. J Biol Chem. 2001;276(3):1701–1706. PubMed PMID: 11096062. | |
Tanida I, Tanida-Miyake E, Komatsu M, Ueno T, Kominami E. Human Apg3p/Aut1p homologue is an authentic E2 enzyme for multiple substrates, GATE-16, GABARAP, and MAP-LC3, and facilitates the conjugation of hApg12p to hApg5p. J Biol Chem. 2002;277(16):13739–13744. PubMed PMID: 11825910. | |
Noda NN, Ohsumi Y, Inagaki F. ATG systems from the protein structural point of view. Chem Rev. 2009;109(4):1587–1598. PubMed PMID: 19236009. | |
Kaiser SE, Mao K, Taherbhoy AM, et al. Noncanonical E2 recruitment by the autophagy E1 revealed by Atg7-Atg3 and Atg7-Atg10 structures. Nat Struct Mol Biol. 2012;19(12):1242–1249. PubMed PMID: 23142976. Pubmed Central PMCID: 3515690. | |
Mizushima N, Yoshimori T, Ohsumi Y. Role of the Apg12 conjugation system in mammalian autophagy. Int J Biochem Cell Biol. 2003;35(5):553–561. PubMed PMID: 12672448. | |
Hanada T, Noda NN, Satomi Y, et al. The Atg12-Atg5 conjugate has a novel E3-like activity for protein lipidation in autophagy. J Biol Chem. 2007;282(52):37298–37302. PubMed PMID: 17986448. | |
Mari M, Griffith J, Rieter E, Krishnappa L, Klionsky DJ, Reggiori F. An Atg9-containing compartment that functions in the early steps of autophagosome biogenesis. J Cell Biol. 2010;190(6):1005–1022. PubMed PMID: 20855505. Pubmed Central PMCID: 3101592. | |
Young AR, Chan EY, Hu XW, et al. Starvation and ULK1-dependent cycling of mammalian Atg9 between the TGN and endosomes. J Cell Sci. 2006;119(Pt 18):3888–3900. PubMed PMID: 16940348. Epub 2006/08/31. eng. | |
Reggiori F, Shintani T, Nair U, Klionsky DJ. Atg9 cycles between mitochondria and the pre-autophagosomal structure in yeasts. Autophagy. 2005;1(2):101–109. PubMed PMID: 16874040. Pubmed Central PMCID: 1762033. | |
Monastyrska I, He C, Geng J, Hoppe AD, Li Z, Klionsky DJ. Arp2 links autophagic machinery with the actin cytoskeleton. Mol Biol Cell. 2008;19(5):1962–1975. PubMed PMID: 18287533. Pubmed Central PMCID: 2366845. | |
Tang HW, Wang YB, Wang SL, Wu MH, Lin SY, Chen GC. Atg1-mediated myosin II activation regulates autophagosome formation during starvation-induced autophagy. EMBO J. 2011;30(4):636–651. PubMed PMID: 21169990. Pubmed Central PMCID: 3041946. | |
Bialik S, Pietrokovski S, Kimchi A. Myosin drives autophagy in a pathway linking Atg1 to Atg9. EMBO J. 2011;30(4):629–630. PubMed PMID: 21326172. Pubmed Central PMCID: 3041959. | |
Reggiori F, Tucker KA, Stromhaug PE, Klionsky DJ. The Atg1-Atg13 complex regulates Atg9 and Atg23 retrieval transport from the pre-autophagosomal structure. Dev Cell. 2004;6(1):79–90. PubMed PMID: 14723849. | |
Orsi A, Razi M, Dooley HC, et al. Dynamic and transient interactions of Atg9 with autophagosomes, but not membrane integration, are required for autophagy. Mol Biol Cell. 2012;23(10):1860–1873. PubMed PMID: 22456507. Pubmed Central PMCID: 3350551. | |
Papinski D, Schuschnig M, Reiter W, et al. Early steps in autophagy depend on direct phosphorylation of Atg9 by the Atg1 kinase. Mol Cell. 2014;53(3):471–483. PubMed PMID: 24440502. Pubmed Central PMCID: 3978657. | |
Webber JL, Tooze SA. Coordinated regulation of autophagy by p38alpha MAPK through mAtg9 and p38IP. EMBO J. 2010;29(1):27–40. PubMed PMID: 19893488. Pubmed Central PMCID: 2808369. | |
Nair U, Jotwani A, Geng J, et al. SNARE proteins are required for macroautophagy. Cell. 2011;146(2):290–302. PubMed PMID: 21784249. Pubmed Central PMCID: 3143362. | |
Puri C, Renna M, Bento CF, Moreau K, Rubinsztein DC. ATG16L1 meets ATG9 in recycling endosomes: additional roles for the plasma membrane and endocytosis in autophagosome biogenesis. Autophagy. 2014;10(1):182–184. PubMed PMID: 24257061. | |
Shpilka T, Weidberg H, Pietrokovski S, Elazar Z. Atg8: an autophagy-related ubiquitin-like protein family. Genome Biol. 2011;12(7):226. PubMed PMID: 21867568. Pubmed Central PMCID: 3218822. Epub 2011/08/27. eng. | |
Tanida I, Sou YS, Minematsu-Ikeguchi N, Ueno T, Kominami E. Atg8L/Apg8L is the fourth mammalian modifier of mammalian Atg8 conjugation mediated by human Atg4B, Atg7 and Atg3. FEBS J. 2006;273(11):2553–2562. PubMed PMID: 16704426. | |
Tanida I, Sou YS, Ezaki J, Minematsu-Ikeguchi N, Ueno T, Kominami E. HsAtg4B/HsApg4B/autophagin-1 cleaves the carboxyl termini of three human Atg8 homologues and delipidates microtubule-associated protein light chain 3- and GABAA receptor-associated protein-phospholipid conjugates. J Biol Chem. 2004;279(35):36268–36276. PubMed PMID: 15187094. | |
Klionsky DJ, Abdalla FC, Abeliovich H, et al. Guidelines for the use and interpretation of assays for monitoring autophagy. Autophagy. 2012;8(4):445–544. PubMed PMID: 22966490. Pubmed Central PMCID: 3404883. | |
Coyle JE, Qamar S, Rajashankar KR, Nikolov DB. Structure of GABARAP in two conformations: implications for GABA(A) receptor localization and tubulin binding. Neuron. 2002;33(1):63–74. PubMed PMID: 11779480. | |
Kouno T, Mizuguchi M, Tanida I, et al. Solution structure of microtubule-associated protein light chain 3 and identification of its functional subdomains. J Biol Chem. 2005;280(26):24610–24617. PubMed PMID: 15857831. | |
Mann SS, Hammarback JA. Molecular characterization of light chain 3. A microtubule binding subunit of MAP1A and MAP1B. J Biol Chem. 1994;269(15):11492–11497. PubMed PMID: 7908909. | |
Wang H, Bedford FK, Brandon NJ, Moss SJ, Olsen RW. GABA(A)-receptor-associated protein links GABA(A) receptors and the cytoskeleton. Nature. 1999;397(6714):69–72. PubMed PMID: 9892355. | |
Nakatogawa H, Ichimura Y, Ohsumi Y. Atg8, a ubiquitin-like protein required for autophagosome formation, mediates membrane tethering and hemifusion. Cell. 2007;130(1):165–178. PubMed PMID: 17632063. Epub 2007/07/17. eng. | |
Weidberg H, Shpilka T, Shvets E, Abada A, Shimron F, Elazar Z. LC3 and GATE-16 N termini mediate membrane fusion processes required for autophagosome biogenesis. Dev Cell. 2011;20(4):444–454. PubMed PMID: 21497758. Epub 2011/04/19. eng. | |
Morvan J, Kochl R, Watson R, Collinson LM, Jefferies HB, Tooze SA. In vitro reconstitution of fusion between immature autophagosomes and endosomes. Autophagy. 2009;5(5):676–689. PubMed PMID: 19337031. Epub 2009/04/02. eng. | |
Xie Z, Nair U, Klionsky DJ. Atg8 controls phagophore expansion during autophagosome formation. Mol Biol Cell. 2008;19(8):3290–3298. PubMed PMID: 18508918. Pubmed Central PMCID: 2488302. Epub 2008/05/30. eng. | |
Xie Z, Nair U, Geng J, Szefler MB, Rothman ED, Klionsky DJ. Indirect estimation of the area density of Atg8 on the phagophore. Autophagy. 2009;5(2):217–220. PubMed PMID: 19088501. Pubmed Central PMCID: 2941343. | |
Alemu EA, Lamark T, Torgersen KM, et al. ATG8 family proteins act as scaffolds for assembly of the ULK complex: sequence requirements for LC3-interacting region (LIR) motifs. J Biol Chem. 2012;287(47):39275–39290. PubMed PMID: 23043107. Pubmed Central PMCID: 3501051. | |
Weidberg H, Shvets E, Shpilka T, Shimron F, Shinder V, Elazar Z. LC3 and GATE-16/GABARAP subfamilies are both essential yet act differently in autophagosome biogenesis. EMBO J. 2010;29(11):1792–1802. PubMed PMID: 20418806. Pubmed Central PMCID: 2885923. | |
Maruyama Y, Sou YS, Kageyama S, et al. LC3B is indispensable for selective autophagy of p62 but not basal autophagy. Biochem Biophys Res Commun. 2014;446(1):309–315. PubMed PMID: 24582747. | |
Lamark T, Kirkin V, Dikic I, Johansen T. NBR1 and p62 as cargo receptors for selective autophagy of ubiquitinated targets. Cell Cycle. 2009;8(13):1986–1990. PubMed PMID: 19502794. | |
Behrends C, Sowa ME, Gygi SP, Harper JW. Network organization of the human autophagy system. Nature. 2010;466(7302):68–76. PubMed PMID: 20562859. Pubmed Central PMCID: 2901998. | |
Novak I, Kirkin V, McEwan DG, et al. Nix is a selective autophagy receptor for mitochondrial clearance. EMBO Rep. 2009;11(1):45–51. PubMed PMID: 20010802. Pubmed Central PMCID: 2816619. Epub 2009/12/17. eng. | |
Noda NN, Kumeta H, Nakatogawa H, et al. Structural basis of target recognition by Atg8/LC3 during selective autophagy. Genes Cells. 2008;13(12):1211–1218. PubMed PMID: 19021777. | |
Rozenknop A, Rogov VV, Rogova NY, et al. Characterization of the interaction of GABARAPL-1 with the LIR motif of NBR1. J Mol Biol. 2011;410(3):477–487. PubMed PMID: 21620860. | |
von Muhlinen N, Akutsu M, Ravenhill BJ, et al. An essential role for the ATG8 ortholog LC3C in antibacterial autophagy. Autophagy. 2013;9(5):784–786. PubMed PMID: 23434839. Pubmed Central PMCID: 3669188. | |
Liu L, Feng D, Chen G, et al. Mitochondrial outer-membrane protein FUNDC1 mediates hypoxia-induced mitophagy in mammalian cells. Nat Cell Biol. 2012;14(2):177–185. PubMed PMID: 22267086. Epub 2012/01/24. eng. | |
Chen G, Han Z, Feng D, et al. A regulatory signaling loop comprising the PGAM5 phosphatase and CK2 controls receptor-mediated mitophagy. Mol Cell. 2014;54(3):362–377. PubMed PMID: 24746696. | |
Wu W, Tian W, Hu Z, et al. ULK1 translocates to mitochondria and phosphorylates FUNDC1 to regulate mitophagy. EMBO Rep. 2014;15(5):566–575. PubMed PMID: 24671035. | |
Popovic D, Akutsu M, Novak I, Harper JW, Behrends C, Dikic I. Rab GTPase-activating proteins in autophagy: regulation of endocytic and autophagy pathways by direct binding to human ATG8 modifiers. Mol Cell Biol. 2012;32(9):1733–1744. PubMed PMID: 22354992. Pubmed Central PMCID: 3347240. | |
Itoh T, Kanno E, Uemura T, Waguri S, Fukuda M. OATL1, a novel autophagosome-resident Rab33B-GAP, regulates autophagosomal maturation. J Cell Biol. 2011;192(5):839–853. PubMed PMID: 21383079. Pubmed Central PMCID: 3051816. Epub 2011/03/09. eng. | |
Pankiv S, Alemu EA, Brech A, et al. FYCO1 is a Rab7 effector that binds to LC3 and PI3P to mediate microtubule plus end-directed vesicle transport. J Cell Biol. 2010;188(2):253–269. PubMed PMID: 20100911. Pubmed Central PMCID: 2812517. | |
Jager S, Bucci C, Tanida I, et al. Role for Rab7 in maturation of late autophagic vacuoles. J Cell Sci. 2004;117(Pt 20):4837–4848. PubMed PMID: 15340014. Epub 2004/09/02. eng. | |
Ichimura Y, Kirisako T, Takao T, et al. A ubiquitin-like system mediates protein lipidation. Nature. 2000;408(6811):488–492. PubMed PMID: 11100732. | |
Nair U, Yen WL, Mari M, et al. A role for Atg8-PE deconjugation in autophagosome biogenesis. Autophagy. 2012;8(5):780–793. PubMed PMID: 22622160. Epub 2012/05/25. Eng. | |
Yu ZQ, Ni T, Hong B, et al. Dual roles of Atg8-PE deconjugation by Atg4 in autophagy. Autophagy. 2012;8(6):883–892. PubMed PMID: 22652539. Pubmed Central PMCID: 3427254. | |
Marino G, Uria JA, Puente XS, Quesada V, Bordallo J, Lopez-Otin C. Human autophagins, a family of cysteine proteinases potentially implicated in cell degradation by autophagy. J Biol Chem. 2003;278(6):3671–3678. PubMed PMID: 12446702. | |
Kumanomidou T, Mizushima T, Komatsu M, et al. The crystal structure of human Atg4b, a processing and de-conjugating enzyme for autophagosome-forming modifiers. J Mol Biol. 2006;355(4):612–618. PubMed PMID: 16325851. | |
Sugawara K, Suzuki NN, Fujioka Y, Mizushima N, Ohsumi Y, Inagaki F. Structural basis for the specificity and catalysis of human Atg4B responsible for mammalian autophagy. J Biol Chem. 2005;280(48):40058–40065. PubMed PMID: 16183633. | |
Betin VM, Lane JD. Caspase cleavage of Atg4D stimulates Atg8 processing and triggers mitochondrial targeting and apoptosis. J Cell Sci. 2009;122(Pt 14):2554–2566. | |
Hemelaar J, Lelyveld VS, Kessler BM, Ploegh HL. A single protease, Apg4B, is specific for the autophagy-related ubiquitin-like proteins GATE-16, MAP1-LC3, GABARAP, and Apg8L. J Biol Chem. 2003;278(51):51841–51850. PubMed PMID: 14530254. | |
Li M, Hou Y, Wang J, Chen X, Shao ZM, Yin XM. Kinetics comparisons of mammalian Atg4 homologues indicate selective preferences toward diverse Atg8 substrates. J Biol Chem. 2011;286(9):7327–7338. PubMed PMID: 21177865. Pubmed Central PMCID: 3044989. | |
Shu CW, Drag M, Bekes M, Zhai D, Salvesen GS, Reed JC. Synthetic substrates for measuring activity of autophagy proteases: autophagins (Atg4). Autophagy. 2010;6(7):936–947. PubMed PMID: 20818167. Pubmed Central PMCID: 3039740. Epub 2010/09/08. eng. | |
Scherz-Shouval R, Sagiv Y, Shorer H, Elazar Z. The COOH terminus of GATE-16, an intra-Golgi transport modulator, is cleaved by the human cysteine protease HsApg4A. J Biol Chem. 2003;278(16):14053–14058. PubMed PMID: 12473658. | |
Kuang E, Okumura CY, Sheffy-Levin S, et al. Regulation of ATG4B stability by RNF5 limits basal levels of autophagy and influences susceptibility to bacterial infection. PLoS Genet. 2012;8(10):e1003007. PubMed PMID: 23093945. Pubmed Central PMCID: 3475677. | |
Scherz-Shouval R, Shvets E, Fass E, Shorer H, Gil L, Elazar Z. Reactive oxygen species are essential for autophagy and specifically regulate the activity of Atg4. EMBO J. 2007;26(7):1749–1760. PubMed PMID: 17347651. | |
Betin VMS, MacVicar TDB, Parsons SF, Anstee DJ, Lane JD. A cryptic mitochondrial targeting motif in Atg4D links caspase cleavage with mitochonrial import and oxidative stress. Autophagy. 2012;8(4):664–676. | |
Marino G, Fernandez AF, Cabrera S, et al. Autophagy is essential for mouse sense of balance. J Clin Invest. 2010;120(7):2331–2344. PubMed PMID: 20577052. Pubmed Central PMCID: 2898610. Epub 2010/06/26. eng. | |
Marino G, Salvador-Montoliu N, Fueyo A, Knecht E, Mizushima N, Lopez-Otin C. Tissue-specific autophagy alterations and increased tumorigenesis in mice deficient in ATG4C/autophagin-3. J Biol Chem. 2007;282(25):18573–18583. PubMed PMID: 17442669. | |
Read R, Savelieva K, Baker K, Hansen G, Vogel P. Histopathological and neurological features of Atg4b knockout mice. Vet Pathol. 2011;48(2):486–494. PubMed PMID: 20634410. | |
Nakatogawa H, Ishii J, Asai E, Ohsumi Y. Atg4 recycles inappropriately lipidated Atg8 to promote autophagosome biogenesis. Autophagy. 2012;8(2):177–186. PubMed PMID: 22240591. Epub 2012/01/14. eng. | |
Takeshige K, Baba M, Tsuboi S, Noda T, Ohsumi Y. Autophagy in yeast demonstrated with proteinase-deficient mutants and conditions for its induction. J Cell Biol. 1992;119(2):301–311. PubMed PMID: 1400575. Pubmed Central PMCID: 2289660. | |
Fujita N, Hayashi-Nishino M, Fukumoto H, et al. An Atg4B mutant hampers the lipidation of LC3 paralogues and causes defects in autophagosome closure. Mol Biol Cell. 2008;19(11):4651–4659. PubMed PMID: 18768752. | |
Molejon MI, Ropolo A, Re AL, Boggio V, Vaccaro MI. The VMP1-Beclin 1 interaction regulates autophagy induction. Sci Rep. 2013;3:1055. PubMed PMID: 23316280. Pubmed Central PMCID: 3542764. | |
De Antoni A, Schmitzova J, Trepte HH, Gallwitz D, Albert S. Significance of GTP hydrolysis in Ypt1p-regulated endoplasmic reticulum to Golgi transport revealed by the analysis of two novel Ypt1-GAPs. J Biol Chem. 2002;277(43):41023–41031. PubMed PMID: 12189143. | |
Lynch-Day MA, Bhandari D, Menon S, et al. Trs85 directs a Ypt1 GEF, TRAPPIII, to the phagophore to promote autophagy. Proc Natl Acad Sci U S A. 2010;107(17):7811–7816. PubMed PMID: 20375281. Pubmed Central PMCID: 2867920. | |
Kakuta S, Yamamoto H, Negishi L, Kondo-Kakuta C, Hayashi N, Ohsumi Y. Atg9 vesicles recruit vesicle-tethering proteins Trs85 and Ypt1 to the autophagosome formation site. J Biol Chem. 2012;287(53):44261–44269. PubMed PMID: 23129774. Pubmed Central PMCID: 3531741. | |
Dunn WA Jr. Studies on the mechanisms of autophagy: formation of the autophagic vacuole. J Cell Biol. 1990;110(6):1923–1933. PubMed PMID: 2351689. | |
Liou W, Geuze HJ, Geelen MJ, Slot JW. The autophagic and endocytic pathways converge at the nascent autophagic vacuoles. J Cell Biol. 1997;136(1):61–70. PubMed PMID: 9008703. | |
Tooze J, Hollinshead M, Ludwig T, Howell K, Hoflack B, Kern H. In exocrine pancreas, the basolateral endocytic pathway converges with the autophagic pathway immediately after the early endosome. J Cell Biol. 1990;111(2):329–345. PubMed PMID: 2166050. | |
Razi M, Chan EY, Tooze SA. Early endosomes and endosomal coatomer are required for autophagy. J Cell Biol. 2009;185(2):305–321. PubMed PMID: 19364919. Pubmed Central PMCID: 2700373. Epub 2009/04/15. eng. | |
Gutierrez MG, Munafo DB, Beron W, Colombo MI. Rab7 is required for the normal progression of the autophagic pathway in mammalian cells. J Cell Sci. 2004;117(Pt 13):2687–2697. PubMed PMID: 15138286. Epub 2004/05/13. eng. | |
Fader CM, Sanchez D, Furlan M, Colombo MI. Induction of autophagy promotes fusion of multivesicular bodies with autophagic vacuoles in k562 cells. Traffic. 2008;9(2):230–250. PubMed PMID: 17999726. | |
Liang C, Lee JS, Inn KS, et al. Beclin1-binding UVRAG targets the class C Vps complex to coordinate autophagosome maturation and endocytic trafficking. Nat Cell Biol. 2008;10(7):776–787. PubMed PMID: 18552835. Pubmed Central PMCID: 2878716. | |
Nickerson DP, Brett CL, Merz AJ. Vps-C complexes: gatekeepers of endolysosomal traffic. Curr Opin Cell Biol. 2009;21(4):543–551. PubMed PMID: 19577915. Pubmed Central PMCID: 2807627. | |
Fader CM, Sanchez DG, Mestre MB, Colombo MI. TI-VAMP/VAMP7 and VAMP3/cellubrevin: two v-SNARE proteins involved in specific steps of the autophagy/multivesicular body pathways. Biochim Biophys Acta. 2009;1793(12):1901–1916. PubMed PMID: 19781582. | |
Dove SK, Dong K, Kobayashi T, Williams FK, Michell RH. Phosphatidylinositol 3,5-bisphosphate and Fab1p/PIKfyve underPPIn endo-lysosome function. Biochem J. 2009;419(1):1–13. PubMed PMID: 19272020. | |
Munafo DB, Colombo MI. Induction of autophagy causes dramatic changes in the subcellular distribution of GFP-Rab24. Traffic. 2002;3(7):472–482. PubMed PMID: 12047555. | |
Tambe Y, Yamamoto A, Isono T, Chano T, Fukuda M, Inoue H. The drs tumor suppressor is involved in the maturation process of autophagy induced by low serum. Cancer Lett. 2009;283(1):74–83. PubMed PMID: 19368996. | |
Rieder SE, Emr SD. A novel RING finger protein complex essential for a late step in protein transport to the yeast vacuole. Mol Biol Cell. 1997;8(11):2307–2327. PubMed PMID: 9362071. Pubmed Central PMCID: 25710. | |
Seals DF, Eitzen G, Margolis N, Wickner WT, Price A. A Ypt/Rab effector complex containing the Sec1 homolog Vps33p is required for homotypic vacuole fusion. Proc Natl Acad Sci U S A. 2000;97(17):9402–9407. PubMed PMID: 10944212. Pubmed Central PMCID: 16876. | |
Wurmser AE, Sato TK, Emr SD. New component of the vacuolar class C-Vps complex couples nucleotide exchange on the Ypt7 GTPase to SNARE-dependent docking and fusion. J Cell Biol. 2000;151(3):551–562. PubMed PMID: 11062257. Pubmed Central PMCID: 2185595. | |
Filimonenko M, Stuffers S, Raiborg C, et al. Functional multivesicular bodies are required for autophagic clearance of protein aggregates associated with neurodegenerative disease. J Cell Biol. 2007;179(3):485–500. PubMed PMID: 17984323. Pubmed Central PMCID: 2064794. | |
Lee JA, Beigneux A, Ahmad ST, Young SG, Gao FB. ESCRT-III dysfunction causes autophagosome accumulation and neurodegeneration. Curr Biol. 2007;17(18):1561–1567. PubMed PMID: 17683935. | |
Rusten TE, Vaccari T, Lindmo K, et al. ESCRTs and Fab1 regulate distinct steps of autophagy. Curr Biol. 2007;17(20):1817–1825. PubMed PMID: 17935992. | |
Hurley JH. The ESCRT complexes. Crit Rev Biochem Mol Biol. 2010;45(6):463–487. PubMed PMID: 20653365. Pubmed Central PMCID: 2988974. | |
Eskelinen EL, Illert AL, Tanaka Y, et al. Role of LAMP-2 in lysosome biogenesis and autophagy. Mol Biol Cell. 2002;13(9):3355–3368. PubMed PMID: 12221139. Pubmed Central PMCID: 124165. Epub 2002/09/11. eng. | |
Eskelinen EL, Schmidt CK, Neu S, et al. Disturbed cholesterol traffic but normal proteolytic function in LAMP-1/LAMP-2 double-deficient fibroblasts. Mol Biol Cell. 2004;15(7):3132–3145. PubMed PMID: 15121881. Pubmed Central PMCID: 452571. Epub 2004/05/04. eng. | |
Huynh KK, Eskelinen EL, Scott CC, Malevanets A, Saftig P, Grinstein S. LAMP proteins are required for fusion of lysosomes with phagosomes. EMBO J. 2007;26(2):313–324. PubMed PMID: 17245426. Pubmed Central PMCID: 1783450. | |
Darsow T, Rieder SE, Emr SD. A multispecificity syntaxin homologue, Vam3p, essential for autophagic and biosynthetic protein transport to the vacuole. J Cell Biol. 1997;138(3):517–529. PubMed PMID: 9245783. | |
Itakura E, Kishi-Itakura C, Mizushima N. The hairpin-type tail-anchored SNARE syntaxin 17 targets to autophagosomes for fusion with endosomes/lysosomes. Cell. 2012;151(6):1256–1269. PubMed PMID: 23217709. | |
Furuta N, Fujita N, Noda T, Yoshimori T, Amano A. Combinational soluble N-ethylmaleimide-sensitive factor attachment protein receptor proteins VAMP8 and Vti1b mediate fusion of antimicrobial and canonical autophagosomes with lysosomes. Mol Biol Cell. 2010;21(6):1001–1010. PubMed PMID: 20089838. Pubmed Central PMCID: 2836953. | |
Atlashkin V, Kreykenbohm V, Eskelinen EL, Wenzel D, Fayyazi A, Fischer von Mollard G. Deletion of the SNARE vti1b in mice results in the loss of a single SNARE partner, syntaxin 8. Mol Cell Biol. 2003;23(15):5198–5207. PubMed PMID: 12861006. Pubmed Central PMCID: 165714. | |
Jiang P, Nishimura T, Sakamaki Y, et al. The HOPS complex mediates autophagosome-lysosome fusion through interaction with syntaxin 17. Mol Biol Cell. 2014;25(8):1327–1337. PubMed PMID: 24554770. Pubmed Central PMCID: 3982997. | |
Chen D, Fan W, Lu Y, Ding X, Chen S, Zhong Q. A mammalian autophagosome maturation mechanism mediated by TECPR1 and the Atg12-Atg5 conjugate. Mol Cell. 2012;45(5):629–641. PubMed PMID: 22342342. Pubmed Central PMCID: 3299828. | |
Burman C, Ktistakis NT. Autophagosome formation in mammalian cells. Semin Immunopathol. 2010;32(4):397–413. PubMed PMID: 20740284. | |
Longatti A, Tooze SA. Vesicular trafficking and autophagosome formation. Cell Death Differ. 2009;16(7):956–965. PubMed PMID: 19373247. | |
Hayashi-Nishino M, Fujita N, Noda T, Yamaguchi A, Yoshimori T, Yamamoto A. A subdomain of the endoplasmic reticulum forms a cradle for autophagosome formation. Nat Cell Biol. 2009;11(12):1433–1437. PubMed PMID: 19898463. | |
Yla-Anttila P, Vihinen H, Jokitalo E, Eskelinen EL. 3D tomography reveals connections between the phagophore and endoplasmic reticulum. Autophagy. 2009;5(8):1180–1185. PubMed PMID: 19855179. | |
Yamamoto A, Masaki R, Tashiro Y. Characterization of the isolation membranes and the limiting membranes of autophagosomes in rat hepatocytes by lectin cytochemistry. J Histochem Cytochem. 1990;38(4):573–580. PubMed PMID: 2319125. | |
Suzuki K, Akioka M, Kondo-Kakuta C, Yamamoto H, Ohsumi Y. Fine mapping of autophagy-related proteins during autophagosome formation in Saccharomyces cerevisiae. J Cell Sci. 2013;126(Pt 11):2534–2544. PubMed PMID: 23549786. | |
Graef M, Friedman JR, Graham C, Babu M, Nunnari J. ER exit sites are physical and functional core autophagosome biogenesis components. Mol Biol Cell. 2013;24(18):2918–2931. PubMed PMID: 23904270. Pubmed Central PMCID: 3771953. | |
Ge L, Melville D, Zhang M, Schekman R. The ER-Golgi intermediate compartment is a key membrane source for the LC3 lipidation step of autophagosome biogenesis. eLife. 2013;2:e00947. PubMed PMID: 23930225. Pubmed Central PMCID: 3736544. | |
Reggiori F, Wang CW, Nair U, Shintani T, Abeliovich H, Klionsky DJ. Early stages of the secretory pathway, but not endosomes, are required for Cvt vesicle and autophagosome assembly in Saccharomyces cerevisiae. Mol Biol Cell. 2004;15(5):2189–2204. PubMed PMID: 15004240. Pubmed Central PMCID: 404015. | |
Ohashi Y, Munro S. Membrane delivery to the yeast autophagosome from the Golgi-endosomal system. Mol Biol Cell. 2010;21(22):3998–4008. PubMed PMID: 20861302. Pubmed Central PMCID: 2982105. | |
Hailey DW, Rambold AS, Satpute-Krishnan P, et al. Mitochondria supply membranes for autophagosome biogenesis during starvation. Cell. 2010;141(4):656–667. PubMed PMID: 20478256. Pubmed Central PMCID: 3059894. | |
Hamasaki M, Furuta N, Matsuda A, et al. Autophagosomes form at ER-mitochondria contact sites. Nature. 2013;495(7441):389–393. PubMed PMID: 23455425. Epub 2013/03/05. eng. | |
Lamb CA, Yoshimori T, Tooze SA. The autophagosome: origins unknown, biogenesis complex. Nat Rev Mol Cell Biol. 2013;14(12):759–774. PubMed PMID: 24201109. | |
Knaevelsrud H, Soreng K, Raiborg C, et al. Membrane remodeling by the PX-BAR protein SNX18 promotes autophagosome formation. J Cell Biol. 2013;202(2):331–349. PubMed PMID: 23878278. Pubmed Central PMCID: 3718966. | |
Moreau K, Ravikumar B, Puri C, Rubinsztein DC. Arf6 promotes autophagosome formation via effects on phosphatidylinositol 4,5-bisphosphate and phospholipase D. J Cell Biol. 2012;196(4):483–496. PubMed PMID: 22351926. Pubmed Central PMCID: 3283994. | |
Dupont N, Chauhan S, Arko-Mensah J, et al. Neutral lipid stores and lipase PNPLA5 contribute to autophagosome biogenesis. Curr Biol. 2014;24(6):609–620. PubMed PMID: 24613307. Pubmed Central PMCID: 4016984. | |
Bejarano E, Yuste A, Patel B, Stout RF Jr, Spray DC, Cuervo AM. Connexins modulate autophagosome biogenesis. Nat Cell Biol. 2014;16(5):401–414. PubMed PMID: 24705551. Pubmed Central PMCID: 4008708. |
 © 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.