Back to Journals » International Journal of Nanomedicine » Volume 11
New silver nanoparticles induce apoptosis-like process in E. coli and interfere with mammalian copper metabolism
Authors Orlov IA, Sankova TP, Babich PS, Sosnin IM, Ilyechova EY, Kirilenko DA, Brunkov PN, Ataev GL, Romanov AE, Puchkova LV
Received 20 July 2016
Accepted for publication 1 November 2016
Published 15 December 2016 Volume 2016:11 Pages 6561—6574
DOI https://doi.org/10.2147/IJN.S117745
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Thomas Webster
Iurii A Orlov,1,2 Tatiana P Sankova,1,2 Polina S Babich,3 Ilya M Sosnin,1,4 Ekaterina Yu Ilyechova,1,5 Demid A Kirilenko,1,6 Pavel N Brunkov,1,6 Gennadii L Ataev,3 Alexey E Romanov,1,4,6 Ludmila V Puchkova1,2,5
1International Research and Education Center “Functional materials and devices of optoelectronics and microelectronics”, ITMO University, 2Department of Biophysics, Peter the Great St Petersburg Polytechnic University, 3Department of Zoology, Herzen State Pedagogical University of Russia, St Petersburg, 4Nanocatalysts and Functional Materials Laboratory, Togliatti State University, Togliatti, 5Department of Molecular Genetics, Research Institute of Experimental Medicine, 6Ioffe Institute, St Petersburg, Russia
Abstract: Silver nanoparticles (SNPs) are new functional materials that are widely used in biomedical and industrial technologies. Two main features that make SNPs valuable are their strong antibacterial effects and low toxicity to eukaryotes. In this study, SNPs were synthesized using a modified method of reducing the metal ions to their atomic state followed by crystallization. SNPs were characterized by UV/vis spectroscopy, X-ray diffractometry, atomic force microscopy, and transmission electron microscopy (TEM). The SNPs were spherically shaped with an average linear dimension of 20 nm. In aqueous solution, the SNPs were beige-yellow in color, and they formed a black color in bacteria-rich growth media. The toxicity and bioavailability of the SNPs were tested using Escherichia coli cells and C57Bl/6 mice. Although the SNPs displayed bactericidal activity, an E. coli cell strain transformed with an expression plasmid carrying a human CTR1 ectodomain with three motives that bind Cu(II), Cu(I), and Ag(I) demonstrated increased resistance to treatment with SNPs. TEM showed that the SNPs were absorbed by the E. coli cell, and flow cytometry showed that the SNPs induced apoptosis-like death. In mice treated with SNPs (daily intraperitoneal injection of 10 µg SNPs/g body weight over 4 days), the ceruloplasmin (Cp) oxidase activity in the blood serum decreased. However, level of Cp gene expression, the relative contents of the Cp protein in the Golgi complex and in the serum did not change. Treatment with SNPs did not influence the activity of superoxide dismutase 1 in the liver and had no apparent toxic effects in mice. These findings expand the scope of application for the use of new SNPs. The data are discussed in a paradigm, in which the effects of SNPs are caused by the interference of silver ions with copper metabolism.
Keywords: silver nanoparticles, bioavailability, apoptosis-like death, mammalian copper metabolism, copper status
Introduction
Recent decades have seen rapidly growing interest in biomedical applications of new functional nanostructural materials based on semiconductors (quantum dots), carbon materials (fullerenes, nanotubes), and metals and their oxides (nanoparticles). A majority of currently produced nanomaterials are silver nanoparticles (SNPs), which are fabricated using physical, chemical, and biological (or green) syntheses.1 Regardless of the synthesis method, SNPs are used as antibacterial, antifungal, antiviral, antibiofilm, and anti-inflammatory drugs, as treatments for medical dressings and instruments, as drug delivery systems, to manufacture medical textiles and underclothes, for water purification filters, among other applications.2–5 The advantage of SNPs is their high antimicrobial activity in combination with low toxicity in eukaryotes. Therefore, in many cases, SNPs may potentially substitute for antibiotics and prevent the development and spread of drug resistance in bacteria. Thus, there is a demand for SNPs with effective antibacterial properties and minimal effects on the cells of higher eukaryotes. At present, there are insufficient data for the rational design of SNPs with predefined biological properties; so the search for SNPs is carried out mostly through empirical means. This research focuses on the synthesis and specification of SNPs, an assessment of their antibacterial properties, and their in vivo effects in mammals.
Materials and methods
Synthesis of SNPs
SNPs were fabricated using a modified method by which silver ions were reduced to their zero oxidation state and subsequently crystallized. Synthesis was performed in distilled water using β-cyclodextrin (Reachim, Moscow, Russia) as a capping agent to create clathrates for SNP formation that protect the forming nanoparticles from agglomeration and other destructive factors. Silver nitrate (Reachim) and plant secondary metabolites (Reachim) were used as the source of silver ions and the reducing agents, respectively. First, 23 mg of β-cyclodextrin powder was dissolved in 100 mL of distilled water and the β-cyclodextrin molecules agglomerated into clathrates. Next, plant secondary metabolites and sodium hydroxide were dissolved in distilled water and added to the solution containing β-cyclodextrin clathrates. The prepared mixture was vigorously blended to homogeneity by a magnetic stirrer at 500 rpm. Then, silver nitrate was added to the mixture, and the colorless solution changed to beige-yellow as a result of crystallization. Synthesis was performed at room temperature for 1 min. In 1 min all silver ions were incorporated into the SNPs, and the synthesis was terminated. To produce SNPs, the following reagent concentrations were used: 230 mg/L β-cyclodextrin, 180 mg/L plant secondary metabolites, 1.8 mL/L 1.0 M NaOH, and 0.3 mL/L 3.0 M AgNO3.
Physicochemical characterization of the SNPs
SNP samples were characterized by UV/vis absorption (Shimadzu UV 1800; Shimadzu, Kyoto, Japan). The structural analysis was performed by X-ray diffraction (XRD; Shimadzu XRD 7000) and transmission electron microscopy (TEM) using a Jeol JEM-2100F microscope (Jeol, Tokyo, Japan; accelerating voltage 200 kV, point-to-point resolution 0.19 nm). The chemical analysis of the SNP solutions was carried out using an energy dispersive X-ray spectroscopy module (INCA, Oxford Instruments, Bristol, UK) attached to TEM (Jeol JEM-2100F). The size of the SNPs was characterized using TEM. Aliquots of the SNP solutions (5 and 10 μg/mL) were dropped onto a lacey carbon film supported by a copper grid and then air-dried before TEM observation. The particle sizes were characterized using conventional bright-field TEM images, and the structures were determined based on selected area electron diffraction patterns obtained from relatively large groups of particles. The mean size and standard deviation were calculated in random fields containing various numbers of particles. The morphology of the SNPs was analyzed by atomic force microscopy (SOLVER P47; ND-MDT, Moscow, Russia).
Used biological objects
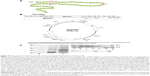
The study was performed in Escherichia coli K802 cells with the genotype F− e14− (McrA−) lacY1 or Δ(lac)6 glnV44 galK2 galT22 rfbD1 metB1 mcrB1 hsdR2 (rK− mK+), strain BL21 (DE3) E. coli cells with the genotype B F− dcm ompT hsdS (rB− mB−) gal λ (DE3), strain BL21 (DE3)/pGEX-4T-1 E. coli cells carrying the glutathione-S-transferase (GST) gene fusion plasmid vector (pGEX-4T-1; Amersham Biosciences, Buckinghamshire, UK), and strain BL21 (DE3)/pNdCTR1 E. coli cells carrying the N-terminal extracellular domain (198 bp) of human copper transporter 1 (CTR1; SLC31A1 gene according to NCBI databases) cloned into pGEX-4T-1 (Figure S1). The cells were grown aerobically in liquid or agar nutrient media based on bovine serum hydrolyzate (Samson-Med, St Petersburg, Russia) at 37°C. An antibiotic (ampicillin; Sigma-Aldrich, St Louis, MO, USA) was used at a concentration of 100 mg/L.
Seven-week-old male C57Bl/6 mice with body weights of 16–19 g were purchased from the Rappolovo nursery (Leningrad, Russia). The animals were maintained in polycarbonate cages with wood shavings in a temperature-controlled facility (23°C–25°C), with a 12:12-h light–dark cycle, 60% humidity with food and water ad libitum. The mice (n=24) were given daily intraperitoneal injections of the SNP solutions (10 μg/g of body weight) in phosphate-buffered saline (PBS; Na/K/Pi buffer, pH 7.4, physiological ionic strength). On the second, third, and fourth days after the first injection, eight mice were sacrificed and blood serum was obtained. The last group of mice was sacrificed at 12 h after the last injection. Thus, in this study, three experimental groups of mice (mice given two, three, and four treatments of SNPs) were used. The cumulative dose of SNPs in the last group was ~40 μg/g of body weight. Control mice of the same age (n=8) were used as the reference group.
The mice were sedated with diethyl ether vapor and euthanized by cervical dislocation, which was performed by skilled personnel. The animals used to isolate total RNA or subcellular fractions were anesthetized with oxybutyrate (1 g/kg of body weight). The blood samples were collected from the ocular vessels, and after clot formation, the serum was separated by centrifugation at 5,000× g for 10 min at 4°C. The liver tissues before isolation of total RNA, or subcellular fractions and silver concentration measurements were stored at −80°C until use.
Procedures involving animals and their care were conducted in conformity with institutional guidelines and are in compliance with national laws (Russian Federation, Ministry of Health N267, June 19, 2003; Guide for the Use of Laboratory Animals, Moscow, 2005). The studies were approved by the local Ethics Committee at the Institute of Experimental Medicine (protocol number N2/13 was approved on June 27, 2013).
Bacterial colony-forming ability measurements
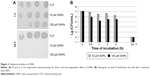
Bacterial cells were washed with sterile water and diluted 1:20 in a solution containing various concentrations of SNPs. Control cell samples were incubated in water. After treatment, the cells were titrated using the ten-fold dilution method to assess cell survival via their colony-forming ability on agar plates. Colonies were counted after 24 h of incubation at 37°C.
TEM analysis of E. coli treated with SNPs
E. coli cells were cultured overnight, diluted 25-fold, and cultured in 200 mL of aqueous media containing 15 μg/mL SNPs for 45 min at 37°C. The cells treated with SNPs were then cultured in water at room temperature for ~2 h. A portion of the cells was centrifuged for 5 min at 2,000× g and then resuspended in water; these cells were named as “washed cells.” The carbon-coated copper grids were dampened with the bacterial suspensions, dried in air at room temperature, and then observed using TEM.
Flow cytometry analysis
E. coli cells were cultured overnight in complex media, collected, and washed once with sterile water. The cells were diluted 100 times and cultured in 25 mL of sterile H2O (control) or 25 mL of sterile H2O with 100 μM of SNPs at 37°C for 4 h. Five milliliter aliquots of the bacterial culture were collected in 2 and 4 h and washed once with 1 mL of cold, filtered 1× PBS. Next, the cells were resuspended in 100 μL of 1× Annexin V-binding buffer (10 mM HEPES, pH 7.4, 140 mM NaCl, 2.5 mM CaCl2) and incubated with 5 μL of FITC-conjugated Annexin V and 2 μL of propidium iodide (PI; BD Pharmingen, San Diego, CA, USA) for 15 min at room temperature in the dark. The samples were diluted by the addition of 400 μL 1× Annexin V-binding buffer, placed on ice, and analyzed by flow cytometry within 15 min. The data were collected using a BD Accuri C6 instrument (Accuri Cytometers, Ann Arbor, MI, USA).
Relative mRNA level measurements
Total RNA extraction, measurements of the RNA concentration and its purity, the design and structure of primers, and parameters of the reverse transcription (RT) and polymerase chain reaction (PCR) procedures have been described previously.6 The PCR products were analyzed on a 1.4% agarose gel stained with ethidium bromide, and the data were processed using ImageJ software. The results were expressed in arbitrary units as a ratio between the amount of the specified mRNA PCR product and the amount of the PCR product of β-actin obtained from the same RNA preparations under similar conditions. Each value was combined from three independent PCR replicas of cDNA samples obtained from three mice. The results are presented as bar charts.
Immunoblotting (Western blot [WB])
For WB analysis, the protein concentrations of the samples were equalized and the proteins were separated by electrophoresis using an 8% polyacrylamide gel with 0.1% sodium dodecyl sulfate according to the Laemmli method. Protein transfer, control of the quality and uniformity of transfer, blocking with 5% nonfat milk, blotting with primary rabbit antibodies against rat ceruloplasmin (Cp) that cross-reacted with murine Cp, and visualization of the immune complexes were performed as described previously.6 Hybond enhanced chemiluminescence (ECL) nitrocellulose membrane, ECL reagent, ECL hyperfilm (GE Healthcare, Piscataway, NJ, USA), and horseradish peroxidase-conjugated goat anti-rabbit secondary antibodies (Abcam, Cambridge, UK) were used. The film was processed using the method reported by Aldridge et al.7 Protein markers with molecular masses ranging from 14.4 to 116 kDa were purchased from Thermo Scientific (Swedesboro, NJ, USA).
Isolation of cytosolic and Golgi complex membrane-enriched fractions
The fractions enriched with Golgi complex membranes were isolated by differential centrifugation. Tissue samples were homogenized (1:9 w/v) in buffer A containing 250 mM sucrose, 100 mM KCl, 5 mM MgCl2, 10 mM Tris-HCl (pH 7.4), 5 mM dithiothreitol, and 0.5 μL/mL protease inhibitor cocktail ( Sigma-Aldrich) using a T10 basic homogenizer (IKA, Staufen, Germany) at a maximum power for three cycles of twenty seconds each. The homogenate was centrifuged at 800× g for 10 min. The 800× g supernatant was centrifuged at 12,000× g for 20 min to separate the mitochondria, lysosomes, and peroxisomes. The resulting supernatant was centrifuged (23,000× g for 1 h) to sediment the Golgi complex membranes. The pellet was resuspended with buffer A and used as the Golgi complex membrane-enriched fraction. The fraction was incubated with Triton X-100 at a final concentration of 1% for 30 min at 0°C, clarified by centrifugation at 15,000× g for 1 h, and used to determine the Cp protein levels by WB. The supernatant obtained from the 23,000× g centrifugation (cytosol) was used to measure superoxide dismutase 1 (SOD1) activity.
Other methods
Specific activities of cuproenzymes were detected using the assay-in-gel method. After non-denaturing polyacrylamide gel electrophoresis gels were stained with o-dianisidine to reveal the Cp oxidase activity8 or with nitro blue tetrazolium to examine SOD activity.9 The total protein concentration was determined using the Bradford assay with bovine serum albumin as a calibration standard. Metal concentrations were measured using graphite furnace atomic absorption spectrometry with a Zeeman correction of nonselective absorption using a ZEEnit 650P spectrometer (Analytik Jena, Jena, Germany). The samples were homogenized in PBS and then dissolved in pure HNO3. All solutions were prepared in deionized water. The reagents used for protein and nucleic acid electrophoresis and the salts were purchased from Sigma-Aldrich (St Louis, MO, USA) and Merck (Darmstadt, Germany).
The data are presented as mean ± standard deviation. The significance of changes was determined by unpaired two-tailed Student’s t-tests; the changes were considered significant at P<0.05 (Statistica 6.0; StatSoft Inc., Tulsa, OK, USA).
Results
SNP specification
SNPs were fabricated by reducing silver ions to atomic state in a molecular host system consisting of β-cyclodextrin clathrates. The physical and chemical properties of the SNPs are summarized in Figure 1. The absorption spectrum of the SNPs presented a highly symmetric band of absorption, with peak maxima at 400 nm (Figure 1A), which indicates the excitation of surface plasmons typical of SNPs. In addition, a broad peak was observed between 350 and 420 nm, indicating the formation of SNPs. Further characterization was performed to confirm the crystalline nature of the SNPs. The XRD spectra for the SNPs revealed four peaks over the entire spectrum of 2θ values, ranging from 37° to 80° (Figure 1B). A comparison of the XRD spectra with the standard ones confirmed that the formed SNPs were nanocrystals, as evidenced by the peaks at 2θ values of 32.3°, 46.3°, 57.5°, and 77.0°, which corresponded to the (111), (200), (220), and (311) planes for silver, respectively (Figure 1C). The fabricated SNPs were spherically shaped (Figure 1D and E). As seen in Figure 1F, the size distribution of the SNPs was uneven, ranging from 15 to 35 nm; most SNPs were ~20 nm in diameter. When stored in darkness at room temperature, the aqueous solution of SNPs was stable, no new species were formed, and the color did not change for 6 months. The SNPs added to the bacteria-rich growth media gradually turned black in color. In the 0.005 M SNP solution, the SNPs were prone to weak, reversible aggregation. Therefore, before use, the SNPs were subjected to ultrasonic treatment using UZDN-2T instrument (Akadempribor, Sumi, Ukraine) at 37 kHz for 5 min without heating. The treatment did not change the biological activities of the SNPs, and TEM showed that the particles did not change in shape or size.
Antibacterial properties of SNPs
The studied SNPs displayed a profound concentration- and time-dependent antibacterial effect against E. coli cells (Figure 2). In this study, a special E. coli strain expressing the extracellular domain of human CTR1 protein (NdCTR1) fused with GST was also used. CTR1 is a transmembrane protein in the plasma membrane, which is an evolutionarily conserved, high-affinity copper importer in eukaryotes.10 The soluble extracellular domain (66 a. a. residues in human CTR1) contains three motifs that normally bind Cu(II) and Cu(I), but can also effectively bind Ag(I). The data, which are presented in Figure 3, demonstrated that the expression of NdCTR1 in E. coli cells rescued these cells from the SNP-induced toxicity in a dose- and time-dependent manner.
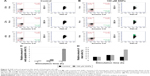
According to the TEM data, the SNPs were adsorbed onto the E. coli cells; the treatment of cells with SNPs did not cause any visible changes in the morphology of the cell wall (Figure 4A and B). The particles did not form large agglomerates and produced an irregular pattern on the cell, with relatively constant density. Washing SNPs did not result in profound desorption or significant loss of the particles (Figure 4C). Flow cytometry showed that the SNPs induced time- and concentration-dependent apoptosis-like E. coli cell death (Figure 5). The data demonstrated that the number of Annexin V-binding sites increased during incubation with SNPs by a factor of five.
SNPs interfere with copper metabolism in the murine liver
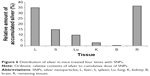
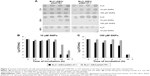
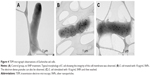
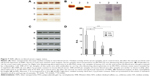
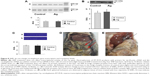
In mice receiving a maximum dose of SNPs, the relative distribution of silver throughout the body was measured. The results indicated that silver was distributed unevenly (Figure 6); the liver, lung, and spleen, which altogether comprised ~10% of the body weight, contained ~60% of the cumulative dose. The highest concentration of silver was found in the liver, followed by the spleen, lung, and kidney; the lowest concentration was found in the brain. Within 2 days following the SNP injection, the oxidase activity of blood serum reduced by 30% (Figure 7A and D) and then dropped progressively, reaching 1/6 of its initial value within 4 days. The relative content of Cp protein, which is responsible for the oxidase activity of blood serum, was determined by WB using antibodies to rat Cp (which cross-react with mouse Cp, Figure 7B). The relative content of the Cp protein did not change following the SNP injection (Figure 7C and D). WB revealed only ~130 kDa polypeptides, which correspond to full-size Cp molecule (Figure 7C). In the liver cells of mice that received four SNP injections, the relative content of mature Cp-mRNA, as measured by RT-PCR (Figure 8A), and the content of Cp protein in the Golgi complexes, as estimated by semiquantitative WB (Figure 8B), did not change significantly. These data indicated that the SNPs only influenced the enzymatic oxidase activity of Cp without affecting Cp synthesis or secretion. In the liver cells of SNP-treated mice, the activity of Cu2+, Zn2+-SOD1, the major cytosolic cuproenzyme, was not affected (Figure 8C). Dark-colored inclusions were observed in the abdominal cavities of the mice that received the largest dose of SNPs. The inclusions resembled beads on a string and were associated with adipose tissue and blood vessels entering the liver. The topological location and geometry of the inclusions were similar in different mice (Figure 8D and E). The “beads” were encapsulated.
Discussion
This study reports the SNPs synthesis, mechanism of their antibacterial effects, and ability of new SNPs to interfere with mammalian copper metabolism. The fabricated preparation of SNPs was a stable, homogeneous, aqueous suspension of spherically shaped, crystalline SNPs with an average linear size of 20 nm (Figure 1). The SNPs apparently corroded in biological media, as the growth media with bacteria containing SNPs gradually blackened due, most likely, to the formation of silver sulfide. This observation is in agreement with data showing the possible corrosion of SNPs in aqueous systems.11
The fabricated SNPs induced the death of bacterial cells in a concentration- and time-dependent manner (Figure 2). The E. coli cells expressing recombinant NdCTR1, which can effectively bind Ag(I),12 displayed increased resistance to SNPs (Figure 3); therefore, it may be assumed that the formation of bioavailable silver ions underlies the antibacterial properties of the SNPs.
The TEM data (Figure 4) indicated that the SNPs were adsorbed onto the E. coli cells and could not be easily totally washed off. It is possible that the corroding SNPs were retained by electrostatic interactions with the lipopolysaccharides and/or glutamate residues of 4-mer peptide, which cross-links the chains of muramic acid to form the cell wall. These findings further confirmed that the morphologic surface of the bacteria interacted with the SNPs.13 No SNP agglomerates like those described by Bao et al14 were observed in the cells. Thus, the results agreed with data that also showed the absence of agglomerates in the bacterial cell cytoplasm.13
Reports of fabricated SNPs without bactericidal properties have not been found. Perhaps, bacteriotoxicity is a universal property of SNPs. Most previous research studies assume that SNPs enter the cells, where they induce the formation of reactive oxygen species (ROS), which subsequently oxidize DNA, lipids, and proteins to produce genotoxic effects and impair protein folding, ultimately resulting in cell death. The increase of ROS levels in various cell types treated with SNPs has been confirmed by the direct measurement of ROS concentrations.15–18 However, the efficient formation of ROS (through the Fenton reaction) is typical for transition metal ions, such as the Fe(II)/Fe(III) and Cu(I)/Cu(II) pairs, but not the Ag(0)/Ag(I) pair. It is possible that the SNPs, which are adsorbed onto the cell surface and have close contact with the cell, are gradually oxidized by electron acceptors (eg, NAD+ and NADP+). The coordination and stabilization resulting from Ag(I) formation by small biomolecules (eg, amino acids) favor oxidation. Then, the Ag(I) ions can be captured by Cu(I) transporting proteins, which all have high affinity to Ag(I). Finally, Ag(I) ions are incorporated into the bacterial copper metabolism. At the same time, it is also possible that SNPs on bacteria surface cause disturbances in cell division, so this could be one more mechanism for their toxicity.
E. coli cells use copper as a catalytic cofactor in enzymes involved in the antioxidant defense system, the respiratory electron transport chain, the shikimate pathway, which is responsible for the biosynthesis of aromatic amino acids, and so on. Copper is presumably taken up by E. coli cells nonspecifically through porins,19 and free copper ions lead to toxic ROS formation.20 However, the cells have a multicomponent homeostatic system for the detoxification and efflux of copper ions that are not recruited by cuproenzymes.21 This system includes the P-type Cu(I)-ATPase CopA22 (a homologue of the mammalian protein ATP7A23), the multi-copper oxidase CueO, which oxidizes periplasmic Cu(I) to Cu(II) and possibly reduces the additional influx of copper into the cytoplasm,24 and the four-part Cus copper proton-driven efflux complex CusCFBA, which accepts Cu ions from CopA and extrudes copper out of the cell.25,26 The expression of the corresponding genes is positively controlled by copper-activated transcription factors.27 All of these proteins are able to recognize and bind both Cu(I) and Ag(I) ions.28
It is possible that the massive influx of Ag(I) into the cells causes competitive displacement of Cu(I) from the copper excretion system, the inclusion of Ag(I) into the active centers of CueO (a Cp paralog29), and the inhibition of its Cu(I)-detoxificating function. For these reasons, in the case of Ag(I) excess, the levels of intracellular Cu(I) that are not enveloped by transport proteins (“unbound” copper) increase. Note that it is the Cu(I) oxidation state that is prone to initiate Fenton reactions and promote ROS formation. The excess of “unbound” copper is accompanied by the deficiency of cuproenzymes, including bacterial Cu2+, Zn2+-SOD, which normally provides protection against ROS in the periplasmic space. Thus, ROS and oxidation products accumulate, causing membrane depolarization (detected with Annexin V) and subsequent cell death (detected with PI); these events that are specific traits of apoptosis-like death30 were observed in the experiments (Figure 5). The results agree with previous experiments that demonstrated the ability of SNPs to induce apoptosis-like death.31 Thus, it is possible that the antimicrobial effect of SNPs in E. coli cells is caused by oxidative stress resulting from ROS production via the Fenton reaction due to the increase of “unbound” Cu(I) and the decrease of protective cuproenzymes (Cu2+, Zn2+-SOD) from Ag(I) intervention in the bacterial copper homeostatic system.
The use of SNPs in medical applications requires testing of their effects in mammals in vivo. It is commonly accepted that SNPs have low toxicity in higher eukaryotes. However, such data are scarce, mostly phenomenological, highly discrepant, and cannot be easily related to each other. It was previously shown that relevant toxic effects (mid-zonal hepatocellular necrosis and gall bladder hemorrhage) develop in mice following a single intravenous injection of 10 nm of SNPs at a dose of 10 μg/g body weight in 24 h.32 However, the 40 and 100 nm SNPs showed mild or negligible toxicity.32 This analysis has demonstrated that the intraperitoneal injection of four such doses of 20 nm SNPs over 4 days did not cause toxic effects. It is possible that the toxic effect depends on the method of injection; alternatively, the SNPs fabricated in this study may have very low toxicity to mammals. In this study, distribution of silver through the murine bodies (Figure 6) was similar as described in previous work.32 In both cases, silver was accumulated by the inner organs, especially the liver, and was practically undetected in the brain. It has also been reported that 100 nm SNPs from ambient air can reach the human brain and may be associated with neurodegeneration.33 Earlier, it has been shown that the concentration of silver atoms in the brain was very low following the intraperitoneal injection of 15 nm SNPs. However, when intranasal injection was used, significant amounts of silver were found in the brain, especially in the olfactory bulbs.34 Perhaps in mammals, the distribution of silver from the SNPs depends on the size of the SNPs and the mode of application. For the first time, it was observed that a fraction of SNPs had accumulated in the extracellular space in the shape of visible, encapsulated granules with relatively constant sizes (Figure 8D and E). The granules may be formed by interactions between the SNPs and extracellular proteins;35 this possibility is exemplified by studies using serum albumin in vitro.36 The mechanism of granule formation and their chemical nature warrant further study.
In mice that received the largest dose of SNPs, a six-fold decrease in blood serum Cp oxidase activity was observed (Figure 7A). Blood serum Cp is a multifunctional, multi-copper blue oxidase synthesized by liver cells. Cp functions as a copper source for non-hepatic cells, it is an indispensable copper source for developing embryos, and it participates in iron transport as ferroxidase. Cp is also a member of the extracellular antioxidant system that can oxidize biogenic amines. There are also suggestions that Cp has other unstudied biological activities.37–39 SNPs did not affect the transcription of the Cp gene, the translation of Cp-mRNA, or Cp secretion (Figure 7C and 8A and B). Moreover, SNPs did not impair the activity of the cytosolic cuproenzyme SOD1 (Figure 8C). The accumulation of silver in the liver and its differential effects on the activity of the two cuproenzymes of hepatic origin (Cp, decreased activity; SOD1, unaffected) might be explained by the different coordination sphere of copper in these two proteins. The copper-binding site of Cp (blue copper site) contains cysteine residues, which are a favorable ligand for Cu(I)/Ag(I), whereas the SOD1 copper site contains only histidine residues.
Despite some level of discord, the data obtained in eukaryotes from different phylogenetic groups generally support the idea that the effects of SNPs can be related to the organization of the copper metabolic system. For instance, it has been shown that the lifelong feeding of fruit flies with food containing SNPs produced no negative effects on the median survival. However, these flies appeared uniformly lighter in body color due to the loss of melanin pigments in their cuticles. Additionally, the fertility of the SNP-fed flies was compromised, and the activities of copper-dependent enzymes (tyrosinase and SOD1) were significantly decreased despite the constant levels of copper present in the tissue.40 In Caenorhabditis elegans, treatment with exogenous SNPs dramatically decreased reproductive potential but increased the expression of Cu2+, Zn2+-SOD3.41 These diverse effects may result from the specific traits of copper metabolism in distant phylogenetic groups.
The data presented herein allow us to think that silver ions, which are produced by the gradual corrosion of SNPs, interfere with copper metabolism. The most profound effect is the insertion of silver into Cp in the liver. It is possible that the chronic application of SNPs will result in deficiency of Cp oxidase activity, which will affect cuproenzyme levels in various mammalian organs. It is also quite probable that the effects of SNPs depend on the specific properties of copper homeostatic systems in different organs. At present, it is difficult to evaluate the influence of SNPs on whole mammalian organisms. An additional obstacle is the absence of data showing the influence of SNPs on nonenzymatic copper functions, including neovascularization,42 NF-kB-, p53- and HIF1-dependent pathways,43–45 other signaling pathways,46–48 and neurotransmission.49 These aspects have not yet been studied in vivo.
Conclusion
In summary, it is concluded that the fabricated SNPs possess profound antibacterial activities and do not display acute toxicity in mammals. The biological activity of the SNPs was mediated by the formation of bioavailable forms of silver during the contact between SNPs and biological fluids and their interference with copper metabolism.
Acknowledgments
This work was supported by Russian Foundation for Basic Research grants (15–04–06770-a and 16–34–60219) and by grants from the Ministry of Education and Science of Russian Federation provision N 220 (14.B25.31.0011 and 6.1278.2014/K). The TEM characterizations were performed using equipment owned by the Joint Research Center “Material science and characterization in advanced technology” (Ioffe Institute, St Petersburg, Russia).
Disclosure
The authors report no conflicts of interest in this work.
References
Ge L, Li Q, Wang M, Ouyang J, Li X, Xing M. Nanosilver particles in medical applications: synthesis, performance, and toxicity. Int J Nanomedicine. 2014;4:2399–2407. | ||
Franci G, Falanga A, Galdiero S, Palomba L, Rai M. Silver nanoparticles as potential antibacterial agents. Molecules. 2015;20(5):8856–8874. | ||
Naqvi SZ, Kiran U, Ali MI, et al. Combined efficacy of biologically synthesized silver nanoparticles and different antibiotics against multidrug-resistant bacteria.Int J Nanomedicine. 2013;8:3187–3195. | ||
Markowska K, Grudniak AM, Wolska KI. Silver nanoparticles as an alternative strategy against bacterial biofilms. Acta Biochim Pol. 2013;60(4):523–530. | ||
Khalili FJ, Jafari S, Eghbal MA. A review of molecular mechanisms involved in toxicity of nanoparticles. Adv Pharm Bull. 2015;5(4):447–454. | ||
Zatulovskaia YA, Ilyechova EY, Puchkova LV. The features of copper metabolism in the rat liver during development. PLoS One. 2015;10(10):e0140797. | ||
Aldridge GM, Podrebarac DM, Greenough WT, Weiler IJ. The use of total protein stains as loading controls: an alternative to high-abundance single protein controls in semi-quantitative immunoblotting. J Neurosci Methods. 2008;172(2):250–254. | ||
Owen JA, Smith H. Detection of ceruloplasmin after zone electrophoresis. Clin Chim Acta. 1961;6(3):441–444. | ||
Vives-Bauza C, Starkov A, Garcia-Arumi E. Measurements of the antioxidant enzyme activities of superoxide dismutase, catalase, and glutathione peroxidase. Methods Cell Biol. 2007;80:379–393. | ||
Öhrvik H, Thiele DJ. How copper traverses cellular membranes through the mammalian copper transporter 1, Ctr1. Ann N Y Acad Sci. 2014;1314:32–41. | ||
Kittler S, Greulich C, Diendorf J, Köller M, Epple M. Toxicity of silver nanoparticles increases during storage because of slow dissolution under release of silver ions. Chemistry of Materials. 2010;22(16):4548–4554. | ||
Bertinato J, Cheung L, Hoque R, Plouffe LJ. Ctr1 transports silver into mammalian cells. Trace Elem Med Biol. 2010;24(3):178–184. | ||
Chwalibog A, Sawosz E, Hotowy A, et al. Visualization of interaction between inorganic nanoparticles and bacteria or fungi. Int J Nanomedicine. 2010;5:1085–1094. | ||
Bao H, Yu X, Xu C, et al. New toxicity mechanism of silver nanoparticles: Promoting apoptosis and inhibiting proliferation. PLoS One. 2015;10(3):e0122535. | ||
Huang J, Cheng J, Yi J. Impact of silver nanoparticles on marine diatom Skeletonema costatum. J Appl Toxicol. 2016;36(10):1343–1354. | ||
Ramalingam B, Parandhaman T, Das SK. Antibacterial effects of biosynthesized silver nanoparticles on surface ultrastructure and nanomechanical properties of gram-negative bacteria viz. Escherichia coli and Pseudomonas aeruginosa. ACS Appl Mater Interfaces. 2016;8(7):4963–4976. | ||
Lee MJ, Lee SJ, Yun SJ, et al. Silver nanoparticles affect glucose metabolism in hepatoma cells through production of reactive oxygen species. Int J Nanomedicine. 2015;11:55–68. | ||
Gurunathan S, Jeong JK, Han JW, Zhang XF, Park JH, Kim JH. Multidimensional effects of biologically synthesized silver nanoparticles in Helicobacter pylori, Helicobacter felis, and human lung (L132) and lung carcinoma A549 cells. Nanoscale Res Lett. 2015;10:35. | ||
Li XZ, Nikaido H, Williams KE. Silver-resistant mutants of Escherichia coli display active efflux of Ag1 and are deficient in porins. J Bacteriol. 1997;179(19):6127–6132. | ||
Gaetke LM, Chow-Johnson HS, Chow CK. Copper: toxicological relevance and mechanisms. Arch Toxicol. 2014;88(11):1929–1938. | ||
Rensing C, Grass G. Escherichia coli mechanisms of copper homeostasis in a changing environment. FEMS Microbiology. 2013;27(2–3):197–213. | ||
Fan B, Rosen BP. Biochemical characterization of CopA, the Escherichia coli Cu(I)-translocating P-type ATPase. J Biol Chem. 2002;277(49):46987–46992. | ||
Inesi G, Pilankatta R, Tadini-Buoninsegni F. Biochemical characterization of P-type copper ATPases.Biochem J. 2014;463(2):167–176. | ||
Singh SK, Roberts SA, McDevitt SF, et al. Crystal structures of multicopper oxidase CueO bound to copper(I) and silver(I): functional role of a methionine-rich sequence. J Biol Chem. 2011;286(43):37849–37857. | ||
Delmar JA, Su CC, Yu EW. Heavy metal transport by the CusCFBA efflux system. Protein Sci. 2015;24(11):1720–1736. | ||
Long F, Su CC, Lei HT, Bolla JR, Do SV, Yu EW. Structure and mechanism of the tripartite CusCBA heavy-metal efflux complex. Philos Trans R Soc Lond B Biol Sci. 2012;367(1542):1047–1058. | ||
Gudipaty SA, McEvoy MM. The histidine kinase CusS senses silver ions through direct binding by its sensor domain. Biochim Biophys Acta. 2014;1844(9):1656–1661. | ||
Randall CP, Gupta A, Jackson N, Busse D, O’Neill AJ. Silver resistance in Gram-negative bacteria: a dissection of endogenous and exogenous mechanisms. J Antimicrob Chemother. 2015;70(4):1037–1046. | ||
Vasin A, Klotchenko S, Puchkova L. Phylogenetic analysis of six-domain multi-copper blue proteins. PLoS Curr. 2013;5:pii:ecurrents.tol.574bcb0f133fe52835911abc4e296141. | ||
Erental A, Kalderon Z, Saada A, Smith Y, Engelberg-Kulka H. Apoptosis-like death, an extreme SOS response in Escherichia coli. MBio. 2014;5(4):e01426-14. | ||
Lee W, Kim KJ, Lee DG. A novel mechanism for the antibacterial effect of silver nanoparticles on Escherichia coli. BioMetals. 2014;27(6):1191–1201. | ||
Recordati C, De Maglie M, Bianchessi S, et al. Tissue distribution and acute toxicity of silver after single intravenous administration in mice: nano-specific and size-dependent effects. Part Fibre Toxicol. 2016;13:12. | ||
Win-Shwe TT, Fujimaki H. Nanoparticles and neurotoxicity. Int J Mol Sci. 2011;12(9):6267–6280. | ||
Sankova TP, Sosnin IM, Karpenko MN, et al. On the biological activity of silver nanoparticles. MPM. 2015;24:289–296. | ||
Duràn N, Silveira CP, Duràn M, Martinez T. Silver nanoparticle protein corona and toxicity: a mini-review. J Nanobiotechnol. 2015;13:55. | ||
Dasgupta N, Ranjan S, Patra D, Srivastava P, Kumar A, Ramalingam C. Bovine serum albumin interacts with silver nanoparticles with a “side-on” or “end on” conformation. Chem Biol Interact. 2016;253:100–111. | ||
Hellman NE, Gitlin JD. Ceruloplasmin metabolism and function. Annu Rev Nutr. 2002;22:439–458. | ||
Ramos D, Mar D, Ishida M, et al. Mechanism of copper uptake from blood plasma ceruloplasmin by mammalian cells. PLoS One. 2016;11(3):e0149516. | ||
Healy J, Tipton K. Ceruloplasmin and what it might do. J Neural Transm. 2007;114(6):777–781. | ||
Armstrong N, Ramamoorthy M, Lyon D, Jones K, Duttaroy A. Mechanism of silver nanoparticles action on insect pigmentation reveals intervention of copper homeostasis. PLoS One. 2013;8(1):e53186. | ||
Roh JY, Sim SJ, Yi J, et al. Ecotoxicity of silver nanoparticles on the soil nematode Caenorhabditis elegans using functional ecotoxicogenomics. Environ Sci Technol. 2009;43(10):3933–3940. | ||
Easter RN, Qilin C, Lai B, Ritman EL, Caruso JA, Zhenyu Q. Vascular metallomics: copper in the vasculature. Vasc Med. 2011;15(1):61–69. | ||
Krohn KA, Link JM, Mason RP. Molecular Imaging of Hypoxia. J Nucl Med. 2008;49(Suppl 2):S129–S148. | ||
Arciello M, Longo A, Viscomi C, et al. Core domain mutant Y220C of p53 protein has a key role in copper homeostasis in case of free fatty acids overload. Biometals. 2015;28(6):1017–1029. | ||
Bartuzi P, Hofker MH, van de Sluis B. Tuning NF-κB activity: a touch of COMMD proteins. Biochim Biophys Acta. 2013;1832(12):2315–2321. | ||
Turski M, Thiele DJ. New roles for copper metabolism in cell proliferation, signaling, and disease. J Biol Chem. 2009;284(2):717–721. | ||
Turski ML, Brady DC, Kim HJ, et al. A novel role for copper in Ras/mitogen-activated protein kinase signaling. Mol Cell Biol. 2012;32(7):1284–1295. | ||
Brady DC, Crowe MS, Turski ML, et al. Copper is required for oncogenic BRAF signalling and tumorigenesis. Nature. 2014;509(7501):492–496. | ||
Opazo KM, Greenough MA, Bush AI. Copper: from neurotransmission to neuroproteostasis. Front Aging Neurosci. 2014;6:143. |
Supplementary material
 © 2016 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2016 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.