Back to Journals » OncoTargets and Therapy » Volume 12
Long noncoding RNA GIHCG induces cancer progression and chemoresistance and indicates poor prognosis in colorectal cancer
Authors Jiang X, Li Q, Zhang S, Song C, Zheng P
Received 26 October 2018
Accepted for publication 20 December 2018
Published 7 February 2019 Volume 2019:12 Pages 1059—1070
DOI https://doi.org/10.2147/OTT.S192290
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Tohru Yamada
Xiaohua Jiang,1,* Qin Li,2,* Shun Zhang,1 Chun Song,1 Ping Zheng2
1Department of Gastrointestinal Surgery, Shanghai East Hospital, School of Medicine, Tongji University, Shanghai 200123, China; 2Department of Gastroenterology, Shanghai East Hospital, School of Medicine, Tongji University, Shanghai 200123, China
*These authors contributed equally to this work
Background: Colorectal cancer (CRC) is one of the leading causes of cancer-related death worldwide, however, the mechanisms of CRC progression remain obscure. The present study investigated the clinical significance and functional role of long noncoding RNA (lncRNA) GIHCG in CRC.
Methods: Expression of GIHCG was detected by quantitative real time polymerase chain reaction (qRT-PCR) in seven CRC cell lines and 110 CRC tissues. Comparison of clinicopathological characteristics in the high GIHCG expression group and the low GIHCG expression group was performed. The overall survival (OS) and progression-free survival (PFS) of the patients were depicted with Kaplan-Meier test and compared with Log-rank test. Univariate and multivariate analyses were carried out to detect the risk factors for poor OS and PFS. In addition, expression of GIHCG was silenced with siRNAs in LoVo cells and overexpressed with pcDNA3.1-GIHCG vector in SW480 cells, respectively. And the Transwell assay, Matrigel assay, colony formation assay and Cell Counting Kit-8 assay (CCK-8) were performed to investigate the role of GIHCG in the migration, invasion and proliferation of CRC cells. Besides, the role of GIHCG in chemoresistance was also detected.
Results: GIHCG was overexpressed in seven CRC cell lines and 110 CRC tissues. High GIHCG expression was correlated with lymphovascular invasion, lymph node metastasis, distant metastasis and advanced TNM stages. Moreover, patients with high GIHCG expression had much poorer OS and PFS rates. Besides, high GIHCG expression was identified as an independent risk factor for poor OS and PFS. The Transwell assay and the Matrigel assay discovered that GIHCG deficiency inhibited cell migration and invasion, while ectopic expression of GIHCG promoted migration and invasion. Besides, the colony formation assay and the CCK-8 assay verified that GIHCG increased cell proliferation ability. By establishing 5-fluorouracil (5-FU) and Oxaliplatin (Oxa)-resistant LoVo cells and SW480 cells, we found chemoresistant CRC cells had much higher expression levels of GIHCG. Also, GIHCG facilitated cell survival under 5-FU or Oxa treatment. Furthermore, silencing of GIHCG notably reduced the improved cell survival rates of 5-FU or Oxa-resistant LoVo cells compared with control cells.
Conclusion: GIHCG contributes to cancer progression and chemoresistance and indicates poor prognosis in CRC. GIHCG may be a promising prognostic biomarker and therapeutic target in CRC.
Keywords: lncRNA, migration, invasion, proliferation, drug-resistance
Introduction
Colorectal cancer (CRC) is the fourth most common cause of cancer-related mortality worldwide, accounting for 6.1% of all types of cancers.1 CRC is a heterogeneous disease, and the pathogenetic mechanisms underlying CRC development result from a wide array of cellular transformation processes. Existing evidence suggests that accumulating genetic and epigenetic mutations in the colorectum drive epithelial dysplasia, uncontrolled cell proliferation, and escape from apoptosis, ultimately resulting in CRC.2–6 Currently identified dysregulated genes involved in CRC mainly include some oncogenes, such as KRAS and MYC, and some oncosuppressive genes, such as APC and TP53; however, the molecular and genetic bases of CRC carcinogenesis and metastasis remain largely unknown.7,8 Besides, despite substantial progress in understanding the molecular mechanisms and treatment for CRC in recent years, the overall survival (OS) rate of CRC patients has not changed dramatically. Therefore, more investigations are needed to discover the mechanisms of CRC progression, which may contribute to developing novel therapeutic targets and diagnostic and prognostic biomarkers.
Long noncoding RNAs (lncRNAs) are generally defined as transcripts longer than 200 nucleotides lacking protein coding potential and transcribed by the RNA polymerase II (RNA Pol II).9–12 More and more lncRNAs are demonstrated to be deregulated in cancer and involved in a wide range of cancer biological steps, such as the hypoxia signaling and epithelial to mesenchymal transition, telomere maintenance, hormone receptor signaling, RNA processing, chromatin state and methylation, insensitivity to growth inhibition, promotion of angiogenesis, and tissue invasion and metastasis.13–16 It has been verified that lncRNAs show a tissue-specific pattern to a greater degree than protein-coding RNAs, and thus lncRNAs are regarded as promising therapeutic targets and diagnostic biomarkers.17–20 lncRNA gradually increased during hepatocarcinogenesis (GIHCG) is a novel lncRNA first identified by Sui et al21 using publicly available microarray data. GIHCG was reported to promote proliferation, migration, and invasion in hepatocellular carcinoma cells.21 Mechanistically, GIHCG was demonstrated to physically associate with EZH2 and the promoter of miR-200b/a/429, recruit EZH2 and DNMT1 to the miR-200b/a/429 promoter regions, upregulate histone H3K27 trimethylation and DNA methylation levels on the miR-200b/a/429 promoter, and dramatically silence miR-200b/a/429 expression.21 Studies have found that GIHCG could also accelerate cancer progression in renal cell carcinoma and tongue squamous cell carcinoma.22,23 Especially, a high level of GIHCG in serum was reported to be correlated with advanced TNM stages and could accurately discriminate renal cell carcinoma patients with healthy controls,10 which provides a potential way for early renal cell carcinoma diagnosis. However, to the best of our knowledge, the functional role and clinical significance of GIHCG in CRC have not been verified.
The objective of this study was to explore the functional role of GIHCG in CRC progression and chemoresistance. We found a much elevated expression level of GIHCG in CRC cells and samples. Increased GIHCG expression was demonstrated to promote CRC clinical progression and correlate with poor prognosis. In addition, upregulation of GIHCG accelerated migration, invasion, proliferation, and chemoresistance of CRC cells. Collectively, these results defined the role of GIHCG in CRC, which may shed light on identifying GIHCG as a novel prognostic biomarker and therapeutic target in CRC.
Materials and methods
Cell culture and development of chemoresistant cells
The human CRC cell lines SW620, HT29, HCT8, HCT116, LoVo, SW480, and DLD1, and normal human colon epithelial cell line (HCoEpic) were all purchased from the Type Culture Collection of the Chinese Academy of Sciences (Shanghai, China). Cells were maintained in either RPMI-1640 medium or Dulbecco’s Modified Eagle’s Medium (DMEM) supplemented with 10% fetal bovine serum (FBS) (Thermo Fisher Scientific, Waltham, MA, USA), 100 U/mL penicillin and 100 μg/mL streptomycin (Thermo Fisher Scientific) at 37°C in a humidified atmosphere containing 5% CO2.
Silencing and ectopic expression of GIHCG were achieved by three siRNAs specially targeting GIHCG (siGIHCG.1, siGIHCG.2, and siGIHCG.3) (GENEWIZ, Suzhou, Jiangsu, China) and pcDNA3.1-GIHCG vector (GENEWIZ), respectively. Scrambled siRNA control (siNC) and pcDNA3.1 empty vector (Vector) were used as controls. Cells were harvested 24 hours post-transfection for subsequent analysis.
Development of 5-fluorouracil (5-FU)-resistant LoVo and SW480 cells was achieved by exposing parental LoVo and SW480 cells to an initial dose of 0.2 μg/mL 5-FU (Abcam, Boston, MA, USA) in RMPI 1640 plus 10% FBS. The surviving population of cells was recovered for 2 weeks. The cells that survived initial 5-FU treatment were then exposed to sequentially increasing concentration (0.2 μg/mL) of 5-FU. Finally, the 5-FU concentration was increased to the clinically relevant plasma concentration of 2.0 μg/mL, and the cells were continuously cultured in 2.0 μg/mL 5-FU, unless otherwise indicated. The development of Oxaliplatin (Oxa) (Abcam)-resistant LoVo cells and SW480 cells was performed similar to that of the 5-FU-resistant cells, except that the initial concentration of Oxa was 1.0 μg/mL, and the final concentration was 4.0 μg/mL. The developed LoVo cells and SW480 cells resistant to 5-FU and Oxa were named LoVo-5-FU-R, SW480-5-FU-R, LoVo-Oxa-R, and SW480-Oxa-R.
Clinical data and tissue collection
A total of 110 CRC samples and paired tumor adjacent tissues were obtained from CRC patients who underwent radical resection in the Shanghai East Hospital (Shanghai, China). All patients gave written informed consent approved by the local ethics committee of the Shanghai East Hospital, which was conducted in accordance with the Declaration of Helsinki. The samples were obtained directly from surgery after removal of a necessary amount of tissue for routine pathology examination. The excluding criteria were as follows: patients who had received preoperative chemotherapy and/or radiation of rectal cancers, patients with hereditary CRC syndromes, and patients with two or more malignancies. The collected samples were frozen and stored in liquid nitrogen until use.
Follow-up was conducted in the out-patient clinic, and the frequency was once every 3 months during the first 2 years, every 6 months during the next 3 years, and yearly thereafter. Cancer progression was dated from the first evidence of relapse based on physical, histological, or imaging data. Results presented in our study were based on follow-up, ending on January 20, 2018.
Quantitative real time polymerase chain reaction (qRT-PCR) assay
Total RNA in CRC cells or samples was extracted with TRIzol RNA Isolation Reagent (Thermo Fisher Scientific) according to the manufacturer’s protocol. The first strand of cDNA for GIHCG was synthesized from 1 μg of RNA using the High-Capacity cDNA Reverse Transcription Kit (Thermo Fisher Scientific). qRT-PCR was carried out using the SYBR® Premix DimerEraser Kit (TaKaRa, Dalian, China) on an Applied Biosystems 7,500 Real-Time PCR System. GAPDH served as an internal control to normalize the expression of GIHCG. The 2−ΔΔCT method was employed to calculate the relative expression of GIHCG. The primers used in this study were as follows: GIHCG forward: 5′-CTTTCAAGAAGTTTGGCTGTC-3′; reverse: 5′-GCTCATTCAACGGATAAGTC-3′. GAPDH, forward: 5′-GGGAAACTGTGGCGTGAT-3′; reverse: 5′-GAGTGGGTGTCGCTGTTGA-3′.
Transwell assay and Matrigel assay
CRC cells were transfected and incubated for 24 hours. Then, the transfected cells were trypsinized and suspended and used for Transwell assay and Matrigel assay. The Matrigel assay was conducted using 24-well BioCoat Matrigel invasion chambers (Corning International, New York, NY, USA) according to the manufacturer’s protocols. Briefly, suspended cells in 250 μL of FBS free medium were seeded in the upper chamber, and the lower chamber was filled with 750 μL of medium with 10% FBS. After 24 hours incubation, the cells remaining in the upper chamber were removed with cotton swabs, and the cells that had invaded through the Matrigel were stained and photographed with an optical microscope (Olympus Corporation, Tokyo, Japan). The Transwell assay was performed as the Matrigel assay, except that the chamber was not pre-coated with Matrigel.
Cell proliferation assays
Colony formation assay and Cell Counting Kit-8 (CCK-8) assay were conducted to evaluate the proliferation ability of CRC cells. In the colony formation assay, transfected CRC cells were planted in 6-well plates (500 cells per well) and then subjected to incubation for 10 days. Subsequently, formed colonies were fixed and stained. Colonies with diameters greater than 1 mm were counted under a light microscope (Olympus Corporation).
In CCK-8 (Dojindo Laboratories, Kumamoto, Japan) assay, transfected cells were seeded in 96-well plates, and the CCK-8 solution (10 μL) was added at indicated time points (0 hour, 24 hours, 48 hours, 72 hours, and 96 hours). Then, the cells were incubated for an additional 2 hours. The number of cells was qualified by monitoring the changes in the absorbance at 450 nm.
Determination of cell survival rate
Cell survival rate was measured by CCK-8 assay. Transfected cells (1×103 cells/well) were seeded into 96-well plates and supplemented 100 μL medium containing 10% FBS. Cells were treated with different concentrations of 5-FU (0 μg/mL, 4 μg/mL, 8 μg/mL, 16 μg/mL, 32 μg/mL, and 64 μg/mL) or Oxa (0 μg/mL, 1 μg/mL, 5 μg/mL, 10 μg/mL, 30 μg/mL, and 60 μg/mL) or with 10 μg/mL of 5-FU or Oxa treatment for different time periods (0 hour, 24 hours, 48 hours, 72 hours, and 96 hours). The survival cell rate was calculated by measuring the absorbance at 450 nm.
Statistical analyses
Data are expressed as mean±SD. Comparisons between values from two groups were performed using Student’s t-tests (two-tailed) or chi-squared test, and multiple groups by ANOVA (Bonferroni post-hoc). Survival curves were depicted using the Kaplan–Meier method and compared by Log-rank test. Univariate and multivariate Cox regression analyses were employed to identify the independent prognostic factors. P<0.05 was deemed to indicate statistical significance. The above statistical analyses were performed by SPSS Statistics 19.0 for Windows (IBM Corporation, Armonk, NY, USA).
Results
Expression of GIHCG is upregulated, and increased expression of GIHCG indicates CRC progression
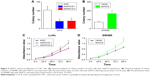
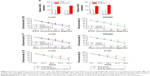
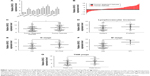
qRT-PCR results showed that HCoEpic displayed a notably lower GIHCG expression level than seven CRC cell lines (Figure 1A). To validate this result, expression of GIHCG was measured in CRC tissues and paired tumor adjacent tissues (n=110) (Figure 1B). Intriguingly, the CRC tissues also exhibited a much higher expression level of GIHCG than tumor adjacent tissues (Figure 1C). In addition, CRC tissues with lymphovascular invasion, lymph node metastasis, distant metastasis, or advanced TNM stages (TNM III/IV) displayed significantly increased GIHCG expression levels than CRC tissues without lymphovascular invasion (Figure 1D), lymph node metastasis (Figure 1E), distant metastasis (Figure 1F), or with less advanced TNM stages (TNM I/II) (Figure 1G), respectively. These results indicated that GIHCG may correlate with cancer progression in CRC.
Overexpression of GIHCG predicts poor prognosis in CRC
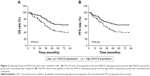
To better understand the role of GIHCG in CRC, the CRC tissues were dichotomized to the high GIHCG expression group and the low GIHCG expression group, with the mean GIHCG expression level serving as the cutoff value. Statistical analyses of GIHCG expression and CRC clinicopathological features revealed that the high GIHCG expression group had much more CRC cases with lymphovascular invasion, lymph node metastasis, distant metastasis, and advanced TNM stages than the low GIHCG expression group (Table 1). Besides, the OS rate and the progression-free survival (PFS) rate of the patients in the high GIHCG expression group had an obviously poorer OS rate (Figure 2A) and a remarkably shorter PFS period (Figure 2B) than the patients in the low GIHCG expression group.
To further define the oncogenic role of GIHCG in CRC, the risk factors for poor survival were statistically analyzed with univariate and multivariate COX regression analyses. Univariate analysis identified five risk factors for both poor OS and poor PFS, which included high GIHCG expression, poor or undifferentiated histological types, lymph node metastasis, distant metastasis, and advanced TNM stages (TNM III/IV) (Table 2). Further analyzing these factors with multivariate analysis revealed high GIHCG expression, poor or undifferentiated histological types, and distant metastasis as independent risk factors for both poor OS and poor PFS (Table 3). Taken together, high GIHCG expression accelerates the progression of CRC, and GIHCG may be utilized as a potential prognostic biomarker in CRC.
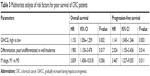  | Table 3 Multivariate analysis of risk factors for poor survival of CRC patients |
GIHCG induces migration, invasion, and proliferation in CRC cells
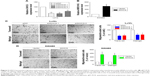
To determine the function of GIHCG in CRC cells, expression of GIHCG was silenced and overexpressed in LoVo cells (Figure 3A) and SW480 cells (Figure 3B), respectively. The Transwell assay and the Matrigel assay discovered that GIHCG deficiency in LoVo cells significantly repressed cell migration ability and invasion (Figure 3C). Accordingly, overexpression of GIHCG markedly promoted the migration and invasion in SW480 cells (Figure 3D). Moreover, the colony formation assay found that interference of GIHCG reduced the colonies in LoVo cells (Figure 4A), while upregulation of GIHCG notably increased the colonies in SW480 cells (Figure 4B). In the CCK-8 assay, GIHCG silencing inhibited cell proliferation (Figure 4C), and overexpression of GIHCG accelerated cell proliferation (Figure 4D).
GIHCG contributes to chemoresistance in CRC cells
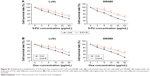
The drug resistance of LoVo-5-FU-R cells, SW480-5-FU-R cells, LoVo-Oxa-R cells, and SW480-Oxa-R cells was validated by detecting cell survival rate when treated with different concentrations of 5-FU or Oxa (Figure S1A and B). Expression of GIHCG was evaluated by qRT-PCR in LoVo-5-FU-R cells, SW480-5-FU-R cells, LoVo-Oxa-R cells, SW480-Oxa-R cells, and corresponding control cells, and the results showed that 5-FU-resistant or Oxa-resistant CRC cells had a much higher GIHCG expression level than corresponding control cells (Figure 5A). Furthermore, knockdown of GIHCG reduced the cell survival rate of LoVo cells under 5-FU treatment, while SW480 cells with GIHCG upregulation showed an obviously improved survival rate (Figure 5B). Accordingly, GIHCG interference significantly inhibited cell survival of LoVo cells under Oxa treatment, and ectopic expression of GIHCG facilitated cell survival of SW480 cells (Figure 5C). In addition, silencing of GIHCG notably reduced the improved cell survival rates of LoVo-5-FU-R cells and LoVo-Oxa-R cells compared with control cells (Figure 5D). Collectively, GIHCG contributes to chemoresistance in CRC cells.
Discussion
Our study first demonstrated the increased expression of GIHCG in CRC cells and tissues, and the significance of GIHCG in promoting CRC clinical progression was also detected. Interestingly, overexpression of GIHCG was correlated with lymphovascular invasion, lymph node metastasis, distant metastasis, and advanced TNM stages, all of which are pivotal factors in predicting the prognosis of CRC patients. As shown in Figure 2A and B, upregulation of GIHCG was confirmed to predict poor OS and PFS in CRC patients. Moreover, high GIHCG expression was identified as an independent risk factor for poor prognosis. Expression of lncRNAs is tissue-specific and relatively stable in body fluid, so they can be detected easily and noninvasively from cancer patients in whole blood, plasma, serum, urine, saliva, and gastric juice samples. These features make them promising biomarkers in cancer diagnosis and prognosis.17,24,25 A number of studies have reviewed the prognostic value of lncRNAs in cancers.25–27 The prognostic value of GIHCG has been verified in hepatocellular carcinoma,21 renal cell carcinoma,22 and tongue cell carcinoma.23 Our results further defined the significance of GIHCG in prognosis. The statistical analyses revealed a close relationship of GIHCG expression and CRC clinicopathological characteristics, which underlies the pivotal role of GIHCG in CRC progression.
Our study first discovered the function of GIHCG contributing to chemoresistance in cancers. We found that GIHCG exhibited much higher expression levels in 5-FU and Oxa resistant CRC cells. Also, the expression level of GIHCG was significantly correlated with cell survival ability under 5-FU and Oxa treatment. In addition, GIHCG deficiency suppressed the elevated cell survival rate of 5-FU-R cells and Oxa-R cells under 5-FU and Oxa treatments, respectively. Drug-resistance remains one of the deadlocks for the low survival rates of CRC patients. Since the 1950s, 5-FU-based chemotherapy has been the mainstay of therapy for patients with CRC.28,29 Oxa, combined with 5-FU and leucovorin, has been the first-line chemotherapy strategy for metastatic CRC.30 However, nearly half of metastatic CRC patients are resistant to 5-FU-based chemotherapies.31 Therefore, it is of paramount importance to elucidate causes underlying this chemoresistance to circumvent it, and to discover more efficient ways in cancer treatment. lncRNAs based therapy provides a potential method for reversing chemoresistance.32 Our results first revealed the function of GIHCG in chemoresistance, and this evidence makes GIHCG a potential target in conquering the chemoresistance of CRC. Combined with traditional chemotherapeutic regimes, CRC patients may achieve a better prognosis.
Conclusion
This study demonstrated the role of GIHCG in accelerating the proliferation, migration, invasion, and chemoresistance in CRC. However, the mechanisms of GIHCG promoting CRC progression were not studied in this study, which deserves further explorations. More investigations from multicenters with more samples are also needed to utilize GIHCG as a prognostic biomarker and therapeutic target in CRC.
Acknowledgment
This study was supported by Shanghai Pudong New Area Science and Technology Development Fund Innovation Fund Project (Number: PKJ2014-Y12), Shanghai Pudong New Area Health Systems Academic Leader Fund Project (Number: PWRD2014-04), and the National Natural Science Foundation (Number: 81672733).
Disclosure
The authors report no conflicts of interest in this work.
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global Cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. | ||
Arends JW. Molecular interactions in the Vogelstein model of colorectal carcinoma. J Pathol. 2000;190(4):412–416. | ||
Lasry A, Zinger A, Ben-Neriah Y. Inflammatory networks underlying colorectal cancer. Nat Immunol. 2016;17(3):230–240. | ||
Strum WB. Colorectal adenomas. N Engl J Med. 2016;374(11):1065–1075. | ||
Thrumurthy SG, Thrumurthy SS, Gilbert CE, Ross P, Haji A. Colorectal adenocarcinoma: risks, prevention and diagnosis. BMJ. 2016;354:i3590. | ||
West NR, McCuaig S, Franchini F, Powrie F. Emerging cytokine networks in colorectal cancer. Nat Rev Immunol. 2015;15(10):615–629. | ||
Yang Y, Junjie P, Sanjun C, Ma Y. Long non-coding RNAs in colorectal cancer: progression and future directions. J Cancer. 2017;8(16):3212–3225. | ||
Fearon ER. Molecular genetics of colorectal cancer. Annu Rev Pathol Mech Dis. 2011;6(1):479–507. | ||
Ransohoff JD, Wei Y, Khavari PA. The functions and unique features of long intergenic non-coding RNA. Nat Rev Mol Cell Biol. 2018;19(3):143–157. | ||
Akhade VS, Pal D, Kanduri C. Long noncoding RNA: genome organization and mechanism of action. Adv Exp Med Biol. 2017;1008:47–74. | ||
Jarroux J, Morillon A, Pinskaya M. History, discovery, and classification of lncRNAs. Adv Exp Med Biol. 2017;1008:1–46. | ||
Kwok ZH, Tay Y. Long noncoding RNAs: lincs between human health and disease. Biochem Soc Trans. 2017;45(3):805–812. | ||
Evans JR, Feng FY, Chinnaiyan AM. The bright side of dark matter: lncRNAs in cancer. J Clin Invest. 2016;126(8):2775–2782. | ||
Bartonicek N, Maag JL, Dinger ME. Long noncoding RNAs in cancer: mechanisms of action and technological advancements. Mol Cancer. 2016;15(1):43. | ||
Kondo Y, Shinjo K, Katsushima K. Long non-coding RNAs as an epigenetic regulator in human cancers. Cancer Sci. 2017;108(10):1927–1933. | ||
Tang Y, Cheung BB, Atmadibrata B, et al. The regulatory role of long noncoding RNAs in cancer. Cancer Lett. 2017;391:12–19. | ||
Bolha L, Ravnik-Glavač M, Glavač D. Long noncoding RNAs as biomarkers in cancer. Disease Markers. 2017;2017:1–14. | ||
Xue M, Zhuo Y, Shan B. MicroRNAs, long noncoding RNAs, and their functions in human disease. Methods Mol Biol. 2017;1617:1–25. | ||
Slaby O, Laga R, Sedlacek O. Therapeutic targeting of non-coding RNAs in cancer. Biochem J. 2017;474(24):4219–4251. | ||
Rao AKDM, Rajkumar T, Mani S. Perspectives of long non-coding RNAs in cancer. Mol Biol Rep. 2017;44(2):203–218. | ||
Sui CJ, Zhou YM, Shen WF, et al. Long noncoding RNA GIHCG promotes hepatocellular carcinoma progression through epigenetically regulating miR-200b/a/429. J Mol Med. 2016;94(11):1281–1296. | ||
He ZH, Qin XH, Zhang XL, Yi JW, Han JY. Long noncoding RNA GIHCG is a potential diagnostic and prognostic biomarker and therapeutic target for renal cell carcinoma. Eur Rev Med Pharmacol Sci. 2018;22(1):46–54. | ||
Ma L, Wang Q, Gong Z, Xue L, Zuo Z. Long noncoding RNA GIHCG enhanced tongue squamous cell carcinoma progression through regulating miR-429. J Cell Biochem. 2018;119(11):9064–9071. | ||
Chandra Gupta S, Nandan Tripathi Y. Potential of long non-coding RNAs in cancer patients: from biomarkers to therapeutic targets. Int J Cancer. 2017;140(9):1955–1967. | ||
Jiang C, Li X, Zhao H, Liu H. Long non-coding RNAs: potential new biomarkers for predicting tumor invasion and metastasis. Mol Cancer. 2016;15(1):62. | ||
Bhan A, Soleimani M, Mandal SS. Long noncoding RNA and cancer: a new paradigm. Cancer Res. 2017;77(15):3965–3981. | ||
Deng H, Wang JM, Li M, et al. Long non-coding RNAs: new biomarkers for prognosis and diagnosis of colon cancer. Tumour Biol. 2017;39(6):1010428317706332. | ||
Salonga D, Danenberg KD, Johnson M, et al. Colorectal tumors responding to 5-fluorouracil have low gene expression levels of dihydropyrimidine dehydrogenase, thymidylate synthase, and thymidine phosphorylase. Clin Cancer Res. 2000;6(4):1322–1327. | ||
Showalter SL, Showalter TN, Witkiewicz A, et al. Evaluating the drug-target relationship between thymidylate synthase expression and tumor response to 5-fluorouracil. Is it time to move forward? Cancer Biol Ther. 2008;7(7):986–994. | ||
Yaffee P, Osipov A, Tan C, Tuli R, Hendifar A. Review of systemic therapies for locally advanced and metastatic rectal cancer. J Gastrointest Oncol. 2015;6(2):185–200. | ||
Douillard JY, Cunningham D, Roth AD, et al. Irinotecan combined with fluorouracil compared with fluorouracil alone as first-line treatment for metastatic colorectal cancer: a multicentre randomised trial. The Lancet. 2000;355(9209):1041–1047. | ||
Luo J, Qu J, Wu DK, Lu ZL, Sun YS, Qu Q. Long non-coding RNAs: a rising biotarget in colorectal cancer. Oncotarget. 2017;8(13):22187–22202. |
Supplementary material
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.