Back to Journals » International Journal of Nanomedicine » Volume 9 » Issue 1
In vivo MR detection of fluorine-labeled human MSC using the bSSFP sequence
Authors Ribot E, Gaudet J, Chen Y, Gilbert K, Foster P
Received 13 December 2013
Accepted for publication 15 February 2014
Published 8 April 2014 Volume 2014:9(1) Pages 1731—1739
DOI https://doi.org/10.2147/IJN.S59127
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 4
Emeline J Ribot,1 Jeffrey M Gaudet,1,2 Yuhua Chen,1 Kyle M Gilbert,1 Paula J Foster1,2
1Imaging Research Laboratories, Robarts Research Institute, London, ON, Canada; 2Department of Medical Biophysics, University of Western Ontario, London, ON, Canada
Abstract: Mesenchymal stem cells (MSC) are used to restore deteriorated cell environments. There is a need to specifically track these cells following transplantation in order to evaluate different methods of implantation, to follow their migration within the body, and to quantify their accumulation at the target. Cellular magnetic resonance imaging (MRI) using fluorine-based nanoemulsions is a great means to detect these transplanted cells in vivo because of the high specificity for fluorine detection and the capability for precise quantification. This technique, however, has low sensitivity, necessitating improvement in MR sequences. To counteract this issue, the balanced steady-state free precession (bSSFP) imaging sequence can be of great interest due to the high signal-to-noise ratio (SNR). Furthermore, it can be applied to obtain 3D images within short acquisition times. In this paper, bSSFP provided accurate quantification of samples of the perfluorocarbon Cell Sense-labeled cells in vitro. Cell Sense was internalized by human MSC (hMSC) without adverse alterations in cell viability or differentiation into adipocytes/osteocytes. The bSSFP sequence was applied in vivo to track and quantify the signals from both Cell Sense-labeled and iron-labeled hMSC after intramuscular implantation. The fluorine signal was observed to decrease faster and more significantly than the volume of iron-associated voids, which points to the advantage of quantifying the fluorine signal and the complexity of quantifying signal loss due to iron.
Keywords: bSSFP, fluorine MRI, mesenchymal stem cell, mouse, cell tracking
Introduction
Cellular therapy is a mainstay in the treatment of oncologic and hematologic diseases and it offers hope for a very large range of other diseases and disorders. Among the various stem cell populations used for cell therapy, adult mesenchymal stem cells (MSC, also referred to as mesenchymal stromal cells) have emerged as a major new cell technology with a diverse spectrum of potential clinical applications.1,2 At least 92 clinical trials are currently using MSC worldwide (http://clinicaltrials.gov/).
The results of preclinical and clinical studies paint a promising picture for stem cell-based therapies, and consequently, there are tremendous expectations for stem cell research. However, many important questions persist relating to the in vivo fate of transplanted stem cells. Translational research in experimental animal models is essential, with a critical emphasis on developing methods for monitoring the viability, as well as the temporal and spatial homing of these cells to target tissue.
Magnetic resonance imaging (MRI) has emerged as an excellent method for tracking cells in vivo. Many cell tracking studies have used iron oxide nanoparticle-based contrast agents to label cells for detection with MRI. Iron-labeled cells appear as distinct regions of signal hypointensity (signal loss) in images. The sensitivity for detection of iron-labeled cells is very high. Even single iron-labeled cells can be visualized in vivo under some conditions.3,4 A challenge encountered with iron-based cell tracking is that other endogenous sources of signal loss also appear in images that are sensitive to iron (for example due to blood, hemosiderin, bone, and air) making it difficult to unambiguously identify regions containing labeled cells.5 In addition, iron-labeled cell quantification is difficult. We, and others, have shown that the contrast generated by iron-labeled cells increases with the amount of iron/voxel but that this is only linear at low iron loadings; the change in contrast reaches a saturation plateau at higher iron loadings.6,7 When quantifying the presence of iron-labeled stem cells over time, most studies measure the “signal void volume”8,9 or the “number of black pixels”,10,11 and present the change relative to the first imaging time point.
Perfluorocarbon (PFC) nanoemulsion formulations have also been used, to a more limited extent, for fluorine-19 (19F) MRI cell tracking.12 One of the key advantages of these agents is the very high specificity; since this atom is mainly absent from the body, there is no background 19F signal in MR images. In addition, the fluorine signal can be accurately quantified from the MR images by comparing the 19F signal in the tissue of interest to an external reference containing a known amount of fluorine atoms. Finally, unlike iron nanoparticles, perfluorocarbon nanoemulsions are biologically inert. Common enzymes do not cleave the bond between carbon and fluorine.13 Fluorine-based compounds are already used in patients as blood substitutes and their safety is well-established.14,15 Furthermore, the number of fluorine atoms that are internalized by cells after labeling is several orders of magnitude lower than what is typically used in artificial blood substitutes.
Examples of 19F MR imaging performed to date include: monitoring inflammation (tumor models, rheumatoid arthritis, lung injuries, and after ischemia in the lung or brain),16–19 detection of T cell and dendritic cell migration,20,21 and tracking the fate of transplanted hematopoietic and neural stem cells.22–24
The main limitation of 19F cell tracking is its relatively low sensitivity when compared to cellular imaging with iron nanoparticles. In this paper, we compare 19F- and iron-based methods for MSC tracking in vivo. Our specific goals were: 1) to evaluate the use of a balanced steady-state free precession (bSSFP) sequence for 19F cell tracking; 2) to demonstrate that primary human MSC (hMSC) could be labeled with a sufficient quantity of the 19F agent, Cell Sense, to allow for cell detection, without altering cell viability or differentiation; and 3) to compare the information obtained from imaging iron- and 19F-labeled hMSC after intramuscular (IM) injection in healthy mice.
Materials and methods
Human mesenchymal stromal cell culture and labeling
hMSC came from bone marrow donated by healthy young adult volunteers after written informed consent according to a protocol approved by the University Health Network Research Ethics Board (Toronto, ON, Canada).25 MSC were grown until passage 4. The cells were cultured in Dulbecco’s Modified Eagle’s Medium low glucose, 2 mM of L-glutamine, 12.5% fetal bovine serum, and 100 u/mL of penicillin and streptomycin. The cells were passaged twice before labeling and injection into mice.
hMSC were labeled with both iron-based and 19F-based agents. The iron-based agent was Molday ION Rhodamine-B (MIRB), an ultra-small iron oxide nanoparticle (BioPal, Worcester, MA, USA). MIRB has a colloidal size of 35 nm, a zeta potential of +31 mV and is cross-linked and labeled with Rhodamine B, allowing visualization by MRI and fluorescence microscopy. The 19F-based agent was Cell Sense (CS-1000), a perfluoropolyether (PFPE) nanoparticle (Celsense Inc., Pittsburg, PA, USA). Cell Sense is an aqueous colloidal suspension (ie, nanoemulsion) of a PFPE perfluorocarbon polymer, having a total fluorine content of 145 mg/mL. The average nanoemulsion droplet size is 180 nm. Cell Sense is formulated with excipients that expedite entry of the nanomaterial into all cell types, regardless of their ability to phagocytose. The PFPE molecule used in Cell Sense is both lipophobic and hydrophobic and stable at low pH.20 In some cases, a red fluorescent version of this PFPE was used (CS-ATM-DM Red).
Cell Sense was added to the cell culture medium at a concentration of 2.5 mg/mL and incubated with the hMSC for 24 hours. MIRB was incubated with hMSC at a concentration of 50 μg/mL for 24 hours.26 After this incubation period, the cells were washed three times with Hank’s balanced salt solution (HBSS [Life Technologies, Carlsbad, CA, USA]), harvested with Trypsin, spun down, and counted.
To evaluate the amount of 19F internalized by the hMSC, the Cell Sense-labeled cells were fixed overnight with 4% paraformaldehyde (PFA) then lysed by sonication using 200 μL of 1% Triton X-100 (Sigma-Aldrich, St Louis, MO, USA) and incubation for 4 hours at room temperature in a sonication bath. This solution was then placed into a glass nuclear magnetic resonance (NMR) tube (5 mm diameter), to which 200 μL of D2O and 100 μL of 0.1% trifluoroacetic acid (TFA) was added. 19F-spectroscopy was then performed at 400 MHz on a vertical scanner using the following parameters: TR =7 seconds; rBW =18859 Hz; NEX =100; line broadening =5 Hz, spectral range from −68 ppm to −93 ppm. The ratio of the areas under the TFA and the Cell Sense-labeled hMSC peaks were calculated to determine the amount of fluorine atoms per cell, as described by Srinivas et al.21
hMSC differentiation assay
To assess the effect of Cell Sense labeling on hMSC differentiation, hMSC were cultured in six-well plates until they reached 70% or 100% confluency for adipogenic and osteogenic differentiation assays, respectively. Then, three wells were incubated with 2.5 mg/mL of Cell Sense nanoparticles for 24 hours and the other three wells were used for the unlabeled hMSC condition. The wells were washed three times with HBSS before adipogenic or osteogenic media was added (StemXVivoTM Osteogenic/Adipogenic Base Media supplemented with StemXVivoTM Osteogenic Supplement or StemXVivoTM Adipogenic Supplement; R&D Systems, Inc., Burlington, ON, Canada). Fresh media was replaced every 3–4 days. After 3 weeks, the cells were stained with either Oil red O, to identify lipid vesicles, or Alizarin Red S to identify extracellular calcification drops.
The cells were examined using a Zeiss AXIO Imager A1 microscope (Carl Zeiss Canada Ltd, Toronto, ON, Canada). The number of adipocytes per condition (unlabeled and Cell Sense-labeled hMSC) were counted in five fields of view (FOV) per well (three wells per condition), and the mean of these taken. Osteogenesis was evaluated by qualitatively comparing the number of extracellular calcification drops in each well.
MRI sequence
All MRI was performed on a 9.4 T Varian small-animal MRI scanner equipped with a custom-built dual-tuned proton/fluorine birdcage coil (diameter =2.2 cm, length =5.1 cm; Agilent, Santa Clara, CA, USA). The coil was tuned to 400.2 MHz for proton imaging and 376.8 MHz for fluorine imaging. A 3D bSSFP sequence was evaluated for fluorine detection using cell samples containing different numbers of Cell Sense-labeled hMSC (2×105; 4×105; 6×105; 8×105, and 1×106) pelleted after fixation in 4% PFA. The bSSFP parameters were: TR =2.7 milliseconds; FA =70°, rBW =25 kHz; FOV =32×24×36 mm; matrix =32×24×18; resolution =1×1×2 mm; NEX =950; 1 hour 17 minutes. Afterwards, NEX was reduced by half (475) to examine the influence of this parameter on the detection limit (acquisition time was thus shorter at 38 minutes).
In vivo MRI of implanted hMSC
Cell Sense-labeled hMSC at 1.5×106 and MIRB-labeled hMSC at 1.5×106 (in 100 μL HBSS/mouse) were injected IM into the right and left hind limb, respectively, in six female nude mice (nu/nu Foxn1 mice, 6–7 weeks old; Charles River Laboratories, Wilmington, MA). These injections were performed while under inhalational anesthesia (isoflurane, 1%) that lasted less than 5 minutes per mouse. For MRI, the mice were anesthetized with an intraperitoneal injection of pentobarbital (65 mg/kg) to avoid detecting isoflurane on the 19F MR images. Four of the six mice were imaged on days 0 (4–6 hours after the injection), 3, and 12 after injection. Two of the six mice were imaged on days 0, 3, 12, 17, and 26 days post-injection. A reference tube containing 2.6×1016 19F atoms/μL was always inserted into the coil with the mice.
The bSSFP parameters were the same as for the hMSC pellets but with NEX =750 and a scan time of around 1 hour. The corresponding proton images were also acquired using bSSFP, for anatomical localization of the 19F signal and to visualize the iron-labeled hMSC, with the following parameters: TR =8 milliseconds; FA =30°; rBW =125 kHz; resolution =200×200×200 μm; eight phase cycles; NEX =2; 46 minutes.
All experiments were approved by the Western University Animal Use Committee.
Analysis
The 19F images were analyzed using Voxel Tracker™ software (Celsense Inc.) and the fluorine signal was quantified by comparison with the signal generated by the reference tube containing a known amount of Cell Sense nanoparticles (2.6×1016 19F/μL). The amount of fluorine atoms measured at day 0 was set to 100% and for each mouse, the amounts measured at day 3 and day 12 were normalized to day 0. The signal void volume due to the presence of MIRB-labeled hMSC was measured from the 1H images using the image analysis program Osirix (Pixmeo SARL, Bernex, Switzerland).27 The volume of the void measured at day 0 was set to 100% and for each mouse, the void volumes measured at day 3 and day 12 were normalized to day 0. For both fluorine and iron quantifications, regions of interest (ROIs) were hand-drawn around the voxels containing fluorine or void signals. ROIs were drawn on every MR slice where the signals appeared. Student’s t-tests were performed.
Immunohistochemistry staining
At endpoint, the mice were sacrificed and perfused with 4% formalin. The hind limb muscles were extracted and cryoprotected with increasing concentrations of sucrose before being frozen and cut into 16 μm sections using a cryostat. Non-specific binding was blocked using 10% normal goat serum (Vector Laboratories, Burlingame, CA, USA) and 0.1% Triton-X-100 (Sigma-Aldrich) in PBS for 1 hour at room temperature. Sections were washed with PBS. Then they were incubated overnight at 4°C with the mouse anti-human Golgi zone (Chemicon International, Temecula, CA, USA) diluted at 1/500 in blocking solution (as described by Gordon et al).28 Slides were mounted in Vectashield containing 4,6-diamidino-2-phenylindole (DAPI; Vector Laboratories).
Results
Labeling of hMSC with Cell Sense
After an incubation of 24 hours, hMSC internalized high amounts of the Cell Sense agent, without the need for transfection agents, as visualized by fluorescence microscopy (Figure 1). The Cell Sense particles are located within the cytoplasm and more precisely into vesicles, as seen by the granulated shape of the fluorescence in Figure 1C. NMR experiments demonstrated that the hMSC internalized 7.1±1.7×1010 19F atoms/cell on average.
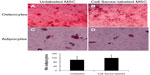
Labeling the hMSC with Cell Sense did not affect differentiation into osteocytes or adipocytes (Figure 2). The amount of calcification drops secreted in the extracellular medium appeared similar for unlabeled (Figure 2A) and labeled hMSC (Figure 2B). There was no significant difference between the number of adipocytes counted for unlabeled and Cell Sense-labeled hMSC (Figure 2C and D).
In vitro detection of Cell Sense-labeled hMSC using bSSFP
The minimum number of Cell Sense-labeled hMSC required for detection using a 3D-bSSFP sequence was analyzed in vitro with Cell Sense-labeled hMSC pellets imaged at 9.4 T. Two slices that cut through the 2×105, 6×105, 1×106 pellets (top row, left) and through the 4×105, 8×105 pellets (top row, right) are shown in Figure 3A. All cell pellets were easily detected with 1 hour scans. The quantification of the fluorine signal (Figure 3B) confirmed that the relationship between the cell numbers and the amount of fluorine atoms detected by the bSSFP sequence is linear (with a correlation factor R2 of 0.82).
The same bSSFP sequence was performed on the pellets, with half of the signal averages (and so half the scan time) (Figure 3A, lower row). All of the cell pellets were still visible except for 2×105 hMSC (dashed circle). Thus, all further images were performed using the higher number of averages (750), but limiting scan time to 1 hour for the in vivo experiments, as is usually performed for small animal imaging.20,23,24
In vivo detection of Cell Sense- and MIRB-labeled hMSC
Each nude mouse received injections of Cell-Sense- and MIRB-labeled hMSC in their left and right thigh muscles, respectively. Figure 4A shows the proton image of an axial slice located at the level of the groin for a representative mouse, acquired on the day of the injection (day 0). The injection site of the Cell Sense-labeled hMSC could be detected on the proton image as an area slightly more hyper-intense than the surrounding muscle (red arrow), due to saline. The corresponding fluorine image is shown in Figure 4B, where only the end of the reference tube and the injected labeled cells were detected. Figure 4C shows the overlay between the proton and the fluorine images, allowing for precise location of the injected cells in the mouse body.
As shown in Figure 4D–F, the fluorine signal can be tracked over time in the muscle (D at day 3, E at day 12, F at day 26 post-injection). The total amount of 19F atoms localized at the injection site (shown at the top of each image) decreased over time, likely due to hMSC death and clearance from the mouse body.
The MIRB-labeled hMSC injected on the opposite side could also be easily detected through the visualization of the area of signal loss in the muscle (Figure 4D–F, blue arrows). It should be noted, however, that the bones in the mouse legs also appeared as signal loss (yellow arrow). Endogenous sources of signal loss such as this can confuse the interpretation of contrast attributed to the presence of iron-labeled cells. For both Cell Sense-labeled and MIRB-labeled hMSC, the injected cells were visible at the site of their injection over the period of time that the mice were scanned (26 days) and were not detected in other locations or organs.
The signal due to the implanted hMSC was quantified for both cell labels at each imaging time point. For MIRB-labeled cells, the volume of the void area induced by the presence of MIRB-labeled hMSC was measured (Figure 5A). There was a significant 20% decrease in the void volume noticeable at day 12 (P=0.044 between day 0 and day 12). For Cell Sense-labeled cells, the fluorine signal was quantified by comparing the signal within the ROI to the signal from the external reference tube (Figure 5B). The SNR of the 19F-hMSC was higher than 7 at all imaging time points, allowing for accurate detection for at least 26 days after the injections. The number of 19F atoms also significantly decreased with time (P=0.004 between day 0 and day 12; P=0.025 between day 3 and day 12). At day 12, the fluorine signal was approximately 50% of that at day 0. This decrease was greater than the one measured from the MIRB-labeled cells.
Microscopy of muscle tissue
Mice were sacrificed 26 days after the cell injections. Fluorescence microscopy demonstrated that the location of the hMSC corresponded with the location of the red fluorescence-tagged Cell Sense nanoparticles (Figure 6A). Immunohistochemistry staining against human Golgi verified the presence of the hMSC at the injection site, next to the muscle fibers (Figure 6B).
Discussion
In this study, we show that efficient labeling of hMSCs with a PFPE-based marker is possible without impacting cell viability or differentiation. This result agrees with other studies that demonstrate the ability of hMSC to internalize iron nanoparticles such as MIRB without cytotoxicity or affecting stem cell differentiation.26,29 Others have also shown that Cell Sense does not impact the phenotype or functioning of dendritic cells12 or differentiation in other stem cell types.22,23 We have also successfully demonstrated in vivo detectability of hMSC labeled with the only fluorine-based contrast agent that has been FDA-approved for cellular imaging, using the 3D-bSSFP imaging sequence. This is the first study to monitor the fate of hMSC using 19F MRI.
Cell detection with bSSFP
The image SNR is key to the ability to detect PFPE-labeled cells by 19F MRI. One important finding from this work was that 19F-labeled human stem cells can be detected and tracked over time with the bSSFP imaging sequence at high magnetic field. It has already been demonstrated that bSSFP is the most SNR efficient pulse sequence in use.30 Proof of principle that bSSFP was advantageous for the detection of 19F-labeled stem cells has been previously reported by Partlow et al.31 In this paper, a liquid PFC was developed and used to label stem/progenitor cells in culture. A single mouse then received an IM injection of 4 million fixed PFC-labeled cells and was imaged after euthanasia at 1.5 T with 2D bSSFP. To the best of our knowledge, this is the only other published report on the use of bSSFP sequence to detect 19F-labeled cells after transplantation.
Two other groups have utilized the high SNR engendered by the 3D bSSFP sequence along with the intravenous injection of the perfluorocarbon V-Sense to study inflammation.32–36 However, these studies used a chemical shift imaging (CSI) approach with bSSFP to allow for visualization of the different spatial distributions of chemically shifted PFC compounds. Overall, the vast majority of studies that perform cell tracking with 19F MRI have employed spin echo sequences.16–19,37–40
Using the 3D bSSFP sequence, we injected and detected 1.5×106 Cell Sense-labeled hMSC in vivo on day 0 using a spatial resolution of 2 mm3 (2 μL). If we assume that on day 0 all of the injected cells are present and evenly distributed, then we detected 30,000 cells per voxel. The literature reports a wide range of detection thresholds for 19F-labeled cells, and the various studies use a range of parameters to perform the cell labeling and MRI. This sensitivity of detection will vary depending on MRI factors (magnetic field strength, pulse sequence, voxel size, radiofrequency coil) and on cell labeling efficiency, which is influenced both by the type of label used and by the cell-labeling protocol. For example, Partlow et al injected and detected 4 million cells in 50 μL at 1.5 T ex vivo.31 The voxel size was 2.5×2.5×35 mm (~220 μL); therefore, they detected 4 million cells in a voxel. They did this in a far shorter imaging time of 7 minutes compared to our 40 minutes. Higher resolution imaging would have resulted in fewer cells/voxel, and consequently, increased scan time.
In vivo detection of Cell Sense-labeled hMSC over time
Our in vivo data demonstrated that 1.5×106 hMSC were easily detected soon after IM injection (within hours on day 0). Three days post-implantation, the number of labeled cells at the implantation site was not significantly different from day 0. Between days 3 and 12 post-implantation, we observed a significant decrease in the signal. A continual gradual decline in the 19F signal was observed until day 26 post-implantation. The MRI signal persisted for at least 26 days and fluorescence microscopy validated the presence of Cell Sense-positive hMSC remaining at the site of cell implantation. The fact that the fluorine signal was still easily detectable and quantifiable as long as 26 days post-injection is likely because the hMSC were implanted in healthy tissue in immunocompromised mice. In diseased tissue and immunocompetent recipients, the rate of cell death would be expected to be greater.
Only a few other studies have used MRI to detect 19F-labeled stem cells in vivo, and only three studies have performed imaging at multiple time points to monitor their fate after transplantation. Boehm-Sturm et al first demonstrated that the fluorine signal changes by less than 20% from day 2 and day 6 after the implantation of 150,000 human neural stem cells (NSCs) directly into the striatum in healthy mice.23 Ruiz-Cabello et al imaged mouse NSCs implanted into normal mouse striatum and showed that the 19F signal remained constant for 1 week post-implantation and was diminished but still visible for up to 2 weeks.41 Histopathology showed that the labeled cells remained viable and that labeled cells had migrated from the site of implantation into the brain parenchyma. Most recently, Bible et al used 19F MRI to monitor the fate of human NSCs suspended in a bioscaffold transplanted into the lesion cavity in a rat model of stroke.24 The area covered by the 19F signal decreased between days 1 and 7. Our results with hMSCs are in agreement with these other longitudinal imaging studies that demonstrate the robust early detection of transplanted cells followed by a decline in the 19F signal that may be due to cell division or cell death. However, they contrast with the study by Helfer et al, where 14×106 Cell Sense-labeled human hematopoietic stem cells (HSCs) either trapped in matrigel or suspended in PBS were imaged after being implanted into the rat hind limb muscle.22 Surprisingly, 2 hours post-implantation, a 19F signal was only detected in the rat implanted with the matrigel containing labeled cells, although eight times more labeled cells were injected compared to our study. This observation may suggest a notable difference in the in vivo fate of intramuscular (IM) transplanted HSC.
Comparison of signal from 19F- versus iron-labeled cells
This is the first study to compare the change in signal due to iron- versus 19F-labeled hMSCs in the same mice over time in vivo. Our results show that the volume of the region of signal loss due to iron-labeled cells diminished more slowly over time compared to the signal due to 19F-labeled cells. This discrepancy is likely related to the fact that bystander cells can be labeled secondarily through the release of labels from dead stem cells;42 although the bystander cells will contain much lower amounts of the label. Because of the large differences in sensitivity of iron or fluorine signal, iron-labeled bystander cells will still produce signal loss, whereas 19F-labeled bystander cells may be undetectable. Several studies have pointed out that the signal loss produced by iron-labeled cells can greatly overestimate the presence of the viable transplanted cell population.29,42–44
The signal void caused by the presence of iron-labeled cells is similar to hypointense signal produced by bone, air, and hemorrhage. This is a major hurdle to the use of MRI to reliably identify iron-labeled cells after their transplantation. A major advantage of 19F MRI is that the detection of the probe is specific to the labeled cells because of the lack of endogenous 19F in the body. Therefore, unlike when using iron oxide nanoparticles, the signal is highly specific and interpretation is straightforward.
Use of the bSSFP sequence can be limited by banding artifacts that get more numerous at high magnetic fields. However, this artifact can be suppressed with shortened TRs (such as that used in this study) or by employing phase cycling.
Conclusion
In conclusion, we have shown that the bSSFP imaging sequence can be employed to detect and track fluorine-labeled human stem cells in vivo on 3D images. Furthermore, this work highlights the discrepancy in signal changes over time between 19F- and iron-labeled cells. 19F MRI provides an essential tool for researchers to study the main issues related to the use of hMSC using preclinical small animal models at high magnetic field.
Acknowledgments
We would like to acknowledge Dr A Keating and Dr X Wang for supplying human MSC.
Disclosure
The authors report no conflicts of interest in this work.
References
Chagastelles PC, Nardi NB, Camassola M. Biology and applications of mesenchymal stem cells. Science Progress. 2010;93:113–127. | |
Da Silva Meirelles L, Nardi NB. Methodology, biology and clinical applications of mesenchymal stem cells. Front Biosci. 2009;14:4281–4498. | |
Heyn C, Ronald JA, Mackenzie LT, et al. In vivo magnetic resonance imaging of single cells in mouse brain with optical validation. Magn Reson Med. 2006;55:23–29. | |
Shapiro EM, Sharer K, Skrtic S, Koretsky AP. In vivo detection of single cells by MRI. Magn Reson Med. 2006;55:242–249. | |
Van Den Bos EJ, Baks T, Moelker AD, et al. Magnetic resonance imaging of haemorrhage within reperfused myocardial infarcts: possible interference with iron oxide-labelled cell tracking. Eur Heart J. 2006;27:1620–1626. | |
Heyn C, Bowen CV, Rutt BK, Foster PJ. Detection threshold of single SPIO-labeled cells with FIESTA. Magn Reson Med. 2005;53:312–320. | |
Bonetto F, Srinivas M, Heerschap A, et al. A novel 19F agent for detection and quantification of human dendritic cells using magnetic resonance imaging. Int J Cancer. 2011;129:365–373. | |
Hill JM, Dick AJ, Raman VK, et al. Serial cardiac magnetic resonance imaging of injected mesenchymal stem cells. Circulation. 2003;108:1009–1014. | |
Stuckey DJ, Carr CA, Phil D, et al. Iron particles for noninvasive monitoring of bone marrow stromal cell engraftment into, and isolation of viable engrafted donor cells from the heart. Stem Cells. 2006;24:1968–1975. | |
Long CM, Van Laarhoven HW, Bulte JW, Levitsky HI. Magnetovaccination as a novel method to assess and quantify dendritic cell tumor antigen capture and delivery to lymph nodes. Cancer Res. 2009;69:3180–3187. | |
Cromer Berman S, Galpoththawela C, Gilad AA, Bulte JWM, Walczak P. Long-term MR cell tracking of neural stem cells grafted in immunocompetent versus immunodeficient mice reveals distinct differences in contrast between live and dead cells. Magn Reson Med. 2011;65:564–574. | |
Ahrens ET, Flores R, Xu H, Morel PA. In vivo imaging platform for tracking immunotherapeutic cells. Nature Biotech. 2005;23:983–987. | |
O’Hagan D. Understanding organofluorine chemistry. An introduction to the C–F bond. Chem Soc Rev. 2008;37:308–319. | |
Krafft MP. Fluorocarbons and fluorinated amphiphiles in drug delivery and biomedical research. Adv Drug Deliv Rev. 2001;47:209–228. | |
Bulte JWM. Hot spot MRI emerges from the background. Nat Biotechnol. 2005;23(8):945–946. | |
Ebner B, Behm P, Jacoby C, et al. Early assessment of pulmonary inflammation by 19F MRI in vivo. Circ Cardiovasc Imaging. 2010;3:202–210. | |
Flögel U, Ding Z, Hardung H, et al. In vivo monitoring of inflammation after cardiac and cerebral ischemia by fluorine magnetic resonance imaging. Circulation. 2008;118:140–148. | |
Balducci A, Wen Y, Zhang Y, et al. A novel probe for the non-invasive detection of tumor-associated inflammation. Onco Immunol. 2013;2:1–11. | |
Balducci A, Helfer BM, Ahrens ET, O’Hanlon CF, Wesa AK. Visualizing arthritic inflammation and therapeutic response by fluorine-19 magnetic resonance imaging (19F MRI). J Inflammation. 2012;9:24–34. | |
Helfer BM, Balducci A, Nelson AD, et al. Functional assessment of human dendritic cells labeled for in vivo 19F magnetic resonance imaging cell tracking. Cytother. 2010;12:238–250. | |
Srinivas M, Morel PA, Ernst LA, Laidlaw DH, Ahrens ET. Fluorine-19 MRI for visualization and quantification of cell migration in a diabetes model. Magn Reson Med. 2007;58:725–734. | |
Helfer BM, Balducci A, Sadeghi Z, et al. 19F MRI tracer preserves in vitro and in vivo properties of hematopoietic stem cells. Cell Transplant. 2013;22:87–97. | |
Boehm-Sturm P, Mengler L, Wecker S, Hoehn M, Kallur T. In vivo tracking of human neural stem cells with 19F magnetic resonance imaging. PLoS One. 2011;6:e29040. | |
Bible E, Dell’Acqua F, Solanky B, et al. Non-invasive imaging of transplanted human neural stem cells and ECM scaffold remodeling in the stroke-damaged rat brain by 19F- and diffusion-MRI. Biomaterials. 2012;33:2858–2871. | |
Dayan V, Yannarelli G, Billia F, et al. Mesenchymal stromal cells mediate a switch to alternatively activated monocytes/macrophages after acute myocardial infarction. Basic Res Cardiol. 2011;106:1299–1310. | |
McFadden C, Mallett CL, Foster PJ. Labeling of multiple cell lines using a new iron oxide agent for cell tracking by MRI. Contrast Media Mol Imaging. 2011;6(6):514–522. | |
Rosset A, Spadola L, Ratib O. OsiriX: An open-source software for navigating in multidimensional DICOM images. J Digital Imaging. 2004;17:205–216. | |
Gordon D, Pavlovska G, Uney JB, Wraith DC, Scolding NJ. Human mesenchymal stem cells infiltrate the spinal cord, reduce demyelination, and localize to white matter lesions in experimental autoimmune encephalomyelitis. J Neuropathol Exp Neurol. 2010;69:1087–1095. | |
Noad J, Gonzalez-Lara LE, Broughton HC, et al. MRI tracking of transplanted iron-labeled mesenchymal stromal cells in an immune-compromised mouse model of critical limb ischemia. NMR Biomed. 2013;26(4):458–467. | |
Scheffler K, Lehnhardt S. Principles and applications of balanced SSFP techniques. Eur Radiol. 2003;13:2409–2418. | |
Partlow KC, Chen J, Brant JA, et al. 19F magnetic resonance imaging for stem/progenitor cell tracking with multiple unique perfluorocarbon nanobeacons. FASEB J. 2007;21:1647–1654. | |
Hertlein T, Sturm V, Kircher S, et al. Visualization of abscess formation in a murine thigh infection model of Staphylococcus aureus by 19F-magnetic resonance imaging (MRI). PLoS One. 2011;6(3):e18246. | |
Weise G, Basse-Luesebrink TC, Wessig C, Jakob PM, Stoll G. In vivo imaging of inflammation in the peripheral nervous system by 19F MRI. Exp Neurol. 2011;229:494–501. | |
Weise G, Basse-Lusebrink TC, Kleinschnitz C, Kampf T, Jakob PM, Stoll G. In vivo imaging of stepwise vessel occlusion in cerebral photothrombosis of mice by 19F MRI. PLoS One. 2011;6(12):e28143. | |
Kampf T, Fischer A, Basse-Lüsebrink TC, et al. Application of compressed sensing to in vivo 3D 19F CSI. J Magn Reson. 2010;207:262–273. | |
Hertlein T, Sturm V, Jakob P, Ohlsen K. 19F Magnetic resonance imaging of perfluorocarbons for the evaluation of response to antibiotic therapy in a Staphylococcus aureus infection model. PLoS One. 2013;8(5):e64440. | |
Kadayakkara DK, Ranganathan S, Young WB, Ahrens ET. Assaying macrophage activity in a murine model of inflammatory bowel disease using fluorine-19 MRI. Lab Invest. 2012;92:636–645. | |
Van Heeswijk RB, De Blois J, Kania G, et al. Selective in vivo visualization of immune-cell infiltration in a mouse model of autoimmune myocarditis by fluorine-19 cardiac magnetic resonance. Circ Cardiovasc Imaging. 2013;6:277–284. | |
Weibel S, Basse-Luesebrink TC, Hess M, et al. Imaging of intratumoral inflammation during oncolytic virotherapy of tumors by 19F-magnetic resonance imaging (MRI). PLoS One. 2013;8(2):e56317. | |
Hitchens TK, Ye Q, Eytan DF, Janjic JM, Ahrens ET, Ho C. 19F MRI detection of acute allograft rejection with in vivo perfluorocarbon labeling of immune cells. Magn Reson Med. 2011;65:1145–1154. | |
Ruiz-Cabello J, Walczak P, Kedziorek DA, et al. In vivo “hot spot” MR imaging of neural stem cells using fluorinated nanoparticles. Magn Reson Med. 2008;60:1506–1511. | |
Pawelczyk E, Jordan EK, Balakumaran A, et al. In vivo transfer of intracellular labels from locally implanted bone marrow stromal cells to resident tissue macrophages. PLoS One. 2009;4:e6712. | |
Terrovitis J, Stuber M, Youssef A, et al. Magnetic resonance imaging overestimates ferumoxide-labeled stem cell survival after transplantation in the heart. Circulation. 2008;117:1555–1562. | |
Gonzalez-Lara LE, Xu X, Hofstetrova K, et al. The use of cellular magnetic resonance imaging to track the fate of iron-labeled multipotent stromal cells after direct transplantation in a mouse model of spinal cord injury. Mol Imaging Biol. 2011;13:702–711. |
 © 2014 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms
and incorporate the Creative Commons Attribution
- Non Commercial (unported, 3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2014 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms
and incorporate the Creative Commons Attribution
- Non Commercial (unported, 3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.