Back to Archived Journals » Current Biomarker Findings » Volume 5
Improving tuberculosis diagnostics with biomarkers
Authors Shu C, Wang J, Lee L, Yu C ![]() , Luh K
, Luh K
Received 7 January 2015
Accepted for publication 14 February 2015
Published 7 May 2015 Volume 2015:5 Pages 13—19
DOI https://doi.org/10.2147/CBF.S41651
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 4
Editor who approved publication: Dr Hung Khong
Chin-Chung Shu,1,2 Jann-Yuan Wang,2 Li-Na Lee,2,3 Chong-Jen Yu,2 Kwen-Tay Luh3
1Department of Traumatology, 2Department of Internal Medicine, 3Department of Laboratory Medicine, National Taiwan University Hospital, Taipei, Taiwan
Abstract: Although many laboratory methods have been developed to expedite the diagnosis of active tuberculosis (TB) and Mycobacterium tuberculosis (Mtb) infection, delays in diagnosis remain a major problem in clinical practice. Biomarkers may contribute favorably or unfavorably to TB diagnosis in a clinical suspect TB case with inconclusive diagnostic findings. A good understanding of the effectiveness and practical limitations of these biomarkers is important to improve diagnosis. This review summarizes currently used biomarkers, mainly as validation, and focuses on latent TB infection, active pulmonary TB, and tuberculous pleural effusion.
Keywords: tuberculosis, biomarker, diagnosis, latent tuberculosis infection, pleural effusion
Introduction
Tuberculosis (TB) remains one of the most common infectious diseases and a global health problem.1 The World Health Organization estimates nine million new cases and 1.5 million deaths from TB in 2013.1 Although implementing directly observed treatment short-course can rapidly decrease the infectivity of patients with active TB, the disease is still transmissible before treatment. Moreover, delayed diagnosis and treatment worsen the outcomes.2 At the same time, TB diagnosis is not always easy, especially in the early stages, because of variable and nonspecific presentations. In patients with smear-negative TB, early detection is even more difficult because their clinical samples are paucibacillary. The problem of smear-negative TB, as well as smear-positive TB, warrants particular attention because patients with smear-negative TB have been reported to be responsible for about 17% of TB transmission.3,4
As regards advances in laboratory measures, the rapid isolation of Mycobacterium tuberculosis (Mtb) from clinical specimens is a milestone in shortening the diagnosis waiting time. Previously, mycobacterial culture, the gold standard of TB diagnosis, was available only after 4–8 weeks with solid culture medium.5 By applying the fluorometric BACTEC technique (BACTEC Mycobacteria Growth Indicator Tube [MGIT] 960 system, Becton-Dickinson, Franklin Lakes, NJ, USA), the results of mycobacterial culture can be made available earlier but still need 1–3 weeks.6 Moreover, this automatic system can detect 960 samples simultaneously and greatly reduce routine laboratory work.
However, isolates of both Mtb and nontuberculous mycobacteria (NTM) can be culture-positive for Mycobacteria in the MGIT system. This emphasizes that a fluorescence signal detected and reported by the MGIT 960 system only implies that some microorganisms are multiplying and consuming oxygen in the culture tube. Thus, when the clinical sample of a patient is culture-positive by the fluorometric MGIT 960 system, careful clinical judgment and further identification of the Mtb antigen or nucleic acid amplification test (NAAT) should be considered. For patients suspected of a diagnosis other than TB and not in urgent need of treatment, anti-TB treatment should be deferred until TB is confirmed by conventional identification or other molecular assays.
Another milestone is the rapid detection of Mtb from clinical specimens. Deoxyribonucleic acid, ribonucleic acid, and Mtb protein can be used as targets for detection. Because the number of Mtb bacilli in any clinical specimen is usually much lower than the threshold of any detection method, amplification should always be the first step for rapid and direct detection. As such, a rapid detection method using proteins as the target can only be applied to an enriched sample, such as a cultured liquid medium. For example, Capilia TB assay (TAUNS, Numazu, Japan), an immuno-chromatographic assay detecting MPB64 (a secreted mycobacterial protein only found in Mtb isolates and some strains of Mycobacterium bovis BCG [Bacillus Calmette–Guérin]), is an easy and rapid diagnostic tool for culture confirmation of Mtb in liquid medium, with positive and negative predictive values of 98.6% and 97.9%, respectively.7
At present, NAAT is the main means of rapid Mtb detection from clinical specimens. In general, NAATs have high positive and negative predictive values for smear-positive respiratory specimens. Thus, they are frequently used to confirm or invalidate TB diagnosis. For patients with smear-negative respiratory specimens, NAATs are used only if pulmonary TB is highly suspected. For sputum screening, the multiplex nested polymerase chain reaction for detecting the Mtb gene is used by combining with immuno-chromatography test for convenience, with 98% specificity.8 Furthermore, the Xpert MTB/RIF (Cepheid, Sunnyvale, CA, USA) non-laboratory-based molecular assay has the potential to improve the diagnosis of TB, especially in HIV-infected or smear-negative but culture-positive pulmonary TB.9 In clinical practice, the use of Xpert reportedly reduces the frequency and impact of unnecessary empirical treatment.10 Unfortunately, NAATs for Mtb have not been approved for use in specimens other than respiratory samples. Moreover, the performance of NAAT can be compromised under several clinical conditions including inadequate specimens, laboratory contamination, inappropriate processing, and the presence of an amplification inhibitor.11–14
Aside from diagnostic advances in the rapid isolation and detection of Mtb, other diagnostic aids for the early identification of TB are needed in view of the aforementioned limitation of current diagnostic tools, and TB can be a life-threatening disease. For improving TB diagnosis, the role of biomarkers is to provide early suspicion, especially while waiting for the isolation of Mtb or for negative findings of TB suspects (Figure 1). Biomarkers represent human response to Mtb bacilli and thereby alert clinicians for further workup of TB in clinical practice. However, the diagnostic biomarkers cannot stand alone, and the final decision of TB treatment must be based on the patient’s immune status, the probability of infection, and the urgency to administer the treatment. For a better understanding of the clinical application of the biomarkers, this review focuses on the usefulness and the limitation of biomarkers with regard to latent TB infection (LTBI), active pulmonary TB, and tuberculous pleural effusion (Figure 2).
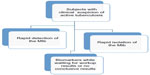  | Figure 1 The role of biomarkers in practice for the clinical suspicion of active pulmonary tuberculosis. |
Biomarkers for latent TB infection
To prevent further TB transmission, control should focus on LTBI.15 Previous studies demonstrated that about 10% of LTBI patients develop the active disease and the risk is even higher in immunocompromised hosts.16 In humans, the T-cell response is the most important defense mechanism against Mtb, and it develops in the first few weeks after the infection. At present, tuberculin skin test and interferon-γ (IFN-γ) release assays (IGRAs) are the most common tests for diagnosing LTBI.17
IGRA is the T-cell response assayed using a purified protein derivative or other Mtb-specific antigens.18 A positive result indicates Mtb infection rather than an active disease, although the former is always the first step toward the latter. Clinical information, radiographic findings, and results of laboratory studies should all be used to judge whether the presence of T-cell response is due to active disease or merely Mtb infection.
In the last decade, two in vitro T-cell-based IGRAs have been developed and commercialized for detecting the host response to Mtb.19 After culturing peripheral blood mononuclear cells with Mtb-specific antigens, the IFN-γ produced by activated T cells is measured by enzyme-linked immunosorbent assay in QuantiFERON-TB Gold-in-tube (QFT-GIT) (Cellestis, Carnegie, Australia) and by the enzyme-linked immuno-spotting technique in T-SPOT TB (Oxford Immunotec, Oxford, UK). Both assays require only one visit and are free of a “booster effect” during repeat testing. The turnaround time is short, with results available the next day.
IGRAs use the two Mtb-specific antigens, early secretory antigenic target-6 kD and culture filtrate protein 10, which are encoded by the region of difference 1 (RD1) of the Mtb genome.20,21 Currently, all available BCG vaccine strains do not have RD1, which they have lost since 1908 during passage.22 This region is also absent in NTM except in Mycobacterium kansasii, Mycobacterium marinum, and Mycobacterium szulgai. Because the antigens used in IGRA are more specific than purified protein derivative, cross-reactivity (leading to false positive) from NTM and BCG is low. Although IGRA is promising, it cannot replace the tuberculin skin test in current perspectives, especially for non-BCG-vaccinated populations.23
Because there is no gold standard for the diagnosis of LTBI, comparing the performance of IGRAs with that of the tuberculin skin test is difficult. Usually, sensitivity is estimated from studies of patients with active TB or via correlation between the proportion of the contact persons with positive immune response and the gradients of exposure. Recent meta-analysis studies demonstrate that T-SPOT TB is slightly more sensitive than QFT-GIT and tuberculin skin test.23,24 In terms of specificity, the two IGRAs are significantly better than the tuberculin skin test, especially in BCG-vaccinated subjects.
The most important limitation of IGRA is that it does not differentiate active TB from mere Mtb infection. A positive result of IGRA means that the patient is very likely to have Mtb infection, but this does not mean that the pulmonary lesion is due to active TB. On the other hand, a negative IGRA result cannot totally exclude the possibility of active TB in a clinical suspect. This is because active TB itself can cause immunosuppression,12 which in turn can give a false-negative result in the immunoassays.
Evidence so far suggests that it is better to apply IGRA only in contact surveillance and for the diagnosis of latent infection. However, its use in other risk populations is still debatable.25–27 For example, in health care workers with positive IGRA, the clinical significance is questionable because there is a high proportion of IGRA reversion (from positive to negative results) beyond the 8% of interexperiment variation.10,12,28 Similarly, the reversion rate is around 45.9% within 6 months in the dialysis population.27 Although the correlation between reversion and occurrence of active TB in the long run is unclear, the persistent LTBI status by IGRA may be a surrogate and is therefore important for further management.29
In a recent report that serially followed up 204 dialysis patients for 1 year, the QFT-GIT-positive population is heterogeneous, and subpopulations have different reversion rates.27 The QFT-GIT response, difference in IFN-γ level between the TB-antigen tube and negative control, is an independent factor for persistent QFT-GIT positivity.27 Furthermore, higher IFN-γ and interleukin (IL)-10 are also significantly associated with persistent LTBI status, diagnosed by QFT-GIT (Shu et al, unpublished data, 2015). Those with high titer of QFT-GIT, together with high serum IL-10 and IFN-γ, will be the target population for surveillance or preventive therapy.
Biomarkers for pulmonary TB
Blood biomarkers
A diagnosis of TB should be kept in mind when a clinician manages patients with pneumonia, especially in intermediate or high-prevalence TB areas. In general, the levels of serum inflammatory markers are lower in patients with pulmonary TB than in patients with community-acquired pneumonia. The markers reported include procalcitonin (PCT), IL-10, and C-reactive protein.30–32 Although the levels of serum inflammatory markers are significantly different between pulmonary TB and pneumonia, the low levels of inflammatory markers are nonspecific in clinical practice. Thus, an integrated model that includes LTBI testing, inflammatory markers, and clinical or radiographic findings may be better than a single test.
Because patients with LTBI will be a risk population for the development of active TB,33,34 biomarker measurements under a positive LTBI status make more sense. In the pathogenesis of TB, macrophages are the first line of defense as the TB bacilli enter the airways.35 An important mechanism for persistent infection in human tissue is via necrosis of Mtb-infected macrophages as the dominant form of cell death instead of apoptosis,36,37 because apoptosis of infected macrophages will result in the removal of intracellular bacilli and more efficient induction of inflammation than necrosis.38 For apoptosis-associated biomarkers, a recent study shows that decoy receptor (DcR) 3 and prostaglandin E2 (PGE2) levels increase from the LTBI group defined by IGRA to the TB group, but lipoxin decreases from LTBI to the TB group.39 In the extended survey, Fas ligand (FasL), an initiator of extrinsic pathway of apoptosis, is decreased from LTBI to TB status, like lipoxin (Shu et al, unpublished data).
DcR3 is a receptor of the tumor necrosis factor (TNF) receptor superfamily, existing as a soluble receptor for FasL, and is considered an immunomodulator.40 The two lipid markers, PGE2 and lipoxin, antagonize macrophage apoptosis in Mtb infection.37,41 Therefore, apoptosis-associated serum biomarkers change along with the status of Mtb infection and are independent factors discriminating active TB from LTBI. Aggressive mycobacteriologic study to detect active TB is necessary for IGRA-positive subjects who have high DcR3 and PGE2, but low lipoxin and FasL.
Previous investigation of T-cell responses in Mtb infection shows that the presence of single-positive TNF-α Mtb-specific CD4+ T cells is a strong predictor of diagnosis of active disease versus latent infection.42 A validation study shows that the sensitivity and specificity of the flow cytometry-based assay are 67% and 92%, respectively, with positive and negative predictive values of 80% and 92.4%, respectively. As such, the proportion of single-positive TNF-α Mtb-specific CD4+ T cells may be a new reference for the rapid diagnosis of active TB.43 Other studies also show similar findings and suggest combined measurement of T-cell phenotype and function as a highly discriminatory biomarker of TB disease activity. This combined approach requires validation in large-scale prospective studies.42
Otherwise, serum cytokines like IL-2, IL-9, IL-13, IL-17, and TNF-α have been reported as adjunctive biomarkers for active TB from LTBI and NTM disease.44,45 Whole blood cells stimulated by Mtb antigen have higher TNF-α production and IL-12 in TB patients compared with LTBI controls.46 Again, these markers warrant further validation before clinical application.32
In regard to serologic tests for antibodies for Mtb infection, a number of commercial antibody-based TB diagnostic tests have been developed and are on sale, although clinical validation is usually absent and current test performance is inconsistent.47
Sputum biomarkers
Unlike serum biomarkers, which are easily influenced by other systemic inflammation or infection, chemicals or materials in the sputum are more direct from the disease site in pulmonary TB. An attractive test for urease in respiratory specimens has been investigated. The point of urease-based diagnostics is that humans lack urease enzymes.48 Since 2009, a group of investigators have started studying the benefits of rabbit urease breath test for TB diagnosis and treatment monitoring.48 The specificity of the urease breath test may be increased by introducing intravenous or inhaled 13C-urea tracer, thus preventing gastric contaminations by Helicobacter species.
In other reports, pulmonary TB may alter volatile organic compounds (VOCs) in breath because Mycobacteria and oxidative stress due to mycobacterial infection both generate distinctive VOCs.49 The validation analysis identifies patients with positive sputum culture with 82.6% sensitivity and 100% specificity.49 On the other hand, TB stearic acid (TBSA) is reported to be another marker, and the literature shows that TBSA exists in all sputum specimens from TB patients, but is not found in sputum samples from persons without TB.50 The possibility of using VOCs and TBSA in sputum examination also required large-scale validation.
Urine biomarkers
Urine represents a clinical sample that is easy to collect from both adults and children. It has been used extensively to evaluate several antigen and DNA detection assays.51 Commercially available assays can detect lipoarabinomannan (LAM) in the urine of TB patients. Although the sensitivity of this test is disappointing in non-HIV-infected patients, moderate sensitivity and high specificity are observed in HIV-infected patients with advanced immunodeficiency.52,53 Although the sensitivity of many TB diagnostic tests declines in HIV-infected patients with advanced immunodeficiency, the sensitivity of urine LAM test paradoxically increases, whereas CD4 lymphocyte counts decrease.54 Currently, a low cost and point-of-care TB diagnostic test in HIV patients is now being developed with urine antigen detection.55
Biomarkers for tuberculous pleural effusion
Tuberculous PE accounts for approximately 5% of all forms of TB and is the second most common extrapulmonary TB.56 Early diagnosis is often difficult because the mycobacterial culture of PE or pleural tissue requires weeks to obtain.57,58 Moreover, only 40%–63% of tuberculous PE cases have positive mycobacterial culture from PE.58–60 Thus, treatment can be delayed, resulting in increased mortality.61
Biomarkers of immune response in PE have been proposed to assist in the rapid diagnosis of tuberculous PE while managing patients with PE of unknown etiology or while waiting for the results of mycobacterial culture. Pleural IFN-γ 62,63 and IFN-induced protein 10 (IP-10),64 as well as adenosine deaminase (ADA),65,66 have been reported to be useful markers for diagnosing tuberculous PE. Moreover, soluble FasL is released from T lymphocytes and exerts cytotoxic effects on Mtb-hidden macrophages, thereby helping in the diagnosis of tuberculous PE.67,68
PCT, both serum and pleural, is lower in patients with tuberculous PE than in those with parapneumonic effusion. Measurement of serum and pleural PCT is useful in differentiating parapneumonic effusion from tuberculous PE.69 In a recent study,70 logistic regression analysis revealed that INF-γ ≥75 pg/mL, ADA ≥40 IU/mL, DcR3 ≥9.3 ng/mL, and soluble TNF receptor 1 (TNF-sR1) ≥3.2 ng/mL were independent factors associated with tuberculous PE. The predicted probability based on the four predictors had an area under the receiver operating characteristic curve of 0.920, with 82.9% sensitivity and 86.7% specificity. The four-marker model slightly reduced the specificity compared with the single test of pleural IFN-γ or ADA, but much elevated the sensitivity of diagnosis for tuberculous PE. For the purpose of avoiding delayed diagnosis, the four-factor model is suggested.
Conclusion
The rapid isolation and detection of Mtb have improved the diagnosis of TB in recent decades. However, delayed diagnosis remains frequent because the isolation of Mtb still requires 1–3 weeks, and rapid detection is mainly useful in smear-positive cases. Biomarkers play an important role in this situation and improve clinical practice by helping decision-making. T-cell responses to Mtb like IGRAs are good diagnostic tools to define Mtb infection, which is the high-risk group in future TB reactivation. The use of biomarkers in patients with Mtb infection may further identify more specific populations who have current active TB disease or who will develop active TB in the future. For helping TB diagnosis, integrating biomarkers into clinical practice is suggested for a clinical TB suspect with negative preliminary workup and pending mycobacterial culture.
Acknowledgments
The authors thank the staff of the Eighth Core Lab of the Department of Medical Research of National Taiwan University Hospital for their technical support. The authors also thank the Department of Medical Research of National Taiwan University Hospital.
Funding sources
This study was funded by grants from the Research Center for Biotechnology and Medicine Policy in Taiwan, the Center for Disease Control, Ministry of Health and Welfare, Taiwan (MOHW103-CDC-C-114-112302), and the Ministry of Science and Technology, Taiwan (103-2325-B-002-005 and 103-2314-B-002-152-MY2). The funders had no role in the study design, data collection and analysis, decision to publish, and preparation of the manuscript.
Disclosure
The authors declare no financial, professional, or other personal conflict of interests of any nature or kind to any related product, service, and/or company.
References
World Health Organization. Global Tuberculosis Report 2014. Geneva, Switzerland: World Health Organization; 2014. | |
Wang JY, Hsueh PR, Jan IS, et al. Empirical treatment with a fluoroquinolone delays the treatment for tuberculosis and is associated with a poor prognosis in endemic areas. Thorax. 2006;61:903–908. | |
Behr MA, Warren SA, Salamon H, et al. Transmission of Mycobacterium tuberculosis from patients smear-negative for acid-fast bacilli. Lancet. 1999;353:444–449. | |
Hernandez-Garduno E, Cook V, Kunimoto D, Elwood RK, Black WA, FitzGerald JM. Transmission of tuberculosis from smear negative patients: a molecular epidemiology study. Thorax. 2004;59:286–290. | |
Somoskovi A, Kodmon C, Lantos A, et al. Comparison of recoveries of mycobacterium tuberculosis using the automated BACTEC MGIT 960 system, the BACTEC 460 TB system, and Lowenstein–Jensen medium. J Clin Microbiol. 2000;38:2395–2397. | |
Kuo SHS, Shi WY, Lin T, et al. Taiwan Tuberculosis Control Report 2009. 4th ed. Taipei, Taiwan (ROC): Department of Health, Centers for Disease Control; 2009. | |
Wang JY, Lee LN, Lai HC, et al. Performance assessment of the Capilia TB assay and the BD ProbeTec ET system for rapid culture confirmation of Mycobacterium tuberculosis. Diagn Microbiol Infect Dis. 2007;59:395–399. | |
Soo PC, Horng YT, Hsueh PR, et al. Direct and simultaneous identification of Mycobacterium tuberculosis complex (MTBC) and Mycobacterium tuberculosis (MTB) by rapid multiplex nested PCR-ICT assay. J Microbiol Methods. 2006;66:440–448. | |
Scott LE, McCarthy K, Gous N, et al. Comparison of Xpert MTB/RIF with other nucleic acid technologies for diagnosing pulmonary tuberculosis in a high HIV prevalence setting: a prospective study. PLoS Med. 2011;8:e1001061. | |
Davis JL, Kawamura LM, Chaisson LH, et al. Impact of GeneXpert MTB/RIF on patients and tuberculosis programs in a low-burden setting. A hypothetical trial. Am J Respir Crit Care Med. 2014;189: 1551–1559. | |
Richeldi L, Barnini S, Saltini C. Molecular diagnosis of tuberculosis. Eur Respir J Suppl. 1995;20:689s–700s. | |
Lopez B, Aguilar D, Orozco H, et al. A marked difference in pathogenesis and immune response induced by different Mycobacterium tuberculosis genotypes. Clin Exp Immunol. 2003;133:30–37. | |
Wang SX, Sng LH, Tay L. Preliminary study on rapid identification of Mycobacterium tuberculosis complex isolates by the BD ProbeTec ET system. J Med Microbiol. 2004;53:57–59. | |
Wang JY, Lee LN, Hsu HL, Hsueh PR, Luh KT. Performance assessment of the DR. MTBC screen assay and the BD ProbeTec ET system for direct detection of Mycobacterium tuberculosis in respiratory specimens. J Clin Microbiol. 2006;44:716–719. | |
Rose DN. Benefits of screening for latent Mycobacterium tuberculosis infection. Arch Intern Med. 2000;160:1513–1521. | |
Horsburgh CR Jr. Priorities for the treatment of latent tuberculosis infection in the United States. N Engl J Med. 2004;350:2060–2067. | |
Diel R, Loddenkemper R, Meywald-Walter K, Niemann S, Nienhaus A. Predictive value of a whole blood IFN-gamma assay for the development of active tuberculosis disease after recent infection with Mycobacterium tuberculosis. Am J Respir Crit Care Med. 2008;177: 1164–1170. | |
Boussiotis VA, Tsai EY, Yunis EJ, et al. IL-10-producing T cells suppress immune responses in anergic tuberculosis patients. J Clin Invest. 2000;105:1317–1325. | |
Pai M, Riley LW, Colford JM Jr. Interferon-gamma assays in the immunodiagnosis of tuberculosis: a systematic review. Lancet Infect Dis. 2004;4:761–776. | |
Berthet FX, Rasmussen PB, Rosenkrands I, Andersen P, Gicquel B. A Mycobacterium tuberculosis operon encoding ESAT-6 and a novel low-molecular-mass culture filtrate protein (CFP-10). Microbiology. 1998;144 (Pt 11):3195–3203. | |
Sorensen AL, Nagai S, Houen G, Andersen P, Andersen AB. Purification and characterization of a low-molecular-mass T-cell antigen secreted by Mycobacterium tuberculosis. Infect Immun. 1995;63:1710–1717. | |
Mahairas GG, Sabo PJ, Hickey MJ, Singh DC, Stover CK. Molecular analysis of genetic differences between Mycobacterium bovis BCG and virulent M. bovis. J Bacteriol. 1996;178:1274–1282. | |
Mazurek GH, Jereb J, Vernon A, et al. Updated guidelines for using Interferon Gamma Release Assays to detect Mycobacterium tuberculosis infection – United States, 2010. MMWR Recomm Rep. 2010;59: 1–25. | |
Menzies D, Pai M, Comstock G. Meta-analysis: new tests for the diagnosis of latent tuberculosis infection: areas of uncertainty and recommendations for research. Ann Intern Med. 2007;146:340–354. | |
Ringshausen FC, Nienhaus A, Schablon A, Schlosser S, Schultze-Werninghaus G, Rohde G. Predictors of persistently positive Mycobacterium-tuberculosis-specific interferon-gamma responses in the serial testing of health care workers. BMC Infect Dis. 2010;10:220. | |
Dyrhol-Riise AM, Gran G, Wentzel-Larsen T, Blomberg B, Haanshuus CG, Morkve O. Diagnosis and follow-up of treatment of latent tuberculosis; the utility of the QuantiFERON-TB Gold In-tube assay in outpatients from a tuberculosis low-endemic country. BMC Infect Dis. 2010;10:57. | |
Shu CC, Wu VC, Yang FJ, et al. Dynamic changes in positive interferon-gamma release assay in a dialysis population: an observational cohort study. J Infect. 2013;67:529–535. | |
Metcalfe JZ, Cattamanchi A, McCulloch CE, Lew JD, Ha NP, Graviss EA. Test variability of the QuantiFERON-TB gold in-tube assay in clinical practice. Am J Respir Crit Care Med. 2013;187:206–211. | |
Fong KS, Tomford JW, Teixeira L, et al. Challenges of interferon-gamma release assay conversions in serial testing of health-care workers in a TB control program. Chest. 2012;142:55–62. | |
Niu WY, Wan YG, Li MY, Wu ZX, Zhang LG, Wang JX. The diagnostic value of serum procalcitonin, IL-10 and C-reactive protein in community acquired pneumonia and tuberculosis. Eur Rev Med Pharmacol Sci. 2013;17:3329–3333. | |
Kang YA, Kwon SY, Yoon HI, Lee JH, Lee CT. Role of C-reactive protein and procalcitonin in differentiation of tuberculosis from bacterial community acquired pneumonia. Korean J Intern Med. 2009;24:337–342. | |
Walzl G, Ronacher K, Hanekom W, Scriba TJ, Zumla A. Immunological biomarkers of tuberculosis. Nat Rev Immunol. 2011;11:343–354. | |
Harries A. What is the additional yield from repeated sputum examinations by microscopy and culture? In: Frieden TR, editor. Toman’s Tuberculosis Case Detection, Treatment and Monitoring. Geneva, Switzerland: World Health Organization; 2004:46–50. | |
Yang WC, Hwang SJ. Incidence, prevalence and mortality trends of dialysis end-stage renal disease in Taiwan from 1990 to 2001: the impact of national health insurance. Nephrol Dial Transplant. 2008;23:3977–3982. | |
Pai M, Joshi R, Dogra S, et al. Serial testing of health care workers for tuberculosis using interferon-gamma assay. Am J Respir Crit Care Med. 2006;174:349–355. | |
Newberry WM, Sanford JP. Defective cellular immunity in renal failure: depression of reactivity of lymphocytes to phytohemagglutinin by renal failure serum. J Clin Invest. 1971;50:1262–1271. | |
Papay P, Eser A, Winkler S, et al. Predictors of indeterminate IFN-gamma release assay in screening for latent TB in inflammatory bowel diseases. Eur J Clin Invest. 2011;41:1071–1076. | |
Targeted tuberculin testing and treatment of latent tuberculosis infection. American Thoracic Society. MMWR Recomm Rep. 2000;49:1–51. | |
Shu CC, Wu MF, Hsu CL, et al. Apoptosis-associated biomarkers in tuberculosis: promising for diagnosis and prognosis prediction. BMC Infect Dis. 2013;13:45. | |
Lange B, Vavra M, Kern WV, Wagner D. Indeterminate results of a tuberculosis-specific interferon-gamma release assay in immunocompromised patients. Eur Respir J. 2010;35:1179–1182. | |
Shu CC, Wang JT, Lee CH, Wang JY, Lee LN, Yu CJ. Predicting results of mycobacterial culture on sputum smear reversion after anti-tuberculous treatment: a case control study. BMC Infect Dis. 2010;10:48. | |
Pollock KM, Whitworth HS, Montamat-Sicotte DJ, et al. T-cell immunophenotyping distinguishes active from latent tuberculosis. J Infect Dis. 2013;208:952–968. | |
Harari A, Rozot V, Bellutti Enders F, et al. Dominant TNF-alpha+ Mycobacterium tuberculosis-specific CD4+ T cell responses discriminate between latent infection and active disease. Nat Med. 2011;17: 372–376. | |
Hur YG, Kang YA, Jang SH, et al. Adjunctive biomarkers for improving diagnosis of tuberculosis and monitoring therapeutic effects. J Infect. Epub November 5, 2014. | |
Chegou NN, Heyckendorf J, Walzl G, Lange C, Ruhwald M. Beyond the IFN-gamma horizon: biomarkers for immunodiagnosis of infection with Mycobacterium tuberculosis. Eur Respir J. 2014;43: 1472–1486. | |
Sutherland JS, de Jong BC, Jeffries DJ, Adetifa IM, Ota MO. Production of TNF-alpha, IL-12(p40) and IL-17 can discriminate between active TB disease and latent infection in a West African cohort. PloS One. 2010;5:e12365. | |
Steingart KR, Flores LL, Dendukuri N, et al. Commercial serological tests for the diagnosis of active pulmonary and extrapulmonary tuberculosis: an updated systematic review and meta-analysis. PLoS Med. 2011;8:e1001062. | |
Habel JE, Bursey EH, Rho BS, et al. Structure of Rv1848 (UreA), the Mycobacterium tuberculosis urease gamma subunit. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2010;66:781–786. | |
Phillips M, Cataneo RN, Condos R, et al. Volatile biomarkers of pulmonary tuberculosis in the breath. Tuberculosis. 2007;87:44–52. | |
Cha D, Cheng D, Liu M, Zeng Z, Hu X, Guan W. Analysis of fatty acids in sputum from patients with pulmonary tuberculosis using gas chromatography-mass spectrometry preceded by solid-phase microextraction and post-derivatization on the fiber. J Chromatogr A. 2009;1216:1450–1457. | |
Green C, Huggett JF, Talbot E, Mwaba P, Reither K, Zumla AI. Rapid diagnosis of tuberculosis through the detection of mycobacterial DNA in urine by nucleic acid amplification methods. Lancet Infect Dis. 2009;9:505–511. | |
Minion J, Leung E, Talbot E, Dheda K, Pai M, Menzies D. Diagnosing tuberculosis with urine lipoarabinomannan: systematic review and meta-analysis. Eur Respir J. 2011;38:1398–1405. | |
Gounder CR, Kufa T, Wada NI, et al. Diagnostic accuracy of a urine lipoarabinomannan enzyme-linked immunosorbent assay for screening ambulatory HIV-infected persons for tuberculosis. J Acquir Immune Defic Syndr. 2011;58:219–223. | |
Lawn SD, Edwards DJ, Kranzer K, Vogt M, Bekker LG, Wood R. Urine lipoarabinomannan assay for tuberculosis screening before antiretroviral therapy diagnostic yield and association with immune reconstitution disease. AIDS. 2009;23:1875–1880. | |
Lawn SD, Kerkhoff AD, Vogt M, Wood R. Diagnostic accuracy of a low-cost, urine antigen, point-of-care screening assay for HIV-associated pulmonary tuberculosis before antiretroviral therapy: a descriptive study. Lancet Infect Dis. 2012;12:201–209. | |
Sharma SK, Mohan A. Extrapulmonary tuberculosis. Indian J Med Res. 2004;120:316–353. | |
Luh KT, Chiang CY, Lee JJ, et al. Taiwan Guidelines for TB Diagnosis and Treatment. 4th ed. Taipei, Taiwan (ROC): Center for Disease Control, Executive Yuan; 2011. | |
Light RW. Update on tuberculous pleural effusion. Respirology. 2010;15:451–458. | |
Gopi A, Madhavan SM, Sharma SK, Sahn SA. Diagnosis and treatment of tuberculous pleural effusion in 2006. Chest. 2007;131:880–889. | |
Ruan SY, Chuang YC, Wang JY, et al. Revisiting tuberculous pleurisy: pleural fluid characteristics and diagnostic yield of mycobacterial culture in an endemic area. Thorax. 2012;67:822–827. | |
Lin MT, Wang JY, Yu CJ, Lee LN, Yang PC. Mycobacterium tuberculosis and polymorphonuclear pleural effusion: incidence and clinical pointers. Respir Med. 2009;103:820–826. | |
Keng LT, Shu CC, Chen JY, et al. Evaluating pleural ADA, ADA2, IFN-gamma and IGRA for diagnosing tuberculous pleurisy. J Infect. 2013;67:294–302. | |
Liu YC, Shin-Jung Lee S, Chen YS, Tu HZ, Chen BC, Huang TS. Differential diagnosis of tuberculous and malignant pleurisy using pleural fluid adenosine deaminase and interferon gamma in Taiwan. J Microbiol Immunol Infect. 2011;44:88–94. | |
Dheda K, Van-Zyl Smit RN, Sechi LA, et al. Clinical diagnostic utility of IP-10 and LAM antigen levels for the diagnosis of tuberculous pleural effusions in a high burden setting. PloS One. 2009;4:e4689. | |
Khan FY, Hamza M, Omran AH, et al. Diagnostic value of pleural fluid interferon-gamma and adenosine deaminase in patients with pleural tuberculosis in Qatar. Int J Gen Med. 2013;6:13–18. | |
Garcia-Zamalloa A, Taboada-Gomez J. Diagnostic accuracy of adenosine deaminase and lymphocyte proportion in pleural fluid for tuberculous pleurisy in different prevalence scenarios. PloS One. 2012;7:e38729. | |
Budak F, Uzaslan EK, Cangur S, Goral G, Oral HB. Increased pleural soluble fas ligand (sFasL) levels in tuberculosis pleurisy and its relation with T-helper type 1 cytokines. Lung. 2008;186:337–343. | |
Wu SH, Li CT, Lin CH, Chu JJ, Cheng ML, Lin KH. Soluble Fas ligand is another good diagnostic marker for tuberculous pleurisy. Diagn Microbiol Infect Dis. 2010;68:395–400. | |
Lee SH, Lee EJ, Min KH, et al. Procalcitonin as a diagnostic marker in differentiating parapneumonic effusion from tuberculous pleurisy or malignant effusion. Clin Biochem. 2013;46:1484–1488. | |
Shu CC, Wang JY, Hsu CL, et al. Diagnostic role of inflammatory and anti-inflammatory cytokines and effector molecules of cytotoxic T lymphocytes in tuberculous pleural effusion. Respirology. 2015;20:147–154. |
 © 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms
and incorporate the Creative Commons Attribution
- Non Commercial (unported, 3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms
and incorporate the Creative Commons Attribution
- Non Commercial (unported, 3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.

