Back to Journals » Breast Cancer: Targets and Therapy » Volume 8
Identification and prioritization of candidate genes for symptom variability in breast cancer survivors based on disease characteristics at the cellular level
Received 12 May 2015
Accepted for publication 20 July 2015
Published 8 March 2016 Volume 2016:8 Pages 29—37
DOI https://doi.org/10.2147/BCTT.S88434
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Professor Pranela Rameshwar
Theresa A Koleck,1 Yvette P Conley2
1School of Nursing, 2Department of Human Genetics, School of Nursing and Graduate School of Public Health, University of Pittsburgh, Pittsburgh, PA, USA
Abstract: Research is beginning to suggest that the presence and/or severity of symptoms reported by breast cancer survivors may be associated with disease-related factors of cancer. In this article, we present a novel approach to the identification and prioritization of biologically plausible candidate genes to investigate relationships between genomic variation and symptom variability in breast cancer survivors. Cognitive dysfunction is utilized as a representative breast cancer survivor symptom to elucidate the conceptualization of and justification for our cellular, disease-based approach to address symptom variability in cancer survivors. Initial candidate gene identification was based on genes evaluated as part of multigene expression profiles for breast cancer, which are commonly used in the clinical setting to characterize the biology of cancer cells for the purpose of describing overall tumor aggressiveness, prognostication, and individualization of therapy. A list of genes evaluated within five multigene expression profiles for breast cancer was compiled. In order to prioritize candidate genes for investigation, genes used in each profile were compared for duplication. Twenty-one genes (BAG1, BCL2, BIRC5, CCNB1, CENPA, CMC2, DIAPH3, ERBB2, ESR1, GRB7, MELK, MKI67, MMP11, MYBL2, NDC80, ORC6, PGR, RACGAP1, RFC4, RRM2, and SCUBE2) are utilized in two or more profiles, including five genes (CCNB1, CENPA, MELK, MYBL2, and ORC6) used in three profiles. To ensure that the parsimonious 21 gene set is representative of the more global biological hallmarks of cancer, an Ingenuity Pathway Analysis was conducted. Evaluation of genes known to impact pathways involved with cancer development and progression provide a means to evaluate the overlap between the biological underpinnings of cancer and symptom development within the context of cancer.
Keywords: breast neoplasms, biological markers, genes, signs and symptoms, cognition
Introduction
Many cancer survivors experience a variety of disruptive and burdensome symptoms, including fatigue, pain, altered sleep, mood dysregulation, and cognitive dysfunction long into survivorship.1–3 Although our ability to describe the duration, frequency, and severity of symptoms related to cancer and cancer treatments has vastly improved, our understanding of the mechanisms that influence symptom variability and our ability to personalize symptom prediction for an individual cancer survivor and intervene effectively remain limited. While still in its infancy, research is beginning to suggest that the presence and/or severity of symptoms reported by cancer survivors may not be solely the result of cancer treatments, but may be associated with disease-related factors of cancer and/or host characteristics that predispose an individual to cancer as well as a particular symptom.4–6
Breast cancer survivors have been the focus of a large proportion of cancer survivor symptom research. Studies conducted with breast cancer survivors on the symptom of cognitive dysfunction especially, are contributing an increasing amount of evidence in support of our hypothesis that disease-related factors of cancer and/or host characteristics that influence cancer development or progression contribute to the presence and severity of symptoms experienced by cancer survivors.
Cognitive dysfunction research in breast cancer survivors has traditionally concentrated on the direct neurotoxic effect of chemotherapeutic agents on the brain. Often referred to as “chemo brain” or “chemo fog”, short- and long-term cognitive changes have been well-documented in women with breast cancer receiving chemotherapy.7 With the increasing use of antiestrogen therapies for prevention of breast cancer recurrence, the influence of estrogen and the use of antiestrogen therapies on cognitive decline in breast cancer survivors have become an additional focus of research on treatment-related cognitive changes.8–14
However, more recent research demonstrates that cognitive changes may actually occur in breast cancer survivors prior to initiation of adjuvant chemotherapy and/or antiestrogen therapy. In a study of 109 women with breast cancer scheduled to receive chemotherapy, Hermelink et al15 found that group mean scores were significantly poorer than test norms on five out of twelve cognitive tests before the start of treatment. In addition, 33 survivors scored in the lower fifth percentile of test norms on two or more cognitive tests unrelated to depression, anxiety, or self-reported cognitive problems. Similarly, Wefel et al16 reported that 29 out of 84 breast cancer survivors diagnosed with stage 1–3a breast cancer were classified as cognitively impaired (ie, multiple cognitive tests with z-scores ≤-1.5 or a single test with z-score ≤-2.0) before receiving chemotherapy compared to normative data.
Even more compelling are findings from studies comparing the cognitive function of breast cancer survivors to healthy controls. Ahles et al17 found that women with invasive breast cancer had poorer overall cognitive performance compared to women with noninvasive breast cancer and healthy controls. Bender et al18 also reported pretreatment differences in cognitive function in the domains of verbal learning and memory and attention between women with breast cancer prescribed antiestrogen therapy with or without chemotherapy and healthy controls matched on age and years of education. Although not statistically significant, Schagen et al19 found that 16.4% of survivors prescribed chemotherapy and 29.8% of survivors with stage 1 breast cancer who were prescribed no systemic treatment displayed cognitive impairment before initiation of adjuvant treatment compared to 10% of healthy controls.
While multiple factors potentially predict cognitive function in women with breast cancer prior to adjuvant therapy, we hypothesize that these pretreatment findings suggest that disease-related factors inherent in breast cancer and/or host characteristics that predispose an individual to cancer as well as cognitive dysfunction may be a major determinant of cognitive changes in women with breast cancer. Additionally, only a subset of breast cancer survivors appears to be affected by cognitive dysfunction. We further hypothesize that heterogeneity in the biology of breast cancers at the cellular level could account for a significant proportion of reported discrepancies in cognitive function among survivors.
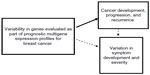
One common clinical tool used to evaluate the underlying biology of breast cancer cells is the prognostic multigene expression profile for breast cancer. These profiles enhance knowledge received from traditional tumor features and utilize predictive algorithms of tumor gene expression levels to individualize treatment through estimation of adjuvant therapy benefit and distant cancer recurrence risk. Thus, each prognostic multigene expression profile is comprised of genes that play an important role in breast cancer aggressiveness and progression, and, consequently, represent ideal candidates for a genetic association study exploring our hypotheses (Figure 1).
In this article, we specifically aim to present a novel approach, based on genes examined in prognostic multigene expression profiles for breast cancer, to the identification and prioritization of biologically plausible candidate genes for future investigations of the association between genetic variation and symptoms experienced by breast cancer survivors. We will focus on 1) characteristics of five different prognostic multigene expression profiles for breast cancer, 2) prioritization of candidate genes replicated in two or more profiles, 3) biological functions of our identified, parsimonious candidate gene set, and 4) a discussion of the potential expanded clinical utility of prognostic multigene expression profiles for breast cancer, and, more generally, cancer symptom prediction.
Methods
Selected breast-cancer-related prognostic multigene expression profiles
Prognostic multigene expression profiles use tumor gene expression levels to evaluate the underlying biology of breast cancer cells and predict long-term outcomes and potential benefit of additional adjuvant therapy. Several groups have developed prognostic multigene expression profiles for breast cancer: the eleven-gene expression signature (marketed as the Breast Cancer IndexSM by bioTheranostics, Inc., San Diego, CA, USA),28 the 14-gene prognostic expression signature (described in Tutt et al23), the 21-gene breast cancer assay (marketed as the Oncotype DX® Breast Cancer Assay by Genomic Health®, Inc., Redwood City, CA, USA),29,30 the 50-gene breast cancer prognostic gene signature assay (marketed as the Prosigna® Breast Cancer Prognostic Gene Signature Assay by NanoString® Technologies, Inc., Seattle, WA, USA) based on the PAM50 Breast Cancer Intrinsic Classifier,31 and the 70-gene breast cancer recurrence assay (marketed as the MammaPrint® 70 Gene Breast Cancer Recurrence Assay by Agendia®, Irvine, CA, USA).32,33
The number of cancer genes utilized in each profile varies greatly, ranging from 7 in the eleven-gene expression signature to 70 in the 70-gene breast cancer recurrence assay. All of the prognostic multigene expression profiles provide predictions of 5- and/or 10-year distant breast cancer recurrence risk, except the 14-gene prognostic expression signature, which is specifically intended for prediction of distant metastasis. With the exception of the 50-gene breast cancer prognostic gene signature assay, which is not purposed to assist in the selection of optimal therapy, results from the remaining prognostic multigene expression profiles are intended to guide clinical treatment decisions, relaying the benefit of additional adjuvant chemotherapy and/or antiestrogen therapy. Table 1 compares important characteristics of the five prognostic multigene expression profiles, including number of genes evaluated, clinical utility, and patient eligibility.
  | Table 1 Prognostic multigene expression profiles |
Identification and prioritization of candidate genes
A list of genes evaluated within each of the five prognostic multigene expression profiles was compiled. Lists of genes were obtained from the following locations in March 2014: the eleven-gene expression profile (Jerevall et al),28 the 14-gene prognostic expression signature (Tutt et al),23 the 21-gene breast cancer assay (http://breast-cancer.oncotypedx.com), the 50-gene breast cancer prognostic gene signature assay (https://genome.unc.edu/pubsup/breastGEO/), and 70-gene breast cancer recurrence assay (Tian et al).40 Genes utilized in two or more profiles were prioritized for selection.
Pathway analysis of gene set
In order to ensure adequate representation of the biological capabilities of cancer in the parsimonious high priority candidate gene set, a gene-gene pathway analysis was conducted, using QIAGEN’s Ingenuity® Pathway Analysis software (IPA®, QIAGEN Redwood City, CA, USA, www.qiagen.com/ingenuity), to evaluate functional networks. Both direct and indirect relationships were analyzed. All molecules and/or relationships were considered.
Results
Among the five included prognostic gene expression profiles, 127 unique genes were identified. Twenty-one genes (BAG1, BCL2, BIRC5, CCNB1, CENPA, CMC2, DIAPH3, ERBB2, ESR1, GRB7, MELK, MKI67, MMP11, MYBL2, NDC80, ORC6, PGR, RACGAP1, RFC4, RRM2, and SCUBE2) are utilized in two or more of the profiles (Table 2). Five genes (CCNB1, CENPA, MELK, MYBL2, and ORC6) are used in three profiles. Primary functions of the identified candidate genes are provided in Table 2.
  | Table 2 Genes utilized in two or more prognostic multigene expression profiles as indicated by X |
The pathway analysis revealed that the main molecular and cellular functions of the parsimonious, high priority gene set are cell cycle, cellular development, cellular growth and proliferation, cell death and survival, and gene expression. Canonical pathways containing one or more of the identified 21 genes are displayed in Figure 2. Three unique networks were identified (Figure 3). The main associated diseases and functions of the three networks are 1) cancer, organismal injury and abnormalities, and reproductive system disease; 2) DNA replication, recombination, and repair, connective tissue disorders, and dental disease; and 3) cellular development, reproductive system development and function, and molecular transport. The pathway analysis also identified a number of plausible upstream transcription regulators of the identified 21 gene set, including TP53, CDKN1A, CDKN2A, E2F1, and E2F4.
  | Figure 2 Overlapping canonical pathways map representing shared biology among the identified candidate genes. |
  | Figure 3 Gene–gene networks generated by pathway analysis. |
Discussion and conclusion
In the future, we envision a holistic, personalized health care environment, in which breast cancer survivors receive not only a refined cancer diagnosis and prognosis based on the results of prognostic multigene expression profiles, but genetically tailored preclinical symptom prediction and proactive symptom management as well. Inspired by findings related to reported changes in cognitive function in women with breast cancer, this project, intended to identify and prioritize biologically plausible candidate genes, represents an initial and integral step in establishing a relationship between genetic variation and symptom variability in breast cancer survivors.
Driven by our hypothesis that symptom variability in breast cancer survivors is related to variation in the biology of cancer cells, we employed the innovative approach detailed in this article to select candidate genes based on prognostic multigene expression profiles for breast cancer. While we believe that all 127 unique genes evaluated as part of the included prognostic multigene expression profiles for breast cancer should be considered as candidates to test the proposed hypothesis, our project revealed considerable overlap in genes represented in the profiles with 21 genes replicated in two or more profiles. Five of the 21 replicated genes are used in three profiles. Because each prognostic multigene expression profile was developed to individualize breast cancer prognostication based on associations between breast cancer tumor gene expression levels and clinically relevant cancer outcomes, including recurrence and metastasis, replication of genes utilized in multiple profiles may be particularly important in describing the heterogeneity of breast cancer tumor cell biology and, consequently, should be prioritized for evaluation.
Nevertheless, by limiting a future investigation to variation in genes replicated in two or more profiles, we risk the inadvertent elimination of one or more of the biologic capabilities that enable tumor growth and metastatic dissemination. Eight biologic hallmarks of cancer have been identified and include resisting cell death, deregulating cellular energetics, sustaining proliferative signaling, evading growth suppressors, avoiding immune destruction, enabling replicative immortality, activating invasion and metastasis, and inducing angiogenesis.41 To ensure that our parsimonious, high priority gene set broadly incorporated these eight hallmarks, an Ingenuity® Pathway Analysis was conducted. The results of our pathway analysis revealed that the main molecular and cellular functions of the gene set were cell cycle, cellular development, cellular growth and proliferation, cell death and survival, and gene expression. We also found that the majority of the canonical pathways the genes in our identified gene set are contained within are related to cancer/cellular signaling. Furthermore, the genes identified by the analysis as potential upstream regulators of the gene set, TP53, CDKN1A, CDKN2A, E2F1, and E2F4, all play important roles in cell cycle control and tumor suppression. Due to the overlap between the molecular and cellular functions of the gene set and the hallmarks of cancer, we believe that the biologic hallmarks of cancer are represented in the prioritized gene set.
Interestingly, a further examination of genes, molecules, gene products, and gene complexes/interactions in the identified networks revealed minimal overlap with genes in the prognostic multigene expression profiles. We were surprised to find that only six genes (CCNE1, CCNE2, FLT1, MCM6, MMP9, and PRC1), beyond those contained in the inputted 21 gene set, are utilized in one of the five included prognostic multigene expression profiles. While unintentional and not the purpose of this project, we believe that the remaining network genes could be considered as potential candidates to develop new prognostic multigene expression profiles for breast cancer, to increase the sensitivity/specificity of current profiles, and/or as a means to potentially expand patient eligibility criteria.
Although conceptualized within the context of cognitive dysfunction, the identified genes would be ideal candidates for investigations of symptom variability in other disruptive and burdensome symptoms associated with breast cancer, its treatments, or both as well. The identified 21 gene set would be especially relevant for symptoms previously found to be associated with disease-related factor of breast cancer, such as fatigue, which has been predicted by tumor size and lymph involvement.42,43 Moreover, the novel approach we employed to select candidate genes for investigations of variability in breast cancer symptoms can also be applied to other cancers that have biologically based commercially available prognostic multigene expression profiles, such as colon and prostate cancer, and associated symptoms.
Genetic variability within the proposed candidate gene set can be evaluated in a number of ways, including evaluation of polymorphisms, gene expression levels, protein levels, and epigenetic changes in both the host and tumor tissue. We recognize that symptom variability, especially at the time of diagnosis, may be driven by tumor gene expression, the consequences of tumor expression and related protein production on the rest of the body, and subsequent removal and treatment of the primary tumor and secondary sites. Thus, we recommend that future symptomatology studies focus on relationships between tumor gene expression/protein levels and symptom variability in cancer survivors. Significant results from tumor gene expression studies in particular, could greatly and directly expand the clinical utility of currently available prognostic multigene expression profiles. For instance, a modified version of the gene expression algorithms could potentially generate a range of Cognitive Decline Risk Scores or Fatigue Risk Scores, based on the treatment regimen that is ultimately selected, other clinicopathologic tumor features, and baseline patient characteristics for each breast cancer survivor. However, relationships between host DNA and symptom variability are desirable as well because host DNA variation remains constant over time and is not tissue specific. Prediction of symptoms using host DNA would be especially advantageous for unusable tumor tissue, survivors with a clear prognosis and treatment regimen recommendation based on other clinicopathologic tumor features who would not be eligible for further prognostic multigene expression profile testing, or post hoc symptom prediction after tissue RNA has degraded and quantified gene expression level accuracy would be questionable.
However, we would like to acknowledge that this approach is not without limitations. Of particular interest, the multigene expression profiles from which candidate genes were selected, with the exception of the 70-gene breast cancer recurrence assay, all require positive breast cancer tumor estrogen or progesterone receptor status as an eligibility criterion. Consequently, genetic variation important for symptom variability in women with hormone receptor negative and triple-negative breast cancers may not be adequately captured in the prioritized gene set. In other words, the clinical applicability of breast cancer survivor symptom prediction may be limited to women with tumors that reflect the eligibility criteria of the five multigene expression profiles used to generate the candidate gene list. Additional candidate genes should be considered for other subsets of breast cancers not emphasized in the currently available multigene expression profiles.
Undoubtedly, the results of future investigations of symptom variability in breast cancer survivors based on disease characteristics at the cellular level, such as genetic variability in the high priority 21 gene set identified in this article, have the potential to substantially impact both the care of breast cancer survivors and the practice of health care providers alike, extending the clinical utility of prognostic multigene expression profiles for breast cancer and providing the patient and the provider with a means of weighing not only cancer prognosis and recurrence, but also the quality of life due to burdensome symptoms, into treatment decisions as well.
Acknowledgments
This project was supported by the Targeted Research and Academic Training for Nurses in Genomics (T32NR009759) training program and the Cognitive Function and Breast Cancer: Genomics and Disease Characteristics (F31NR014590) study. The authors acknowledge Dr Susan Cohen, PhD, FNP-BC, FAAN, and members of the University of Pittsburgh Cancer Survivorship Research Seminar for thoughtfully reviewing this manuscript. Portions of this project were presented at the International Society of Nurses in Genetics (ISONG) Silver Anniversary Conference in 2013.
Disclosure
The authors report no conflicts of interest in this work.
References
Shi Q, Smith TG, Michonski JD, Stein KD, Kaw CK, Cleeland CS. Symptom burden in cancer survivors one year after diagnosis: a report form the American Cancer Society’s Studies of Cancer Survivors. Cancer. 2012;117(12):2779–2790. | |
Burkett VS, Cleeland CS. Symptom burden in cancer survivorship. J Cancer Surviv. 2007;1(2):167–175. | |
Wu HS, Harden JK. Symptom burden and quality of life in survivorship: a review of the literature. Cancer Nurs. 2015;38(1):E29–E54. | |
Spiegel D, Giese-Davis J. Depression and cancer: mechanisms and disease progression. Biol Psychiatr. 2003;54(3):269–282. | |
Barsevick A, Frost M, Zwinderman A, Hall P, Halyard M; GENEQOL Consortium. I’m so tired: biological and genetic mechanisms of cancer-related fatigue. Qual Life Res. 2011;19(10):1419–1427. | |
Vardy J, Wefel JS, Ahles T, Tannock IF, Schagen SB. Cancer and cancer-therapy related cognitive dysfunction: an international perspective from the Venice cognitive workshop. Ann Oncol. 2008;19(4):623–629. | |
Falleti MG, Sanfilippo A, Maruff P, Weih L, Phillips KA. The nature and severity of cognitive impairment associated with adjuvant chemotherapy in women with breast cancer: a meta-analysis of the current literature. Brain Cogn. 2005;59(1):60–70. | |
Bender CM, Sereika SM, Brufsky AM, et al. Memory impairments with adjuvant anastrozole versus tamoxifen in women with early-stage breast cancer. Menopause. 2010;14(6):995–998. | |
Castellon SA, Ganz PA, Bower JE, Petersen L, Abraham L, Greendale GA. Neurocognitive performance in breast cancer survivors exposed to adjuvant chemotherapy and tamoxifen. J Clin Exp Neuropsychol. 2004;26(7):955–969. | |
Breckenridge LM, Bruns GL, Todd BL, Feuerstein M. Cognitive limitations associated with tamoxifen and aromatase inhibitors in employed breast cancer survivors. Psychooncology. 2012;53:43–53. | |
Lejbak L, Vrbancic M, Crossley M. Endocrine therapy is associated with low performance on some estrogen-sensitive cognitive tasks in postmenopausal women with breast cancer. J Clin Exp Neuropsychol. 2010;32(8):836–846. | |
Schilder CM, Seynaeve C, Beex LV, et al. Effects of tamoxifen and exemestane on cognitive functioning of postmenopausal patients with breast cancer: results from the neuropsychological side study of the tamoxifen and exemestane adjuvant multinational trial. J Clin Oncol. 2010;28(8):1294–1300. | |
Shilling V, Jenkins V, Fallowfield L, Howell T. The effects of hormone therapy on cognition in breast cancer. J Steroid Biochem Mol Biol. 2003;86(3–5):405–412. | |
Jenkins V, Shilling V, Fallowfield L, Howell A, Hutton S. Does hormone therapy for the treatment of breast cancer have a detrimental effect on memory and cognition?: a pilot study. Psychooncology. 2004;13(1):61–66. | |
Hermelink K, Untch M, Lux MP, et al. Cognitive function during neoadjuvant chemotherapy for breast cancer: results of a prospective, multicenter, longitudinal study. Cancer. 2007;109(9):1905–1913. | |
Wefel JS, Lenzi R, Theriault R, Buzdar AU, Cruickshank S, Meyers CA. “Chemobrain” in breast carcinoma?: a prologue. Cancer. 2004; 101(3):466–475. | |
Ahles TA, Avenue L, York N, Saykin AJ, Mcdonald BC, Kaufman PA. Cognitive function in breast cancer patients prior to adjuvant treatment. Breast Cancer Res Treat. 2011;110(1):143–152. | |
Bender CM, Sereika SM, Ryan CM, Brufsky AM, Puhalla S, Berga SL. Does lifetime exposure to hormones predict pretreatment cognitive function in women before adjuvant therapy for breast cancer? Menopause. 2013;20(9):1–8. | |
Schagen SB, Muller MJ, Boogerd W, Mellenbergh GJ, van Dam FS. Change in cognitive function after chemotherapy: a prospective longitudinal study in breast cancer patients. J Natl Cancer Inst. 2006;98(23):1742–1745. | |
Zhang Y, Schnabel CA, Schroeder BE, et al. Breast cancer index identifies early-stage estrogen receptor-positive breast cancer patients at risk for early- and late-distant recurrence. Clin Cancer Res. 2013;19(15):4196–4205. | |
Sgroi DC, Carney E, Zarrella E, et al. Prediction of late disease recurrence and extended adjuvant letrozole benefit by the HOXB13/IL17BR biomarker. J Natl Cancer Inst. 2013;105(14):1036–1042. | |
Sgroi DC, Sestak I, Cuzick J, et al. Prediction of late distant recurrence in patients with oestrogen-receptor positive breast cancer: a prospective comparison of the breast-cancer index (BCI) assay, 21-gene recurrence score, and IHC4 in the TransATAC study population. Lancet Oncol. 2013;14(11):1067–1076. | |
Tutt A, Wang A, Rowland C, et al. Risk estimation of distant metastasis in node-negative, estrogen receptor-positive breast cancer patients using an RT-PCR based prognostic expression signature. BMC Cancer. 2008;8:339. | |
Genomic Health®, Inc. Oncotype DX® Breast Cancer Assay Invasive Sales Aid. Available from: http://breast-cancer.oncotypedx.com/en-US/Professional-Invasive/~/media/ODX-Breast/Images/Invasive/InvasiveSalesAid_11-2.pdf. Accessed November 3, 2016. | |
Genomic Health®, Inc. Oncotype DX® Breast Cancer Assay DCIS Sales Aid. Available from: http://breast-cancer.oncotypedx.com/en-US/Professional-DCIS/~/media/ODX-Breast/Files/Downloads/DCIS_Sales_Aid.pdf. Accessed November 13, 2015. | |
NanoString Technologies®, Inc. Package Insert Prosigna® Breast Cancer Prognostic Gene Signature Assay. Available from: http://prosigna.com/docs/Prosigna_Packet_Insert_US.pdf. Accessed October 5, 2015. | |
Agendia®. Mammaprint® 70-Gene Breast Cancer Recurrence Assay. Available from: http://www.agendia.com/healthcare-professionals/breast-cancer/mammaprint/. Accessed November 12, 2015. | |
Jerevall PL, Ma XJ, Li H, et al. Prognostic utility of HOXB13:IL17BR and molecular grade index in early-stage breast cancer patients from the Stockholm trial. Br J Cancer. 2011;104(11):1762–1769. | |
Paik S, Shak S, Tang G, et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med. 2004;351(27):2817–2826. | |
Paik S, Tang G, Shak S, et al. Gene expression and benefit of chemotherapy in women with node-negative, estrogen receptor-positive breast cancer. J Clin Oncol. 2006;24(23):3726–3734. | |
Parker JS, Mullins M, Cheang MCU, et al. Supervised risk predictor of breast cancer based on intrinsic subtypes. J Clin Oncol. 2009;27(8):1160–1167. | |
Buyse M, Loi S, van’t Veer L, et al. Validation and clinical utility of a 70-gene prognostic signature for women with node-negative breast cancer. J Natl Cancer Inst. 2006;98(17):1183–1192. | |
van’t Veer LJ, Dai H, van de Vijver MJ, et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature. 2002;415(6871):530–536. | |
Horn D, Zhou W, Trevisson E, et al. The conserved mitochondrial twin Cx9C protein Cmc2 Is a Cmc1 homologue essential for cytochrome coxidase biogenesis. J Biol Chem. 2010;285(20):15088–15099. | |
Vulsteke V, Beullens M, Boudrez A, et al. Inhibition of spliceosome assembly by the cell cycle-regulated protein kinase MELK and involvement of splicing factor NIPP1. J Biol Chem. 2004;279(10):8642–8647. | |
Lin ML, Park JH, Nishidate T, Nakamura Y, Katagiri T. Involvement of maternal embryonic leucine zipper kinase (MELK) in mammary carcinogenesis through interaction with Bcl-G, a pro-apoptotic member of the Bcl-2 family. Breast Cancer Res. 2007;9(1):R17. | |
Beullens M, Vancauwenbergh S, Morrice N, et al. Substrate specificity and activity regulation of protein kinase MELK. J Biol Chem. 2005;280(48):40003–40011. | |
Lin YC, Chen CC, Cheng CJ, Yang RB. Domain and functional analysis of a novel breast tumor suppressor protein, SCUBE2. J Biol Chem. 2011;286(30):27039–27047. | |
NCBI Resource Coordinators. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 2015; 43(Database issue):D6-17. | |
Tian S, Roepman P, van’t Veer LJ, Bernards R, de Snoo F, Glas AM. Biological functions of the genes in the mammaprint breast cancer profile reflect the hallmarks of cancer. Biomark Insights. 2010;5:129–138. | |
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144(5):646–674. | |
Goldstein D, Bennett BK, Webber K, et al. Cancer-related fatigue in women with breast cancer: outcomes of a 5-year prospective cohort study. J Clin Oncol. 2012;30(15):1805–1812. | |
Wielgus KK, Berger AM, Hertzog M. Predictors of fatigue 30 days after completing anthracycline plus taxane adjuvant chemotherapy for breast cancer. Oncol Nurs Forum. 2009;36(1):38–48. |
 © 2016 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2016 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.