Back to Journals » Cancer Management and Research » Volume 12
Evaluation of PCR-Reverse Dot Blot Human Papillomavirus Genotyping Test in Predicting Residual/Recurrent CIN 2+ in Posttreatment Patients in China
Authors Zhang Q , Dong B, Chen L , Lin T, Tong Y , Lin W , Lin H , Gao Y, Lin F, Sun P
Received 6 November 2019
Accepted for publication 13 March 2020
Published 1 April 2020 Volume 2020:12 Pages 2369—2379
DOI https://doi.org/10.2147/CMAR.S237490
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Bilikere Dwarakanath
Qiaoyu Zhang,1,* Binhua Dong,1,* Lihua Chen,1 Tingting Lin,2 Yao Tong,1 Wenyu Lin,1 Haifeng Lin,1 Yuqin Gao,1 Fen Lin,1 Pengming Sun1,3
1Laboratory of Gynecologic Oncology, Fujian Provincial Maternity and Children’s Health Hospital, Affiliated Hospital of Fujian Medical University, Fuzhou, People’s Republic of China; 2Department of Gynecology, The First Affiliated Hospital of Xiamen University, Xiamen, People’s Republic of China; 3Department of Gynecology, Fujian Provincial Maternity and Children’s Health Hospital, Affiliated Hospital of Fujian Medical University, Fuzhou, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Pengming Sun
Laboratory of Gynecologic Oncology, Fujian Provincial Maternity and Children’s Hospital, Affiliated Hospital of Fujian Medical University, Fuzhou, People’s Republic of China
Tel +86 591 8755 8732
Fax +86 591 8755 1247
Email [email protected]
Objective: To assess the clinical value of the PCR-reverse dot blot human papillomavirus genotyping test during follow-up of patients with CIN grade 2 or worse (CIN 2+).
Methods: Four hundred patients with CIN 2+ receiving treatment from January 2008 to January 2017 were included in our study. Postoperative follow-up procedures comprised HPV examination and cervical cytology every 3– 6 months for the first 2 years and then followed up every 6– 12 months. A pathology examination was performed when there was a positive funding for HPV 16/18 or an abnormal ThinPrep cytology test (TCT) with or without positive for HR-HPV according to the American Society for Coloscopy and Cervical Pathology (ASCCP) guidelines.
Results: The median follow-up period was 27.10± 12.47 months (ranging from 3 to 50 months). During follow-up, 12.00% (48/400) of the women developed residual/recurrent disease. The highest risk in CIN 2+ and CIN 3+ residual/recurrence was HPV-16/-18 (hazard ratio (HR)=12.898, 95% CI= 6.849– 24.289; HR= 20.726, 95% CI= 9.64– 44.562, respectively). Among the different follow-up methods, type-specific (TP) HR-HPV persistent infection showed the highest cumulative incidence risk (CIR) (84.62%, 95% CI=73.29– 95.94) and HR (5.38, 95% CI= 2.596– 11.149) during the 4-year follow-up period. At the CIN 2+ and CIN 3+ endpoints, TP-HPV testing had relatively high sensitivity (84.62%, 95% CI=73.29– 95.94 and 89.28%, 95% CI= 77.83– 100.00, respectively) and specificity (78.07%, 95% CI= 72.70– 83.44 and 75.73%, 95% CI= 70.30– 81.17, respectively). However, at the CIN 2+/CIN 3+ endpoint, TCT follow-up had a sensitivity of 60.42%/62.16% (95% CI=46.58– 72.25/46.54– 77.79) and specificity of 90.18%/88.72% (95% CI=86.95– 93.41/85.35– 92.10).
Conclusion: TP HR-HPV follow-up can provide a reliable and sensitive clinical reference for CIN 2+ postoperative patients.
Keywords: papillomavirus, genotype, cell biology, histology, postoperative
Introduction
Cervical cancer is the third most frequent cancer among women globally. There are approximately 97,500 newly diagnosed cervical cancer cases and 30,500 related deaths in China. Invasive cervical cancer usually develops from cervical intraepithelial neoplasia (CIN) over a long period of time.1 Most invasive cervical cancer progresses from high-grade cervical lesions. To prevent cancer progression from cervical lesions, conization of the lesion or even hysterectomy is needed after diagnosis. However, 15% or more of high-grade cervical lesions (CIN grade 2 or worse) remain after treatment making the development of recurrent CIN grade 2 or worse (CIN 2+) more likely, most cases of CIN 2+ are diagnosed within 2 years of treatment.2 Thus, developing a follow-up strategy for patients to avoid residual cervical lesions and/or prevent recurrence is important. The 2013 American College of Obstetricians and Gynecologists (ACOG) guidelines recommend that women be treated for CIN 2+ during follow-up visits at 12 months and 24 months by cytology and high-risk human papillomavirus (HR-HPV) cotesting.3 However, the success of this follow-up strategy depends on a professional pathologist. In most resource-limited countries, the number of pathologists is low, and the high economic burden and complex infrastructural demand may hinder the use of this strategy. In contrast to most developed countries, in many resource-limited countries, such as China, many patients are not aware of the importance of postoperative follow-up, and the regular examination coverage rate after treatment is still low. In addition, most cervical cancer screening is opportunistic.4 Thus, more attention should be paid to the sensitivity and negative predictive value (NPV) during the postoperative period.
Vitale et al5 proposed P16INK4a as a progression/regression tumour marker in low-grade cervical lesions (LSILs) and showed its excellent clinical effect. Valenti et al6 showed that some markers, such as p16, E-cadherin, Ki67, pRb and p53, were able to predict which cervical intraepithelial lesions had a greater chance of developing. Rossetti et al7 suggested sentinel lymph nodes (SLNs) for the detection of early-stage cervical cancer. By studying cervical cancer biological markers can not only reveal early cervical lesions but also predict the recurrent and residual conditions of patients after cervical lesion removal, preventing fertility loss at a young age and reducing the economic and psychological burden caused by unnecessary surgical treatment. However, the above methods are inconvenient and may cause large economic burden, which is not suitable for resource-limited countries. In our previous study,8 the PCR-reverse dot blot (PCR-RDB) Yaneng® Human Papillomavirus Genotyping Kit (Yaneng Biotech, Shenzhen, China), which detects 18 HR-HPV types (HPV-16, −18, −31, −33, −35, −39, −45, −51, −52, −53, −56, 58, −59, −66, −68, −73, −82, and −83) and 5 low-risk (LR)-HPV types (HPV-6, −11, −42, −43, and −81), was evaluated in a hospital-based population and showed a reliable and sensitive clinical reference for cervical screening. The PCR-RDB is largely used in China. Kang et al9 stated that the PCR-RDB HPV genotyping test could be applied in cervical cancer primary screening for hospital-based opportunistic screening. However, there is no adequate research on whether the PCR-RDB HPV test is suitable for the posttreatment of high-grade cervical lesions. Previously, several studies supported the prognostic role of HPV testing, with its relatively high sensitivity and specificity compared with those of other methods. Soderlund-Strand et al10 reported that HR-HPV persistent infection predicts recurrent or residual disease in posttreatment surveillance more accurately than cotesting and improves the specificity of the prediction. However, most of these articles were based on Hybrid Capture® 2 (HC2) HPV DNA (Qiagen, Hilden, Germany), Cervista® (Hologic, Massachusetts, USA), Aptima® (Hologic, Massachusetts, USA), and Cobas® HPV tests (Roche Diagnostics, Indianapolis, USA). Four types of HPV testing methods have been approved by the US Food and Drug Administration (FDA). Only the Cobas® HPV test is capable of genotyping. A few studies have reported the clinical validation of type-specific HPV genotyping in high-grade cervical lesion follow-up. It was unclear which persistent infection HPV genotype requires the most attention. The specificity of HR-HPV genotyping was relatively low. In addition, most of these studies were based on Western and current postoperative follow-up guidelines, which were based on the characteristics of residual or recurrent disease in Western countries. Studies that focus on postoperative follow-up strategies for high-grade lesions or worse that were conducted in East Asia are few, especially in resource-limited countries. As many factors may be associated with residual disease or recurrence in postoperative patients, including ethnic differences and geographical differences, whether the guidelines made by Western countries are suited for East Asian patients remains unknown.
In this study, we investigated the risk of residual or recurrent disease in posttreatment patients with CIN 2+ cervical lesions in China. Moreover, the feasibility of detecting the type-specific HPV persistent infection by PCR-RDB HPV assay as a CIN 2+ postoperative follow-up method was also assessed.
Materials and Methods
Patients
According to the sample size calculation formula (N=Z0.052×(P×(1-P))/E2('N' is sample size; ‘Z0.05ʹ is 1.96; 'E' is the error value; 'P' is the probability value)), we arrived at the conclusion that 59 patients would be needed at least. Between January 2008 and January 2017, 400 patients diagnosed with CIN 2+ by colposcopy and cervical biopsy with full data in Fujian Provincial Maternity and Children’s Health Hospital, affiliated hospital of Fujian Medical University were enrolled in this retrospective cohort study finally. The included participants met the following inclusion criteria: 1) patients with CIN 2+ underwent cervical conization; 2) all patients were followed up from the day of surgery, and the postoperative follow-up procedures comprised HPV examination and vaginal stump cytology every 3 to 6 months as necessary for the first 2 years and every 6 to 12 months for up to 4 postoperative years postoperatively, with pathology examination performed when there were abnormal findings (HPV 16/18 positivity or ThinPrep cytology test (TCT) ≥ASCUS with or without HR-HPV positivity); 3) patients had no immune system diseases, sexually transmitted diseases or related drug histories; 4) there was no history of cervical disease treatment or chemoradiotherapy; and 5) all patients that reported the presence of cancer subsequently underwent a second excisional procedure. The exclusion criteria were as follows: 1) patients with a large amount of missing data or 2) history of complete hysterectomy before the first follow-up period. The study was approved by the Ethics Committee of the Affiliated Hospital of Fujian Medical University (2014–45), and all individuals in this study provided written informed consent.
Liquid-Based Cytology
Cytological specimens’ results were independently evaluated by two experienced cytopathologists. The cervical samples would be analyzed again when the diagnosis was different. According to the Bethesda system,11 the results include negative for intraepithelial lesion or malignancy (NILM), atypical squamous cells of undetermined significance (ASC-US), low-grade squamous intraepithelial lesion (LSIL), atypical squamous cells, not possible to exclude high-grade squamous intraepithelial lesion (ASC-H), high-grade squamous intraepithelial lesion (HSIL), squamous cervical cancer (SCC), atypical glandular cells (AGC), and adenocarcinoma in situ (AIS).
HPV Genotyping
The YaNeng® PCR-RDB HPV genotyping kit (Yaneng Biosciences, Shenzhen, China) can detect 23 HPV types (16, 18, 31, 33, 35, 39, 45, 51, 52, 53, 56, 58, 59, 66, 68, 73, 82, 83, 6, 11, 42, 43, and 81). This method is permitted by the China Food and Drug Administration (CFDA) with number 20,020,515. The experiment was conducted in accordance with the product instructions.8
Follow-Up Content and Related Definitions
All subjects were tested by HPV genotyping before diagnosis. Only 59 patients had no TCT results before surgery. Progression to CIN 2+ over less than 1 year indicated residual disease in posttreatment patients. Progression to CIN 2+ over more than 1 year indicated recurrence in posttreatment patients. In cervical cancer patients, recurrence was defined as progression to VaIN 2 or worse over more than 1 year. The timespan for residual or recurrent disease was recorded from the time of conization at baseline until CIN 2+ lesions were first detected. A lesion was considered to have undergone remission/regression when 1) HPV testing in clinical follow-up was negative or 2) at least two ensuing cytology and/or cervical biopsy samples obtained at least six months apart were normal and with HPV negative registered in the latter period. HR-HPV persistent infection was defined as any type of HR-HPV infection detected at least 2 times during follow-up. Type-specific HPV persistent infection was defined as the detection of the same HR-HPV type after treatment at least 2 times during the follow-up period or the same HR-HPV type preoperatively and postoperatively.
Statistical Analysis
The hazard ratio (HR) with the 95% confidence interval (95% CI) was estimated for groups of HR-HPV types using multivariate Cox proportional hazard regression. The data were analysed using the IBM SPSS statistical package version 22.0 (IBM Corporation, Armonk, NY, USA) and Stata 12.0 software program (StataCorp LP, College Station, TX, USA). P<0.05 or P<0.001 was regarded as statistically significant.
Results
Characteristics of the Study Population and HPV Status
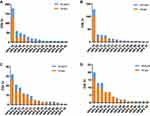
A total of 400 patients were included in the study. Patients who did not undergo follow-up evaluations were excluded. Among the included patients, 145 had CIN 2, 173 had CIN 3, 5 had AIS, 68 had SCC and 9 had atypical adenocarcinoma (ADC). The median age of this population was 41.48±9.10 years (ranging from 21 to 71 years). Table 1 shows the demographic characteristics of the patients. The median follow-up period was 27.10±12.47 months (ranging from 3 to 50 months). The HR-HPV-positive rate of this population was 90.50% (362/400) before surgery, with prevalence rates of 71.27% (258/362) for single-type HR-HPV, 21.82% (79/362) for double-type HR-HPV and 6.91% (25/362) for multiple-type HR-HVP. The most frequent HPV type was HPV-16, followed by HPV-52, −58, −18 and −33 (Figure 1).
 |
Table 1 Clinical Characteristics of Patients |
Characteristics of Postoperative Patients
During the follow-up period, 12.00% (48/400) of the women had residual/recurrent disease. Of these patients, 12.5% (6/48) progressed to invasive cervical cancer. The median residual/recurrence time was 15.06±12.71 months (ranging from 3 to 50 months). The HR-HPV-positive rates were 47.57% (127/267), 20.33% (73/359), and 17.28% (56/324) at the first, second and third follow-up evaluations, respectively. The type-specific (TP)-HPV-positive rate was 31.09% (83/267), 13.93% (50/359), and 12.35% (40/324) at the first, second and third follow-up visits, respectively. The HR-HPV- and TP-HPV-positive rates differed between the different age groups. Among patients younger than 30 years, the HR-HPV-positive rates were 31.82% (7/22), 16.67% (5/30) and 10.34% (3/29) at the first, second and third follow-up evaluations, respectively, and the TP-HPV-positive rates were 13.64% (3/22), 10% (3/30) and 6.90% (2/29) at the first, second and third follow-up evaluations, respectively. Among women aged 31 to 54 years, the positive rates at the first, second and third follow-up evaluations were 44.44% (96/216), 18.27% (55/301) and 16.18% (44/272), respectively, for HR-HPV and 28.70% (62/216), 11.30% (34/301) and 11.40% (31/272), respectively, for TP-HPV. Among women 55 years of age or older, the positive rates at the first, second and third follow-up evaluations were 82.76% (24/29), 46.43% (13/28) and 39.13 (9/23), respectively, for HR-HPV and 62.07% (18/29), 46.43% (13/28) and 30.43% (7/23), respectively, for TP-HPV. As age increased, both the HR-HPV and TP-HPV prevalence rates also increased. The most frequent HR-HPV genotype at the first follow-up evaluation was HPV-16, followed by HPV-58, −52, 18 and −53 (Figure 1).
The Risk for Residual/Recurrent Disease After Surgery
The 4-year cumulative risk for residual/recurrent CIN 2+ was highest in the HPV-16/-31/-33/-35/-52/-58 group (cumulative incidence risk [CIR]= 76.92%, 95% CI= 63.70–90.15), followed by the HPV-16/-18 (CIR= 64.1%, 95% CI= 49.05–79.16), single HPV-16 (CIR= 51.28%, 95% CI= 35.59–66.97), HPV-18/-39/-45/-59/-68 (CIR= 17.95%, 95% CI= 4.06–26.71) and single HPV-18 groups (CIR= 12.82%, 95% CI= 2.33–23.31). The CIN 2+ cumulative risk for the HPV-16/-31/-33/-35/-52/-58 group was significantly different from that of the single HPV-16, single HPV-18 and HPV-18/-39/-45/-59/-68 groups (all P<0.05). However, the CIN 2+ cumulative risk for the HPV-16/-31/-33/-35/-52/-58 group was not significantly different from that of the HPV-16/-18 group (P=0.231). For CIN 3+, HPV-16/18 and the HPV-16/31/33/35/52/58 groups (both CIR= 73.33%, 95% CI=57.51–89.16) had the highest cumulative risk, followed by single HPV-16 (CIR= 56.67%, 95% CI= 38.93–74.40), HPV-18/39/45/59/68 (CIR= 20.00%, 95% CI= 5.69–34.31), and single HPV-18 groups (CIR= 16.67%, 95% CI= 3.33–30.00). There was no significant difference between the HPV-16/18 and HPV-16 groups (P=0.176)
Using multivariate Cox regression analysis, the various HR-HPV risk groups showed different 4-year HRs for residual/recurrent CIN 2+ and CIN 3+ (Table 2 and Supplement Table 1). The HPV-16/18 group had the highest HR for residual/recurrent CIN 2+ and CIN 3+ (HR= 12.898, 95% CI= 6.849–24.289; HR= 20.726, 95% CI=9.64–44.562, respectively).
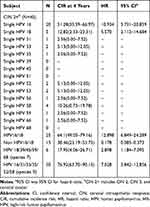 |
Table 2 The Cumulative Risk and Hazard Ratio for Type-Specific HR-HPV Prevalence at First Follow-Up |
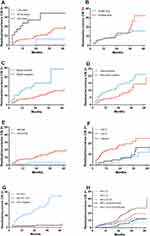
Multivariate Cox survival analysis demonstrated that margin status, age, preoperative pathology and HR-HPV infection status were all associated with the risk of residual/recurrent CIN 2+ (Figure 2).
HPV, TCT and TCT Combined with HPV (Cotesting) in the Detection of Residual/Recurrent Disease
The different TCT statuses and HPV statuses showed different CIRs and HRs at the first follow-up evaluation, and the results are shown in Table 3. TP-HPV showed the highest CIR and HR at the 4-year follow-up evaluation, namely, 84.62% (95% CI=73.29–95.94) and 5.38 (95% CI= 2.596–11.149), respectively.
 |
Table 3 The Cumulative Risk and Hazard Ratio for the 4-Year Follow-Up |
As shown in Table 4, HR-HPV testing showed the highest sensitivity and NPV, namely, 89.74% (95% CI=80.22–99.26) and 97.14% (95% CI= 94.38–99.90), respectively, at the CIN 2+ endpoint and 96.43% (95% CI= 89.55–100.00) and 99.29% (95% CI= 97.89–100.00), respectively, at the CIN 3+ endpoint. However, the specificity was low at both the CIN 2+ endpoint and CIN 3+ endpoint, namely, 59.65% (95% CI= 53.28–66.02) and 58.16% (95% CI= 51.91–64.41), respectively. None of the TP-HPV tests reached the highest specificity of 91.67% (95% CI= 88.08–95.25) or 92.05% (95% CI= 88.62–95.48) at either the CIN 2+ or CIN 3+ endpoints, respectively; the sensitivity was only 5.13% (95% CI= 0.00–12.05) and 7.14% (95% CI= 0.00–16.68) at the two endpoints. TP-HPV testing had high sensitivity and specificity compared with HR-HPV testing, namely, 84.62% (95% CI=73.29–95.94) and 78.07% (95% CI= 72.70–83.44), respectively, at the CIN 2+ endpoint and 89.28% (95% CI= 77.83–100.00) and 75.73% (95% CI= 70.30–81.17), respectively, at the CIN 3+ endpoint. The performance of each method was evaluated at the first follow-up period. The threshold analyses showed that HR-HPV testing and TP-HPV testing had similar sensitivity for predicting postoperative high-grade residual/recurrent disease, but the latter had a higher specificity and was the most suitable for predicting postoperative high-grade residual/recurrent disease.
 |
Table 4 Compare Different Methods in High-Grade Cervical Lesions or Worse |
Discussion
Cervical cancer is the second most common gynaecologic malignancy among women in developing countries, with approximately 85% of the cervical cancer cases ending in death.12 Persistent HR-HPV infection plays a major role in the development of cervical cancer. Many published studies confirmed that the HPV type in the excised cone or in pretreatment specimens is the same as that in posttreatment specimens by HPV testing with increased specificity.13–16 However, controversy remains. Some studies have reported a degree of sensitivity loss associated with the persistence of HPV type specificity,14,17 while others have not shown this correlation.9,18 These differences may be the result of differences in HPV testing methods,19 geographical differences, and population differences, among others. In this study, although the sensitivity of the type-specific HR-HPV persistence was slightly lower than that of HR-HPV, its specificity was greatly improved. Our study confirmed that type-specific HR-HPV testing improves the postoperative detection of recurrent CIN 2+ lesions.
A previous study found that differences in the recurrence of CIN 2+ cervical lesions at different ages exist. A meta-analysis suggested20 that most CIN 2 lesions, particularly in young women (<30 years), regress spontaneously. Another study suggested21 that a negative oncogenic HPV test or negative HPV test plus cytology correlates with a low remaining lifetime cervical cancer risk for unvaccinated women with a cervix after the age of 55 years. These studies have shown a lower probability of progression in patients less than 30 years of age and over 55 years of age. However, these conclusions are both drawn from cervical cancer screening. Whether this difference was also present during the postoperative follow-up period is still uncertain. Our study showed a similar result during the follow-up period: the recurrence rate was highest among patients aged 30–54 years (77.08%) and was lower in patients younger than 30 years (2.08%) and older than 55 years (20.83%). Many factors may cause this difference, such as nonspecific immunity of the vaginal tract, which could contribute to a reduction in the duration of HR-HPV infection, and the basis for this difference requires further investigation.
In view of the difference in the recurrence or residual rates of patients among different age groups, the management of cervical lesions and even cervical cancer should be personalized, taking into account the performance status of the patient, particularly in cases of older women. It is worth noting that in this study, older women had relatively high recurrence or residual rates after treatment. Pallis et al estimated22 that the number of older patients with cancer will be increasing in the coming years. According to previous studies of older women with gynaecological cancer, although age is considered as a poor prognostic factor, the cause appears to be related not only to disease status but also to treatment strategy and intensity.23,24 Many studies have demonstrated that older patients can benefit from treatment to manage their gynaecological cancers.25,26 Therefore, proper management is needed to address the upcoming outbreak of older cancer patients. Specific difficulties of cervical screening in older women have long been documented with the loss of efficiency of TCT examination in this population.27–29 Thus, a more sensitive and specific method is needed. Our results showed that type-specific HR-HPV testing was a highly sensitive and specific method. This method has great clinical value for the postoperative follow-up of older women.
In this study, we found that compared with single HPV infection, multiple HPV infection showed a higher CIN 2+/CIN 3+ residual/recurrence rate. HPV-16 was the most common oncogenic genotype found in both preoperative and postoperative women, with the highest 4-year CIR (51.28%, 95% CI=35.59–66.97) and HR (10.934, 95% CI=5.731–20.859). Rositch et al30 demonstrated that the clinical HR-HPV persistence differed by HPV type. Molano et al31 also suggested that HPV-16 was associated with cervical malignancy and tended to persist longer in cervical tissue than in other types. There was no significant difference between the HPV16/18 and HPV 16 groups, which had similarly high 4-year CIRs and HRs (CIR=64.1%, 95% CI=49.05–79.16, HR=12.898, 95% CI=6.849–24.289 and CIR= 51.28%, 95% CI=35.59–66.97, HR=10.934, 95% CI=5.731–20.859, respectively). The assumption was confirmed by the fact that single HPV-16 has a similar CIN 2+/CIN 3+ progression risk when evaluated in combination with other HPV types.
The remaining 16.67% (8/48) of HR-HPV-negative patients suffered from residual or recurrent CIN 2+. This result was in contrast to that of a previous study.18 However, HR-HPV negativity in the follow-up period does not imply that HR-HPV is not involved in the aetiology of CIN 2+ residual lesions or recurrence, since HR-HPV-negative cases may have been transiently infected with HR-HPV before HPV testing was conducted and because of the false-positive rate of HPV detection method.
In the former studies, most of CIN 2 + postoperative follow-up were based on four types of the HPV detection methods approved by the FDA. These methods does not discriminate individual HPV genotypes. Our study found different HR-HPV genotypes in patients with CIN 2 + have different residual or recurrent risks after surgery. In China, HPV 16, 18 and 58 have the highest residual or recurrent risk. Besides, the study found type-specific HR-HPV follow-up is a relatively sensitive and specific method in China and can provide more clinical reference value in predicting recurrence or residual disease in posttreatment patients with CIN 2+. In the future, HPV genotyping tests for CIN 2 + postoperative patients can help to recognize patients with TP HR-HPV infection in time. Individualized follow-up strategies for these patients not only can recognize residual or recurrent patients in time but also reduce the financial and psychological burden of patients after surgery. In summary, the strengths of this article are two sides. First, this article is the first to perform PCR-RDB HPV testing during the follow-up of postoperative women with CIN 2+. Second, this study is dedicated to evaluating methods not only for high-grade lesions but also for early cervical cancer during postoperative follow-up (including AIS and SCC of A1 stage). However, this study had several limitations. The population came from the Fujian Provincial Maternity and Children’s Health Hospital, affiliated Hospital of Fujian Medical University, and there was no multicentre sampling; thus, selection bias may exist. Additionally, the sample size is small, and a larger population is needed for further study.
Acknowledgments
The authors would like to thank the Fujian Province Cervical Lesions Screening Cohorts Investigators for providing the data to the present study. Above all, they are grateful to the patients who made this study possible. This work was supported by grants from Fujian Provincial Health and Family Planning Commission Innovation Project (grant no. 2009-CXB-33 and 2019-CX-7) and Fujian Provincial Natural Science Foundation of China (grant no. 2017J01232), and Fujian Provincial Maternity and Children’s Hospital Natural Science Foundation (grant no.13-27 and 19-07).
Disclosure
The authors report no conflicts of interest in this work.
References
1. Chen W, Zheng R, Baade PD, et al. Cancer statistics in China, 2015. CA Cancer J Clin. 2016;66(2):115–132. doi:10.3322/caac.21338
2. Kocken M, Helmerhorst TJ, Berkhof J, et al. Risk of recurrent high-grade cervical intraepithelial neoplasia after successful treatment: a long-term multi-cohort study. Lancet Oncol. 2011;12(5):441–450. doi:10.1016/S1470-2045(11)70078-X
3. American College of Obstetricians and Gynecologists. Practice Bulletin No. 140: management of abnormal cervical cancer screening test results and cervical cancer precursors. Obstet Gynecol. 2013;122(6):1338–1367. doi:10.1097/01.AOG.0000438960.31355.9e
4. Gao K, Eurasian M, Zhang J, et al. Can genomic amplification of human telomerase gene and C-MYC in liquid-based cytological specimens be used as a method for opportunistic cervical cancer screening? Gynecol Obstet Invest. 2015;80(3):153–163. doi:10.1159/000371760
5. Vitale SG, Valenti G, Rapisarda AMC, et al. P16INK4a as a progression/regression tumour marker in LSIL cervix lesions: our clinical experience. Eur J Gynaecol Oncol. 2016;37(5):685–688.
6. Valenti G, Vitale SG, Tropea A, Biondi A, Laganà AS. Tumor markers of uterine cervical cancer: a new scenario to guide surgical practice? Updates Surg. 2017;69(4):441–449. doi:10.1007/s13304-017-0491-3
7. Rossetti D, Vitale SG, Tropea A, Biondi A, Lagana AS. New procedures for the identification of sentinel lymph node: shaping the horizon of future management in early stage uterine cervical cancer. Updates Surg. 2017;69(3):383–388. doi:10.1007/s13304-017-0456-6
8. Sun P, Song Y, Ruan G, et al. Clinical validation of the PCR-reverse dot blot human papillomavirus genotyping test in cervical lesions from Chinese women in the Fujian province: a hospital-based population study. J Gynecol Oncol. 2017;28(5):e50. doi:10.3802/jgo.2017.28.e50
9. Kang WD, Oh MJ, Kim SM, Nam JH, Park CS, Choi HS. Significance of human papillomavirus genotyping with high-grade cervical intraepithelial neoplasia treated by a loop electrosurgical excision procedure. Am J Obstet Gynecol. 2010;203(1):
10. Soderlund-Strand A, Kjellberg L, Dillner J. Human papillomavirus type-specific persistence and recurrence after treatment for cervical dysplasia. J Med Virol. 2014;86(4):634–641. doi:10.1002/jmv.23806
11. Verma I, Jain V, Kaur T. Application of bethesda system for cervical cytology in unhealthy cervix. J Clin Diagn Res. 2014;8(9):OC26–30. doi:10.7860/JCDR/2014/9620.4893
12. Goodman A. HPV testing as a screen for cervical cancer. BMJ (Clinical Research Ed). 2015;350(jun301):h2372.
13. Choi H, Kang W, Kim S. Significance of human papillomavirus genotyping with high-grade cervical intraepithelial neoplasia treated with a loop electrosurgical excision procedure. Gynecol Oncol. 2010;116(3):S30–S30.
14. Brismar S, Bo J, Borjesson M, Arbyn M, Andersson S. Follow-up after treatment of cervical intraepithelial neoplasia by human papillomavirus genotyping. Am J Obstetr Gynecol. 2009;201(1):
15. Kang WD, Kim SM. Human papillomavirus genotyping as a reliable prognostic marker of recurrence after loop electrosurgical excision procedure for high-grade cervical intraepithelial neoplasia (CIN2-3) especially in postmenopausal women. Menopause (New York, NY). 2016;23(1):81–86. doi:10.1097/GME.0000000000000488
16. Banister CE, Messersmith AR, Cai B, et al. Disparity in the persistence of high-risk human papillomavirus genotypes between African American and European American women of college age. J Infect Dis. 2015;211(1):100–108. doi:10.1093/infdis/jiu394
17. Kreimer AR, Guido RS, Solomon D, et al. Human papillomavirus testing following loop electrosurgical excision procedure identifies women at risk for posttreatment cervical intraepithelial neoplasia grade 2 or 3 disease. Cancer Epidemiol Biomarkers Prev. 2006;15(5):908–914. doi:10.1158/1055-9965.EPI-05-0845
18. Heymans J, Benoy IH, Poppe W, Depuydt CE. Type-specific HPV geno-typing improves detection of recurrent high-grade cervical neoplasia after conisation. Int J Cancer. 2011;129(4):903–909. doi:10.1002/ijc.v129.4
19. Arbyn M, Ronco G, Anttila A, et al. Evidence regarding human papillomavirus testing in secondary prevention of cervical cancer. Vaccine. 2012;30(Suppl 5):F88–99. doi:10.1016/j.vaccine.2012.06.095
20. Tainio K, Athanasiou A, Tikkinen KAO, et al. Clinical course of untreated cervical intraepithelial neoplasia grade 2 under active surveillance: systematic review and meta-analysis. BMJ (Clinical Research Ed). 2018;360:k499. doi:10.1136/bmj.k499
21. Malagon T, Kulasingam S, Mayrand MH, et al. Age at last screening and remaining lifetime risk of cervical cancer in older, unvaccinated, HPV-negative women: a modelling study. Lancet Oncol. 2018;19(12):1569–1578. doi:10.1016/S1470-2045(18)30536-9
22. Pallis AG, Fortpied C, Wedding U, et al. EORTC elderly task force position paper: approach to the older cancer patient. Eur J Cancer (Oxford, England: 1990). 2010;46(9):1502–1513. doi:10.1016/j.ejca.2010.02.022
23. Jørgensen TL, Teiblum S, Paludan M, et al. Significance of age and comorbidity on treatment modality, treatment adherence, and prognosis in elderly ovarian cancer patients. Gynecol Oncol. 2012;127(2):367–374. doi:10.1016/j.ygyno.2012.07.001
24. Chéreau E, Ballester M, Selle F, Rouzier R, Dara E. Ovarian cancer in the elderly: impact of surgery on morbidity and survival. Eur J Surg Oncol. 2011. 37(6):0–542.
25. Vitale SG, Capriglione S, Zito G, et al. Management of endometrial, ovarian and cervical cancer in the elderly: current approach to a challenging condition. Arch Gynecol Obstet. 2019;299(2):299–315. doi:10.1007/s00404-018-5006-z
26. Shimamoto K, Saito T, Kitade S, et al. A study of treatments and outcomes in elderly women with cervical cancer. Eur J Obstet Gynecol Reprod Biol. 2018;228:174–179. doi:10.1016/j.ejogrb.2018.06.032
27. Rebolj M, vanBallegooijen M, Lynge E, et al. Incidence of cervical cancer after several negative smear results by age 50: prospective observational study. BMJ. 2009;338.
28. Gustafsson L, Sparén P, Gustafsson M, Pettersson B, Adami HO. Low efficiency of cytologic screening for cancer in situ of the cervix in older women. Int J Cancer. 1996;63(6):804–809. doi:10.1002/ijc.2910630610
29. Rositch AF, Silver MI, Burke A, et al. The correlation between human papillomavirus positivity and abnormal cervical cytology result differs by age among perimenopausal women. J Low Genit Tract Dis. 2013;17(1):38–47. doi:10.1097/LGT.0b013e3182503402
30. Rositch AF, Koshiol J, Hudgens MG, et al. Patterns of persistent genital human papillomavirus infection among women worldwide: a literature review and meta-analysis. Int J Cancer. 2013;133(6):1271–1285. doi:10.1002/ijc.v133.6
31. Molano M, Van den Brule A, Plummer M, et al. Determinants of clearance of human papillomavirus infections in Colombian women with normal cytology: a population-based, 5-year follow-up study. Am J Epidemiol. 2003;158(5):486–494. doi:10.1093/aje/kwg171
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.


