Back to Journals » Cancer Management and Research » Volume 10
Diagnostic and prognostic values of the mRNA expression of excision repair cross-complementation enzymes in hepatitis B virus-related hepatocellular carcinoma
Authors Yang L, Xu M, Cui CB, Wei PH, Wu SZ, Cen ZJ , Meng XX, Huang QG, Xie ZC
Received 3 July 2018
Accepted for publication 7 September 2018
Published 5 November 2018 Volume 2018:10 Pages 5313—5328
DOI https://doi.org/10.2147/CMAR.S179043
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 4
Editor who approved publication: Dr Antonella D'Anneo
Lu Yang,1,* Ming Xu,2,* Chuan-Bao Cui,1 Peng-Hai Wei,1 Shu-Zhi Wu,1 Zuo-Jie Cen,1 Xing-Xing Meng,1 Qiong-Guang Huang,1 Zhi-Chun Xie1
1Department of Epidemiology, Guangxi Medical University, Nanning 530021, Guangxi Zhuang Autonomous Region, People’s Republic of China; 2Department of Human Anatomy and Histology and Embryology, Qilu Medical University, Zibo 255213, Shandong Province, People’s Republic of China
*These authors contributed equally to this work
Background: The current study aims at using the whole genome expression profile chips for systematically investigating the diagnostic and prognostic values of excision repair cross-complementation (ERCC) genes in hepatitis B virus (HBV)-related hepatocellular carcinoma (HCC).
Materials and methods: Whole genome expression profile chips were obtained from the GSE14520. The receiver-operating characteristic (ROC) curve, survival analysis, and nomogram were used to investigate the diagnostic and prognostic values of ERCC genes. Investigation of the potential function of ERCC8 was carried out by gene set enrichment analysis (GSEA) and genome-wide coexpression analysis.
Results: ROC analysis suggests that six ERCC genes (ERCC1, ERCC2, ERCC3, ERCC4, ERCC5, and ERCC8) were dysregulated and may have potential to distinguish between HBV-related HCC tumor and paracancerous tissues (area under the curve of ROC ranged from 0.623 to 0.744). Survival analysis demonstrated that high ERCC8 expression was associated with a significantly decreased risk of recurrence (adjusted P=0.021; HR=0.643; 95% CI=0.442–0.937) and death (adjusted P=0.049; HR=0.631; 95% CI=0.399–0.998) in HBV-related HCC. Then, we also developed two nomograms for the HBV-related HCC individualized prognosis predictions. GSEA suggests that the high expression of ERCC8 may have involvement in the energy metabolism biological processes. As the genome-wide coexpression analysis and functional assessment of ERCC8 suggest, those coexpressed genes were significantly enriched in multiple biological processes of DNA damage and repair.
Conclusion: The present study indicates that six ERCC genes (ERCC1, ERCC2, ERCC3, ERCC4, ERCC5, and ERCC8) were dysregulated between HBV-related HCC tumor and paracancerous tissues and that the mRNA expression of ERCC8 may serve as a potential biomarker for the HBV-related HCC prognosis.
Keywords: HBV, hepatocellular carcinoma, diagnosis, prognosis, ERCC
Introduction
Liver cancer is one of the most frequent malignant tumors occurring in China; besides, it has emerged as the third leading cause of cancer death in China.1 According to cancer statistics in China in 2015, liver cancer is most frequently observed in males as well as in rural areas. Moreover, the temporal tendency of incidence rates in liver cancer is decreased.1 The histological type of most primary liver cancers is hepatocellular carcinoma (HCC).2 Hepatitis B virus (HBV) infection is identified as a risk factor for HCC, especially in China, where the infection rate of HBV is quite high. HBV is a single-stranded DNA virus, which can be integrated into the host genome, subsequently may activate the cellular proto-oncogenes or suppress growth-regulating genes in cis,3 and induce chromosomal instability or insertional mutations.4 HBV-induced DNA damage is one of the potential mechanisms that contribute to the hepatocarcinogenesis of HCC.5
The excision repair cross-complementation (ERCC) genes include ERCC1, ERCC2, ERCC3, ERCC4, ERCC5, ERCC6, and ERCC8, constituting a set of proteins that are involved in DNA repair. Nucleotide excision repair (NER) is considered as an extensively pivotal excision mechanism, which removes DNA damage caused by various kinds factors, and a key DNA repair pathway in eukaryotic cells, whereas the ERCC genes constitute the key genes in the NER pathway.6,7 As revealed by the investigations, the genetic variation or mutation of the NER pathway genes is likely to impact the cancer risk by affecting the DNA repair efficacy, and some nonsynonymous coding single-nucleotide polymorphisms or mutations in DNA repair genes or their regulatory sequences induce the phenotypic changes and are involved in cancer development.8 Previous reports of the ERCC genes, suggesting they were associated with multiple cancer prognoses,9–12 and polymorphisms of the ERCC genes also have significant association with the HCC development, progression, survival, and cancer susceptibility.5,13–15 However, the systematic investigation of the diagnostic and prognostic values of the expression level of ERCC genes in HCC continue to be unclear. Therefore, the aim of the present study was to use the whole genome expression data set from the Gene Expression Omnibus (GEO; https://www.ncbi.nlm.nih.gov/geo/) public database for the purpose of systematically investigating the diagnostic and prognostic values of ERCC genes in HBV-related HCC.
Materials and methods
Data processing
In order to perform the present study, the inclusion criteria of public data set were set as follows: 1) the patients were HBV-related HCC; 2) the information of clinical outcome and features should be available; and 3) the data are the whole genome expression profile data. Through the retrieval of the public database, only the data set of GSE14520 caters to the inclusion criteria. GSE14520 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE14520; accessed May 13, 2018), a data set from the GEO public database, was included in the present study. Calculation of the expression profile chip data set of GSE14520 was carried out with the use of the match probes package in the R platform; subsequent to that, the robust multi-array average (RMA) method in the R affy package was used to obtain probe set expression summaries.16,17 Details of the experiment protocols and data processing are available at https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSM362992. Moreover, just the data set of the Affymetrix HT Human Genome U133A Array (platform GPL3921) was included in the current study to avoid the batch effect. These patients having a clearly HBV infection were included in the current study, and those patients without complete prognostic information and other clinical features were excluded. Owing to the fact that the population included in the current study originates from the public databases and the present study does not contain any studies performed by any of the authors with human participants or animals, there is no need for approval by additional ethics committees.
Expression distribution and survival analysis of ERCC genes
Investigation of the expression distribution of ERCC genes was carried out with the help of the independent samples t-test and the Metabolic gEne RApid Visualizer (MERAV, http://merav.wi.mit.edu/, accessed May 13, 2018) online tool.18 Receiver-operating characteristic (ROC) curves were put to use for the investigation of the potential application values in distinct HBV-related HCC tumors and adjacent normal liver tissues. The Kaplan–Meier method, with a log-rank test, and the univariate and multivariate Cox proportional hazards regression models were employed for the purpose of comparing the prognosis between different groups. Stratified analysis was employed for the assessment of the prognostic values of prognostic ERCC genes in different strata of HBV-related HCCs. Joint effect survival analyses between prognostic ERCC genes and serum alpha-fetoprotein (AFP), which is a recognized prognostic biomarker in HCC, have also been carried out for investigating the combined effect on HCC prognosis prediction. The high- and low-expression groups of the ERCC family genes were cutoff in accordance with the median values.19–22
Nomogram construction
To investigate the individualized prognostic prediction and evaluation system, we also developed a nomogram for the evaluation of the prognostic risk of each patient, which was based on the prognostic clinical indicators and prognostic-related ERCC genes of Cox regression model. Besides that, evaluation of the nomogram was using the concordance index (c-index). The nomogram was carried out by an rms package (https://CRAN.R-project.org/package=rms), together with its dependent packages in the R platform, with visualization using gplots.23,24
Gene set enrichment analysis (GSEA)
In a bid to figure out the potential mechanism of prognostic-related ERCC genes participating in the HBV-related HCC prognosis, we further performed a GSEA (http://software.broadinstitute.org/gsea/index.jsp) to discover the potential biological processes and pathways involved in the different expression levels of ERCC genes.25,26 The Molecular Signatures Database of GSEA used the c2 (c2.all.v6.1.symbols.gmt) and c5 (c5.all.v6.1.symbols.gmt) reference gene sets.27 Results with a P-value below 0.05 and false discovery rate (FDR) below 0.25 were considered statistically significant.
Coexpression analysis
To further evaluate the function of ERCC8 in HBV-related HCC, we further carried out a genome-wide coexpression analysis of ERCC8 in the HBV-related HCC of the GSE14520 cohort. The Pearson’s correlation coefficient was used to evaluate the coexpression genes, and the Database for Annotation, Visualization and Integrated Discovery (DAVID; https://david.ncifcrf.gov/home.jsp, accessed May 13, 2018) v6.828,29 was used for the purpose of functional assessment.
Statistical analysis
FDR in the GSEA was used to estimate the probability that the normalized enrichment score represents a false-positive finding, which was calculated with the help of the Benjamini and Hochberg method. A P-value <0.05 was regarded as statistically significant. All the statistical analyses were performed using SPSS v20.0 (IBM Corporation, Armonk, NY, USA) and R3.3.1 (https://www.r-project.org/).
Results
Study population
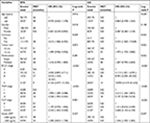
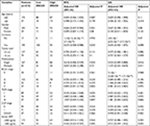
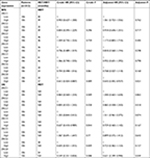
A total of 212 HBV-related HCC patients were included in the present study, coupled with 204 adjacent normal liver tissue specimens. Both the demographic and clinical features have been presented in Table 1. The clinical features including gender, cirrhosis, Barcelona Clinic Liver Cancer (BCLC) stage, TNM stage, and Cancer of the Liver Italian Program (CLIP) stage were significantly associated with the recurrence-free survival (RFS) of the HBV-related HCC, whereas the clinical features including cirrhosis, tumor size, BCLC stage, TNM stage, CLIP stage, and serum AFP were significantly correlated with overall survival (OS).
Expression distribution and survival analysis of ERCC genes
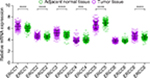
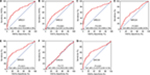
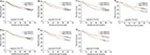
Through the comparison of the expression of ERCC genes in the HBV-related HCC tumor tissues and adjacent normal liver tissues, we observed that ERCC1, ERCC2, ERCC3, and ERCC8 were significantly upregulated in the HBV-related HCC tumor tissues, whereas ERCC4 and ERCC5 were significantly downregulated in the HBV-related HCC tumor tissues (Figure 1). These dysregulated genes are also observable in liver cancer using the MERAV online tool (Figure 2A–G). Thereafter, we performed the investigation of the potential application values of the ERCC genes in distinguishing the HBV-related HCC tumor tissues and adjacent normal liver tissues using the ROC curve. As the ROC analysis suggests, six ERCC genes (ERCC1, ERCC2, ERCC3, ERCC4, ERCC5, and ERCC8) could be a biomarker used to distinguish between HBV-related HCC tumor tissues and the adjacent normal liver tissues; the area under the curve of the ROC was in the range between 0.623 and 0.744 (Figure 3A–G).
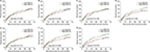
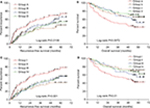
Survival analysis of ERCC family genes has been presented in Figures 4A–G and 5A–G; these findings provide the demonstration that the high ERCC8 expression significantly reduced the risk of recurrence (adjusted P=0.021; HR=0.643; 95% CI=0.442–0.937, Table 2, Figure 4G) and death (adjusted P=0.049; HR=0.631; 95% CI=0.399–0.998, Table 2, Figure 5G) in the HBV-related HCC. Stratification analysis was also used to assess the prognostic values of ERCC8 in the HBV-related HCC. The high expression of ERCC8 significantly reduced the risk of recurrence in the patients having cirrhosis, serum AFP >300 ng/mL, single nodules, and in the male patients (Table 3), whereas the high expression of ERCC8 significantly reduced the risk of death in the patients having single nodules, together with the male patients (Table 3). The joint effect survival analysis of ERCC8 and serum AFP suggests that the patients with high ERCC8 expression and a serum AFP ≤300 ng/mL (adjusted P=0.035; HR=0.583; 95% CI=0.354–0.962, Table 4, Figure 6A) or >300 ng/mL (adjusted P=0.004; HR=0.426; 95% CI=0.237–0.768, Table 4, Figure 6A) showed a significantly decreased risk of recurrence in the HBV-related HCC compared with patients having AFP >300 ng/mL and low ERCC8 expression. In contrast, the patients with high ERCC8 expression and AFP ≤300 ng/mL showed a significantly reduced risk of death in the HBV-related HCC (adjusted P=0.028; HR=0.499; 95% CI=0.268–0.927, Table 4, Figure 6B) compared with patients having AFP >300 ng/mL and low ERCC8 expression.
Thereafter, we also performed the verification of these joint effect survival analyses between ERCC8 and AFP mRNA expression level and demonstrated that the AFP mRNA expression also performed well in the HCC prognosis prediction. Patients with high ERCC8 expression and low (adjusted P=0.003; HR=0.443; 95% CI=0.259–0.757, Table 4, Figure 6C) or high (adjusted P=0.002; HR=0.418; 95% CI=0.238–0.734, Table 4, Figure 6C) AFP level showed a significantly decreased risk of recurrence in the HBV-related HCC compared with patients who had high AFP and low ERCC8 expression, as well as in patients with low AFP and ERCC8 expression (adjusted P=0.004; HR=0.425; 95% CI=0.238–0.757, Table 4, Figure 6C). With respect to OS, we observed that the patients with high ERCC8 expression and low AFP expression showed a significantly reduced risk of death in the HBV-related HCC (adjusted P=0.035; HR=0.487; 95% CI=0.249–0.952, Table 4, Figure 6D) compared with patients who had high AFP and low ERCC8 expression.
Nomogram construction
Thereafter, we performed the investigation of the potential application of ERCC8 mRNA expression in the individualized prognosis prediction. Those clinical features that were significantly correlated to RFS or OS were included in the development of nomogram for RFS and OS, correspondingly. The nomogram of the HBV-related HCC RFS was developed in accordance with the gender, cirrhosis, TMN stage, BCLC stage, CLIP stage, and ERCC8 expression (c-index [95% CI]=0.68 [0.63–0.74], Figure 7A), whereas the nomogram of HBV-related HCC OS was developed by tumor size, cirrhosis, TMN stage, BCLC stage, CLIP stage, AFP, and ERCC8 expression (c-index [95% CI]=0.72 [0.66–0.79], Figure 7B). In accordance with the nomogram for RFS and OS, we observed that the contribution of ERCC8 expression in the HBV-related HCC prognosis prediction was lower compared with some conventional clinical indicators; however, the expression of ERCC8 contributed much more than the serum AFP in the nomogram of the HBV-related HCC OS (Figure 7B).
GSEA of ERCC8
Investigation by GSEA using the c2 reference gene set indicates that the high expression of ERCC8 was significantly enriched in the NF-kappa-B and Wnt signaling pathway, as well as the P53 independent G1-S DNA damage checkpoint, TCA cycle and respiratory electron transport, and oxidative phosphorylation (Figure 8A–F, Table S1). GSEA with c5 indicates that the high expression of ERCC8 was significantly correlated to metabolically related biological processes, for instance, nucleoside triphosphate and cellular amine metabolic processes, and the generation of precursor metabolites and energy (Figure 8G–I, Table S2). In comparison, using the results of the c2 and c5 reference gene set, we observed that the high expression of ERCC8 may be involved in energy metabolism biological processes, which may have an impact on the prognosis of the HBV-related HCC.
Genome-wide coexpression analysis and function assessment of ERCC8
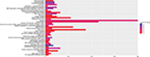
For the purpose of further exploring the function of ERCC8 in the HBV-related HCC, a genome-wide coexpression analysis was carried out. With respect to the multiple probes corresponding to one gene, we used the mean values of multiple probes to present the expression values of this gene in a genome-wide coexpression analysis. A total of 273 genes with |Pearson’s correlation coefficient|>0.4 and P<0.05 were taken into consideration as coexpressed with ERCC8 in the HBV-related HCC (Figure 9, Table S3). Thereafter, the functional assessment of the ERCC8-coexpressed genes was performed using DAVID v6.8. Gene Ontology (GO) term analysis suggested that ERCC8 and its coexpressed genes together are significantly enriched in the MAPK cascade, damaged DNA binding, DNA synthesis involved in DNA repair, and DNA damage checkpoints (Figure 10). In contrast, the Kyoto Encyclopedia of Genes and Genomes suggested that ERCC8 and its coexpressed genes were significantly enriched in the ribosome-signaling pathway (hsa03010, P=0.003, gene count=8).
Discussion
Through the review of the literature, we observed that extensive previous studies have focused on the investigation of the association between the ERCC gene polymorphisms and the risk of cancers30,31 or on the ERCC gene polymorphisms and cancer prognosis.32,33 Besides that, the ERCC gene polymorphisms have also been reported for having correlation with the prognosis of patients treated with chemotherapy.34–37 However, there are rare reports of the systematic investigations of the expression of ERCC in cancer diagnosis and prognosis.9,38,39
Previous studies substantiated that polymorphisms in ERCC2 and ERCC5 confer susceptibility to the acute myeloid leukemia (AML)40 and that polymorphisms in ERCC2 and XPC were significantly associated with the AML survival.41 Polymorphisms in ERCC5 also make contribution toward the susceptibility of lung cancer and squamous cell carcinomas of the oropharynx, larynx, and esophagus;42 head and neck;43 prostate cancer;44 HCC;15 and bladder cancer (BC).31 A study performed by Sakano et al also found that the genetic variants of ERCC5 contribute to the tumor invasiveness in BC.45 In addition, the polymorphisms ERCC2-rs1799793 and ERCC5-rs17655 may also contribute to the recurrence risk of squamous cell carcinoma of the oropharynx.37
There have also been reports of the prognostic values of ERCC genetic variants in multiple cancers. An investigation carried out by Liu et al revealed the fact that ERCC1-rs11615 is likely to be a useful genetic marker for the prediction of the osteosarcoma prognosis and that the CC genotype is correlated with a better clinical outcome.46 The work performed by Liu et al indicated that the XRCC1-399A/A genotype was significantly correlated with a better clinical outcome.47 Previous studies demonstrated that genetic variants of ERCC2 and ERCC5 may be the independent prognostic factors in the patients with melanoma32,48 and osteosarcoma.49 In addition, polymorphisms of ERCC5 were also correlated with the response to chemotherapy and the prognosis of osteosarcoma.50,51 Furthermore, polymorphisms of ERCC5 have also been reported to have an influence in multiple cancer prognosis, such as lung cancer,33,47 and colorectal cancer (CRC).52,53
The interindividual differences in the patients treated with chemotherapy mainly affected the efficiency of the DNA damage–repair system in the cells and then affected the clinical outcomes. Extensive studies have demonstrated that the ERCC genes were involved in the responses to chemotherapy and the prognosis of the patients with CRC36,54–56 as well as advanced non-small cell lung cancer (NSCLC).57,58 The ERCC5 polymorphism His46His was reported to be associated with the susceptibility of platinum-based chemotherapy in the advanced NSCLC35 and significantly affected clinical outcomes.58 The ERCC5 promoter polymorphism rs751402 was significantly associated with the chemotherapy response in the advanced NSCLC, and the AA genotype may contribute to increase the effect of chemotherapy response.59 Hu et al substantiated that the rs2296147 T allele and rs873601 G allele were associated with the better progression-free survival (PFS) and OS in the advanced NSCLC patients undergoing the platinum-based chemotherapy57; furthermore, the results of the rs2296147 T allele in OS of the advanced NSCLC patients undergoing platinum-based chemotherapy are also verifiable in another cohort.60 However, the rs1047768 TT genotype, which showed a significantly shortened median PFS and OS in the advanced NSCLC patients undergoing platinum-based chemotherapy in the study of Zhang et al,60 is unable to be verified in the cohort collected by Hu et al.57 With respect to the ERCC genetic variation in CRC response to platinum-based chemotherapy and therapeutic outcomes, multiple studies demonstrated that the ERCC5 polymorphisms may have an impact on the susceptibility and therapeutic outcomes to the colorectal cancer patients, who received oxaliplatin chemotherapy.36,54–56
By reviewing the above literature, we found that a considerable amount of literature has been published on the genetic variation of ERCC genes in cancer susceptibility, prognosis, and therapeutic outcomes; however, the expression of ERCC genes in cancer diagnostic and prognostic values were rarely mentioned. Several investigations verified the fact that the low expression of ERCC1 was significantly correlated with a better clinical outcome in patients with breast cancer,38 NSCLC,61 and ovarian cancer,9 as well as in advanced NSCLC treated with platinum-based chemotherapy.10 A systematic investigation of the ERCC genes in the ovarian cancer prognosis was carried out by Zhao et al using the Kaplan–Meier Plotter online tool and revealed that the high expression of ERCC8 and ERCC1 had correlations with a poor OS, whereas the high expression of ERCC4 was associated with a better OS.9 In our current study, we observed that the high expression of ERCC8 was significantly correlated with a better RFS and OS in the HBV-related HCC after hepatectomy, and our current study was the first to investigate the potential prognostic application values of ERCC8 in the HBV-related HCC. Comprehensive survival analysis of stratification analysis and joint effect survival analysis indicates that ERCC8 may be served as an independent prognostic indicator for the HBV-related HCC; the nomograms indicate that the combination of ERCC8 and traditional prognostic indicators may have an application value in the individualized prognosis prediction. It is worth noting that the results of ERCC8 obtained in our current study were contradictory to the results of the study carried out by Zhao et al in the ovarian cancer prognosis.9 The possible causes of the above inconsistency results suggested that the survival analysis in the study by Zhao et al was derived from the Kaplan–Meier method, which was a univariate survival analysis and that the results obtained from such kinds of statistical methods may be unreliable. However, the key advantage of the results of the current study was assessed by a multivariate Cox proportional hazards regression model, and the statistical method was more reliable than the Kaplan–Meier method in survival analysis. Another possible reason may be that ERCC8 plays different roles in both the ovarian cancer and HBV-related HCC. However, these results still require to be further verified in a larger sample, and the hypotheses also need further experimental validation.
The current study has some limitations that need to clarification. First, since the data set is from the public database, the clinical information in the GSE14520 was constrained. Consequently, we are incapable of attaining complete clinical information with which to carry out a comprehensive prognostic analysis; the incomplete clinical information, such as postoperative treatment, may influence prognosis. Second, the raw values of serum AFP level in GSE14520 were not provided as well, accordingly, we were unable to carry out a joint effect survival analysis in the serum with an AFP cutoff 400 ng/mL, which is a recognized cutoff value. Third, through the retrieval of the public database, only the GSE14520 data set caters to the requirements of the current study. That is why the results of the present study are derived from a single cohort, and additional cohorts to verify the results are needed.
Despite these limitations, our current study is the first to systematically investigate the diagnostic and prognostic values of ERCC genes in the HBV-related HCC. As our findings suggest, the dysregulated ERCC genes may have a potential application in the HBV-related HCC diagnosis and ERCC8 may be a potential novel indicator for the HBV-related HCC prognosis prediction. We also developed two nomograms for the individualized prognosis prediction and investigated the potential mechanisms and functions of ERCC8 in the HBV-related HCC using the GSEA and genome-wide coexpression analysis approaches.
Conclusion
The present study indicates that six ERCC genes (ERCC1, ERCC2, ERCC3, ERCC4, ERCC5, and ERCC8) were dysregulated between the HBV-related HCC tumor and paracancerous tissues, and mRNA expression of ERCC8 may act as a potential biomarker for the HBV-related HCC prognosis.
Acknowledgments
This work was supported in part by the National Natural Science Foundation of China (Grant NO.81460512) and Shandong Province Education Department Colleges and Universities Science and Technology Plan Project (J17KB088). The authors thank the contributors of the GSE14520 for sharing the HBV-related HCC expression profile data set on open access. In addition, we would like to acknowledge the helpful comments on this article received from our reviewers.
Disclosure
The authors report no conflicts of interest in this work.
References
Chen W, Zheng R, Baade PD, et al. Cancer statistics in China, 2015. CA Cancer J Clin. 2016;66(2):115–132. | ||
Tang D, Nagano H, Nakamura M, et al. Clinical and pathological features of Allen’s type C classification of resected combined hepatocellular and cholangiocarcinoma: a comparative study with hepatocellular carcinoma and cholangiocellular carcinoma. J Gastrointest Surg. 2006;10(7):987–998. | ||
Mckillop IH, Moran DM, Jin X, Koniaris LG. Molecular pathogenesis of hepatocellular carcinoma. J Surg Res. 2006;136(1):125–135. | ||
Pang R, Tse E, Poon RT. Molecular pathways in hepatocellular carcinoma. Cancer Lett. 2006;240(2):157–169. | ||
Jung SW, Park NH, Shin JW, et al. Polymorphisms of DNA repair genes in Korean hepatocellular carcinoma patients with chronic hepatitis B: possible implications on survival. J Hepatol. 2012;57(3):621–627. | ||
Le May N, Egly J-M, Coin F. True lies: the double life of the nucleotide excision repair factors in transcription and DNA repair. J Nucleic Acids. 2010;2010:1–10. | ||
Hou SM, Fält S, Angelini S, et al. The XPD variant alleles are associated with increased aromatic DNA adduct level and lung cancer risk. Carcinogenesis. 2002;23(4):599–603. | ||
Karahalil B, Bohr VA, Wilson DM. Impact of DNA polymorphisms in key DNA base excision repair proteins on cancer risk. Hum Exp Toxicol. 2012;31(10):981–1005. | ||
Zhao M, Li S, Zhou L, Shen Q, Zhu H, Zhu X. Prognostic values of excision repair cross-complementing genes mRNA expression in ovarian cancer patients. Life Sci. 2018;194:34–39. | ||
Qin X, Yao W, Li W, et al. ERCC1 and BRCA1 mRNA expressions are associated with clinical outcome of non-small cell lung cancer treated with platinum-based chemotherapy. Tumour Biol. 2014;35(5):4697–4704. | ||
Sun JM, Ahn MJ, Park MJ, et al. Expression of excision repair cross-complementation group 1 as predictive marker for nasopharyngeal cancer treated with concurrent chemoradiotherapy. Int J Rad Oncol Biol Phys. 2011;80(3):655–660. | ||
Grimminger PP, Shi M, Barrett C, et al. TS and ERCC-1 mRNA expressions and clinical outcome in patients with metastatic colon cancer in CONFIRM-1 and -2 clinical trials. Pharmacogenomics J. 2012;12(5):404–411. | ||
Li LM, Zeng XY, Ji L, et al. [Association of XPC and XPG polymorphisms with the risk of hepatocellular carcinoma] XPC hé XPG duō tài xìng yǔ gān xìbāo ái fēngxiǎn de guānxì. Zhonghua Gan Zang Bing Za Zhi. 2010;18(4):271–275. Chinese. | ||
Sakurada T, Yoshikawa M, Sunaga M, et al. Expression of drug-resistant factor genes in hepatocellular carcinoma patients undergoing chemotherapy with platinum complex by arterial infusion. Pharmaceutics. 2010;2(3):300–312. | ||
Yoon AJ, Kuo WH, Lin CW, Yang SF. Role of ERCC5 polymorphism in risk of hepatocellular carcinoma. Oncol Lett. 2011;2(5):911–914. | ||
Roessler S, Long EL, Budhu A, et al. Integrative genomic identification of genes on 8p associated with hepatocellular carcinoma progression and patient survival. Gastroenterology. 2012;142(4):957–966. | ||
Roessler S, Jia HL, Budhu A, et al. A unique metastasis gene signature enables prediction of tumor relapse in early-stage hepatocellular carcinoma patients. Cancer Res. 2010;70(24):10202–10212. | ||
Shaul YD, Yuan B, Thiru P, et al. MERAV: a tool for comparing gene expression across human tissues and cell types. Nucleic Acids Res. 2016;44(D1):D560–D566. | ||
Liao X, Huang K, Huang R, et al. Genome-scale analysis to identify prognostic markers in patients with early-stage pancreatic ductal adenocarcinoma after pancreaticoduodenectomy. Onco Targets Ther. 2017;10:4493–4506. | ||
Liao X, Huang R, Liu X, et al. Distinct prognostic values of alcohol dehydrogenase mRNA expression in pancreatic adenocarcinoma. Onco Targets Ther. 2017;10:3719–3732. | ||
Liao X, Liu X, Yang C, et al. Distinct diagnostic and prognostic values of minichromosome maintenance gene expression in patients with hepatocellular carcinoma. J Cancer. 2018;9(13):2357–2373. | ||
Huang R, Liao X, Li Q. Identification and validation of potential prognostic gene biomarkers for predicting survival in patients with acute myeloid leukemia. Onco Targets Ther. 2017;10:5243–5254. | ||
Liao X, Zhu G, Huang R, et al. Identification of potential prognostic microRNA biomarkers for predicting survival in patients with hepatocellular carcinoma. Cancer Manag Res. 2018;10:787–803. | ||
Liao X, Yang C, Huang R, et al. Identification of potential prognostic long non-coding RNA biomarkers for predicting survival in patients with hepatocellular carcinoma. Cell Physiol Biochem. 2018;48(5):1854–1869. | ||
Subramanian A, Tamayo P, Mootha VK, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci. 2005;102(43):15545–15550. | ||
Mootha VK, Lindgren CM, Eriksson KF, et al. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet. 2003;34(3):267–273. | ||
Liberzon A, Birger C, Thorvaldsdóttir H, Ghandi M, Mesirov JP, Tamayo P. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 2015;1(6):417–425. | ||
Dennis G, Sherman BT, Hosack DA, et al. DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome Biol. 2003;4(5):P3. | ||
Huang Daw, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4(1):44–57. | ||
Qiu L, Wang Z, Shi X, Wang Z. Associations between XPC polymorphisms and risk of cancers: a meta-analysis. Eur J Cancer. 2008;44(15):2241–2253. | ||
García-Closas M, Malats N, Real FX, et al. Genetic variation in the nucleotide excision repair pathway and bladder cancer risk. Cancer Epidemiol Biomarkers Prev. 2006;15(3):536–542. | ||
Li C, Yin M, Wang LE, et al. Polymorphisms of nucleotide excision repair genes predict melanoma survival. J Invest Dermatol. 2013;133(7):1813–1821. | ||
Matakidou A, El Galta R, Webb EL, et al. Genetic variation in the DNA repair genes is predictive of outcome in lung cancer. Hum Mol Genet. 2007;16(19):2333–2340. | ||
Schöffski P, Taron M, Jimeno J, et al. Predictive impact of DNA repair functionality on clinical outcome of advanced sarcoma patients treated with trabectedin: a retrospective multicentric study. Eur J Cancer. 2011;47(7):1006–1012. | ||
Sun X, Li F, Sun N, et al. Polymorphisms in XRCC1 and XPG and response to platinum-based chemotherapy in advanced non-small cell lung cancer patients. Lung Cancer. 2009;65(2):230–236. | ||
Kweekel DM, Antonini NF, Nortier JW, Punt CJ, Gelderblom H, Guchelaar HJ. Explorative study to identify novel candidate genes related to oxaliplatin efficacy and toxicity using a DNA repair array. Br J Cancer. 2009;101(2):357–362. | ||
Song X, Sturgis EM, Jin L, Wang Z, Wei Q, Li G. Variants in nucleotide excision repair core genes and susceptibility to recurrence of squamous cell carcinoma of the oropharynx. Int J Cancer. 2013;133(3):695–704. | ||
Goyal S, Parikh RR, Green C. Clinicopathologic significance of excision repair cross-complementation 1 expression in patients treated with breast-conserving surgery and radiation therapy. Int J Rad Oncol, Biol, Phys. 2010;76(3):679–684. | ||
Sasaki S, Watanabe T, Nakayama H. Analysis of the mRNA expression of chemotherapy-related genes in colorectal carcinoma using the Danenberg tumor profile method. J Oncol. 2013;2013:386906–3869067. | ||
Ba˘nescu C, Iancu M, Trifa AP, et al. Influence of XPC, XPD, XPF, and XPG gene polymorphisms on the risk and the outcome of acute myeloid leukemia in a Romanian population. Tumour Biol. 2016;37(7):9357–9366. | ||
Strom SS, Estey E, Outschoorn UM, Garcia-Manero G. Acute myeloid leukemia outcome: role of nucleotide excision repair polymorphisms in intermediate risk patients. Leuk Lymphoma. 2010;51(4):598–605. | ||
Cui Y, Morgenstern H, Greenland S, et al. Polymorphism of Xeroderma Pigmentosum group G and the risk of lung cancer and squamous cell carcinomas of the oropharynx, larynx and esophagus. Int J Cancer. 2006;118(3):714–720. | ||
Ma H, Yu H, Liu Z, Wang LE, Sturgis EM, Wei Q. Polymorphisms of XPG/ERCC5 and risk of squamous cell carcinoma of the head and neck. Pharmacogenet Genomics. 2012;22(1):50–57. | ||
Zhang XJ, Liu P, Zhu F. Polymorphisms of DNA repair-related genes with susceptibility and prognosis of prostate cancer. Genet Mol Res. 2014;13(2):4419–4424. | ||
Sakano S, Kumar R, Larsson P, et al. A single-nucleotide polymorphism in the XPG gene, and tumour stage, grade, and clinical course in patients with nonmuscle-invasive neoplasms of the urinary bladder. BJU Int. 2006;97(4):847–851. | ||
Liu X, Zhang Z, Deng C, Tian Y, Ma X. Meta-analysis showing that ERCC1 polymorphism is predictive of osteosarcoma prognosis. Oncotarget. 2017;8(37):62769–62779. | ||
Liu D, Wu J, Shi GY, Zhou HF, Yu Y. Role of XRCC1 and ERCC5 polymorphisms on clinical outcomes in advanced non-small cell lung cancer. Genet Mol Res. 2014;13(2):3100–3107. | ||
Schrama D, Scherer D, Schneider M, et al. ERCC5 p.Asp1104His and ERCC2 p.Lys751Gln polymorphisms are independent prognostic factors for the clinical course of melanoma. J Invest Dermatol. 2011;131(6):1280–1290. | ||
Li J, Liu S, Wang W, et al. ERCC polymorphisms and prognosis of patients with osteosarcoma. Tumour Biol. 2014;35(10):10129–10136. | ||
Bai SB, Chen HX, Bao YX, Luo X, Zhong JJ. Predictive impact of common variations in DNA repair genes on clinical outcome of osteosarcoma. Asian Pac J Cancer Prev. 2013;14(6):3677–3680. | ||
Sun XH, Hou WG, Zhao HX, Zhao YL, Ma C, Liu Y. Single nucleotide polymorphisms in the NER pathway and clinical outcome of patients with bone malignant tumors. Asian Pac J Cancer Prev. 2013;14(3):2049–2052. | ||
Negandhi AA, Hyde A, Dicks E, et al. MTHFR Glu429Ala and ERCC5 His46His polymorphisms are associated with prognosis in colorectal cancer patients: analysis of two independent cohorts from Newfoundland. PLoS One. 2013;8(4):e61469. | ||
Wang F, Zhang SD, Xu HM, et al. XPG rs2296147 T>C polymorphism predicted clinical outcome in colorectal cancer. Oncotarget. 2016;7(10):11724–11732. | ||
Liu D, Wu HZ, Zhang YN, et al. DNA repair genes XPC, XPG polymorphisms: relation to the risk of colorectal carcinoma and therapeutic outcome with oxaliplatin-based adjuvant chemotherapy. Mol Carcinog. 2012;51(Suppl 1):E83–E93. | ||
Chen J, Luo X, Xie G, et al. Functional analysis of SNPs in the ERCC5 promoter in advanced colorectal cancer patients treated with oxaliplatin-based chemotherapy. Medicine. 2016;95(19):e3652. | ||
Sun K, Gong A, Liang P. Predictive impact of genetic polymorphisms in DNA repair genes on susceptibility and therapeutic outcomes to colorectal cancer patients. Tumour Biol. 2015;36(3):1549–1559. | ||
Hu W, Pan J, Zhao P, Yang G, Yang S. Genetic polymorphisms in XPG could predict clinical outcome of platinum-based chemotherapy for advanced non-small cell lung cancer. Tumour Biol. 2014;35(6):5561–5567. | ||
Jin ZY, Zhao XT, Zhang LN, Wang Y, Yue WT, Xu SF. Effects of polymorphisms in the XRCC1, XRCC3, and XPG genes on clinical outcomes of platinum-based chemotherapy for treatment of non-small cell lung cancer. Genet Mol Res. 2014;13(3):7617–7625. | ||
He C, Duan Z, Li P, Xu Q, Yuan Y. Role of ERCC5 promoter polymorphisms in response to platinum-based chemotherapy in patients with advanced non-small-cell lung cancer. Anticancer Drugs. 2013;24(3):300–305. | ||
Zhang T, Sun J, Lv M, et al. XPG is predictive gene of clinical outcome in advanced non-small-cell lung cancer with platinum drug therapy. Asian Pac J Cancer Prev. 2013;14(2):701–705. | ||
Jeong YH, Lee CK, Jo K, et al. Correlation analysis and prognostic impact of (18)F-FDG PET and excision repair cross-complementation group 1 (ERCC-1) expression in non-small cell lung cancer. Nucl Med Mol Imaging. 2015;49(2):108–114. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.