Back to Journals » Drug Design, Development and Therapy » Volume 9
Construction of interference vector targeting Ep-CAM gene and its effects on colorectal cancer cell proliferation
Authors Qi Y, Zhou F, Zhang L, Liu L, Xu H, Guo H
Received 15 February 2015
Accepted for publication 10 March 2015
Published 14 May 2015 Volume 2015:9 Pages 2647—2652
DOI https://doi.org/10.2147/DDDT.S82917
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Prof. Dr. Wei Duan
Yanmei Qi,1 Fengqiang Zhou,2 Lu Zhang,2 Lei Liu,2 Hong Xu,2 Huiguang Guo2
1Department of Gastroenterology, 2Department of General Surgery, Binzhou People’s Hospital, Binzhou, Shandong, People’s Republic of China
Background: Prior study indicates that abnormal protein expression and functional changes in the development and progression of colorectal cancer is related to gene expression. The aim of this study was to construct an interference plasmid targeting the Ep-CAM gene and to investigate its effects on the proliferation of colorectal cancer cells.
Methods: In this study, HT-29 and HCT-116 colorectal cancer cell lines were selected as cell models. The double-stranded micro (mi)RNA oligo was inserted into the pcDNATM6.2-GW/EmGFPmiR vector, which is an expression of miRNA. Lipofectamine™ 2000 was used to transfer plasmid into the empty plasmid group (transfected pcDNATM6.2-GW/EmGFPmiR-neg) and the interference group (transfected pcDNATM6.2-GW/EmGFPmiR-Ep-CAM-1), respectively. Meanwhile, the nontransferred HT-29 and HCT-116 acts as the blank control group. Reverse transcription polymerase chain reaction (RT-PCR) was used to detect the transfection efficiency. Western blot was used to detect Ep-CAM protein expression. The cell proliferation in each group was detected by using 3-(4,5-Dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay.
Results: The results indicated that the Ep-CAM messenger (m)RNA expression in the interference group was lower significantly compared with that of the empty plasmid group and control group (P<0.01). Western blot analysis results showed that Ep-CAM protein expression was significantly lower in interference group compared with that of the empty plasmid group and the control group (P<0.01). MTT assay results demonstrated that the proliferation ability of cells in the interference group was significantly inhibited compared with the two other groups (P<0.05).
Conclusion: Silencing of Ep-CAM can significantly inhibit the proliferation of colorectal cancer cells.
Keywords: epidermal growth factor, Ep-CAM, colorectal cancer, vector construction
Introduction
Colorectal cancer is a common gastrointestinal malignancy. The morbidity of colorectal cancer has been rising in recent years and is a serious threat to health.1 Abnormal expression and functional changes in the development and progression of colorectal cancer are related to gene and protein expression. The serious study of heterogeneity of colorectal cancer related molecular is an important strategy for the discovery of new cancer therapies and therapeutic targets.2
Colorectal cancer is a common malignant tumor of the digestive tract.3 The development of colorectal cancer always with the characteristics of abnormal expression of proteins or abnormal function of some genes. In-depth study of molecular events in the heterogeneity is an important method to determine the tumor treatment and new therapeutic targets.4,5
Ep-CAM is one of the classic tumor-associated antigens and is mainly involved in the interactions between the matrix and cells and cell differentiation, cell cycle regulation, cell signaling, and the cell migration process.3,6 Recently, scientists discovered that the expression of Ep-CAM was increased and also, that the risk of metastasis and recurrence were also increased in colorectal carcinoma patients.7
In addition, the previous study indicates the relationship between Ep-CAM and tumor is close – many tumors have been found with the expression of Ep-CAM.8 However, a number of reports have confirmed the expression of Ep-CAM in gastric and breast cancer proliferation and migration but also, that it influences the cell cycle of these tumors.9,10 So far, for breast cancer and other tumors, it has been shown that Ep-CAM can be used clinically, for judging prognosis or as a diagnostic marker.11 The interaction between Ep-CAM research and the development of colorectal cancer invasion and metastases are less studied – we found only one article on analysis of the expression of the Ep-CAM gene in colon cancer cell lines.4
According to the previous work, we hypothesized that inhibiting the Ep-CAM gene could be helpful to inhibit the cell proliferation of colorectal carcinoma cells. Therefore, we established the Ep-CAM-interference plasmid, which expresses the micro (mi)RNA blocking the Ep-CAM gene. The expression of the Ep-CAM gene in colorectal carcinoma cells was examined, and the cell proliferation was also detected to confirm the anticancer effects of the Ep-CAM-interference plasmid. To our knowledge, few studies focus on Ep-CAM in tumor metastasis and the mechanism. The present study could provide an effective targeting method for colorectal carcinoma.
Materials and methods
Materials
Colon cancer cell lines HT-29 and HCT-116 were purchased from the Shanghai Cell Biology Research Institute of the Chinese Academy of Sciences, Shanghai, People’s Republic of China. β-catenin and Ep-CAM interference of plasmid were purchased from Shanghai Genechem Co. Ltd. G418 and RNA enzymes were purchased from Sigma-Aldrich Corp (St Louis, MO, USA). Roswell Park Memorial Institute (RPMI) 1640 medium were purchased from HyClone company (Thermo Fisher Scientific Inc, Waltham, MA, USA) Liposome transfection reagent Lipofectamine™ 2000 was purchased from Invitrogen (Life Technologies Corp, Carlsbad CA, USA). RNA extraction reagent TRIzol® was purchased from Shanghai Biological Engineering Co, Ltd (Shanghai, People’s Republic of China). PrimeScript III™ reverse transcription kit was purchased from Takara Biotechnology (Dalian) Co Ltd. (Dalian, People’s Republic of China). DNA fluorescent dye SYBR® Green was purchased from Toyobo Co, Osaka, Japan. Methyl thiazolyl tetrazolium and dimethyl sulfoxide (DMSO) were purchased from Beijing Suolaibao Co (Beijing, People’s Republic of China).
Instruments
The Millicell® cell culture chamber (EMD Millipore, Billerica, MA, USA), culture plates (Corning Inc, Corning, NY, USA), NV2500E incubator (Heracell®; Thermo Fisher Scientific Inc), LightCycler2.0 polymerase chain reaction (PCR) system (F. Hoffman-La Roche Ltd, Basel, Switzerland), the FACSCalibur flow cytometry (BD Biosciences, Franklin Lakes, NJ, USA), and a 20PR-52D type low-speed centrifuge (Hitachi, Tokyo, Japan) were used in this study.
The construction of Ep-CAM-interference plasmid
According to the gene sequence of Genbank accession number BC014785 (Invitrogen; Life Technologies Corp), the design and synthesis of the target gene Ep-CAM single-stranded sequence was as follows: Positive-sense strand: 5′-TGCTGAACTGAAGTACACTGGCATTGGTTTTGGCCACTGACTGACCAATGCCAGTACTTCAGTT-3′. Antisense strand: 5′-CCTGAACTGAAGTACTGGCATTGGTCAGTCAGTGGCCAA AACCAATGCCAGTGTACTTCAGTTC-3′. Recombinant cloning was performed with a vector construction kit. Using the BLOCK-iT™ Pol II miRNAi Expression Vector Kit with EmGFP, the double stranded miRNA oligo was inserted into the miRNA expression vector of pcDNATM6.2-GW/EmGFPmiR. The recombinant plasmid was named pcDNA6.2-GW/EmGFPmiR-Ep-CAM-l.
Ep-CAM plasmid transfection and cell block
The day before, transfection was performed by 0.1% trypsin logarithmic digestion of cells, 2×105/well in a six-well plate, with 10% fetal bovine serum in RPMI 1640 medium at 37°C, 5% CO2. When the cells were grown to confluence of 80%–90%, according to Lipofectamine™ 2000 manual, the empty plasmid group (transfection of pcDNATM6.2-GW/EmGFPmiR-NEG) and interference plasmid group (transfected with pcDNATM6.2-GW/EmGFPmiR-EPCA M-1) were given the plasmid transfection respectively. Forty-eight hours after transfection, 1:10 passage, good adherent cells (600 μg/mL) after G418 screening was performed. Two weeks after transfection, multiple expression of green fluorescent protein was used to choose single resistant clones under fluorescence microscope. The clone that stable expression the pcDNATM6.2-GW/EmGFPmiR-neg cells (empty plasmid group) and pcDNATM6.2-GW/EmGFPmiR-Ep-CAM-1 (interference group) were successfully established. Inhibiting the activity of the Ep-CAM gene, the Ep-CAM content was detected. The interference group was analyzed by using the reverse transcription (RT)-PCR.
Primer sequences design
Primer sequences of Ep-CAM: Ep-CAM, with primers (Jierui Company, Shanghai, People’s Republic of China), was amplified by PCR. The primers were 5′ACCTGCTCTGAGCGAGTGAGAACCT-3′ and 5′GACCAGGATCCAGATCCA GTTG-3′. The amplified fragment length was 337 bp. With β-actin (actin) as internal reference, the upstream primer was 5′-AACAGAGGCACCT CACCCT-3′, and the downstream primer was 5′-TACATGTACATGGCTGG-GGTGTYGAA-3′. The amplified fragment length was 214 bp. The reaction conditions were: 95°C denaturing 1 minute, denaturation 30 seconds at 94°C, 61°C (Ep-CAM)/57°C (β-actin) annealing 40 seconds, 72°C extend for 40 seconds for a total of 35 cycles, 72°C 10 minutes. The reaction product was confirmed by 1.5% agarose gel electrophoresis.
Collecting the cells in each group, according to the TRIzol manual, to extract RNA, complementary (c)DNA was synthesized using the PrimeScript III™ reverse transcription kit. Real-time PCR reaction was finished with SYBR Green reaction conditions. Annealing temperature and primer concentration in the reaction were optimized to obtain the optimal reaction system of β-catenin, cyclin D1, and GAPDH genes.
Testing index and method
The first evaluation of Ep-CAM in colorectal cancer cells was by western blot and RT-PCR. As well, 3-(4,5-Dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) was used to determine the cell proliferation in each group, based on Ep-CAM silence. Firstly, the logarithmic growth phase cells were inoculated with 10% fetal calf serum, and forming the single cell suspension, with 1,000 to 10,000 cells/well inoculated into 96-well plates with pore volume of 200 μL. Cells were cultured for 3–5 days (according to the objective and requirement culture time). Each well was treated with MTT solution (5 mg/mL, phosphate buffered saline pH =7.4 20 μL) for 4 hours. Then the supernatant was discarded. The suspended cells were centrifugated and then discarded into suction hole 150 μL DMSO was added into each hole 10 minute oscillation, which can be fully crystallization. The hole was measured by the enzyme-linked immunosorbent assay (ELISA) and monitor light absorption value (at the wavelength of 490 nm), record the results.
Statistical analysis
For quantitative data, all results were expressed as the mean ± standard error of the mean (SEM). Statistical significance between groups was determined by using SPSS 13 software (SPSS, Chicago, IL, USA). Each experiment was repeated at least three times, and P-value of <0.05 was considered as statistically significant.
Results
Detection of transfection efficiency
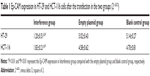
The transfection efficiency was detected by fluorescence quantitative PCR in two cell lines. The Ep-CAM expression in the “interference” group in the transfected HT-29 and HCT-116 cell lines was significantly lower than that in the empty plasmid group and blank control group (Figure 1) (P<0.001). Meanwhile, there were no significant difference between the empty plasmid group and blank control group (Figure 1 and Table 1) (P=0.37).
Ep-CAM protein expression was inhibited the gene silence
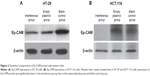
The Ep-CAM protein expression in HT-29 and HCT-116 cell lines was also examined using the western blot assay. The results showed that the Ep-CAM expression in the interference group was significantly lower compared with that of the empty vector group and blank control group (Figure 2) (P<0.05). Furthermore, there was no significant difference for Ep-CAM expression between the empty plasmid group and blank control group (Figure 2) (P<0.05).
Silence of Ep-CAM inhibits the proliferation of colorectal cancer cells
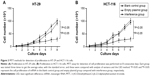
The MTT assay was employed to test the cell proliferation ability of HT-29 and HCT-116 cell lines for 8 days. The results were plotted as a curve. The results showed the cell proliferation in all of the three groups gradually increased with time in the two cell lines. The proliferation ability of cells in the interference group was significantly inhibited compared with the two other groups (Figure 3) (P<0.05). Furthermore, there was no significant difference between the normal group and the group of cell lines transfected with empty plasmid (Figure 3 and Table 2) (P>0.05). From these results, we can see that the silence of Ep-CAM can significantly inhibit the proliferation of colorectal cancer cells.
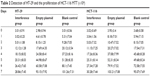  | Table 2 Detection of HT-29 and the proliferation of HCT-116 MTT (×104) |
Discussion
This study used a small RNA-mediated gene silencing method to suppress the Ep-CAM gene in colorectal cancer cell lines and to study its role in colorectal cancer. In this study, we established that the plasmid that could inhibit the Ep-CAM gene in colorectal carcinoma cells. The messenger (m)RNA expression and protein expression of Ep-CAM gene were inhibited in the interference group compared with the empty plasmid and blank control group. Previous study also revealed that the high expression of Ep-CAM is associated with a high degree of malignancy, metastasis, and risk of recurrence.6 Inhibition of the Ep-CAM gene could inhibit the proliferation of the HT-29 and HCT-116 cells efficiently, which is strong evidence for tumor-targeting therapy. In the study of the topic, current research on Ep-CAM is sparse, with no studies on the relationship between pathways.
The role of the Ep-CAM signal transduction pathway was seldomly described in the previous studies.12,13 A previous study on breast cancer showed that Ep-CAM transfected small interfering (si)RNA cell line indicates stronger invasion and migration ability compared to the untransfected cell line. This suggests that Ep-CAM can affect cell migration, invasion, and proliferation by enhancing E-cadherin-mediated cell-to-cell adhesion.14 Another study revealed that intracellular β-catenin and α-catenin mRNA levels are influenced by Ep-CAM siRNA – high expression of Ep-CAM in siRNA cells decreased α-catenin mRNA level and increased the intracellular β-catenin level. The β-catenin decline is due to the cells and cytoplasmic intercellular connection body caused by redistribution, but not reduced the level of transcription.14 Another report on the expression of Ep-CAM and survivin have certain effect to it.15 Ep-CAM appears to have relationships with multiple signal pathways. A previous study indicated that Ep-CAM could coexpress with many genes.16 The classic WNT signal pathway also has shown a correlation with Ep-CAM.17 A previous study showed that high expression of the WNT target genes BAMBI and DKK1 significantly correlated with the expression of Ep-CAM.18 Another study showed that β-catenin has a synergistic effect on colorectal cancer cells treated with Ep-CAM. Furthermore, the β-catenin mildly increased the expression of Ep-CAM.19 Therefore, β-catenin can activate Ep-CAM, and β-catenin may be a target of Ep-CAM transcription – this can be studied in future works.20–22 Future study should explore the function and the promising application of the Ep-CAM gene therapy in the colorectal carcinoma animal model.
Actually, a few studies have reported some different intervention methods or techniques for Ep-CAM gene expression. Naundorf et al23 found that in patients with colon cancer, the administration of edrecolomab, a monoclonal murine immunoglobulin (Ig)G1 antibody directed against Ep-CAM, significantly improved overall survival by reducing the risk of tumor recurrence. However, this antibody rapidly loses its efficacy in humans due to neutralization by antiedrecolomab antibodies. Meanwhile, a fully humanized anti-Ep-CAM antibody has also been developed,24 however, its antigen-dependent cellular cytotoxicity is serious. In our study, we established the plasmid that expresses the anti-Ep-CAM gene, which is safe and noncytotoxic for cells. Meanwhile, anti-Ep-CAM siRNA may also express continuously. Therefore, clinically, this kind of established plasmid could act as a safe, effective, long-acting antitumor agent.
In conclusion, this study successfully constructed the vector, which targets the Ep-CAM, and was imported into the colorectal cancer cells, to investigate the biological effects. Ep-CAM can promote the cell proliferation of colorectal cancer cells.
Acknowledgment
This study was supported by Shandong Province Science and Technology Department Policy Guidance Projects (fund number 2012YD18101).
Disclosure
The authors report no conflicts of interest in this work.
References
Cheng YD, Yang H, Chen GQ, Zhang ZC. Molecularly targeted drugs for metastatic colorectal cancer. Drug Des Devel Ther. 2013;7:1315–1322. | ||
Ceelen WP. Current management of peritoneal carcinomatosis from colorectal cancer. Minerva Chir. 2013;68(1):77–86. | ||
Riedy M. Preventing colorectal cancer. Adv NPs Pas. 2013;4(6): 18–21. | ||
Kumar M, Nagpal R, Hemalatha R, et al. Targeted cancer therapies: the future of cancer treatment. Acta Biomed. 2012;83(3):220–233. | ||
Wang K, Grivennikov SI, Karin M. Implications of anti-cytokine therapy in colorectal cancer and autoimmune diseases. Ann Rheum Dis. 2013;72 Suppl 2:ii100–ii103. | ||
Schnell U, Cirulli V, Giepmans BN. EpCAM: structure and function in health and disease. Biochim Biophys Acta. 2013;1828(8):1989–2001. | ||
Shen X, Wu D, He X, Chen G, Zeng Y. Expression of IGF-IR in breast cancer tissue before and after neoadjuvant chemotherapy. Cancer Cell Research. 2014;3:70–75. | ||
Ou C, Zhang H, Sun Z, Li G, Li X. Toll-like receptors in nonresolving inflammation-related cancer. Zhong Nan Da Xue Xue Bao Yi Xue Ban. 2015;40(2):202–207. | ||
Warneke VS, Behrens HM, Haag J, et al. Members of the EpCAM signalling pathway are expressed in gastric cancer tissue and are correlated with patient prognosis. Br J Cancer. 2013;109(8):2217–2227. | ||
Gadalla SE, Öjemalm K, Vasquez PL, et al. EpCAM associates with endoplasmic reticulum aminopeptidase 2 (ERAP2) in breast cancer cells. Biochem Biophys Res Commun. 2013;439(2):203–208. | ||
Martowicz A, Rainer J, Lelong J, Spizzo G, Gastl G, Untergasser G. EpCAM overexpression prolongs proliferative capacity of primary human breast epithelial cells and supports hyperplastic growth. Mol Cancer. 2013;12:56. | ||
Kowalski M, Entwistle J, Cizeau J, et al. A Phase I study of an intravesically administered immunotoxin targeting EpCAM for the treatment of nonmuscle-invasive bladder cancer in BCGrefractory and BCG-intolerant patients. Drug Des Devel Ther. 2010;4:313–320. | ||
Azer SA. Overview of molecular pathways in inflammatory bowel disease associated with colorectal cancer development. Eur J Gastroenterol Hepatol. 2013;25(3):271–281. | ||
Osta WA, Chen Y, Mikhitarian K, et al. EpCAM is overexpressed in breast cancer and is a potential target for breast cancer gene therapy. Cancer Res. 2004;64(16):5818–5824. | ||
Baeuerle PA, Gires O. EpCAM (CD326) finding its role in cancer. Br J Cancer. 2007;96(3):417–423. | ||
Sankpal NV, Fleming TP, Gillanders WE. EpCAM modulates NF-κB signaling and interleukin-8 expression in breast cancer. Mol Cancer Res. 2013;11(4):418–426. | ||
Farhana L, Dawson MI, Das JK, et al. Adamantyl Retinoid-Related Molecules Induce Apoptosis in Pancreatic Cancer Cells by Inhibiting IGF-1R and Wnt/β-Catenin Pathways. J Oncol. 2012;2012:796729. | ||
Wang K, Li N, Yeung CH, Li JY, Wang HY, Cooper TG. Oncogenic Wnt/β-catenin signalling pathways in the cancer-resistant epididymis have implications for cancer research. Mol Hum Reprod. 2013;19(2):57–71. | ||
Guerra E, Lattanzio R, La Sorda R, et al. mTrop1/Epcam knockout mice develop congenital tufting enteropathy through dysregulation of intestinal E-cadherin/β-catenin. PLoS One. 2012;7(11):e49302. | ||
Yamashita T, Budhu A, Forgues M, Wang XW. Activation of hepatic stem cell marker EpCAM by Wnt-beta-catenin signaling in hepatocellular carcinoma. Cancer Res. 2007;67(22):10831–10839. | ||
Ji J, Yamashita T, Wang XW. Wnt/beta-catenin signaling activates microRNA-181 expression in hepatocellular carcinoma. Cell Biosci. 2011;1(1):4. | ||
MacDonald GC, Rasamoelisolo M, Entwistle J, et al. A phase I clinical study of VB4-845: weekly intratumoral administration of an anti-EpCAM recombinant fusion protein in patients with squamous cell carcinoma of the head and neck. Drug Des Devel Ther. 2009;2:105–114. | ||
Naundorf S, Preithner S, Mayer P, et al. In vitro and in vivo activity of MT201, a fully human monoclonal antibody for pancarcinoma treatment. Int J Cancer. 2002;100(1):101–110. | ||
Hartung G, Hofheinz RD, Dencausse Y, et al. Adjuvant therapy with edrecolomab versus observation in stage II colon cancer: a multicenter randomized phase III study. Onkologie. 2005;28(6–7):347–350. |
 © 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.