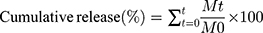Back to Journals » International Journal of Nanomedicine » Volume 15
Biocompatibility, Cytotoxicity, Antimicrobial and Epigenetic Effects of Novel Chitosan-Based Quercetin Nanohydrogel in Human Cancer Cells
Authors Abbaszadeh S , Rashidipour M, Khosravi P, Shahryarhesami S , Ashrafi B , Kaviani M, Moradi Sarabi M
Received 15 May 2020
Accepted for publication 28 July 2020
Published 11 August 2020 Volume 2020:15 Pages 5963—5975
DOI https://doi.org/10.2147/IJN.S263013
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 5
Editor who approved publication: Dr Thomas Webster
Saber Abbaszadeh,1 Marzieh Rashidipour,2 Peyman Khosravi,1 Soroosh Shahryarhesami,3 Behnam Ashrafi,2 Mozhgan Kaviani,4 Mostafa Moradi Sarabi1,2,5,6
1Department of Biochemistry and Genetics, School of Medicine, Lorestan University of Medical Sciences, Khorramabad, Iran; 2Nutritional Health Research Center, Lorestan University of Medical Sciences, Khorramabad, Iran; 3Functional Genome Analysis/B070, German Cancer Research Center (DKFZ), Heidelberg, Germany; 4Department of Internal Medicine, School of Medicine, Lorestan University of Medical Sciences, Khorramabad, Iran; 5Razi Herbal Medicines Research Center, Lorestan University of Medical Sciences, Khorramabad, Iran; 6Hepatitis Research Center, Lorestan University of Medical Sciences, Khorramabad, Iran
Correspondence: Mostafa Moradi Sarabi
Department of Biochemistry and Genetics, School of Medicine, Lorestan University of Medical Sciences, Khorramabad, Iran
Email [email protected]
Background: Previous studies have reported that quercetin (Q) has a potential antibacterial and anticancer activity. However, its application is limited by many important factors including high hydrophobicity and low absorption.
Methodology: In the current study, we synthesized and characterized (Patent) a novel chitosan-based quercetin nanohydrogel (ChiNH/Q). Encapsulation efficiency was confirmed by UV/VIS spectrophotometer. Physicochemical characterization of ChiNH/Q was assessed by PDI, DLS, SEM, FTIR, and XRD. The toxicity of the ChiNH/Q against five strains of the pathogen and HepG2 cells was examined. Moreover, the quantification of ChiNH/Q on genomic global DNA methylation and expression of DNMTs (DNMT1/3A/3B) in HepG2 cancer cells were evaluated by ELISA and real-time PCR, respectively.
Results: Under the SEM-based images, the hydrodynamic size of the ChiNH/Q was 743.6 nm. The changes in the PDI were 0.507, and zeta potential was obtained as 12.1 mV for ChiNH/Q. The FTIR peak of ChiNH/Q showed the peak at 627 cm− 1 corresponded to tensile vibrational of NH2-groups related to Q, and it is the indication of Q loading in the formulation. Moreover, XRD data have detected the encapsulation of ChiNH/Q. The ChiNH/Q showed a potent antimicrobial inhibitory effect and exerted cytotoxic effects against HepG2 cancer cells with IC50 values of 100 μg/mL. Moreover, our data have shown that ChiNH/Q effectively reduced (65%) the average expression level of all the three DNMTs (p< 0.05) and significantly increased (1.01%) the 5-methylated cytosine (5-mC) levels in HepG2 cells.
Conclusion: Our results showed for the first time the bioavailability and potentiality of ChiNH/Q as a potent antimicrobial and anticancer agent against cancer cells. Our result provided evidence that ChiNH/Q could effectively reduce cellular DNMT expression levels and increase genomic global DNA methylation in HepG2 cancer cells. Our results suggest a potential clinical application of nanoparticles as antimicrobial and anticancer agents in combination cancer therapy.
Keywords: chitosan nanohydrogel, quercetin, cytotoxic activity, antimicrobial activity, DNA methylation, gene expression
Introduction
Human hepatocellular carcinoma (HCC) is the fifth leading common cancer in the world and it is the third main cause of cancer-related deaths, worldwide.1,2 In recent years, the role of environmental factors, including flavonoids, has drawn much attention in the treatment of cancer.3,4 Quercetin (3, 3ʹ, 4ʹ, 5, 7-pentahydroxyflavone) (Q) is one of the most important natural dietary polyphenolic flavonoids which is abundantly found in fruits and vegetables. It has been reported that Q could exert distinct antioxidant, anti-inflammatory, and anti-cancer effects and its role in preventing cancer has been addressed previously.5–8 Moreover, it is well documented that Q is a potent bioflavonoid that exerts suppressing effects on the growth of several types of cancers including liver, esophagus, bladder, and colon cancer.5 However, the dualistic character of Q, regarding its role in the proliferation of cancer cell lines, has been investigated, which may affect present views on Q supposed beneficial anti-cancer effect. From a mechanistic viewpoint, it has been suggested that Q may affect the carcinogenic potential of various compounds. Many studies have shown that Q activates the metabolism of polyaromatic hydrocarbons and estrogens, which leads to the formation of catechol and subsequent oxidation metabolites known to be involved in carcinogenesis.9–11 Q already possesses a catechol group, which, due to its oxidant properties, provides the basis for the oxidizing toxic effects of this flavonoid.12 Moreover, many studies pointed at possible adverse health effects caused by Q and reported that Q can be genotoxic and also play a mutagenic role in various cultured cells.13,14 Previous studies have shown that Q is also a potent chemical agent that could cause apoptosis in various cancer cells and it showed high antimicrobial properties.15 Moreover, it has been suggested that the anti-cancer effects of polyphenols resulted from the impact of these flavonoids on epigenetic changes including DNA methylation.16 Alteration in DNA methylation plays an important role in the biology of cancer and it mainly inhibits the expression of tumor suppressor genes in cancer cells, including liver cancer.17–20 DNA methylation is a chemical modification that is mainly generated in the promoter CpG islands of the genome by the action of a family of enzymes known as DNA methyltransferases (DNMTs), including DNMT1, DNMT3A, and DNMT3B.21,22 DNMT1 is the most abundant DNMT which is involved in faithful transmission and the maintenance of DNA methylation in daughter cells during cell divisions. DNMT3A and DNMT3B act as the most potent global de novo methyltransferases that prefer to act on unmethylated CpG islands.22–24 Numerous studies suggested that DNMTs are up-regulated to different levels in various cancers, including HCC.22,25 Moreover, many reports have demonstrated that alteration in regional and global genomic DNA methylation patterns occur in early stages in liver tumorigenesis and might associate with the pathogenesis of HCC.26 In vivo and in vitro studies have demonstrated that polyphenols such as Q could alter hepatocytes metabolism and DNMTs’ activity.27,28 Also, molecular docking studies have demonstrated that Q modulates the activity of DNMTs and can function as a competitive inhibitor of the active site of DNMTs.29 Moreover, it has been suggested that Q could induce apoptosis via re-expression and inhibition of DNA methylation of apoptotic genes in cancer cell lines.30 However, many studies indicated that the application of Q has been limited by its low solubility, low bioavailability, and also it showed poor percutaneous and circulation absorption.7,31,32 Therefore, to solve this problem, many researchers have attempted to improve the bioavailability forms of Q via chemical modifications, including Q-loaded liposomes and nanoparticles.32–34 Quagliariello et al fabricated hyaluronic acid-based Q nano-hydrogel and highlighted the antitumor and anti-inflammatory properties of the Q nanocarriers in the regulation of breast cancer cell proliferation.35 Advantages of using nanoparticles including the delivery of drugs and genes into various specific sites of the body could result in longer shelf-life of the drug at the target site, and improvement of intracellular absorption, and further, the stability of drugs has also been discussed.8,36-38 Furthermore, targeted drug delivery through nanoparticles provides a new therapeutic approach, in which a drug that has been administered through intravenous or oral routes acts effectively, with lower side effects in the patient.39–42 Nanohydrogels have shown different characteristics, making them an excellent vehicle for the delivery of therapeutic agents. The use of the natural polymer chitosan, as a hydrogel scaffold, is very important due to its polymer compatibility, low toxicity, and environmental degradability.43 Chitosan is a linear amino polysaccharide, derived from deacetylation of chitin and various studies have shown that chitosan hydrogels contribute to appropriate delivery of drugs to specific sites.32,44-48 Moreover, it has been suggested that chitosan-derived hydrogels are sensitive to enzymatic degradation and can easily degrade in nature as well as in human body.44–48 In the current study, a novel chitosan-based quercetin containing nanohydrogel (ChiNH/Q) was fabricated (Patent) and after physicochemical characterization, its biocompatibility, cytotoxicity, antimicrobial and epigenetic aspects as a potential antitumor agent were investigated using well-characterized HepG2 human hepatoma cells.
Materials and Methods
All chemicals and reagents were purchased from Merck (Darmstadt, Germany), Sigma Aldrich (Gillingham, UK) and Gibco Invitrogen (Paisley, UK).
Preparation of Chitosan-Nanohydrogel (ChiNH) and Chitosan-Based Quercetin-Containing Hydrogels (ChiNH/Q)
The ChiNH nanoparticles were prepared by the ionic gelation method. Briefly, 1% chitosan solution (0.12%, 0.25%, 0.5% and 1% of chitosan) was prepared in acetic acid (0.2% v/v, Merck, Darmstadt, Germany), with vigorous stirring for 12 hours at pH=4.7. An aqueous solution of 0.5% tripolyphosphate (TPP) (0.12%, 0.25%, 0.5% and 1% of TPP) was also prepared separately. Both solutions were refined using a filter paper to remove insoluble particles (0.45 μ pore size). Then, 5 mL of the TPP solution was added to 20 mL of chitosan solution dropwise, and stirred for an additional 10 minutes. To produce ChiNH/Q, 5 mg of Q (0.62, 1.25, 2.5, and 5 mg of Q) was added to chitosan before the addition of TPP. After 12 hours of continuous stirring, the obtained particles were lyophilized, freeze-dried, and kept at 4ºC until further use. An optimal amount of ingredients (chitosan, TPP, and Q) for hydrogel formation were used.
Encapsulation Efficiency
About 0.02 g of the freeze-dried hydrogel was poured into the test tube, and after addition of 4 mL of 2M hydrochloric acid, the resulting solution was left in an ultrasonic apparatus for 30 min. Finally, the dispersed content was refined with filter paper and its volume was increased up to 10 mL by addition of double-distilled water (ddH2O). To evaluate the amount of Q loaded in the formulation, the sample’s absorbance was measured, using the UV/VIS spectrophotometer at 254 nm, and calculated by the equation below.48,49

Particle Physicochemical Characterization
Size, Polydispersity Index (PDI) and Zeta Potential
Size distribution and zeta potential of the nanoparticles were studied using dynamic light scattering (DLS) and a Malvern Zetasizer Nano-range instrument (Malvern Instruments Ltd., Malvern, UK), respectively. The colloidal emulsions were diluted in ddH2O (1:1000, v/v) and studied in triplicate at 25°C, at a fixed angle, using this instrument.
Scanning Electron Microscopy (SEM)
To conduct the SEM imaging, freeze-dried nanoparticles (left at −50°C for 24 h) were left on a stub, sputter-coated with gold, and examined at 15 kV with a 6300 field emission scanning electron microscope (Hitachi, S-4160).
Fourier-Transform Infrared (FTIR) Spectroscopy
The infrared absorption spectra of the samples including chitosan solution, TPP, ChiNH, and ChiNH/Q were studied using FTIR (BRUKER, TENSOR 27. Germany).
X-Ray Diffraction (XRD) Test
The pattern of the ChiNH/Q and the formulation of ChiNH underwent X-ray diffraction to record possible peaks of the nanogel. The samples were finely pulverized in a glass substrate, using UV-filtered (Ni-filtered Cu Ka radiation) in the range of 2 theta (2θ) at 2–50°C, and the experimental parameters were set as follows: Voltage of 40 kV; current of 20 mA; and angular velocity of 4°/min. The XRD patterns of Q, ChiNH, and ChiNH/Q were recorded using the Dmax 2100 Rigaku diffractometer (Rigaku Americas Co., TX, USA).
Release Kinetics
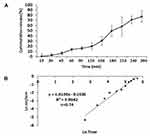
The amount of Q emission was determined, using a dialysis bag. To this end, the dialysis bag was immersed in ddH2O at 25°C for 24 h. About five mL of the prepared formulation solution (equivalent to 1.5 mg of Q) and donor compartment was left in the dialysis bag (12 KD). The loaded bag was soaked in 90 mL distilled water containing 30 mL ethanol as an acceptor compartment. The acceptor compartment agitated at 600 rpm. At specified intervals, 1 mL of the release medium was separated at time intervals of 15, 30, 45, 60, 90, 120, 150, 180, 210, 240, and 300 min for analysis and immediately replaced with 1 mL of the fresh medium (pH = 7.4). The graph for the Q levels was plotted against time. The mechanism of release of the Q from the formulation was investigated by the Korsmeyer–Peppas equation using the following formula:
where Mt is indicating the amount of Q at any given time, M0 is the Q amount at the beginning of the test, t is the release time, K is the kinetic constant for the release, and the value of n is the characteristic power for the release. The release mechanism can be described based on the value of n and indicates the Fickian and non-Fickian (anomalous) behaviors. For spherical particles, n ≤ 0.43 indicates a case I transport (Fickian release), while n = 0.85 indicates a case II transport (non-Fickian release). Intermediate values of n, ie, 0.43 < n < 0.85, indicate a non-Fickian release and the release follows the combination of the mechanisms of diffusion and inflation.50,51 The linear graph was plotted using the Ln 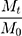 at the final time against Ln(t). The cumulative amount of Q released from the formulation was calculated at different intervals by the following equation and the graph for the cumulative amount was plotted against time.
at the final time against Ln(t). The cumulative amount of Q released from the formulation was calculated at different intervals by the following equation and the graph for the cumulative amount was plotted against time.
pH Test
To evaluate pH changes, 0.5 g of formulations with and without Q was added to 10 mL ddH2O, and the pH was determined throughout 0–90 (0, 1, 2, 7, 14, 30, 45, 60, and 90) days.
HepG2 Cell Culture and MTT Assay
Human hepatoma HepG2 cell line was purchased from the National cell bank of Iran (NCBI, Pasteur Institute, Tehran). Cells were cultured in RPMI 1640 medium supplemented with 10% FBS, 2mM Gln, penicillin (100 U/mL), and streptomycin (100 μg/mL) in humidified 5% CO2, the atmosphere at 37°C temperature. For ChiNH, Q, and ChiNH/Q treatment, cells were seeded into six-well plates at a density of 3 × 104 cells and allowed to attach for 24 h. The viability of HepG2 cells was determined by MTT test using 5 different concentrations of 10, 50, 100, 500 and 1000 µg/mL of ChiNH, Q and ChiNH/Q for 48h based on the method described by Rashidipour et al.52 All experiments were performed in triplicate and the data were expressed as mean ± SD.
Antimicrobial Activity of ChiNH/Q
Microbial Strains and Culture
To evaluate the antimicrobial effects of ChiNH/Q, five strains of the pathogen, including Staphylococcus aureus (S. aureus) (ATCC 12600), Listeria monocytogenes (L. monocytogenes) (PTCC 1297), Escherichia coli (E. coli) (PTCC 1395), Klebsiella pneumoniae (K. pneumoniae) (PTCC 1290) and Candida albicans (C. albicans) (PTCC 5072) were obtained from the National Cell Bank of Iran (NCBI, Pasteur Institute, Tehran).
Antimicrobial effects of broth microdilution were used according to the CLSI standard method.53,54 Briefly, to examine the Minimum Inhibitory Concentration (MIC), ChiNH/Q and blank (1000 µg/mL), Q (5000 µg/mL), Ceftriaxone/Fluconazole (200 µg/mL) sterile media, Mueller-Hinton broth for bacteria and Sabouraud Dextrose broth agar (SDA) for C. albicans were prepared. Then, different concentrations of stocks were added to the wells and the microbial suspension equivalent to Half McFarland (1.5×108 CFU/mL) was added to the wells and kept in a 37°C incubator for 24 hours. After this time, salts of 2,3,5- Triphenyltetrazolium chloride were used to evaluate microbial viability. Pink wells indicate microbial life and non-color wells indicate microbial inhibition.53 After calculating the MIC, 5 µL was cultured on Mueller-Hinton agar (for bacteria) and SDA (for C. albicans) with 24 min incubation, and then incubated for 24 h. At 37°C, the first concentration where no colony was found, reported as the Minimum Bactericidal Concentration and Minimum Fungicidal Concentration (MBC/MFC).
RNA Extraction and Quantitative Real-Time RT-PCR
Total RNA was extracted from HepG2 cells using the TriPure isolation reagent (Roche Applied Science, Germany) according to the manufacturer’s instructions. The integrity of purified RNA was assessed on 2% formaldehyde containing 1.5% agarose, and purified RNA was stored at −80°C until use. The complementary DNAs (cDNAs) were synthesized from 2 µg total RNA by reverse transcriptase (RT) using oligodT primers and M-MuLV RT (MBI Fermentas, Lithuania) in a 20μL final reaction volumes according to the manufacturer’s provider. Transcript levels of mRNA encoding DNMT1, DNMT3A, and DNMT3B were analyzed by quantitative real-time RT-PCR assay using a Corrbet sequence detection system (Rotor gene 6000). SYBR Green-based analysis and Master Mix (ABI, UK) were used and all reactions were carried out in triplicate. The primers information and conditions for DNMTs real-time RT-PCR procedures have been previously described.22 The GAPDH was used as a reference gene and the relative gene expression levels were determined using the 2 –ΔΔCT standard method.55
Analysis of Global Genomic DNA Methylation
Global DNA methylation was quantified in DNA isolated from HepG2 cells using a 5-mC DNA ELISA kit (Zymo Research, Freiburg, Germany) according to the manufacturer’s instructions as previously described.22
Statistical Analysis
SPSS 21 analytic software (SPSS, Inc., Chicago) and Graph Pad Prism (Version 8.01) were used for data analysis. All data were presented as mean ± standard deviation (SD) from three independent experiments. Data were analyzed using one-way ANOVA followed by Tukey’s multiple comparison tests. Differences with p-value ≤0.05 were set as the level of significance.
Results and Discussion
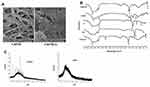
We hypothesized that “a novel quercetin Q nanohydrogel, using Chitosan (ChiNH/Q) would have more biological properties for pathogenic microbial strains and cancer cells than free Q”. Many studies effectively highlight the high antimicrobial properties of Q and ability of Q to modulate epigenetic modifications particularly DNA methylation and restore gene expression. However, the application of Q is limited by several factors including low bioavailability, low solubility, and low circulation absorption, and also application forms of Q. Numerous methods including nanoliposomal and nanoparticle‐based targeted delivery to tumors, are actively being sought to develop the bioavailability of Q. To the best of our knowledge, no previous studies have addressed the bioavailability, cytotoxicity, antimicrobial properties, and effect of Q nanohydrogels on DNA methylation in HCC. Therefore, our study is the first report for the synthesis of ChiNH/Q. Due to the many important characteristics of chitosan, including cationic characters and primary amino groups, which are responsible for many associated properties such as transfection and controlled drug release, it has been suggested that chitosan is the most important polysaccharide for various drug delivery purposes.56 The SEM image of the freeze-dried hydrogel sample was prepared which indicated that ChiNH was successfully synthesized from chitosan (Figure 1A). The compact and dense matrix could be due to the formation of electrostatic bondings between the TPP functional groups and the amino groups in the chitosan. The hydrodynamic size, zeta potential, and PDI of the produced particles were measured (Table 1). The encapsulation efficiency was found to be 62.24% and the percentage of Q in chitosan-based quercetin-containing hydrogels was 1.42%. The size of the ChiNH/Q was 743.6 nm and the size of ChiNH was 992.7 nm. The changes in the PDI, in the ChiNH/Q and ChiNH were 0.507 and 0.537, respectively. These changes indicate a uniform dispersion of particles in the formulation of ChiNH/Q.57–59 Our results demonstrated that ChiNHs and loading Q could affect the particle sizes so that ChiNHs showed the largest size (992.7 nm), whereas, ChiNHs/Q showed the smallest size (743.6 nm) (Table 1). The smaller size of ChiNHs/Q might be due to the crosslinking between the sodium-TPP functional group, the chitosan amine groups, and the hydroxyl functional groups in the Q. For practical purposes, the physical characteristics of nanoparticles including size, shape, structure, and coating have significant effects on the cytotoxicity, distribution, and also cellular uptake of nanoparticles. It has been suggested that the nanoparticles of comparably smaller sizes are better because of the ratio of surface to volume as well as colloidal stability increases as the particle size decreases.57 Zeta potential was determined as 12.1 mV and 3.7 mV for ChiNH/Q and ChiNH, respectively, given the presence of the anionic TPP compounds in nano-hydrogels.
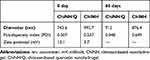 |
Table 1 The Physicochemical Characteristics of ChiNH/Q and ChiNH as a Function of Time (0 and 60 days) |
To investigate the chemical composition of the produced ChiNH in the presence and absence of Q, the FTIRs were separately studied (Figure 1B). FTIR analysis indicated that the formation of bioactive groups surrounded the Q in the hydrogel cavities. The FTIR spectrum of ChiNH/Q presented 6 peaks especially at 1153–1415 cm−1, 1563 cm−1, 1638–1640 cm−1, 2925 cm−1 and 3237–3562 cm−1 for ChiNH/Q (Figure 1B). The broad and strong absorbance peaks of 1163–1216 cm−1 indicate the presence of symmetrical and asymmetric stretching vibration, PO3 group and P=O groups related to chitosan and sodium TPP, respectively.60 The absorption peak at 3570 cm−1 was assigned to the existing N-H stretching of amid groups of chitosan. The signals observed at 1617 cm−1 attributed to amid II (N-H bonding) of chitosan. Absorbance peaks at the wavelengths of 1638–1640 cm−1 and 2925–3237 cm−1 indicated the presence of symmetric and asymmetric tensile vibrational groups of C=O; 1455 cm-1 was assigned to the aromatic C=C stretching and 1137 cm−1 indicated phenolic C-O stretching related to Q. Moreover, in the ChiNH/Q there was a strong absorbance peak at 627 cm−1 corresponding to tensile vibrational of NH2-groups related to Q, and indicating that Q was included in the formulation, and the presence of this peak in the respective spectrum indicating cross-linking created by Q (Figure 1B). These data demonstrate the formation of a chitosan-based quercetin nanohydrogel. Also, based on the FTIR data, the dispersion was potentially lower due to the interaction and loading of the Q inside the formulation cavities.
According to the results of XRD graphs, no specific peak was observed on the patterns of the ChiNH and ChiNH/Q formulations (Figure 1C). As shown in Figure 1C, this observation explains that Q is encapsulated or dispersed among the chitosan-sodium TPP polymers which are linked by an interconnected ionic network, and an amorphous complex is formed through intermolecular interaction in the matrix. Various factors affect the release of nanoparticles, including the particle size, molecular weight of the polymer, and the interaction between the particles and the polymer.60,61 To investigate the releasing mechanism, the final formulation was prepared under optimal conditions, and Q release was calculated at 15, 30, 45, 60, 90, 120, 150, 180, 210, 240, and 300-min intervals using the dialysis bag (Figure 2A). In this test, at specified intervals, 1 mL of the receiving solution was separated for analysis and it immediately replaced with 1 mL of the fresh receiving solution (pH=7.4). The specific concentration of ChiNH/Q was poured into the dialysis bag (12 KD) and left in the container. The value of n was calculated, using the graphed equation of the line. According to Figure 2A and the obtained data, 77.28% of the initial Q was released after 300 minutes. The Korsmeyer–Peppas mathematical model was used to calculate n, which was ultimately calculated at 0.74 (Figure 2B). This indicates that Q release from chitosan-based polymer follows the combination of two processes: penetration and erosion. The constant of release (K) was 1.61 and the correlation coefficient was 0.96 (Figure 2B).52 As a test of stability, pH showed that the H+ concentration in the formulation increased by approximately 1.05 times from day 0 to day 90, which could be due to the presence of chitosan hydrolysis and release of the glucosamine and oligosaccharides (Figure 3).
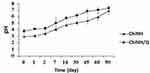 |
Figure 3 The pH of ChiNH and ChiNH/Q formulation throughout 90 days. |
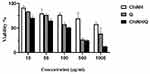
Our study indicated for the first time, the cytotoxicity effect of encapsulated Q by ChiNHs against HepG2 cells using the MTT assay. The optical microscopy was employed to examine the influences of ChiNH, Q and ChiNH/Q on the morphological changes of HepG2 cells (Figure 4). Our data indicated that ChiNH/Q at concentrations of 50, 100, 500, and 1000 µg/mL significantly decreased the HepG2 cell viability in a dose-dependent way, as compared with untreated control cells (P<0.05) (Figure 5). Furthermore, we found that ChiNH/Q significantly increased drug cytotoxicity on HepG2 cells, as compared with free Q (p<0.05) (Figure 5). Moreover, the IC50 values of ChiNH/Q against HepG2 cells were significantly lower than free Q at 50 µg/mL (p<0.05) (Figure 5). Then, 100 µg/mL (lethal-dose) of each of ChiNH/Q, and free Q showed 50.7% and 57% cell viability at 48 hours, respectively. Whereas, 50 µg/mL (sub‐lethal dose) of each of ChiNH/Q and free Q showed 65% and 76.5% cell viability at 48 h, respectively. These data have proved the efficiency of ChiNH/Q as an anticancer agent with low IC50 against HepG2 cells. The mechanisms by which nanoparticles can stop the growth of cancer cells are still not clear. However, experimental studies indicated that Q-nanoparticles induced antioxidant activity and more efficient interaction of Q with proteins.62 Moreover, in vitro studies has been indicated that Q-nanoparticles are a promising candidate for the delivery of the anticancer drug delivery in cancer cells and show impressive cytotoxicity for cancer cells.63
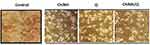 |
Figure 4 The optical microscopy images of the morphological changes of HepG2 cells after treatment with 50 µg/mL of ChiNH, Q and ChiNH/Q for 48 h. |
Our study for the first time evaluated the antimicrobial activity of ChiNH/Q against different pathogenic bacterial strains, including K. pneumoniae PTCC 1290, E. coli PTCC 1395, S. aureus ATCC 12600, L. monocytogenes PTCC 1297 and the yeast C. albicans PTCC 5072, using broth microdilution assay and according to the CLSI standard method.53 According to the results in Table 2, the antibacterial and antifungal results showed that ChiNH/Q suppressed the growth of Gram-positive bacterium L. monocytogenes and C. albicans to greater extents compared to free Q as evidenced by MIC 250 and 1000, respectively, while free Q resulted in MIC equal to 1250 in both of L. monocytogenes and C. albicans (Table 2). These results are likely to be due to the proper enclosure of Q in the nanohydrogel network, and the better effects of encapsulated Q could be attributed to its slower release in the polymer matrix.64 Moreover, our results indicated that for two Gram-negative bacterial strains, including E. coli and K. pneumoniae, the MIC of free Q was 312.5 and 156.25, respectively. These results demonstrated that free Q showed a greater inhibitory effect than ChiNH/Q on these Gram-negative bacteria (Table 2). Also, for Gram-positive bacteria like S. aureus the MIC was >1000 after treating with free Q and ChiNH/Q. These data showed that S.aureus is resistant to all treatments, possibly due to the complexity of the cell wall structure of this bacterium and many studies have shown that Gram-positive bacteria are more resistant to the penetration of nanoparticles.65–68 Since, Gram-positive bacteria such as S. aureus have a much thicker layer of peptidoglycan than Gram-negative bacteria, including E.coli, the penetration of nanoparticles into the wall of Gram-negative bacteria is easier due to the lower thickness.69 Our data indicated that ChiNH/Q showed a potent antimicrobial effect against L. monocytogenes and C.albicans. Moreover, the results are interesting since the values of MIC caused by ChiNH/Q are more close to Ceftriaxone/Fluconazole, as positive control for L. monocytogenes and C. albicans compared to free Q (Table 2). Our results are in line with Milanezi et al that Q-capped gold nanoparticles demonstrated a higher antimicrobial activity than free Q in pathogenic strains.70 It has been suggested that the small size of the nanoparticles makes them suitable targets for antimicrobial and intracellular targeting purposes.71 Moreover, nanogels and microgels as drug carriers are capable of carrying significant amounts of pharmaceutical compounds and protecting them against proteolytic degradation.72 Overall, our data indicated that ChiNH/Q might be a potent antimicrobial agent with a strong inhibitory effect on pathogenic microorganisms and could be used as a potential alternative antibiotic in medical and therapeutic applications.
 |
Table 2 The Antimicrobial Effects of Quercetin-Chitosan Nanohydrogel |
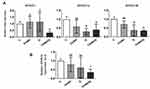
We evaluated for the first time the application of encapsulated Q by ChiNHs on the DNMTs’ gene expression in HepG2 cells. To verify if the DNMTs’expression is influenced by ChiNH, ChiNH/Q, and free Q treatment, we selected a sub-lethal dose of 50 µg/mL and HepG2 cells were incubated with 50 µg/mL of ChiNH/Q, ChiNH, and free Q for 48 h. We observed variable effects of ChiNH, ChiNH/Q, and free Q on all the three DNMT´s gene expressions (DNMT1, DNMT3Aand DNMT3B) in HepG2 cells. As shown in Figure 6A and Table 3, our study showed that treatment of HepG2 cells with ChiNH/Q resulted in a decrease in mRNA transcription levels of DNMT1, DNMT3A, and DNMT3B by 68%, 59%, and 65%, respectively, as compared to control cells (P<0.05) (Table 3). Moreover, ChiNH/Q showed significant effects on the DNMT1 expression level in HepG2 cells, compared to ChiNH-treated cells (P<0.05) (Figure 6A). According to previous reports, the DNMT1 expression is a key element in liver malignancy and significantly correlated with the poor prognosis of HCC.73 Furthermore, experimental studies suggested an indispensable role of DNMT1 as a key genomic regulator of liver growth and regeneration.74 Moreover, the statistical analysis of our study showed that Q similar to ChiNH/Q could suppress the expression level of DNMT3A (70%-reduced) and DNMT3B (63% reduced) in HepG2 cells compared to control cells (P<0.05) (Figure 5A) (Table 3). Interestingly, our results indicated that the average expression level of all three DNMTs (T1/3A/3B) was effectively reduced by ChiNH/Q (65% reduced) compared to control cells (P<0.05) (Figure 6B) (Table 3). We observed a trend for the coordinated pattern of all three DNMTs’ average inhibition expression, using ChiNH/Q, in comparison to free Q (Figure 6B). As expected, our results demonstrated that the decreased DNMTs’ expression level was observed only in ChiNH/Q-treated HepG2 cells compared to ChiNH-treated cells (Figure 6B). Overall, based on these results, downregulation of DNMTs was effectively observed in the ChiNH/Q-treated HepG2 cells, and ChiNH/Q exhibits a greater effect on the reduction of the expression level of DNMTs while compared to free Q. Many studies have reported the increased DNMTs’ expression levels in HCC, while reducing the expression of DNMTs by ChiNH/Q could help to control HCC.75
 |
Table 3 The Effects of ChiNH, Q and ChiNH/Q on DNMTs mRNA Gene Expression Levels in the HepG2 Cancer Cells. The Values are Represented as Mean ± SD |
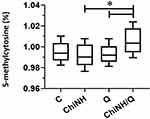
This report demonstrated for the first time, the application of ChiNH/Q on the global genomic DNA methylation profile in HepG2 cells. Our results indicated that overall about 0.995% of the cytosine residues were methylated in the genome of the media-only treated (control) HepG2 cells (Figure 7). Moreover, we found that the level of methylated cytosine in ChiNH/Q-treated cells was significantly higher (1.01%) than those with Q and ChiNH-treated cells (0.993% and 0.992%, respectively) (p<0.05) (Figure 7). As expected, these results indicated that ChiNH/Q effectively increased the global DNA methylation of HepG2 cells, compared to ChiNH and free Q-treated cells. These results suggested that ChiNH/Q always displayed further efficacy, in comparison to free Q in decreasing DNMTs’ gene expression and also increasing the global DNA methylation in HepG2 cancer cells. Many studies also demonstrated that Q can exert anti-cancer effect by regulating epigenetic modifications including, miRNA expression and DNA methylation, and enhance the sensitivity of tumor cells to chemotherapy.76,77 Other studies also reported the anti-inflammatory properties of Q nanoparticles and demonstrate that biodegradable Q nanoparticles increases the cellular uptake of chemotherapy drugs into cancer cells and enhances their cytotoxicity and antitumor properties.78 Global DNA hypomethylation is a common epigenetic change in HCC and plays an important role in elevating chromosomal instability, and initiation and progression of HCC.26,79 Overall, our study successfully explains that ChiNH/Q always shows a higher efficacy than free Q, and ChiNH/Q can effectively reduce cellular DNMTs’ expression level and also increase genomic global DNA methylation in HepG2 cancer cells. By linking polyphenol nanoparticles and DNA methylation, our current study provides some insight into the capacity of nanoparticles in the delivery of anti-tumor agents regarding epigenetic modification of HCC and also can open new insights to clinical potential of nanoparticles in targeted-therapy in cancer.
Conclusions
In conclusion, the formulation of novel Q nanohydrogel using chitosan (ChiNH/Q) detailed in this study was assessed by PDI, DLS, SEM, FTIR, and XRD. Encapsulation efficiency was confirmed by UV/VIS spectrophotometer. The physicochemical characterization of ChiNH/Q showed that the size of the ChiNH/Q was 743.6 nm. The FTIR peak showed that the presence of carboxyl, P=O, C=O, and NH2 groups were important factors in the synthesis of ChiNH/Q, and XRD data detected the encapsulation of Q in the ChiNH. The ChiNH/Q produced in this study exerted significant cytotoxic effects against HepG2 cancer cells with IC50 values of 100 µg/mL. Also, ChiNH/Q applied potent antimicrobial effects against Gram-positive bacteria and C. albicans, and it might be a promising antimicrobial agent in medical applications. Moreover, our results showed that ChiNH/Q increased availability of Q and exhibited a greater effect on the reduction of DNMTs’ gene expression and also ChiNH/Q increased in global DNA methylation in HepG2 cancer cells. Our study provides new insights into the epigenetic mechanisms by which nanohydrogels containing flavonoids, effectively influence the gene expression in HCC. Also, our data indicated that the synthesis of Q nanoparticles was efficient and could be considered as an agent carrier in targeted therapy in liver cancer. However, further investigations including in vitro and in vivo models are needed to evaluate the potential application of the ChiNH/Q in nano-therapy approaches.
Abbreviations
Q, quercetin; ChiNH, chitosan-nanohydogel; ChiNH/Q, chitosan-based quercetin nanohydrogel; PCR, polymerase chain reaction; MIC, minimum inhibitory concentration; SDA, Sabouraud dextrose broth agar; MBC, minimum bactericidal concentration; MFC, minimum fungicidal concentration; SEM, scanning electron microscope; DLS, dynamic light scattering; FTIR, Fourier-transform infrared spectroscopy; XRD, x-ray diffraction; UV/VIS, ultraviolet–visible spectroscopy; ELISA, enzyme-linked immunosorbent assay; DNMTs, DNA methyltransferases; DNMT1; DNA methyltransferase 1; DNMT3A, DNA methyltransferase 3A; DNMT3B, DNA methyltransferase 3B; GAPDH, glyceraldehyde-3-phosphate dehydrogenase; miRNA, microRNA.
Acknowledgments
The work described in this manuscript was part of the dissertation of Saber Abbaszadeh, submitted to the Lorestan University of Medical Sciences in partial fulfillment of the requirements for the MSc in biochemistry. This work was supported by a grant from the Vice Chancellor for Research, Lorestan University of Medical Sciences (grant number A-10-1477-5).
Author Contributions
All authors made a significant contribution to the work reported, whether that is in the conception, study design, execution, acquisition of data, analysis and interpretation, or in all these areas; took part in drafting, revising or critically reviewing the article; gave final approval of the version to be published; have agreed on the journal to which the article has been submitted; and agree to be accountable for all aspects of the work.
Disclosure
The authors declare that there is no conflict of interest.
References
1. El-Serag HB, Rudolph KL. Hepatocellular carcinoma: epidemiology and molecular carcinogenesis. Gastroenterology. 2007;132(7):2557–2576. doi:10.1053/j.gastro.2007.04.061
2. Abd-Rabou AA, Shalby AB, Ahmed HH. Anti-cancer activity of quercetin, gallic acid, and ellagic acid against HEPG2 and HCT 116 cell lines: in vitro. Int J Pharm Biol Sci. 2016;4(5):B584–B592. doi:10.22376/ijpbs.2016.7.4.b584-592
3. Cueva C, Moreno-Arribas MV, Martín-Álvarez PJ, et al. Antimicrobial activity of phenolic acids against commensal, probiotic and pathogenic bacteria. Res Microbiol. 2010;161(5):372–382. doi:10.1016/j.resmic.2010.04.006
4. Naghdi N. Folklore medicinal plants used in liver disease: a review. Int J Green Pharm. 2018;12(03).
5. Zhao J-L, Zhao J, Jiao H-J. Synergistic growth-suppressive effects of quercetin and cisplatin on HepG2 human hepatocellular carcinoma cells. Appl Biochem Biotechnol. 2014;172(2):784–791. doi:10.1007/s12010-013-0561-z
6. Formica J, Regelson W. Review of the biology of quercetin and related bioflavonoids. Food Chem Toxicol. 1995;33(12):1061–1080. doi:10.1016/0278-6915(95)00077-1
7. Tian H, Zhang J, Zhang H, et al. Low side-effect and heat-shock protein-inhibited chemo-phototherapy nanoplatform via co-assembling strategy of biotin-tailored IR780 and quercetin. Chem Eng J. 2020;382:123043. doi:10.1016/j.cej.2019.123043
8. Jiang W, Zhang H, Wu J, et al. CuS@MOF-based well-designed quercetin delivery system for chemo-photothermal therapy. ACS Appl Mater Interfaces. 2018;10(40):34513–34523. doi:10.1021/acsami.8b13487
9. Bolton JL, Pisha E, Zhang F, et al. Role of quinoids in estrogen carcinogenesis. Chem Res Toxicol. 1998;11(10):1113–1127. doi:10.1021/tx9801007
10. Penning TM, Burczynski ME, Hung C-F, et al. Dihydrodiol dehydrogenases and polycyclic aromatic hydrocarbon activation: generation of reactive and redox active o-quinones. Chem Res Toxicol. 1999;12(1):1–18. doi:10.1021/tx980143n
11. Cavalieri E, Rogan E, Chakravarti D. The role of endogenous catechol quinones in the initiation of cancer and neurodegenerative diseases. Methods Enzymol. 2004;382:293–319. doi:10.1016/S0076-6879(04)82017-2
12. Cao G, Sofic E, Prior RL. Antioxidant and prooxidant behavior of flavonoids: structure-activity relationships. Free Radic Biol Med. 1997;22(5):749–760. doi:10.1016/s0891-5849(96)00351-6
13. Hatcher JF, Bryan GT. Factors affecting the mutagenic activity of quercetin for Salmonella typhimurium TA98: metal ions, antioxidants and pH. Mutat Res. 1985;148(1–2):13–23. doi:10.1016/0027-5107(85)90203-9
14. van der Hoeven JC, Bruggeman IM, Debets FM. Genotoxicity of quercetin in cultured mammalian cells. Mutat Res. 1984;136(1):9–21. doi:10.1016/0165-1218(84)90130-7
15. Granato M, Rizzello C, Gilardini Montani MS, et al. Quercetin induces apoptosis and autophagy in primary effusion lymphoma cells by inhibiting PI3K/AKT/mTOR and STAT3 signaling pathways. J Nutr Biochem. 2017;41:124–136. doi:10.1016/j.jnutbio.2016.12.011
16. Lee WJ, Shim J-Y, Zhu BT. Mechanisms for the inhibition of DNA methyltransferases by tea catechins and bioflavonoids. Mol Pharmacol. 2005;68(4):1018–1030. doi:10.1124/mol.104.008367
17. Morales-Ruiz T, García-Ortiz MV, Devesa-Guerra I, et al. DNA methylation reprogramming of human cancer cells by expression of a plant 5-methylcytosine DNA glycosylase. Epigenetics. 2018;13(1):95–107. doi:10.1080/15592294.2017.1414128
18. Kurayoshi K, et al. The key role of E2F in tumor suppression through specific regulation of tumor suppressor genes in response to oncogenic changes. In: Gene Expression and Regulation in Mammalian Cells-Transcription Toward the Establishment of Novel Therapeutics. 2018:143.
19. Wu H, Coskun V, Tao J, et al. Dnmt3a-dependent nonpromoter DNA methylation facilitates transcription of neurogenic genes. Science. 2010;329(5990):444–448. doi:10.1126/science.1190485
20. Oh B-K, Kim H, Park H-J, et al. DNA methyltransferase expression and DNA methylation in human hepatocellular carcinoma and their clinicopathological correlation. Int J Mol Med. 2007;20(1):65–73.
21. Challen GA, Sun D, Mayle A, et al. Dnmt3a and Dnmt3b have overlapping and distinct functions in hematopoietic stem cells. Cell Stem Cell. 2014;15(3):350–364. doi:10.1016/j.stem.2014.06.018
22. Sarabi MM, Naghibalhossaini F. Association of DNA methyltransferases expression with global and gene-specific DNA methylation in colorectal cancer cells. Cell Biochem Funct. 2015;33(7):427–433. doi:10.1002/cbf.3126
23. Robertson KD, Uzvolgyi E, Liang G, et al. The human DNA methyltransferases (DNMTs) 1, 3a and 3b: coordinate mRNA expression in normal tissues and overexpression in tumors. Nucleic Acids Res. 1999;27(11):2291–2298. doi:10.1093/nar/27.11.2291
24. Jahangiri R, Jamialahmadi K, Gharib M, et al. Expression and clinicopathological significance of DNA methyltransferase 1, 3A and 3B in tamoxifen-treated breast cancer patients. Gene. 2019;685:24–31. doi:10.1016/j.gene.2018.10.060
25. Gailhouste L, Liew LC, Yasukawa K, et al. Differentiation therapy by epigenetic reconditioning exerts antitumor effects on liver cancer cells. Mol Ther. 2018;26(7):1840–1854. doi:10.1016/j.ymthe.2018.04.018
26. Toh TB, Lim JJ, Chow EK-H. Epigenetics of hepatocellular carcinoma. Clin Transl Med. 2019;8(1):13. doi:10.1186/s40169-019-0230-0
27. Liu CM, Ma J-Q, Xie W-R, et al. Quercetin protects mouse liver against nickel-induced DNA methylation and inflammation associated with the Nrf2/HO-1 and p38/STAT1/NF-κB pathway. Food Chem Toxicol. 2015;82:19–26. doi:10.1016/j.fct.2015.05.001
28. Kanwal R, Datt M, Liu X, et al. Dietary flavones as dual inhibitors of DNA methyltransferases and histone methyltransferases. PLoS One. 2016;11(9):e0162956. doi:10.1371/journal.pone.0162956
29. Kedhari Sundaram M, Hussain A, Haque S, et al. Quercetin modifies 5ʹCpG promoter methylation and reactivates various tumor suppressor genes by modulating epigenetic marks in human cervical cancer cells. J Cell Biochem. 2019;120(10):18357–18369. doi:10.1002/jcb.29147
30. Alvarez MC, Maso V, Torello CO, et al. The polyphenol quercetin induces cell death in leukemia by targeting epigenetic regulators of pro-apoptotic genes. Clin Epigenetics. 2018;10(1):139. doi:10.1186/s13148-018-0563-3
31. Hung C-F, Fang C-L, Al-Suwayeh SA, et al. Evaluation of drug and sunscreen permeation via skin irradiated with UVA and UVB: comparisons of normal skin and chronologically aged skin. J Dermatol Sci. 2012;68(3):135–148. doi:10.1016/j.jdermsci.2012.09.005
32. Abdel-Wahhab MA, Aljawish A, El-Nekeety AA, et al. Chitosan nanoparticles and quercetin modulate gene expression and prevent the genotoxicity of aflatoxin B1 in rat liver. Toxicol Rep. 2015;2:737–747. doi:10.1016/j.toxrep.2015.05.007
33. Nan W, Ding L, Chen H, et al. Topical use of quercetin-loaded chitosan nanoparticles against ultraviolet B radiation. Front Pharmacol. 2018;9:826. doi:10.3389/fphar.2018.00826
34. Hatahet T, Morille M, Hommoss A, et al. Quercetin topical application, from conventional dosage forms to nanodosage forms. Eur J Pharm Biopharm. 2016;108:41–53. doi:10.1016/j.ejpb.2016.08.011
35. Quagliariello V, Iaffaioli RV, Armenia E, et al. Hyaluronic acid nanohydrogel loaded with quercetin alone or in combination to a macrolide derivative of rapamycin RAD001 (Everolimus) as a new treatment for hormone-responsive human breast cancer. J Cell Physiol. 2017;232(8):2063–2074. doi:10.1002/jcp.25587
36. Jain KK. Nanotechnology in clinical laboratory diagnostics. Clinica chimica acta. 2005;358(1–2):37–54. doi:10.1016/j.cccn.2005.03.014
37. Sahoo SK, Dilnawaz F, Krishnakumar S. Nanotechnology in ocular drug delivery. Drug Discov Today. 2008;13(3–4):144–151. doi:10.1016/j.drudis.2007.10.021
38. Park K. Facing the truth about nanotechnology in drug delivery. ACS Nano. 2013;7(9):7442–7447. doi:10.1021/nn404501g
39. Misra R, Acharya S, Sahoo SK. Cancer nanotechnology: application of nanotechnology in cancer therapy. Drug Discov Today. 2010;15(19–20):842–850. doi:10.1016/j.drudis.2010.08.006
40. Caruso G, Raudino G, Caffo M, et al. Nanotechnology platforms in diagnosis and treatment of primary brain tumors. Recent Pat Nanotechnol. 2010;4(2):119–124. doi:10.2174/187221010791208786
41. Kumari A, Yadav SK, Yadav SC. Biodegradable polymeric nanoparticles based drug delivery systems. Colloids Surf B Biointerfaces. 2010;75(1):1–18. doi:10.1016/j.colsurfb.2009.09.001
42. Lim E-K, Chung BH, Chung SJ. Recent advances in pH-sensitive polymeric nanoparticles for smart drug delivery in cancer therapy. Curr Drug Targets. 2018;19(4):300–317. doi:10.2174/1389450117666160602202339
43. de Oliveira Pedro R, Hoffmann S, Pereira S, et al. Self-assembled amphiphilic chitosan nanoparticles for quercetin delivery to breast cancer cells. Eur J Pharm Biopharm. 2018;131:203–210. doi:10.1016/j.ejpb.2018.08.009
44. Zhu D, Tao W, Zhang H, et al. Docetaxel (DTX)-loaded polydopamine-modified TPGS-PLA nanoparticles as a targeted drug delivery system for the treatment of liver cancer. Acta Biomater. 2016;30:144–154. doi:10.1016/j.actbio.2015.11.031
45. Curcio M, Cirillo G, Parisi OI, et al. Quercetin-imprinted nanospheres as novel drug delivery devices. J Funct Biomater. 2012;3(2):269–282. doi:10.3390/jfb3020269
46. Bhattarai N, Gunn J, Zhang M. Chitosan-based hydrogels for controlled, localized drug delivery. Adv Drug Deliv Rev. 2010;62(1):83–99. doi:10.1016/j.addr.2009.07.019
47. Li J, Mooney DJ. Designing hydrogels for controlled drug delivery. Nat Rev Mater. 2016;1(12):16071. doi:10.1038/natrevmats.2016.71
48. Yadollahi M, Farhoudian S, Barkhordari S, et al. Facile synthesis of chitosan/ZnO bio-nanocomposite hydrogel beads as drug delivery systems. Int J Biol Macromol. 2016;82:273–278. doi:10.1016/j.ijbiomac.2015.09.064
49. Popa N, Novac O, Profire L, et al. Hydrogels based on chitosan–xanthan for controlled release of theophylline. J Mater Sci Mater Med. 2010;21(4):1241–1248. doi:10.1007/s10856-009-3937-4
50. Keawchaoon L, Yoksan R. Preparation, characterization and in vitro release study of carvacrol-loaded chitosan nanoparticles. Colloids Surf B Biointerfaces. 2011;84(1):163–171. doi:10.1016/j.colsurfb.2010.12.031
51. Stulzer HK, Tagliari MP, Parize AL, et al. Evaluation of cross-linked chitosan microparticles containing acyclovir obtained by spray-drying. Mater Sci Eng C. 2009;29(2):387–392. doi:10.1016/j.msec.2008.07.030
52. Rashidipour M, Maleki A, Kordi S, et al. Pectin/chitosan/tripolyphosphate nanoparticles: efficient carriers for reducing soil sorption, cytotoxicity and mutagenicity of paraquat and enhancement of its herbicide activity. J Agric Food Chem. 2019;67:5736–5745. doi:10.1021/acs.jafc.9b01106
53. Haddouchi F, Chaouche TM, Zaouali Y, et al. Chemical composition and antimicrobial activity of the essential oils from four Ruta species growing in Algeria. Food Chem. 2013;141(1):253–258. doi:10.1016/j.foodchem.2013.03.007
54. Wayne P. NCCLS (National Committee for Clinical Laboratory Standards) Method for Dilution Antimicrobial Susceptibility Tests of Bacteria That Grow Aerobically. Approved Standard. M100-S12. 2002.
55. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods. 2001;25(4):402–408. doi:10.1006/meth.2001.1262
56. Abd Elgadir M, Uddin MS, Ferdosh S, et al. Impact of chitosan composites and chitosan nanoparticle composites on various drug delivery systems: a review. J Food Drug Anal. 2015;23(4):619–629. doi:10.1016/j.jfda.2014.10.008
57. Bhattacharjee S. DLS and zeta potential - what they are and what they are not? J Control Release. 2016;235:337–351. doi:10.1016/j.jconrel.2016.06.017
58. Bhattacharjee S. In relation to the following article “DLS and zeta potential - what they are and what they are not?” Journal of Controlled Release, 2016, 235, 337–351. J Control Release. 2016;238:311–312. doi:10.1016/j.jconrel.2016.07.002
59. Chacko RT, Ventura J, Zhuang J, et al. Polymer nanogels: a versatile nanoscopic drug delivery platform. Adv Drug Deliv Rev. 2012;64(9):836–851. doi:10.1016/j.addr.2012.02.002
60. Ashrafi B, Rashidipour M, Marzban A, et al. Mentha piperita essential oils loaded in a chitosan nanogel with inhibitory effect on biofilm formation against S. mutans on the dental surface. Carbohydr Polym. 2019;212:142–149. doi:10.1016/j.carbpol.2019.02.018
61. Abyadeh M, Karimi Zarchi AA, Faramarzi MA, et al. Evaluation of factors affecting size and size distribution of chitosan-electrosprayed nanoparticles. Avicenna J Med Biotechnol. 2017;9(3):126–132.
62. Amanzadeh E, Esmaeili A, Abadi REN, et al. Quercetin conjugated with superparamagnetic iron oxide nanoparticles improves learning and memory better than free quercetin via interacting with proteins involved in LTP. Sci Rep. 2019;9(1):6876. doi:10.1038/s41598-019-43345-w
63. Rahimi S, Khoee S, Ghandi M. Preparation and characterization of rod-like chitosan-quinoline nanoparticles as pH-responsive nanocarriers for quercetin delivery. Int J Biol Macromol. 2019;128:279–289. doi:10.1016/j.ijbiomac.2019.01.137
64. Beyki M, Zhaveh S, Khalili ST, et al. Encapsulation of Mentha piperita essential oils in chitosan–cinnamic acid nanogel with enhanced antimicrobial activity against Aspergillus flavus. Ind Crops Prod. 2014;54:310–319. doi:10.1016/j.indcrop.2014.01.033
65. Feng QL, Wu J, Chen GQ, et al. A mechanistic study of the antibacterial effect of silver ions on Escherichia coli and Staphylococcus aureus. J Biomed Mater Res. 2000;52(4):662–668. doi:10.1002/1097-4636(20001215)52:4<662::aid-jbm10>3.0.co;2-3
66. Kim TH, Kim M, Park HS, et al. Size-dependent cellular toxicity of silver nanoparticles. J Biomed Mater Res A. 2012;100(4):1033–1043. doi:10.1002/jbm.a.34053
67. Mukha IP, Eremenko AM, Smirnova NP, et al. [Antimicrobial activity of stable silver nanoparticles of a certain size]. Prikl Biokhim Mikrobiol. 2013;49(2):215–223. doi:10.7868/s0555109913020128
68. Cavassin ED, de Figueiredo LFP, Otoch JP, et al. Comparison of methods to detect the in vitro activity of silver nanoparticles (AgNP) against multidrug resistant bacteria. J Nanobiotechnology. 2015;13:64. doi:10.1186/s12951-015-0120-6
69. Slavin YN, Asnis J, Häfeli UO, et al. Metal nanoparticles: understanding the mechanisms behind antibacterial activity. J Nanobiotechnology. 2017;15(1):65. doi:10.1186/s12951-017-0308-z
70. Milanezi FG, Meireles LM, de Christo Scherer MM, et al. Antioxidant, antimicrobial and cytotoxic activities of gold nanoparticles capped with quercetin. Saudi Pharm J. 2019;27(7):968–974. doi:10.1016/j.jsps.2019.07.005
71. Wang L, Hu C, Shao L. The antimicrobial activity of nanoparticles: present situation and prospects for the future. Int J Nanomedicine. 2017;12:1227–1249. doi:10.2147/IJN.S121956
72. Nordstrom R, Andrén OCJ, Singh S, et al. Degradable dendritic nanogels as carriers for antimicrobial peptides. J Colloid Interface Sci. 2019;554:592–602. doi:10.1016/j.jcis.2019.07.028
73. Saito Y, Kanai Y, Nakagawa T, et al. Increased protein expression of DNA methyltransferase (DNMT) 1 is significantly correlated with the malignant potential and poor prognosis of human hepatocellular carcinomas. Int J Cancer. 2003;105(4):527–532. doi:10.1002/ijc.11127
74. Kaji K, Factor VM, Andersen JB, et al. DNMT1 is a required genomic regulator for murine liver histogenesis and regeneration. Hepatology. 2016;64(2):582–598. doi:10.1002/hep.28563
75. Oh BK, Kim H, Park H-J, et al. DNA methyltransferase expression and DNA methylation in human hepatocellular carcinoma and their clinicopathological correlation. Int J Mol Med. 2007;20(1):65–73.
76. Tang SM, Deng X-T, Zhou J, et al. Pharmacological basis and new insights of quercetin action in respect to its anti-cancer effects. Biomed Pharmacother. 2020;121:109604. doi:10.1016/j.biopha.2019.109604
77. Kim DH, Khan H, Ullah H, et al. MicroRNA targeting by quercetin in cancer treatment and chemoprotection. Pharmacol Res. 2019;147:104346. doi:10.1016/j.phrs.2019.104346
78. Serri C, Quagliariello V, Iaffaioli RV, et al. Combination therapy for the treatment of pancreatic cancer through hyaluronic acid-decorated nanoparticles loaded with quercetin and gemcitabine: a preliminary in vitro study. J Cell Physiol. 2019;234(4):4959–4969. doi:10.1002/jcp.27297
79. Eden A, Gaudet F, Waghmare A, et al. Chromosomal instability and tumors promoted by DNA hypomethylation. Science. 2003;300(5618):455. doi:10.1126/science.1083557
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.