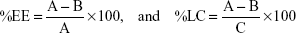Back to Journals » International Journal of Nanomedicine » Volume 11
Aptamer-hybrid nanoparticle bioconjugate efficiently delivers miRNA-29b to non-small-cell lung cancer cells and inhibits growth by downregulating essential oncoproteins
Authors Perepelyuk M, Maher C, Lakshmikuttyamma A, Shoyele S
Received 13 April 2016
Accepted for publication 23 May 2016
Published 28 July 2016 Volume 2016:11 Pages 3533—3544
DOI https://doi.org/10.2147/IJN.S110488
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Carlos Rinaldi
Maryna Perepelyuk, Christina Maher, Ashakumary Lakshmikuttyamma, Sunday A Shoyele
Department of Pharmaceutical Science, College of Pharmacy, Thomas Jefferson University, Philadelphia, PA, USA
Abstract: MicroRNAs (miRNAs) are potentially attractive candidates for cancer therapy. However, their therapeutic application is limited by lack of availability of an efficient delivery system to stably deliver these potent molecules intracellularly to cancer cells while avoiding healthy cells. We developed a novel aptamer-hybrid nanoparticle bioconjugate delivery system to selectively deliver miRNA-29b to MUC1-expressing cancer cells. Significant downregulation of oncoproteins DNMT3b and MCL1 was demonstrated by these MUC1 aptamer-functionalized hybrid nanoparticles in A549 cells. Furthermore, downregulation of these oncoproteins led to antiproliferative effect and induction of apoptosis in a superior version when compared with Lipofectamine 2000. This novel aptamer-hybrid nanoparticle bioconjugate delivery system could potentially serve as a platform for intracellular delivery of miRNAs to cancer cells, hence improving the therapeutic outcome of lung cancer.
Keywords: aptamer, nanoparticles, microRNA, lung cancer, targeted delivery
Introduction
Non-small-cell lung cancer (NSCLC) is one of the leading causes of cancer mortality in the US and the world, with a 5-year survival rate of only 15% for all stages combined.1,2 There has not been any significant decrease in the overall 5-year mortality rate for >4 decades despite the investment of billions of dollars into research.3 This could be attributed to lack of precise process of screening, heterogeneity of diseases, and, most importantly, the lack of novel therapies coming through.
MicroRNAs (miRNAs) are a novel RNA-interference technology that is currently being explored as a promising strategy in lung cancer treatment. They are small in size (~22 nucleotides), and they regulate gene expression at the post-transcription level through RNA interference. They may be seen as a new tool in the box for cancer treatment. Members of miRNA-29 family have been known to play an important role in cancer by regulating cell differentiation, proliferation, apoptosis, invasion, and migration.4,5 However, clinical application of this relatively novel therapeutic faces many challenges, including degradation in serum, rapid blood clearance, stimulation of immune response, off-target effects, and poor cellular uptake.6 Recently, our group developed a novel targeted hybrid nanoparticle delivery system.7–9 This delivery system demonstrated its ability to protect loaded oligonucleotide against serum nuclease, reduce rate of blood clearance, efficient cellular uptake, and selective delivery to lung cancer in a mouse model.7–9 This hybrid nanoparticle delivery system was able to achieve all the aforementioned abilities because of the uniqueness of its makeup. Our hybrid nanoparticle technology benefits from the incorporation of human immunoglobulin G (human IgG) and poloxamer-188 (polyoxyethylene–polyoxypropylene block copolymer), to achieve stable and efficient delivery of nucleic acid-based therapeutics.7 Human IgG is the main encapsulating component of these hybrid nanoparticles. Human IgG antibody also helps to reduce/eliminate well-documented immunogenic reaction experienced with most nanoparticle formulations, by “deceiving” the body to believe that the nanoparticles are natural components of the blood.7 Poloxamer-188, a nonionic triblock copolymer, helps to circumvent the reticuloendothelial system during systemic circulation, since it has been previously used as a stealth polymer by preventing macrophage uptake during circulation.10,11
In the present study, we aim to optimize cellular uptake of miRNA-29b in lung cancer cells using our novel targeted hybrid nanoparticle system to selectively deliver miRNA-29b to lung cancer cells. Furthermore, we demonstrate the cancer-suppressing and apoptotic ability of encapsulated miRNA-29b in lung cancer cell lines (A549). We hypothesize that our novel targeted hybrid nanoparticles will efficiently deliver loaded miRNA-29b to lung cancer cells while avoiding normal cells by conjugation of mucin 1 (MUC1) aptamer to the nanoparticles. Furthermore, we hypothesize that selectively delivered miRNA-29b will downregulate oncoproteins myeloid cell leukemia sequence 1 (MCL1) and DNA methyltransferase 3B (DNMT3B) in lung cancer, hence inhibiting cancer cell growth.
miRNA-29b is the most abundantly expressed of all miRNA-29 families in normal cells.12 Previous studies have demonstrated that miRNA-29b plays an important role in cancer by regulating cell proliferation, differentiation, and apoptosis.5,6,13,14 It is usually silenced or downregulated in lung cancer and other types of cancer.6,15 In contrast, forced expression in acute myeloid leukemia and lung cancer resulted in significant reduction of DNMT3B at both protein and mRNA levels by direct interaction.15 Downregulation of DNMT3B led to inhibition of cell proliferation and apoptosis of lung cancer cells.15,16 Furthermore, MCL1, a potent antiapoptotic protein of the BCL-2 family, has been shown to be downregulated by miRNA-29b in cancer cells.15 miRNA-29b is therefore a very attractive candidate for miRNA-based therapeutics due to its potent tumor suppressant capabilities.
To actively and selectively deliver the encapsulated miRNA-29b to lung cancer cells, these nanoparticles were functionalized with MUC1 aptamer.
MUC1 is a transmembrane protein that is aberrantly overexpressed in NSCLCs.17 MUC1 has been shown to be overexpressed in >80% of lung adenocarcinoma.17 It has also been shown that targeting MUC1 function inhibits mutant K-ras signaling in NSCLC cells.17 We aim to take advantage of this overexpression in NSCLC to achieve an optimized targeting of miRNA-29b to NSCLC by conjugating MUC1 aptamer to the surface of our hybrid nanoparticles.
Previous studies have shown that MUC1 aptamer-functionalized nanoparticles improved the efficacy of drug delivery to breast cancer cells.18
Experimental section
Materials
Human IgG was purchased from Equitech-Bio Inc. (Kerrville, TX, USA). Poloxamer-188, RNase-free water, 4,6-diamidino-2-phenylindole (DAPI), and fetal bovine serum (FBS) were obtained from Thermo Fisher Scientific (Waltham, MA, USA). Aptamer against MUC1 (5′-GCA GTT GAT CCT TTG GAT ACC CTG G-3′) was designed and purchased from GE Healthcare Bio-Sciences Corp. (Piscataway, NJ, USA). A C12 spacer was attached to the aptamer at the 3′-end to form the following sequence: 5′-GCA GTT GAT CCT TTG GAT ACC CTG G-C12H25-3′. One aptamer was modified with 3′-NH2 and 5′-FITC, whereas the other was only modified with 3′-NH2. The aptamers contained a 12-carbon spacer. siGLO-Green (6-FAM-labeled) was obtained from GE Healthcare Bio-Sciences Corp. HCl was obtained from Thermo Fisher Scientific. MiRIDIAN mimic with miR-29b and miRIDIAN mimic NC were obtained from GE Healthcare Bio-Sciences Corp. Pierce RIPA lysis buffer was purchased from Thermo Fisher Scientific. Rabbit antihuman DNMT3B and rabbit antihuman MCL1 antibody were obtained from Thermo Fisher Scientific. Mouse antihuman β-tubulin antibody was purchased from Sigma-Aldrich (St Louis, MO, USA).
Cell culture
Adenocarcinoma cell line A549 and normal lung fibroblast cell line MRC-5 were obtained from American Type Culture Collection (Rockville, MD, USA). A549 cell was originally derived from a 58-year-old male Caucasian, while MRC-5 was originally derived from a 14-week gestation male Caucasian. A549 cells were maintained in F12 K medium supplemented with 10% FBS and 1% antibiotics. MRC-5 was maintained in Eagle’s Minimum Essential Medium supplemented with 10% FBS. Both cells were kept in a humidified air atmosphere with 5% carbon dioxide.
Preparation of nanoparticles
Human IgG was diluted in 0.01N HCl to make up IgG concentration of 1 mg/mL (ie, 10 mg) in 10 mL of 0.01 N HCl. A total of 0.73 mg of miRNA-29b was then added. The mixture was stirred on a magnetic stirrer until all the components were fully dissolved. The mixture was titrated with 0.01 N NaOH up to a pH value close to 7. Nanoparticles were spontaneously formed at the pH value very close to 7. The nanoparticles were allowed to mix on the magnetic stirrer for ~10 minutes. The colloidal suspension was centrifuged using a microcentrifuge (Eppendorf centrifuge 5418) at 2,000 rpm for 5 minutes. The supernatant was either decanted or kept for the measurement of unencapsulated miRNA-29b to be used to calculate encapsulation efficiency and loading capacity. Nanoparticles were then rinsed three times with double distilled deionized water. Nanoparticles were subsequently suspended in 0.2% v/v poloxamer-188, with gentle shaking for 10 minutes in order to coat nanoparticles with poloxamer-188. Nanoparticles were centrifuged and the supernatant decanted before being rinsed thrice with double distilled deionized water. Particles were then loaded into a freeze dryer (Labconco Freeze Zone 4.6), and lyophilization was performed for 48 hours.
Conjugation of aptamer to prepared nanoparticles
A total of 50 μL of poloxamer-free nanoparticle suspension (10 μg/mL in DNase, RNase-free water) was mixed with 100 μL of 40 mM EDC and 100 μL of 10 mM NHS for 15 minutes at room temperature with gentle stirring. This helps to activate the carboxyl groups on nanoparticles for aptamer conjugation. A total of 50 μL of 1 μg/mL aptamer in DNase, RNAse-free water was then added to the nanoparticles and mixed gently for 2 hours at room temperature. Nanoparticles were subsequently centrifuged at 4,000 rpm and 10°C for 5 minutes using 30 kDa cutoff centrifugal ultrafilters (EMD Millipore, Billerica, MA, USA) to exclude unreacted EDC and NHS. Aptamer-functionalized nanoparticles were then suspended in 0.2% v/v poloxamer-188, with gentle shaking for 10 minutes in order to coat them with poloxamer-188.
Characterization of nanoparticles
Particle size and surface charge (zeta potential) of the nanoparticles were measured using photon correlation spectroscopy (ZetaSizer Nano ZS; Malvern Instruments, Malvern, UK). Nanoparticles were dispersed in deionized water followed by sonication for ~5 minutes. A scattering angle (θ) of 173° was used to measure intensity autocorrelation. The Z-average and polydispersity index were recorded in triplicate. For zeta potential, samples were taken in a universal dip cell (Malvern Instruments) and the zeta potential recorded in triplicate.
We evaluated the shape of nanoparticles using scanning electron microscopy. Suspensions of nanoparticles were dropped on an aluminum stub and allowed to dry at room temperature. Samples were then coated with a thin layer of palladium. Coated samples were imaged using a Zeiss Supra 50 V system (Carl Zeiss Meditec AG, Jena, Germany).
Fourier transform-infrared spectroscopy
A single-reflection-attenuated total reflectance with a diamond internal reflection crystal installed in an iS10 FT-IR spectrometer (Thermo Fisher Scientific) was used to obtain spectra from different samples of nanoparticles. Lyophilized powders were placed on the surface of the attenuated total reflectance crystal after background spectra had been collected. Spectra were collected after 64 scans at 4 cm−1 resolution. Data were analyzed using OMNIC software.
miRNA release study
Release of miRNA from MUC1 aptamer-functionalized hybrid nanoparticles was measured at pH values 5, 6.6, and 7.4. Nanoparticles (3 mg) were dispersed in 0.5 mL of a buffered solution in a tubular cellulose dialysis membrane secured tightly at both ends. This was then incubated at 37°C in 5 mL buffered solution reservoirs while the reservoir was gently agitated. The amount of miRNA released at different time points was analyzed and quantified for percentage cumulative release using ion-pair HPLC.
Ion-pair HPLC
miRNA was quantitatively determined using a Waters 2695 separation module combined with a Waters 2998 photodiode array detector Alliance HPLC system (Waters, Milford, MA, USA). Using a Clarity 3 μm Oligo-RP column (Phenomenex, Torrance, CA, USA) with a column dimension of 50×2.0 mm, 1 μL of miRNA sample was injected into the HPLC using 20 mM triethylamine–acetic acid (pH 7) and 5%–12% acetonitrile, gradient elusion as mobile phase. A flow rate of 0.2 mL/min was used for analysis. UV detection was performed at 269 nm, and Empower Pro software was used to record chromatograms.
Fluorescence microscopy
A549 cells were seeded at a density of 2×104 in eight-well coated glass slides (Discovery Labware, Tewksbury, MA, USA) followed by incubation for 48 hours. Cells were washed with PBS and incubated with either FITC-MUC1 aptamer-functionalized miRNA-29b-loaded hybrid nanoparticles or MUC1 aptamer-functionalized siGLO-FAM-loaded hybrid nanoparticles dispersed in the Opti-Mem medium at a concentration of 100 μg/mL for 2 hours or 4 hours. Cells were then washed twice with PBS, fixed using 2% paraformaldehyde, and incubated at room temperature for 20 minutes. Cells washed with PBS were then blocked with 5% BSA for 30 minutes at room temperature. Cells were stained with either Lysotracker-Red or AlexaFluoro-555-labeled wheat germ agglutinin (WGA-AF-555) before being stained with DAPI to visualize nucleus. Leica DMI 6000B fluorescence microscope (Leica Microsystems, Wetzlar, Germany) was then used to observe cells after being mounted.
Reverse transcriptase polymerase chain reaction
Qiagen RNAeasy kit (Qiagen, NV, Venlo, the Netherlands) was used to isolate total RNA and Verzo cDNA kit (Thermo Fisher Scientific) was used for reverse transcription. Polymerase chain reaction (PCR) was carried out in 25 μL reaction mixtures. These reaction mixtures contained 1.0 μL of cDNA, 1× Qiagen buffer, 0.2 mM of dNTP mixture, 0.2 μM of each primer, and 1.5 U of HotStar Taq (Qiagen, Valencia, CA, USA). One cycle reaction at 95°C for 15 minutes was followed by 35 amplification cycles (94°C for 1 minute, 66°C for 1 minute, and 72°C for 1 minute) for MUC1. For β-actin PCR, the same conditions except for the annealing temperature (59°C) were used. Primer sequences of MUC1 are forward 5′-TCTCAAGCAGCCAGCGCCTGCCTG-3′, reverse 5′-TCCCCAGGTGGCAGCTGAACC-3′ and β-actin, forward 5′-CCAAGGCCAACCGCGAGAAGAT-3′, and reverse 5′-TTGCTCGAAGTCCAGGGCGA-3′.
Flow cytometry
Approximately 1 million cells per well (A549 and MRC-5) were seeded in a six-well plate and incubated for 48 hours. Cells were then treated with 100 μg/mL of siGLO-FAM-loaded equivalent nanoparticles resuspended in Opti-MEM medium and incubated for 4 hours. Corresponding cells were pretreated with five times excess of free MUC1 aptamer, 60 minutes prior to being treated with the MUC1 aptamer-functionalized siGLO-FAM-loaded hybrid nanoparticles. Cells were washed with PBS. The cells were then detached by trypsinization and centrifuged at 1,000 rpm for 5 minutes. The pellet was washed and resuspended in PBS. A 0.75 μm cell strainer was used to filter the cells before being analyzed by flow cytometry (BDFACS caliber) to prevent cell aggregates from blocking the tube lines of the instrument. A total of 10,000 cells were measured in each sample.
Cell death detection ELISA
Cell death detection ELISA was carried out according to manufacturer’s instructions (Hoffman-La Roche Ltd., Basel, Switzerland). Briefly, A549 cells were seeded at a density of 2×104/well for 24 hours. Next day they were treated with MUC1 aptamer-functionalized miR-29b-loaded hybrid nanoparticles, MUC1 aptamer-functionalized negative control miRNA-loaded hybrid nanoparticles, and miR-29b in Lipofectamine 2000 Transfection Agent (Ambion, Austin, TX, USA) in Opti-MEM medium. Cells were washed in PBS and lysed in 200 μL/well of incubation buffer for 30 minutes at room temperature. The lysates were centrifuged at 4,000× g for 10 minutes. Supernatant was used right away in the experiment. Plastic wells of the ELISA kit were incubated with coating solution (containing antihistone antibodies) overnight at 4°C. Next day it was substituted by incubation buffer for 30 minutes and washed three times with washing solution. A total of 100 μL of homogenate was placed into the wells and incubated for 90 minutes at room temperature with mild shaking. It was washed three times and incubated with conjugate solution containing anti-DNA peroxidase antibodies for 90 minutes. Wells were washed three times and exposed to ABTS substrate for 30 minutes until the color developed. Absorption values were read using a BioTek Epoch Microplate Spectrophotometer (BioTek Instruments Inc., Winooski, VT, USA) with Gen5 1.10 software.
Western blot analysis
One million A549 cells per well were seeded in culture dish and incubated for 48 hours. Cells were treated with MUC1 aptamer-functionalized miR-29b-loaded hybrid nanoparticles (at an equivalent concentration of 100 nM of miRNA-29b), MUC1 aptamer-functionalized negative control miRNA-loaded hybrid nanoparticles as well as miR-29b in Lipofectamine 2000 reagent in Opti-MEM medium for 3 days. Cells were lysed in Pierce® RIPA buffer (Thermo Fisher Scientific) with addition of Pierce™ Protease Inhibitor Mini Tablets (Thermo Fisher Scientific) for 30 minutes on ice and centrifuged for 10 minutes at 10,000× g at +4°C. The protein concentration was determined with the Coomassie Plus™ (Bradford) Assay Kit (Thermo Fisher Scientific). A total of 120 μg of total proteins was separated on NuPAGE 4%–12% Bis–Tris gels (Thermo Fisher Scientific) with NuPAGE MES SDS running buffer (Thermo Fisher Scientific) and subsequently transferred onto a nitrocellulose membrane with a pore size of 0.22 μm (Thermo Fisher Scientific). The membrane with proteins was blocked according to manufacturer’s instructions (Thermo Fisher Scientific) for 1 hour at room temperature and probed with primary antibodies overnight at 4°C DNMT3B (1:1,000, Thermo Fisher Scientific) and MCL1 (1:500; Thermo Fisher Scientific). β-Tubulin was used as a housekeeping gene (1:2,000, Sigma-Aldrich). Next day the membranes were washed three times for 5 minutes in wash buffer according to manufacturer’s instructions (Thermo Fisher Scientific) and incubated with secondary goat antimouse (Molecular Probes, Eugene, OR) or goat antirabbit (1:1,000, Thermo Fisher Scientific) secondary antibodies, conjugated with horseradish peroxidase at a dilution of 1:1,000.
Immune complexes were detected with chemiluminescent substrate, Pierce ECL Western Blotting substrate (Thermo Fisher Scientific), in a dark room on tabletop processor SRX-101A (Konica Minolta, Tokyo, Japan).
Cell viability
MTT assay was used to determine the effect of MUC1 aptamer-functionalized miRNA-29b-loaded hybrid nanoparticles on the proliferation of A549 cells. Cells (1×104) per well were seeded in 96-well plates and incubated at 37°C in a humidified atmosphere with 5% carbon dioxide for 24 hours. The cells were then treated with different concentrations of miRNA-29b-loaded hybrid nanoparticles in Opti-MEM for 7 hours before being replaced with F12 K medium supplemented with 10% FBS and 1% antibiotics. This was then incubated for 72 hours. miRNA-29b transfected with NeoFX transfection agent was used as control in this experiment. Approximately 10 μL of 12 mmol/L MTT reagent was then added to each well. This was then incubated at 37°C for 4 hours. The medium was aspirated and 50 μL of sterile dimethyl sulfoxide was added to each well and mixed thoroughly with pipette. The cells were then incubated at 37°C for 10 minutes. The plate was read at 540 nm and 650 nm.
Statistical analysis
Results are presented as mean ± SD, unless otherwise indicated. Statistically significant difference between two groups was determined by two-tailed Student’s t-test. A P-value of 0.05 was taken as statistically significant.
Results
Nanoparticle characterization
Particle size and zeta potential analysis by photon correlation spectroscopy, shown in Table 1, demonstrated that nonfunctionalized hybrid nanoparticles were ~240 nm in size, while MUC1 aptamer-functionalized hybrid nanoparticles were ~595 nm in size. Nonfunctionalized hybrid nanoparticles were negatively charged with a zeta potential of −2.1, while the conjugation of MUC1 aptamer increased the zeta potential of these nanoparticles to +4.1.
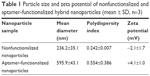  | Table 1 Particle size and zeta potential of nonfunctionalized and aptamer-functionalized hybrid nanoparticles (mean ± SD, n=3) |
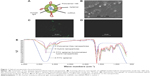
The schematic of the MUC1 aptamer-functionalized hybrid nanoparticle is shown in Figure 1A. Scanning electron micrograph in Figure 1B reveals that the hybrid nanoparticles were spherical in shape with smooth surface.
miRNA encapsulation efficiency (EE) and loading capacity (LC) were measured using ion-pair HPLC analysis of the filtrates obtained by centrifuging the nanoparticles formed to determine the amount of unencapsulated miRNA-29b. EE and LC were calculated using the following equation:
|
|
where A = total amount of miRNA, B = free miRNA, and C = weight of hybrid nanoparticles in grams.
EE was calculated to be 98.8%±0.4%, whereas LC was calculated to be 8.6%±0.1%.
Conjugation of MUC1 aptamer to the surface of the nanoparticles was verified using fluorescence microscopy. Figure 1C demonstrates the successful conjugation of FITC-MUC1 aptamer to the nanoparticles by the presence of fluorescent green color in the micrograph. In contrast, nonfunctionalized hybrid nanoparticles in Figure 1D showed the absence of fluorescent green color, confirming the lack of FITC-MUC1 aptamer in the nanoparticles. We also used Fourier transform-infrared (FT-IR) to confirm the successful conjugation of MUC1 aptamer to the nanoparticles. Figure 1E shows the FT-IR spectra generated from functionalized and nonfunctionalized hybrid nanoparticles. A distinctive and conspicuous difference between these sets of spectra bears testament to the successful conjugation of MUC1 aptamer to the hybrid nanoparticles.
In vitro release study
The release of miRNA-29b from MUC1 aptamer-functionalized hybrid nanoparticles was compared at pH values 5, 6.6, and 7.4. Figure 2 demonstrates a limitation in the release of miRNA-29b at pH values 6.6 and 7.4 when compared with the release profile at pH 5.
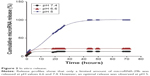  | Figure 2 In vitro release. |
Expression of MUC1 protein in NSCLC cells
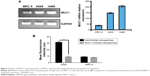
Reverse transcriptase PCR was used to confirm the expression of MUC1 in two NSCLC cells, A549 and H460, to ensure that these cells actually express MUC1. Figure 3A demonstrates the relatively higher expression of MUC1 in both cancer cells when compared with normal lung fibroblast cell line MRC-5.
Cellular uptake of MUC1 aptamer-functionalized hybrid nanoparticles
Internalization of nanoparticles was evaluated using both flow cytometry and fluorescence microscopy. Flow cytometry was used to compare nanoparticle uptake by A549 and MRC-5. siGLO-FAM (green) was used as a model small double-stranded RNA labeled with FAM (a green dye). MUC1 aptamer conjugated to siGLO-loaded hybrid nanoparticles used in this experiment was bereft of FITC so as to avoid double fluorescence. As shown in Figure 3B, internalization of MUC1 aptamer-functionalized siGLO-FAM-loaded nanoparticles in A549 cells was significantly higher (P≤0.001) in noninhibited cells when compared with cells pretreated with MUC1 aptamer. Furthermore, uptake of nanoparticles by MRC-5 was much lower than the uptake by A549 cells.
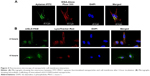
Fluorescence microscopy was used to study the interaction of FITC-MUC1 aptamer-functionalized hybrid nanoparticles with the cell membrane of A549 cells. Figure 4A demonstrates the presence of FITC-MUC1 aptamer-functionalized hybrid nanoparticles (green color) on the cell membrane and inside the cytosol of A549 cells after 2 hours of incubation. Cell membrane of cells was stained with WGA-Alexa Fluor 555 (red color) to enable the identification of the boundaries of cells. Colocalization of both green and red colors on the membrane of the cells suggests a possible interaction between the FITC-MUC1 aptamer-functionalized hybrid nanoparticles and the membrane of the cells possibly due to the presence of MUC1 on the membrane.
Intracellular trafficking of internalized nanoparticles was also monitored using fluorescence microscopy. Late endosomes/lysosomes were labeled with LysoTracker-Red to enable the monitoring of the fate of internalized nanoparticles. Figure 4B demonstrates maximum level of colocalization of siGLO-FAM and LysoTracker-Red after 2 hours of incubation. However, after 4 hours, the siGLO-FAM was observed to be leaving the late endosomes and moving into the cytoplasm as indicated by the numerous green colored dots surrounding the nucleus.
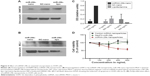
MUC1 aptamer-functionalized miRNA-29b-loaded hybrid nanoparticles downregulate DNMT3B and MCL1 proteins and induce apoptosis in A549 cells
Figure 5A demonstrates the knockdown efficiency of MUC1 aptamer-functionalized miRNA-29b-loaded nanoparticles against DNMT3B. These nanoparticles produced superior downregulation of DNMT3B when compared with Lipofectamine 2000 transfected miRNA-29b and the negative control miRNA-loaded nanoparticles. Figure 5B further demonstrates the superior transfection efficiency of MUC1 aptamer-functionalized miRNA-29b-loaded nanoparticles in comparison to lipofectamine by effectively downregulating MCL1. In contrast, both lipofectamine and MUC1 aptamer-functionalized negative control miRNA-loaded hybrid nanoparticles did not achieve the same level of downregulation.
Superior induction of apoptosis was observed in A549 cells treated with MUC1 aptamer-functionalized miRNA-29b-loaded hybrid nanoparticles when compared with nontreated cells, MUC1 aptamer-functionalized negative control miRNA-loaded hybrid nanoparticles-treated cells and lipofectamine-transfected miRNA-29b (Figure 5C).
Antiproliferative effect of MUC1 aptamer-functionalized miRNA-29b-loaded nanoparticles was evaluated using MTT assay and compared with that of NeoFX-transfected miRNA-29b and negative control miRNA-loaded nanoparticles in A549 cells. As demonstrated in Figure 5D, MUC1 aptamer-functionalized miRNA-29b-loaded nanoparticles were significantly more cytotoxic to A549 cells than NeoFX-transfected miRNA-29b and negative control miRNA-loaded miRNA nanoparticles.
Discussion
Translational application of miRNA-based therapeutics is limited by a lack of smart nanoparticle delivery system to selectively deliver these important molecules intracellularly to cancer cells.
Recently, our group developed a novel hybrid nanoparticles delivery system for safe and effective delivery of nucleic acid-based therapeutics to lung cancer.7–9 These nanoparticles were able to protect loaded nucleic acid against serum nuclease both in vitro and in vivo.7,9 Furthermore, these hybrid nanoparticles were able to successfully deliver siRNA against mutant K-ras to the cytosol of A549 lung cancer cell lines without eliciting immunogenic/inflammatory response in murine macrophages, RAW 264.7.7,8 A significant downregulation of both protein and mRNA of K-ras was achieved.7 In the present study, we were able to show that these nanoparticles could be functionalized with MUC1 aptamer to enable selective delivery of miRNAs to NSCLC cells. miRNA-29b was selected as a model miRNA since it has been previously reported that miRNA-29b, a tumor suppressor miRNA, is aberrantly expressed in NSCLC and its perturbation is related to tumor development and progression.4–6,19 Hence, miRNA-29b has been identified as an attractive candidate for miRNA-based therapeutics in NSCLC.
Hybrid nanoparticles comprising human IgG and poloxamer-188 were prepared using our previously reported isoelectric point (pI)-based nanoprecipitation technology.7,8,20 This technology is based on the fact that proteins (human IgG) have minimum solubility but maximum precipitation at their pI. Nanoparticles produced using this technology are characteristically spherical in morphology as seen in Figure 1B. The particles were negatively charged possibly because of the presence of residual miRNA on the surface of the nanoparticles. However, the charge on the particles became positive following the conjugation of aptamer to the surface of the nanoparticles. The positive charge could be attributed to the presence of an amino group NH2 in the aptamer to facilitate the conjugation reaction. The loss of the lone pair of electrons on the amino group resulted in the positive charges. Both fluorescence microscopy and FT-IR analysis were performed to confirm the successful conjugation of MUC1 aptamer to the preformed hybrid nanoparticles. As shown in Figure 1C, FITC-labeled MUC1 aptamer present on the surface of the hybrid nanoparticles led to the presence of green color in the MUC1 aptamer-functionalized hybrid nanoparticles. In contrast, nonfunctionalized hybrid nanoparticles did not show any green color in Figure 1D. This method of confirmation of successful aptamer conjugation to nanoparticles has been previously reported by Farokhzad et al.21 FT-IR data in Figure 1E demonstrate a distinctive difference between the spectra of MUC1 aptamer-functionalized hybrid nanoparticles and the nonfunctionalized hybrid nanoparticles. Nonfunctionalized hybrid nanoparticles had a peak at 3,274 cm−1, which can be attributed to the group stretching vibration of both NH2 and OH in the human IgG.22,23 Following the conjugation of MUC1 aptamer to the hybrid nanoparticles, this peak became sharper and more intense due to the presence of additional NH2 and OH stretching vibration attributable to the conjugation of NH2 aptamer to the hybrid nanoparticles. Both MUC1 aptamer-functionalized and nonfunctionalized nanoparticles show a peak at ~1,541 cm−1, which could be attributed to the presence of amide II carbonyl stretch from human IgG in both nanoparticles.24 Furthermore, the peak at ~1,660 cm−1 found in the spectra for MUC1 aptamer-functionalized hybrid nanoparticles but absent in the corresponding nonfunctionalized hybrid nanoparticles is attributed to conjugated amide stretching due to the conjugation of the amino group from the aptamer to the carboxyl groups present in human IgG.
Figure 2 demonstrates a limitation in the release of miRNA from MUC1 aptamer-functionalized hybrid nanoparticles at pH values 6.6 and 7.4 when compared with the release profile at pH 5. This demonstrates the pH-sensitive nature of these nanoparticle delivery systems. At pH values of 6.6 and 7.4, very limited amount of the loaded miRNA (<20%) was released throughout the study period. The limited release of siRNA at pH values 6.6 and 7.4 could be attributed to the reduced/limited solubility of human IgG at these pH values. Proteins are known to have limited solubility at pH values close to their pI.22 Since the pI of human IgG is 7, its solubility at neutral pH values is quite limited. This makes it difficult for the encapsulated miRNA to be released at this pH value, hence possibly limiting its release extracellularly in blood and tumor microenvironment. However, at acidic pH value of 5, an optimal miRNA release of ~100% was obtained due to the solubility of human IgG at this pH. An optimum release of the loaded miRNA is very desirable at pH 5, as pH 5 represents the acidic condition of endosome/lysosome.25,26
Uptake of MUC1 aptamer-functionalized nanoparticles, as demonstrated in Figure 3B, is significantly higher in A549 cells in comparison to MRC-5 cells. The differential internalization of nanoparticles between these cell lines could be attributed to the differential expression of MUC1 on the surface of these cells. While A549 is an adenocarcinoma cell line that is known to aberrantly overexpress MUC1,17 MRC-5 is a normal lung fibroblast cell line with limited expression of MUC1 as shown in Figure 3A. To further confirm the involvement of MUC1 in the internalization of these nanoparticles, cells were pretreated with free MUC1 aptamer prior to their treatment with the MUC1 aptamer-functionalized hybrid nanoparticles to further elucidate the role of MUC1 in uptake of nanoparticles. In Figure 3B, the significantly lower uptake of nanoparticles by A549 cells following pretreatment with free MUC1 aptamer could be attributed to competitive binding of this free MUC1 aptamer to MUC1 on the membrane of the cells, limiting the uptake of MUC1 aptamer-functionalized hybrid nanoparticles. Figure 4A further demonstrates the interaction between the cell membrane of A549 cells and FITC-MUC1 aptamer conjugated to the hybrid nanoparticles. Presence of green color on the membrane of the A549 cells suggests the binding of these nanoparticles to the membrane prior to internalization.
One of the challenges facing the translational application of nanomedicine is the lack of effective endosomal escape following internalization by host cells. Stable nucleic acid lipid particles, which to date are the most advanced delivery system for nucleic acid, have demonstrated some limitation in recent studies. It was revealed that ~70% of the internalized nucleic acid undergoes exocytosis (endocytic recycling) through egress of the lipid nanoparticles from late endosomes and lysosomes.27 It is therefore important to design nanoparticle delivery system with the ability to escape the recycling pathways. To demonstrate the ability of our hybrid nanoparticles to escape endocytic recycling, we monitored intracellular trafficking of internalized nanoparticles using fluorescence microscopy. In Figure 4B, colocalization of nanoparticles (green) and endosome (red) indicates the presence of nanoparticles in the late endosome after 2 hours of incubation.7 However, release of siGLO-FAM (green) can be seen after 4 hours of incubation, indicating the escape of encapsulated siGLO-FAM from late endosomes into the cytosol of the cells. This was possible due to the buffering capacity of the hybrid nanoparticles in the endosome. The solubility of IgG and poloxamer-188 at acidic pH 5, as demonstrated by the in vitro release data in Figure 2, makes it possible for the dissolved IgG in the endosome to activate the proton pump that raises osmotic pressure in the endosome subsequently leading to the swelling and subsequent escape of siGLO-FAM from the endosomes into the cytoplasm.27,28
To evaluate the ability of internalized miRNA-29b to downregulate target oncoproteins DNMT3B and MCL1, expression of these proteins was monitored using Western blot. As demonstrated in Figure 5A and B, both target oncoproteins were downregulated in A549 cells following treatment with MUC1 aptamer-functionalized miRNA-29b-loaded hybrid nanoparticles in a superior version when compared with lipofectamine-transfected miRNA-29b. DNMT3B has been identified as a member of the DNA methyltransferase family accounting for inactivation of tumor suppressor genes in many cancer cells.29 miRNA-29b has been reported to exert its tumor-suppressive role by directly targeting DNMT3b in cancer cells.5,6,29 Furthermore, miRNA-29b has been reported to be downregulated in malignant cells, whereas MCL1 was upregulated.5,6,30 MCL1 is a potent antiapoptic protein of the BCL-2 family. Downregulation of both DNMT3B and MCL1 by miRNA-29b has been shown to inhibit cancer cell growth and promote apoptosis.29,30 However, the challenge facing the clinical application of miRNA-29b to cancer therapy is the availability of an efficient and safe delivery system to enhance intracellular delivery of this potent molecule to cancer cells. Figure 5A and B demonstrates the superior efficiency of MUC1 aptamer-functionalized miRNA-29b-loaded hybrid nanoparticles in downregulating oncoproteins DNMT3b and MCL1 in NSCLC cells over lipofectamine-delivered miRNA-29b. The ability of this treatment modality to induce apoptosis in A549 cells due to downregulation of DNMT3B and MCL1 was also demonstrated in Figure 5C. MUC1 aptamer miRNA-29b-loaded hybrid nanoparticles also demonstrated superior antiproliferative effect in A549 cells in comparison to NeoFX-delivered miRNA-29b. NeoFX is another commonly used, commercially available nucleic acid transfection agent.
Conclusion
MUC1 aptamer-functionalized hybrid nanoparticle delivery system was successfully fabricated. Our results demonstrated that this delivery system can efficiently deliver miRNAs effectively to cancer cells in a superior version compared with commercially available transfection agents. We also demonstrated that direct downregulation of DNMT3b and MCL1 led to antiproliferative effect in A549 cells. This novel aptamer-hybrid nanoparticle bioconjugate delivery system could potentially serve as a platform for intracellular delivery of miRNAs to cancer cells, hence improving the therapeutic outcome of lung cancer.
Acknowledgments
The authors acknowledge the support of the Science Center, Philadelphia and Thomas Jefferson University, toward this project through the award of QED grant number: S1402.
Disclosure
The authors report no conflicts of interest in this work.
References
Howlader N, Noone AM, Krapcho M, et al. Seer Cancer Statistics Review 1975–2009 (Vintage 2009 Population). Bethesda: National Cancer Institute; 2009. Available from: http://seer.cancer.gov/csr/1975_2009_pops09/. Accessed January 4, 2016. | ||
Jemal A, Siegel R, Ward E, Murray T, Xu J, Thun MJ. Cancer statistics, 2007. CA Cancer J Clin. 2007;57:43–66. | ||
Marshall E. Cancer research and the $90 billion metaphor. Science. 2011;331(6024):1540–1541. | ||
Kriegel AJ, Liu Y, Fang Y, Ding X, Liang M. The miR-29 family: genomics, cell biology and relevance to renal and cardiovascular injury. Physiol Genomics. 2012;44(4):237–244. | ||
Schmitt MJ, Margue C, Behrmann I, Kreis S. miRNA-29: a microRNA family with tumor-suppressing and immune-modulating properties. Curr Mod Med. 2012;13(4):572–585. | ||
Yadava PK. Nucleic Acid Therapeutics: Current Targets For Antisense. Mol Biol Today. 2000;1(1):1–16. | ||
Lakshmikuttyamma A, Sun Y, Lu B, Undieh AS, Shoyele SA. Stable and efficient transfection of siRNA for mutated KRAS silencing using novel hybrid nanoparticles. Mol Pharm. 2014;11(12):4415–4424. | ||
Dim N, Perepelyuk M, Gomes O, et al. Novel targeted siRNA-loaded hybrid nanoparticles: preparation, characterization and in vitro evaluation. J Nanobiotechnology. 2015;13:61. | ||
Perepelyuk M, Thangavel C, Liu Y, et al. Biodistribution and pharmacokinetic study of siRNA-loaded anti-NTSR1-mAb-functionalized novel hybrid nanoparticles in metastatic orthotopic murine lung cancer model. Mol Ther Nucleic Acids. 2016;5:e282. | ||
Jain D, Athawale R, Bejay A, Shrikhande S, Goel PN, Gude RP. Studies on stabilization mechanism and stealth effect of 681 poloxamer 188 onto PLGA nanoparticles. Colloids Surf B Biointerfaces. 2013;109:59–67. | ||
Zhang WL, Liv JP, Chen ZQ. Stealth tanshinone IIA-684 loaded solid lipid nanoparticles: effects of poloxamer 188 coating on in vitro phagocytosis and in vivo pharmacokinetics in rats. Acta Pharm Sin. 2009;44(12):1422–1428. | ||
Yan B, Guo Q, Fu F-J, et al. The role of miR-29b in cancer: regulation, function, and signaling. Onco Target Ther. 2015;8:539–548. | ||
Wu Y, Crawford M, Mao Y, et al. Therapeutic delivery of microRNA-29b by cationic lipoplexes for lung cancer. Mol Ther Nucleic Acids. 2013;2:e84. | ||
Garzon R, Heaphy CE, Havelange V, et al. MicroRNA 29b function in acute myeloid leukemia. Blood. 2009;114(26):5331–5341. | ||
Wang Y, Zhang X, Li H, Yu J, Ren X. The role of miRNA-29 family in cancer. Eur J Cell Biol. 2013;92(3):123–128. | ||
Pandey M, Sultana S, Gupta KP. Involvement of epigenetics and microRNA-29b in the urethane induced inception and establishment of mouse lung tumors. Exp Mol Pthol. 2014;96(1):61–70. | ||
Kharbanda A, Rajabi H, Jin C, Alam M, Wong K-K, Kufe D. Muc 1 confers EMT and KRAS independence in mutant KRAS lung cancer cells. Oncotarget. 2014;5(19):8893–8904. | ||
Yu C, Hu Y, Duan J, et al. Novel aptamer-nanoparticle bioconjugates enhances delivery of anticancer to MUC1-positive cancer cells in vitro. PLoS One. 2011;6(9):e24077. | ||
Liu H, Wang B, Lin J, Zhao L. microRNA-29b: an emerging player in human cancer. Asian Pac J Cancer Prev. 2014;15(21):9059–9064. | ||
Srinivasan AR, Shoyele SA. Self –associated IgG1 submicron particles for pulmonary drug delivery: effects of non-ionic surfactants on size, shape, stability and aerosol performance. AAPS PharmSciTech. 2013;14(1):200–210. | ||
Farokhzad OC, Jon S, Khademhosseini A, Tran T-NT, LaVan DA, Langer R. Nanoparticle-aptamer bioconjugate: a new approach for targeting prostate cancer. Cancer Res. 2004;64(21):7668–7672. | ||
Srinivasan AR, Shoyele SA. Influence of surface modification and the pH on the release mechanisms and kinetics of erlotinib from antibody-functionalized chitosan nanoparticles. Ind Eng Chem Res. 2014;53:2987–2993. | ||
Guo M, Diao P, Cai S. Hydrothermal growth of well aligned ZnO nanorod arrays: dependence of morphology and alignment ordering upon preparing conditions. J Solid State Chem. 2005;178:1864–1873. | ||
AbdElhady M. Preparation and characterization of chitosan/zinc oxide nanoparticles for imparting antimicrobial and UV protection to cotton fabric. Int J Carbohydr Chem. 2012;2012:6. | ||
Juliano RL, Ming X, Nakagawa O. Cellular uptake and intracellular trafficking of antisense and siRNA oligonucleotides. Bioconjug Chem. 2013;23(2):147–157. | ||
Luzio JP, Pryor PR, Bright NA. Lysosomes: fusion and function. Nat Rev Mol Cell Biol. 2008;8(8):622–632. | ||
Sahay G, Querbes W, Alabi C, et al. Efficiency of siRNA delivery by lipid nanoparticles is limited by endocytic recycling. Nat Biotechnol. 2013;31(7):653–658. | ||
Dhar S, Gu FX, Langer R, Farokhzad OC, Lippard SJ. Targeted delivery of cisplatin to prostate cancer cells by aptamer functionalized Pt(IV) prodrug-PLGA-PEG nanoparticles. Proc Natl Acad Sci U S A. 2008;105(45):17356–17361. | ||
Yan B, Guo Q, Nan X-X, et al. Microribonucleic acid 29b inhibits cell proliferation and invasion and enhances cell apoptosis and chemotherapy effects of cisplatin via targeting of DNMT3B and AKT3 in prostate cancer. Onco Target Ther. 2015;8:557–565. | ||
Mott JL, Kobayashi S, Bronk SF, Goves GJ. mir-29 regulates mcl-1 protein expression and apoptosis. Oncogene. 2007;26(42):6133–6140. |
 © 2016 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2016 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.