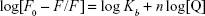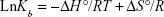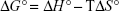Back to Journals » International Journal of Nanomedicine » Volume 14
Albumin binding, anticancer and antibacterial properties of synthesized zero valent iron nanoparticles
Authors Sedaghat Anbouhi T, Mokhtari Esfidvajani E, Nemati F, Haghighat S, Sari S, Attar F, Pakaghideh A, Sohrabi MJ, Mousavi SE, Falahati M
Received 23 September 2018
Accepted for publication 22 November 2018
Published 28 December 2018 Volume 2019:14 Pages 243—256
DOI https://doi.org/10.2147/IJN.S188497
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Thomas Webster
Tabassom Sedaghat Anbouhi,1,* Elnaz Mokhtari Esfidvajani,2,* Fahimeh Nemati,1 Setareh Haghighat,3 Soyar Sari,2 Farnoosh Attar,4 Arezoo Pakaghideh,5 Mohammad Javad Sohrabi,6 Seyyedeh Elaheh Mousavi,6 Mojtaba Falahati7
1Department of Biotechnology, Faculty of Advanced Science and Technology, Tehran Medical Sciences, Islamic Azad University, Tehran, Iran; 2Department of Cellular and Molecular Biology, Faculty of Advanced Science and Technology, Tehran Medical Sciences, Islamic Azad University, Tehran, Iran; 3Department of Microbiology, Faculty of Advanced Science and Technology, Tehran Medical Sciences, Islamic Azad University, Tehran, Iran; 4Department of Biology, Faculty of Food Industry and Agriculture, Standard Research Institute (SRI), Karaj, Iran; 5Department of Toxicology and Pharmacology, Faculty of Pharmacy, Pharmaceutical Science Branch, Islamic Azad University (IAUPS), Tehran, Iran; 6Department of Pharmacology, School of Medicine, Tehran University of Medical Sciences, Tehran, Iran; 7Department of Nanotechnology, Faculty of Advanced Science and Technology, Tehran Medical Sciences, Islamic Azad University, Tehran, Iran
*These authors contributed equally to this work
Background: Nanoparticles (NPs) have been emerging as potential players in modern medicine with clinical applications ranging from therapeutic purposes to antimicrobial agents. However, before applications in medical agents, some in vitro studies should be done to explore their biological responses.
Aim: In this study, protein binding, anticancer and antibacterial activates of zero valent iron (ZVFe) were explored.
Materials and methods: ZVFe nanoparticles were synthesized and fully characterized by X-ray diffraction, field-emission scanning electron microscope, and dynamic light scattering analyses. Afterward, the interaction of ZVFe NPs with human serum albumin (HSA) was examined using a range of techniques including intrinsic fluorescence, circular dichroism, and UV–visible spectroscopic methods. Molecular docking study was run to determine the kind of interaction between ZVFe NPs and HSA. The anticancer influence of ZVFe NPs on SH-SY5Y was examined by MTT and flow cytometry analysis, whereas human white blood cells were used as the control cell. Also, the antibacterial effect of ZVFe NPs was examined on Pseudomonas aeruginosa (ATCC 27853), Escherichia coli (ATCC 25922), and Staphylococcus aureus (ATCC 25923).
Results: X-ray diffraction, transmission electron microscope, and dynamic light scattering analyses verified the synthesis of ZVFe NPs in a nano-sized diameter. Fluorescence spectroscopy analysis showed that ZVFe NPs spontaneously formed a complex with HSA through hydrogen bonds and van der Waals interactions. Also, circular dichroism spectroscopy study revealed that ZVFe NPs did not change the secondary structure of HSA. Moreover, UV–visible data presented that melting temperature (Tm) of HSA in the absence and presence of ZVFe NPs was almost identical. Molecular dynamic study also showed that ZVFe NP came into contact with polar residues on the surface of HSA molecule. Cellular assays showed that ZVFe NPs can induce cell mortality in a dose-dependent manner against SH-SY5Y cells, whereas these NPs did not trigger significant cell mortality against normal white bloods in the concentration range studied (1–100 µg/mL). Antibacterial assays showed a noteworthy inhibition on both bacterial strains.
Conclusion: In conclusion, it was revealed that ZVFe NPs did not induce a substantial influence on the structure of protein and cytotoxicity against normal cell, whereas they derived significant anticancer and antibacterial effects.
Keywords: zero valent iron, nanoparticle, spectroscopy, docking, anticancer, antibacterial activity
Introduction
In the past few years, there has been a rapid interest in the application of nanoparticles (NPs) in the biochemical sensing,1 diagnosis,2 and therapy systems.3 NPs are expected to play an important role in the formation of several technological,4 biological,5 and biomedical innovations.6 To date, a large number of NPs have been designed and synthesized, especially those made from noble metals NPs.7,8 Through the potential features of pharmacodynamics and pharmacokinetics, metal NPs can be offered as an excellent candidate in the medicinal chemistry.9,10 Hence, synthesis and development of unique anticancer11,12 and antimicrobial drugs derivatives12 might be productive in the design of biologically active agents. Zero valent iron (ZVFe) NPs that show unique optical, electronic, and chemical features provide efficient uses in chemistry, biology, and environmental sciences because of suitable surface bioconjugation with molecular probes and significant optical or magnetic properties.13 Especially in modern biological researches, ZVFe NPs with the correct substrate could be implemented to take advantage of the degradation potentials of microbial communities.14 Also, it has been well documented that ZVFe NPs could increase head and neck cancer cells mortality15,16 and ovarian cancer cells16 through mitochondria-mediated apoptosis or autophagy.
As nanobiomedicine has been developing a growing interest in the application of NPs as anticancer or antimicrobial drugs, determining their adverse effects regarding their impacts on the structural changes of some carrier proteins like human serum albumin (HSA) and cytotoxicity against normal cells become crucially important.17
NPs may become lodged in tissues, and the high durability and reactivity of some NPs raise concerns of their outcomes in the healthy systems.18,19 Indeed, at this time, not enough data exist to evaluate the biomedical exposure for most engineered NPs. Some NPs that accumulate in animal tissues may have the ability to easily pass through the cell membranes or cross the blood–brain barrier and attack the cells. This may be a potential feature for implementations such as targeted drug delivery and other disorders treatments, but could lead to unintended influences in some cases. Thus, the influence of NPs targeted to human is inevitable as they can come into contact with body through biomedical applications. Furthermore, the small size of NPs is similar to that of most biomacromolecules, and this results in the interaction of NPs with proteins. Therefore, it is vital to explore in advance the detailed interaction mechanism between NPs and biological systems such as cell and proteins before using them as medical agents. For example, Maji et al20 showed that plant extract-mediated silver NPs provide potential protein binding and antibacterial and anticancer activities. Karthika et al21 reported the fabrication of silver and gold NPs by plant extract and investigated their DNA/protein interactions, anticancer and antimicrobial activities. Khan and Al-Thabaiti22 revealed that green-fabricated ZVFe NPs have a strong albumin binding and significant antimicrobial activities.
Therefore, we aimed to fabricate ZVFe NPs with chemical method and investigate their interaction with HSA by biophysical and molecular docking studies. Afterwards, their anticancer and antibacterial effects against human neuroblastoma cell line (SH-SY5Y) and three bacterial strains were investigated, respectively.
HSA is known as a soluble protein in the blood circulatory system showing a number of physiological roles including transporters for a variety of organic and inorganic compounds and controlling the osmotic pressure and pH of blood.23 HSA is composed of 585 residues with dominant α-helix conformation, three structural domains (I–III), and numerous efficient drug binding sites.24 HSA has been widely used as one of the most extensively studied proteins, not only because of its biomedical significance, low cost, availability, and wide approval in the pharmaceutical industry, but also due to its ligand-binding properties.25,26
The white blood cells (WBCs) and SH-SY5Y cells27,28 are highly sensitive and commonly used for assessing the NP toxicity and their anticancer effects, respectively.
Also, the antibacterial effect of ZVFe NPs was investigated on Pseudomonas aeruginosa, Escherichia coli, and Staphylococcus aureus.
Experimental
Synthesis of ZVFe NPs
The method of synthesis was modified according to Yuvakkumar et al.29 For the synthesis of ZVFe NPs, an aqueous solution of 0.14 M FeSO4·7H2O was dissolved in a 4/1 (v/v) ethanol/water mixture and stirred well with an electric rod for 15 minutes. Then, sodium borohydride solution (100 mL of 0.2 M) was added dropwise into the mixture with gentle stirring for another 30 minutes. The product was separated by centrifugation at 5,000 rpm for 15 minutes and washed three times with ethanol to remove excess borohydrate. Prepared NPs were finally dried under vacuum overnight.
X-ray diffraction analysis
Powder X-ray diffraction (XRD) analysis of ZVFe NP was carried out using Bruker D8 Advance Diffractometer (Bruker AXS, Karlsruhe, Germany) with Cu Kα radiation (λ=1.54 Å) at 45 kV and 40 mA. ZVFe NP was placed in a glass holder and scanned over a 2θ range of 20°–80°. Scan rate was 0.5 min−1.
Field-emission transmission electron microscope study
Field-emission transmission electron microscope (FETEM) analysis of ZVFe NP was carried out using a Philips JSM-6360LA instrument (Philips, Eindhoven, Netherlands). Samples were prepared by sonication for 30 minutes using a sonicator (Misonix S3000).
Dynamic light scattering study
Hydrodynamic diameter, polydispersity index (PDI), and zeta potential of ZVFe with a concentration of 50 μM was measured by using a dynamic light scattering (DLS; Malvern, He/Ne 633 nm laser) at room temperature. For each sample, three independent measurements were done with five scans for each run.
Fluorescence spectroscopy
Fluorescence emission spectra were recorded at three different temperatures of 298, 310, and 315 K on a Cary Eclipse VARIAN fluorescence spectrophotometer (Hitachi, Japan). All measurements were carried out at an excitation wavelength of 280 nm and spectral bandwidths of 5 nm. Static fluorescence spectra of HSA and ZVFe NP mixtures were determined in 20 mM phosphate buffer, pH 7.5 at a fixed protein concentration of 2 μM, and varying concentrations (1–15 μM) of ZVFe NP. The spectrum of the buffer and ZVFe NP samples were subtracted from those of protein–NP mixture.
Circular dichroism spectroscopy
Circular dichroism (CD) spectra were recorded from 190 to 260 nm at 0.2 nm with a scan of rate of 2 points per second at 298 K, with three scans averaged for each CD spectrum. The data were expressed as ellipticity (mdeg), which was obtained in mdeg directly from the instrument. The far CD records of HSA (3 μM) was measured in the absence and presence of varying concentrations of ZVFe NP (3–30 μM), using a spectropolarimeter (model 215; Aviv, Lakewood, NJ, USA). Each record of HSA was subtracted from that of ZVFe NP sample as control.
Thermal melting measurement
The absorbance of HSA as a function of temperature was measured at 295 nm using a UV–visible (UV–vis) spectrophotometer equipped with a thermal bath. The absorbance of HSA (3 μM) in the absence and presence of ZVFe NP (3 M) was recorded as a function of temperature over the range of 30°C–90°C at a heating rate of 1°C min−1. The melting temperature (Tm) was calculated from the midpoint of the melting curves.
Molecular docking study
The ZVFe NP was clustered in our previous study.30 Briefly, geometry of ZVFe was optimized by the CASTEP module in Materials Studio 5.0 developed by Accelrys Software Inc. Local density approximation functional and ultrasoft pseudo potentials were used to model electron–ion interactions. HEX 6.3 software was used to run molecular docking study. Visualization of the ligand site was performed by using CHIMERA (www.cgl.ucsf.edu/chimera) and PyMOL (http://pymol.sourceforge.net/) tools.
Cell culture
SH-SY5Y cell line was purchased from Pasture Institute (Tehran, Iran) and was cultured in plastic tissue culture flasks in Eagle’s minimum essential medium and Ham’s F12 (1:1) supplemented with 10% FBS and 100 U/mL penicillin–streptomycin–neomycin mixture. The cells were maintained at 37°C in a humidified atmosphere containing 5% CO2. WBCs were separated and cultured based on the method reported in our previous paper.31 Blood samples were obtained after written informed consent was confirmed in accordance with the Declaration of Helsinki, according to protocol approved by the Ethical Committee of Pharmaceutical Sciences Branch, Islamic Azad University of Tehran, Tehran, Iran.
3-(4, 5-Dimethylthiazol-2-yl)-2, 5-diphenyltetrazolium bromide assay
MTT test has been carried out to evaluate cells viability. Following the 24 hours exposure to varying concentrations (1–100 μg/mL) of ZVFe and replacement of the medium, the cells (5×103 per well) were incubated in 37°C for 4 hours with MTT (Sigma-Aldrich, St Louis, MO, USA) solution (0.5 mg/mL). Then, supernatants were gently removed and formazan salt was extracted in DMSO. Measurement of the optical density was performed at 570 nm using a microplate reader (Expert 96, Asys Hitch, Ec Austria). DMSO and NP were used as blank references.
Flow cytometry analysis
Apoptosis induction by ZVFe NPs was assessed by flow cytometric analysis of negative control and ZVFe NPs (IC50 concentration) incubated cells according to the manufacturer’s protocol (BD Biosciences, San Jose, CA, USA). Briefly, SH-SY5Y cells (3×105 per well) were treated with IC50 concentration of ZVFe NPs for 24 hours, collected, washed, and resuspended in 500 μL of 1× Annexin-binding buffer, incubated at room temperature with Annexin V-FITC and PI stain, and analyzed by flow cytometry (BD Biosciences).
Antibacterial assay
The antibacterial potential of synthesized ZVFe NP was analyzed initially by well-diffusion method using following Gram-negative and Gram-positive strains of bacteria.
Totally, three bacterial strains, including P. aeruginosa (ATCC 27853), E. coli (ATCC 25922) as Gram-negative and S. aureus (ATCC 25923) as Gram-positive strains, were used for analyzing the antibacterial potential of ZVFe NP. Bacteria were subcultured from pure cultures of different strains of bacteria on Mueller–Hinton Broth for overnight at 37°C. The turbidity of bacterial culture was adjusted to 0.5 McFerland (1.5×108 CFU/mL) standard. Each bacterial strain was swabbed uniformly onto separate Muller–Hinton agar plates using sterile cotton swabs under sterile condition. Wells with 8 mm diameters were prepared by punching a sterile cork borer onto agar plates and removing the agar to form a well. One hundred microliters of 250 μg/mL (w/v) concentration of ZVFe NP were then applied to wells. After incubation for 24 hours, at 35°C–37°C, the zones of inhibition around the wells were measured in millimeter using a caliper. The minimum inhibitory concentration (MIC) and minimum bactericidal concentration (MBC) of ZVFe NP were determined against bacterial strains by the microdilution method.31
Statistical analysis
All experiments were carried out in triplicate and data were shown as mean ± SD of three independent experiments. The significance of difference between control- and sample-treated groups was calculated by one-way ANOVA followed by Dunnett’s t-test using SPSS version 20 (SPSS Inc., Chicago, IL, USA). P-value <0.05 was nominated as the level of significance.
Results and discussion
ZVFe NP characterization
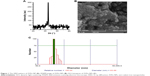
The oxidation state and phase of ZVFe NP were analyzed by employing XRD technique (Figure 1A). It was shown that the Fe NP was in its zero valent state and no appearance of oxide/hydroxide ligand was revealed on the surfaces of the ZVFe NPs. The peak appearing at 2θ value of ~45° is in good agreement with the value for Fe0 reported.32
Particle size was calculated with the XRD by using Scherer equation:32
|
where D is the particle size in Angstrom, λ is the wavelength of radiation, β is the full width at half maximum, and θ is the angle obtained from 2θ corresponding to maximum peak intensity. The mean crystalline dimension of the ZVFe NP was found to be 37.2 nm.
FESEM is widely employed as a potential technique to detect the size distribution and morphology of particles.33 The diameter and morphology of ZVFe NPs are shown in Figure 1B. Figure 1B demonstrated that the diameter of ZVFe NPs was about 30 nm and NPs exhibited homogeneous and nearly spherical shape. By using DLS, we monitored the hydrodynamic radius of ZVFe NPs at room temperature. DLS exhibits a hydrodynamic radius value of 83.88 nm with a PDI value of 0.21 (Figure 1C). Zeta potential value was also obtained from the DLS study. It was determined that the zeta potential value of ZVFe NPs was around −31.91±6.24 mV. These data also confirm the presence of monodispersed ZVFe NPs in the solution and high colloidal stability of the fabricated NPs.
Thermodynamic parameters
Fluorescence spectroscopy is a sensitive and simple tool in the protein conformational changes study. Quenching studies utilizing intrinsic Trp fluorescence of proteins can yield important data regarding the microenvironment of the aromatic fluorophore.34–36 The quenching mechanism can be classified as dynamic quenching (collisional system) or static quenching (complex formation) between fluorophores and quenchers.37 Dynamic and static quenching can be distinguished by their different relations on temperature and viscosity.38 The probability of fluorescence quenching with ligands depends on the rate of collision of the quencher and the ground-state fluorophore (static quenching) or excited fluorophore (dynamic quenching).39
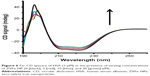
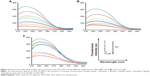
As shown in Figure 2, the fluorescence intensity of HSA decreases as the ZVFe NP concentration increases at 298 K (Figure 2A), 310 K (Figure 2B), and 315 K (Figure 2C). To determine the mode of quenching and NP-induced conformational changes, further analysis was done.
Mode of interaction
Static quenching is invariably related to the temperature, whereas collision quenching is directly related to the temperature. Therefore, fluorescence experiments were performed at three different temperatures (298, 310, and 315 K) to reveal the mode of fluorescence quenching by using the Stern–Volmer equation (Equation 1):40,41
|
|
where F0 and F exhibit the fluorescence intensities of protein in the absence and presence of the quencher (Q), respectively. In addition, the slope of this plot yields Stern–Volmer constant (KSV), which is commonly used to reveal the mode of interaction between ligand and protein. The KSV plot showed straight lines at every studied temperature, which indicates that the Trp residues are embedded at an equal accessible position to the ZVFe NPs (Figure 3). The KSV values for the interaction of HSA and ZVFe NP are summarized in Table 1. As shown in Table 1, when the temperature goes up, the KSV value decreases, exhibiting a probable static quenching mechanism (complex formation) between HSA and ZVFe NPs.
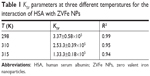  | Table 1 KSV parameters at three different temperatures for the interaction of HSA with ZVFe NPs |
Binding parameters
Several equations can be used for calculating the binding constant of the interaction between a protein and a ligand. One of the most routinely used equations is the Hill equation, since it can determine the correct stoichiometry for the binding sites of protein and ligands.42 To determine the binding parameters between ZVFe NP and HSA, Equation 2 was used:42
|
|
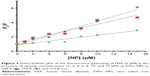
where Kb exhibits the binding constant between protein and NP and n shows the number of binding sites. The linear plot of log[F0 – F/F] vs log[Q] gives rise to the calculation of Kb and n from the y-intercept and slop of Hill equation, respectively (Figure 4). As shown in Table 2, when the temperature of the system increases, n and Kb values reduce, revealing the temperature-induced conformational changes of protein. We infer from these results that the binding site residues in HSA were accessible at 25°C, and buried at around 42°C. Therefore, this binding site moiety is expected to be involved in temperature-induced conformational transitions of HSA.
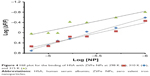  | Figure 4 Hill plot for the binding of HSA with ZVFe NPs at 298 K ( |
  | Table 2 Binding parameters at three different temperatures for the interaction of HSA with ZVFe NPs |
Thermodynamic parameters
In general, intermolecular interacting forces between a ligand and a protein include hydrogen bonding, van der Waals interaction, electrostatic forces, and hydrophobic network.43 The thermodynamic parameters such as enthalpy change (ΔH°), standard entropy change (ΔS°), and standard Gibbs free energy change (ΔG°) of interaction were determined using van’t Hoff equation (Equation 3):43
|
|
where Kb is the binding constant at the corresponding temperature and R is the gas constant. By plotting LnKb vs 1/T, the ΔH° and ΔS° can be determined (Figure 5; Table 3). The temperature-dependent thermodynamic parameters for the HSA–ZVFe NP system are used to characterize the intermolecular bonds between protein and NP. The value of ΔG° was then calculated by Gibbs–Helmholtz equation (Equation 4):44
|
|
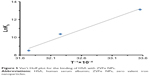  | Figure 5 Van’t Hoff plot for the binding of HSA with ZVFe NPs. |
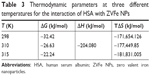  | Table 3 Thermodynamic parameters at three different temperatures for the interaction of HSA with ZVFe NPs |
The sign and magnitude of thermodynamic parameters are associated with the involving forces during protein–ligand interaction. ΔH°>0 and ΔS°>0 exhibit hydrophobic interactions, ΔH°<0 and ΔS°<0 show hydrogen bonds and van der Waals interactions, and ΔH°<0 and ΔS°>0 represent electrostatic interactions.40 Negative values of ΔH° (−53.21 kJ/mol) and ΔS° (−42.44 kJ/mol) showed that the NP-induced conformational changes of protein is induced by an enthalpy-driven manner.40 Likewise, ΔH°<0 and ΔS°<0 are characteristics of hydrogen bonds and van der Waals interactions in aqueous solution. Due to the large surface-to-volume-ratio of NPs, water molecules are substantially adsorbed on their surface and can mediate the establishment of hydrogen bonds between NP and protein.
Water is an extremely good acceptor and donator of hydrogen bonds. So, if a protein is partially unfolded in solution due to its interaction with NP, geometrically perfect hydrogen bonds can be formed with a very polarized molecule, water, on the surface of NP.
Secondary structural studies
Due to the influence of NP on the spectra of HSA, it is crucial to determine the structural changes of HSA in the presence of ZVFe NP. CD is a sensitive technique to monitor the secondary structural changes of protein.45 Far UV–CD (190–260 nm) spectra of HSA and HSA–ZVFe NP are shown in Figure 6. In HSA spectrum, there are two bands in the UV region, at 208 and 222 nm, which are characteristic of the α-helical conformation of HSA. Both transitions at 208 and 222 nm are known as a broad n→π* transition for the peptide bond of the α-helix.46 Figure 6 shows that the spectra obtained for HSA in the presence of various concentrations of ZVFe NPs were also similar to the spectrum of HSA in the absence of NPs. Therefore, it can be concluded that the relative α-helix structure of HSA remains unchanged in the presence of ZVFe NP.
UV–vis spectroscopy study
UV–vis absorption measurements are commonly used to investigate structural changes of proteins and complex formation.47 Thermal behaviors of HSA in the absence and presence of ZVFe NP can provide detailed information about the interaction strength of NPs with proteins. It is well known that, when the temperature in the solution goes up, the folded state of protein shifts toward the denatured structure and generates a hyperchromic influence on the absorption spectra of protein residues. In order to reveal this transition process, the Tm point, which is defined as the temperature, where half of the total native protein are denatured, is usually introduced.
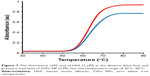
The melting curves of HSA at 295 nm in the absence and presence of ZVFe NPs are presented in Figure 7. Here, the thermal denaturation experiment revealed Tm values of around 64°C and 65°C for HSA in the presence and absence of the ZVFe NP, respectively. These almost identical values of Tm for HSA in the absence and presence of ZVFe NP are in good agreement with CD data. Therefore, it may be concluded that HSA structure remains almost unchanged after interaction with ZVFe NPs.
Molecular docking study
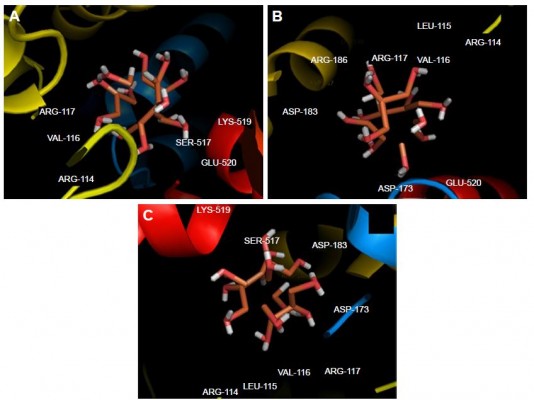
Molecular docking study was carried out to support the experimental data indicating that hydrogen bonds are the main forces in the interaction of HSA with ZVFe NPs. The X-ray crystallographic 3D structure of HSA at 2.5 A resolution (Protein Data Bank [PDB] ID: 1AO6) was downloaded from the online protein data bank Research Collaboratory for Structural Bioinformatics (RCSB) PDB (http://www.pdb.org).48 The molecular docking was performed with designed ZVFe cluster. The resulting E-value was found to be −280.64. The docked complex is shown in Figure 8. The ligand with surrounding active site residues within 3.5 Å is given in Figure 9A–C. The nearest interacting residues are SER-517, ASP-173, ASP-183, ARG-114, LEU-115, VAL-116, ARG-117, LYS-519, GLU-520, LYS-519, and ARG-186. These residues possess hydroxyl, carboxyl, and amine groups, which provide a potential medium to form hydrogen bonds with water molecules on ZVFe NP surface. Indeed, it may be indicated that the type of interaction between NPs and macromolecules depends on a number of factors, such as concentration, pH, temperature, and type of NPs/proteins. Actually, the type of interaction between NPs and proteins can be weakened or strengthened by changing these factors. It seems ZVFe NPs form hydrogen bonds and van der Waals interaction with HSA, which are known as relatively weak forces. Therefore, these interactions do not induce substantial conformational changes on the HSA structure.
  | Figure 8 Molecular docking of HSA with water-coated ZVFe NP. |
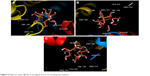  | Figure 9 Different views (A–C) of the ligand and its surrounding site residues. |
MTT assay
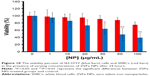
Different cells were employed to reveal the in vitro cytotoxic effect of ZVFe NPs. SH-SY5Y and WBCs were incubated with different concentrations (1, 10, 20, 30, 50, 100 μg/mL) of ZVFe NPs for 24 hours. Cell viability was then assessed by MTT assay. It was observed that ZVFe NPs induced mortality of cancer cells (Figure 10) and growth inhibitions were 95.37%±14.02%, 86.13%±19.27%, 73.31%±11.87% (P<0.05), 64.19%±15.16% (P<0.05), 48.21%±17.62% (P<0.01), and 39.11%±12.51% (P<0.01) in SH-SY5Y cells after treatment with 1, 10, 20, 30, 50, and 100 μg/mL of ZVFe NPs, respectively. However, it was revealed that the WBCs viability was 98%±19.05%, 96%±8.68%, 95%±12.16%, 91%±10.66%, 88%±8.09%, and 75%±3.97% (P<0.05), after treatment with 1, 10, 20, 30, 50, and 100 μg/mL of ZVFe NPs, respectively. As presented in Figure 10, some cytotoxic activity was already revealed at 20 μg/mL dose of ZVFe NPs on SH-SY5Y cells, whereas maximal effect was obtained at a concentration of 100 μg/mL in WBCs. Therefore, it was concluded that ZVFe NPs with concentrations of <100 μg/mL did not induce cytotoxicity on normal cells, so they could be used as potential candidates to act as nontoxic drug carriers or anticancer agents. The IC50 concentration of ZVFe NPs against SH-SY5Y cells and WBCs were calculated to be 47.89±81 μg/mL and >100 μg/mL, respectively. In fact, the nano state of ZVFe intensified the difference between cytotoxicity outcome in cancer and normal cells. Based on these observations, the present paper focused on SH-SY5Y cells for subsequent assays.
Quantification of apoptosis
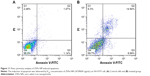
Quantification of apoptosis induction by ZVFe NPs on SH-SY5Y cells (IC50 concentration) was carried out by using flow cytometry. Figure 11 displays the ZVFe NPs-triggered apoptosis in SH-SY5Y cells after 24 hours incubation at IC50 concentration. The flow cytometry analysis results indicated that the rate of early apoptosis (Q3), late apoptosis (Q2), and necrosis (Q1) in control cells were 1.14%, 1.07%, and 2.48%, respectively (Figure 11A). However, treatment of the SH-SY5Y cells with IC50 concentration of ZVFe NPs for 24 hours enhanced the rate of early apoptosis (8.99%, P<0.001), late apoptosis (14.94%, P<0.001), and necrosis (6.3%, P<0. 01; Figure 11B).
Antibacterial assay
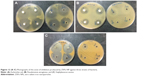
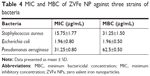
The antibacterial activity of ZVFe NP against one Gram-positive and two Gram-negative strains of bacteria are illustrated in Figure 12. Also, the inhibition zone diameter of ZVFe NP against E. coli, P. aeruginosa, and S. aureus is displayed in Figure 13. The results showed a noteworthy inhibitory effect of ZVFe NPs on both bacterial strains. Table 4 shows the MIC and MBC of ZVFe NP against three strains of bacteria. It was observed that ZVFe NP induced strong antibacterial effect against E. coli, P. aeruginosa, and S. aureus with an MIC of 1.96, 31.25, and 15.75 μg/mL, respectively. The MBC observed in the present study was 1.96 μg/mL for E. coli, 31.25 μg/mL for S. aureus, and 62.5 μg/mL for P. aeruginosa.
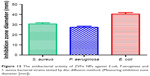  | Figure 13 The antibacterial activity of ZVFe NPs aganist E.coli, P.aeruginosa and S. aureus bacterial strains tested by disc diffusion method. (Measuring inhibition zone diameter [mm]). |
Discussion
In pharmaceutical industry, it is critical to determine and specify the pharmacokinetics and pharmacological influence of drugs like NPs.49,50 It is invariably assumed that the probable influence of drugs (NPs) heavily depends on their binding affinity to carrier proteins such as albumin.51 The interaction of NPs with proteins also affects the rate at which the NPs are delivered to targeted sites of reaction.52 Hence, analyzing the interaction of NPs with albumin is essential not only because the unbound NPs may affect a number of overcritical pharmacokinetic constants including the constant distribution volume, but also because it provides a crucial effect to express the administration regimen dose.53 Hence, it is important to examine that albumin can eventually influence the NP’s absorption, tissue distribution and pharmacodynamics, metabolism, and excretion features. In addition, the application of NPs, which can enter the body through the different routes and can interact with the albumin, could alter the conformation of albumin.
Uncontrolled side effects such as pH, temperature, and other denaturing conditions can influence the binding of an NP to albumin.54 Indeed, conformational changes of protein in the presence of NPs can weaken or strengthen the binding affinity of proteins to NPs. Furthermore, a comprehensive characterization of albumin in the presence of NPs and thermodynamic parameters is also necessary not only for understanding its pivotal physiological features, but also most recently to develop nanosystems as targeted delivery vehicles for several therapeutic and diagnostic NPs. Hence, the data about NP–protein interaction may provide potential and central details in the field of nanomedicine.
We showed that ZVFe NP binds to HSA through hydrogen bonds and does not play a significant role in the structural changes of HSA. The reason that hydrogen bonds do not induce a net destabilizing/stabilizing impact on protein conformation lies in the fact that in order to establish protein–NPs hydrogen bonds, another set of hydrogen bonds on the surface of protein should be disrupted.55 Interaction of protein with NP causes destabilization of protein conformation and exposure of the hydrophobic core. Contacting the dipoles that form the hydrogen bonds with these hydrophobic patches is energetically unfavorable. So, the overall effect of hydrogen bonds in NP-induced conformational alterations of proteins is that of inducing structural destabilization, not denaturation.
Biophysical study of zinc oxide interaction with albumin demonstrated no structural perturbation in the secondary structure of protein; however, some minor tertiary changes were reported.56 Gold NPs were indicated to cause conformational changes in the structure of albumin in a dose-dependent manner,57 whereas no major conformational change was reported for albumin when adsorbed onto the carbon C60 fullerene NP.58 The interaction of magnetic iron oxide NPs with albumin was also investigated by spectroscopic methods.59 The results indicated that the interaction was spontaneous and the electrostatic interactions played key roles in the interaction process. In addition, secondary structural alteration of albumin in the presence of MNPs was shown. It seems that the chemical composition of NPs may play an important role in inducing conformational changes of proteins upon interaction.
One of the crucial challenges in the development of anticancer drugs is increasing the therapeutic effectiveness of anticancer NPs and minimizing their side effects on normal cells. In our MTT assay, ZVFe NPs demonstrated a broad range of cytotoxicity with most mortality effect on SH-SY5Y cells but not on normal WBCs. Although, the mechanism of cancer-specific toxicity of ZVFe NPs is still unknown, this selective killing effect of ZVFe NPs in cancer cells is clinically important. Recently, selectivity in killing a wide spectrum of cancer cells by NPs including zinc oxide NPs,60 iron oxide NPs,61 and copper oxide NPs62 has been reported.
However, oxidative stress induced by ZVFe NPs in human bronchial epithelial cells has been reported.63 Therefore, these conflicting results may suggest that further studies should be done to explore the mechanisms behind the cytotoxic effects of NPs.
The effect of ZVFe NPs on bacterial activity was also investigated. The synthesized ZVFe NPs showed a noteworthy inhibitory effect on tested pathogenic bacteria. Based on these findings, bactericidal action of ZVFe NPs was affective. The different sensitivities of Gram-positive and Gram-negative bacteria against ZVFe NPs could be explained in accordance with the morphological differences between these microorganisms, due to different polarities of their cell wall. Other studies have demonstrated that the size of NPs can also affect their bactericidal activities. For example, Lee et al64 reported that the inactivation of E. coli by ZVFe NPs could be induced by infiltration of the NPs through E. coli membranes. A number of other investigations on zinc oxide65,66 and magnesium oxide NPs67 have also shown that antibacterial activity of NPs depend on their dimension.
In this study, the size of ZVFe NPs was about 30 nm, which enabled the NPs to cross the cell membrane and induce some adverse effects on the viability of bacteria. It may be suggested that ZVFe NPs could interact with intracellular oxygen, causing oxidative stress induction and finally driving cell membrane leakage.64 Herein, the concentration of ZVFe NPs was also a crucial factor to induce bactericidal activity against pathogenic bacteria. This behavior was reported by other studies when they explored the antimicrobial impacts of silver,68 zinc oxide,65 and iron oxide69 NPs on S. aureus and E. coli. Also, it should be noted that the fabrication route of NPs also influences the bactericidal activity. Indeed, different synthetic methods can form different NPs with various physicochemical and antibacterial properties.70
Conclusion
The results of this study indicate that ZVFe NPs potentially bind to HSA through hydrogen bonds and van der Waals interactions and induce marginal structural changes on the HSA structure. Cytotoxicity assay revealed the selective killing of cancer cells by ZVFe NPs. Antibacterial examination showed that ZVFe NPs provide a potential antibacterial effect against E. coli, P. aeruginosa, and S. aureus.
Therefore, it may be concluded that ZVFe NP may be used in medicine as a potential anticancer and/or antibacterial drug. However, it should be noted that some in vivo studies are required to more explore the anticancer and antibacterial activities of ZVFe NPs.
Acknowledgment
The research has been supported by Tehran University of Medical Sciences and Health Services grant no. 96033036163, Tehran, Iran.
Disclosure
The authors report no conflicts of interest in this work.
References
Burns A, Sengupta P, Zedayko T, Baird B, Wiesner U. Core/Shell fluorescent silica nanoparticles for chemical sensing: towards single-particle laboratories. Small. 2006;2(6):723–726. | ||
Kahn N, Lavie O, Paz M, Segev Y, Haick H. Dynamic nanoparticle-based flexible sensors: diagnosis of ovarian carcinoma from exhaled breath. Nano Lett. 2015;15(10):7023–7028. | ||
Chen PC, Mwakwari SC, Oyelere AK. Gold nanoparticles: from nanomedicine to nanosensing. Nanotechnol Sci Appl. 2008;1:45. | ||
Gehrke I, Geiser A, Somborn-Schulz A. Innovations in nanotechnology for water treatment. Nanotechnol Sci Appl. 2015;8:1. | ||
Blanco E, Shen H, Ferrari M. Principles of nanoparticle design for overcoming biological barriers to drug delivery. Nat Biotechnol. 2015;33(9):941–951. | ||
du X, Li X, Xiong L, Zhang X, Kleitz F, Qiao SZ. Mesoporous silica nanoparticles with organo-bridged silsesquioxane framework as innovative platforms for bioimaging and therapeutic agent delivery. Biomaterials. 2016;91:90–127. | ||
Teimouri M, Khosravi-Nejad F, Attar F, et al. Gold nanoparticles fabrication by plant extracts: synthesis, characterization, degradation of 4-nitrophenol from industrial wastewater, and insecticidal activity – a review. J Clean Prod. 2018;184:740–753. | ||
Tian J, Zhao Z, Kumar A, Boughton RI, Liu H. Recent progress in design, synthesis, and applications of one-dimensional TiO2 nanostructured surface heterostructures: a review. Chem Soc Rev. 2014;43(20):6920–6937. | ||
Ling D, Lee N, Hyeon T. Chemical synthesis and assembly of uniformly sized iron oxide nanoparticles for medical applications. Acc Chem Res. 2015;48(5):1276–1285. | ||
Zheng H, Zhang Y, Liu L, et al. One-pot synthesis of metal-organic frameworks with encapsulated target molecules and their applications for controlled drug delivery. J Am Chem Soc. 2016;138(3):962–968. | ||
Praveen PA, Ramesh Babu R, Balaji P, Murugadas A, Akbarsha MA. Laser assisted anticancer activity of benzimidazole based metal organic nanoparticles. J Photochem Photobiol B. 2018;180:218–224. | ||
Patra N, Kar D, Pal A, Behera A. Antibacterial, anticancer, anti-diabetic and catalytic activity of bio-conjugated metal nanoparticles. Adv Nat Sci Nanosci Nanotechnol. 2018;9(3):035001. | ||
Pérez N, Ruiz-Rubio L, Vilas JL, Rodríguez M, Martinez-Martinez V, León LM. Synthesis and characterization of near-infrared fluorescent and magnetic iron zero-valent nanoparticles. J Photochem Photobiol A. 2016;315:1–7. | ||
Lefevre E, Bossa N, Wiesner MR, Gunsch CK. A review of the environmental implications of in situ remediation by nanoscale zero valent iron (nZVI): behavior, transport and impacts on microbial communities. Sci Total Environ. 2016;565:889–901. | ||
Huang K-J, Wu S-R, Shieh D-B. Zero-valent iron nanoparticles inhibited head and neck cancer cells growth: a pilot evaluation and mechanistic characterization. Free Radic Biol Med. 2017;108:S39. | ||
Shieh D-B, Yang L-X, Lee W-T, et al. Zero-valent iron based nanoparticles selectively inhibit cancerous cells through mitochondria-mediated autophagy. Paper presented at: IEEE 17th International Conference on Nanotechnology (IEEE-NANO); July 25–28, 2017; Pittsburg, PA, USA. | ||
Nel AE, Mädler L, Velegol D, et al. Understanding biophysicochemical interactions at the nano-bio interface. Nat Mater. 2009;8(7):543–557. | ||
Baek M, Chung HE, Yu J, et al. Pharmacokinetics, tissue distribution, and excretion of zinc oxide nanoparticles. Int J Nanomedicine. 2012;7:3081. | ||
Park EJ, Lee GH, Yoon C, et al. Tissue distribution following 28 day repeated oral administration of aluminum-based nanoparticles with different properties and the in vitro toxicity. J Appl Toxicol. 2017;37(12):1408–1419. | ||
Maji A, Beg M, Mandal AK, et al. Spectroscopic interaction study of human serum albumin and human hemoglobin with Marsilea quadrifolia leaves extract mediated silver nanoparticles having antibacterial and anticancer activity. J Mol Struct. 2017;1141:584–592. | ||
Karthika V, Arumugam A, Gopinath K, et al. Guazuma ulmifolia bark-synthesized Ag, Au and Ag/Au alloy nanoparticles: photocatalytic potential, DNA/protein interactions, anticancer activity and toxicity against 14 species of microbial pathogens. J Photochem Photobiol B. 2017;167:189–199. | ||
Khan Z, Al-Thabaiti SA. Green synthesis of zero-valent Fe-nanoparticles: catalytic degradation of rhodamine B, interactions with bovine serum albumin and their enhanced antimicrobial activities. J Photochem Photobiol B. 2018;180:259–267. | ||
Kragh-Hansen U. Human serum albumin: a multifunctional protein. In: Otagiri M, Chuang VTG, editors. Albumin in Medicine. Amsterdam, Netherlands: Springer; 2016:1–24. | ||
Gao X, Bi H, Jia J, Tang L. Spectroscopic and in silico study of binding mechanism of cynidine-3-O-glucoside with human serum albumin and glycated human serum albumin. Luminescence. 2017;32(4):640–651. | ||
Laurini E, Marson D, Posocco P, Fermeglia M, Pricl S. Structure and binding thermodynamics of viologen-phosphorous dendrimers to human serum albumin: a combined computational/experimental investigation. Fluid Phase Equilib. 2016;422:18–31. | ||
Liu Z, Chen X. Simple bioconjugate chemistry serves great clinical advances: albumin as a versatile platform for diagnosis and precision therapy. Chem Soc Rev. 2016;45(5):1432–1456. | ||
Fallacara AL, Mancini A, Zamperini C, et al. Pyrazolo[3,4-d]pyrimidines-loaded human serum albumin (HSA) nanoparticles: preparation, characterization and cytotoxicity evaluation against neuroblastoma cell line. Bioorg Med Chem Lett. 2017;27(14):3196–3200. | ||
Valente MJ, Bastos ML, Fernandes E, Carvalho F, Guedes de Pinho P, Carvalho M. Neurotoxicity of β-keto amphetamines: deathly mechanisms elicited by methylone and MDPV in human dopaminergic SH-SY5Y cells. ACS Chem Neurosci. 2017;8(4):850–859. | ||
Yuvakkumar R, Elango V, Rajendran V, Kannan N. Preparation and characterization of zero valent iron nanoparticles. Dig J Nanomater Biostruct. 2011;6(4):1771–1776. | ||
Asl BA, Mogharizadeh L, Khomjani N, et al. Probing the interaction of zero valent iron nanoparticles with blood system by biophysical, docking, cellular, and molecular studies. Int J Biol Macromol. 2018;109:639–650. | ||
Abdlomajid E, Kharazi H, Chalaki M, et al. Titanium oxide nanoparticles fabrication, hemoglobin interaction, white blood cells cytotoxicity, and antibacterial studies. J Biomol Struct Dyn. Epub 2018 Jul 25. | ||
Allabaksh MB, Mandal BK, Kesarla MK, Kumar KS, Reddy PS. Preparation of stable zero valent iron nanoparticles using different chelating agents. J Chem Pharm Res. 2010;2(5):67–74. | ||
Foo Y-T, Chan JE-M, Ngoh G-C, Abdullah AZ, Horri BA, Salamatinia B. Synthesis and characterization of NiO and Ni nanoparticles using nanocrystalline cellulose (NCC) as a template. Ceram Int. 2017;43(18):16331–16339. | ||
Poureshghi F, Ghandforoushan P, Safarnejad A, Soltani S. Interaction of an antiepileptic drug, lamotrigine with human serum albumin (HSA): application of spectroscopic techniques and molecular modeling methods. J Photochem Photobiol B. 2017;166:187–192. | ||
Danesh N, Navaee Sedighi Z, Beigoli S, Sharifi-Rad A, Saberi MR, Chamani J. Determining the binding site and binding affinity of estradiol to human serum albumin and holo-transferrin: fluorescence spectroscopic, isothermal titration calorimetry and molecular modeling approaches. J Biomol Struct Dyn. 2018;36(7):1747–1763. | ||
Naeeminejad S, Assaran Darban R, Beigoli S, Saberi MR, Chamani J. Studying the interaction between three synthesized heterocyclic sulfonamide compounds with hemoglobin by spectroscopy and molecular modeling techniques. J Biomol Struct Dyn. 2017;35(15):3250–3267. | ||
Sinha S, Tikariha D, Lakra J, et al. Interaction of bovine serum albumin with cationic monomeric and dimeric surfactants: a comparative study. J Mol Liq. 2016;218:421–428. | ||
Chen H, Rao H, Yang J, Qiao Y, Wang F, Yao J. Interaction of diuron to human serum albumin: insights from spectroscopic and molecular docking studies. J Environ Sci Health B. 2016;51(3):154–159. | ||
Nunes NM, Pacheco AFC, Agudelo ÁJP, et al. Interaction of cinnamic acid and methyl cinnamate with bovine serum albumin: a thermodynamic approach. Food Chem. 2017;237:525–531. | ||
Zeinabad HA, Kachooei E, Saboury AA, et al. Thermodynamic and conformational changes of protein toward interaction with nanoparticles: a spectroscopic overview. RSC Adv. 2016;6(107):105903–105919. | ||
Zeinabad HA, Zarrabian A, Saboury AA, Alizadeh AM, Falahati M. Interaction of single and multi wall carbon nanotubes with the biological systems: tau protein and PC12 cells as targets. Sci Rep. 2016;6:26508. | ||
Abdullah SMS, Fatma S, Rabbani G, Ashraf JM. A spectroscopic and molecular docking approach on the binding of tinzaparin sodium with human serum albumin. J Mol Struct. 2017;1127:283–288. | ||
Pishkar L, Taheri S, Makarem S, et al. Studies on the interaction between nanodiamond and human hemoglobin by surface tension measurement and spectroscopy methods. J Biomol Struct Dyn. 2017;35(3):603–615. | ||
Esfandfar P, Falahati M, Saboury A. Spectroscopic studies of interaction between CuO nanoparticles and bovine serum albumin. J Biomol Struct Dyn. 2016;34(9):1962–1968. | ||
Hajsalimi G, Taheri S, Shahi F, Attar F, Ahmadi H, Falahati M. Interaction of iron nanoparticles with nervous system: an in vitro study. J Biomol Struct Dyn. 2018;36(4):928–937. | ||
Seelig J, Schönfeld HJ. Thermal protein unfolding by differential scanning calorimetry and circular dichroism spectroscopy two-state model versus sequential unfolding. Q Rev Biophys. 2016;49:e9. | ||
Abbasi-Tajarag K, Divsalar A, Saboury AA, Ghalandari B, Ghourchian H. Destructive effect of anticancer oxali-palladium on heme degradation through the generation of endogenous hydrogen peroxide. J Biomol Struct Dyn. 2016;34(11):2493–2504. | ||
Sugio S, Kashima A, Mochizuki S, Noda M, Kobayashi K. Crystal structure of human serum albumin at 2.5 Å resolution. Protein Eng. 1999;12(6):439–446. | ||
Rafiei P, Haddadi A. Pharmacokinetic consequences of PLGA nanoparticles in docetaxel drug delivery. Pharm Nanotechnol. 2017;5(1):3–23. | ||
Liu Y, Li J, Li Z, Tang X, Zhang Z. Pharmacokinetics of a ternary conjugate based pH-responsive 10-HCPT prodrug nano-micelle delivery system. Asian J Pharm Sci. 2017;12(6):542–549. | ||
Lacroix A, Edwardson TGW, Hancock MA, Dore MD, Sleiman HF. Development of DNA nanostructures for high-affinity binding to human serum albumin. J Am Chem Soc. 2017;139(21):7355–7362. | ||
Naveenraj S, Mangalaraja RV, Krasulyaa O, Syed A, Ameen F, Anandan S. A general microwave synthesis of metal (Ni, Cu, Zn) selenide nanoparticles and their competitive interaction with human serum albumin. New J Chem. 2018;42(8):5759–5766. | ||
Yamamoto S, Maeda N, Nagashima Y, et al. A phase II, multicenter, single-arm study of tri-weekly low-dose nanoparticle albumin-bound paclitaxel chemotherapy for patients with metastatic or recurrent breast cancer. Breast Cancer. 2017;24(6):783–789. | ||
Mohan V, Sengupta B, Acharyya A, Yadav R, Das N, Sen P. Region-specific double denaturation of human serum albumin: combined effects of temperature and GnHCl on structural and dynamical responses. ACS Omega. 2018;3(8):10406–10417. | ||
Newberry RW, Raines RT. A prevalent intraresidue hydrogen bond stabilizes proteins. Nat Chem Biol. 2016;12(12):1084–1088. | ||
Bardhan M, Mandal G, Ganguly T, State S. Steady state, time resolved, and circular dichroism spectroscopic studies to reveal the nature of interactions of zinc oxide nanoparticles with transport protein bovine serum albumin and to monitor the possible protein conformational changes. J Appl Phys. 2009;106(3):034701. | ||
Wangoo N, Suri CR, Shekhawat G. Interaction of gold nanoparticles with protein: a spectroscopic study to monitor protein conformational changes. Appl Phys Lett. 2008;92(13):133104. | ||
Liu S, Sui Y, Guo K, Yin Z, Gao X. Spectroscopic study on the interaction of pristine C60 and serum albumins in solution. Nanoscale Res Lett. 2012;7(1):433. | ||
Yang Q, Liang J, Han H. Probing the interaction of magnetic iron oxide nanoparticles with bovine serum albumin by spectroscopic techniques. J Phys Chem B. 2009;113(30):10454–10458. | ||
Premanathan M, Karthikeyan K, Jeyasubramanian K, Manivannan G. Selective toxicity of ZnO nanoparticles toward Gram-positive bacteria and cancer cells by apoptosis through lipid peroxidation. Nanomedicine. 2011;7(2):184–192. | ||
Ahamed M, Alhadlaq HA, Khan MAM, Akhtar MJ. Selective killing of cancer cells by iron oxide nanoparticles mediated through reactive oxygen species via p53 pathway. J Nanopart Res. 2013;15(1):1225. | ||
Shafagh M, Rahmani F, Delirezh N. CuO nanoparticles induce cytotoxicity and apoptosis in human K562 cancer cell line via mitochondrial pathway, through reactive oxygen species and P53. Iran J Basic Med Sci. 2015;18(10):993. | ||
Keenan CR, Goth-Goldstein R, Lucas D, Sedlak DL. Oxidative stress induced by zero-valent iron nanoparticles and Fe (II) in human bronchial epithelial cells. Environ Sci Technol. 2009;43(12):4555–4560. | ||
Lee C, Kim JY, Lee WI, Nelson KL, Yoon J, Sedlak DL. Bactericidal effect of zero-valent iron nanoparticles on Escherichia coli. Environ Sci Technol. 2008;42(13):4927–4933. | ||
Zhang L, Jiang Y, Ding Y, Povey M, York D. Investigation into the antibacterial behaviour of suspensions of ZnO nanoparticles (ZnO nanofluids). J Nanopart Res. 2007;9(3):479–489. | ||
Yamamoto O. Influence of particle size on the antibacterial activity of zinc oxide. Int J Inorg Mater. 2001;3(7):643–646. | ||
Makhluf S, Dror R, Nitzan Y, Abramovich Y, Jelinek R, Gedanken A. Microwave-assisted synthesis of nanocrystalline MgO and its use as a bacteriocide. Adv Funct Mater. 2005;15(10):1708–1715. | ||
Kim JS, Kuk E, Yu KN, et al. Antimicrobial effects of silver nanoparticles. Nanomedicine. 2007;3(1):95–101. | ||
Tran N, Mir A, Mallik D, Sinha A, Nayar S, Webster TJ. Bactericidal effect of iron oxide nanoparticles on Staphylococcus aureus. Int J Nanomedicine. 2010;5:277. | ||
Ismail RA, Sulaiman GM, Abdulrahman SA, Marzoog TR. Antibacterial activity of magnetic iron oxide nanoparticles synthesized by laser ablation in liquid. Mater Sci Eng C Mater Biol Appl. 2015;53:286–297. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.


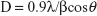

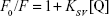

 ), 310 K (
), 310 K ( ), and 315 K (
), and 315 K ( ).
).