Back to Journals » Drug Design, Development and Therapy » Volume 9
Clinicopathological significance and potential drug target of T-cadherin in NSCLC
Authors Wang Z, Wang B, Guo H, Shi G, Hong X
Received 13 September 2014
Accepted for publication 13 October 2014
Published 19 December 2014 Volume 2015:9 Pages 207—216
DOI https://doi.org/10.2147/DDDT.S74259
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Professor Shu-Feng Zhou
Zhidong Wang,1 Bin Wang,1 Huanchen Guo,2 Guoyu Shi,2 Xiuqin Hong3
1Oncology Department, Eighth Hospital of Changsha, Changsha, People’s Republic of China; 2Department of Respiratory Medicine, Shouguang Hospital of Traditional Chinese Medicine, Shouguang, People’s Republic of China; 3Institute of Gerontology, Hunan Geriatric Hospital, Changsha, People’s Republic of China
Background: Previous studies demonstrate that T-cadherin is a candidate tumor suppressor in several types of human tumors, including non-small cell lung cancer (NSCLC). Lack of protein expression of T-cadherin by hypermethylation has been found to play an important role in lung alveolar differentiation regulation and epithelial tumorigenesis. However, the correlation between T-cadherin hypermethylation and clinicopathological characteristics of NSCLC remains unclear. Here we conducted a systematic review and meta-analysis to quantitatively evaluate the effects of T-cadherin hypermethylation on the incidence of NSCLC and clinicopathological characteristics.
Methods: A detailed literature search was carried out for related research publications. Analyses of pooled data were performed. Odds ratio (OR) and hazard ratio (HR) were calculated and summarized, respectively.
Results: Final analysis of 1,172 NSCLC patients from 15 eligible studies was performed. T-cadherin hypermethylation was observed to be significantly higher in NSCLC than in normal lung tissue, based on the pooled OR from nine studies including 532 NSCLC and 372 normal lung tissue samples (OR=8.19, 95% confidence interval [CI]=5.41–12.39, P<0.00001). T-cadherin hypermethylation may also be associated with pathological types. The pooled OR was obtained from four studies including 111patients with squamous cell carcinoma and 106 with adenocarcinoma (OR=0.35, 95% CI=0.19–0.66, P=0.001), which indicated that T-cadherin hypermethylation plays a more important role in the pathogenesis of adenocarcinoma. We did not find that T-cadherin hypermethylation was correlated with the sex or smoking status, clinical stages, or epidermal growth factor receptor (EGFR) mutation status. However, T-cadherin hypermethylation was found to be significantly higher in poorly differentiated NSCLC than in moderately and highly differentiated NSCLC, and NSCLC patients with T-cadherin hypermethylation had a lower survival rate than those without T-cadherin hypermethylation.
Conclusion: The results of this meta-analysis suggest that T-cadherin hypermethylation is associated with an increased risk and worse survival in NSCLC. T-cadherin hypermethylation, which induces the inactivation of T-cadherin gene, plays an important role in the carcinogenesis, cancer progression, as well as clinical outcome.
Keywords: methylation, lung cancer, meta-analysis, EGFR, odds ratio, hazard ratio
Introduction
Non-small cell lung cancer (NSCLC) accounts for approximately 85% of all lung cancer and is the leading cause of cancer-related deaths worldwide.1,2 Despite the advances in early detection, radical surgical cure, and multimodal therapeutic modalities, at diagnosis, there are about 80% of NSCLC cases in advanced stage, and the prognosis remains poor.3 Therefore, investigation of the mechanism of initiation, progression, as well as identification of prognostic markers is still needed for selection of patients with high chance of lung cancer recurrence and to provide better prognosis and individualized treatment. Epigenetic control of gene expression plays an important role in carcinogenesis. Aberrant methylation of CpG dinucleotides is a commonly observed epigenetic modification in human cancer.4–6 Recently, Heller et al identified a large number of tumor-specific methylated genes in NSCLC patients and found that many of them were regulated by methylation and involved in the pathogenesis of NSCLC.7 The DNA methylation signature of NSCLC affects the outcome of certain patients and can be practically determined by user-friendly polymerase chain reaction assays, and it has been shown that analysis of the best DNA methylation biomarkers improved prognostic accuracy beyond standard staging.8 Interestingly, patterns of DNA methylation can divide NSCLC into two phenotypically distinct subtypes of tumors and provide proof of principle that differences in DNA methylation can be used as a platform for predictive biomarker discovery and development.9 Detection of aberrant gene promoter methylation as a tool for the diagnosis of tumors or its use as prognostic marker has been widely described for many different cancers, including NSCLC.10
Cadherins function as membrane receptors mediating outside-in signals, activating small guanosine-5′-triphosphate (GTP)ases and the β-catenin/Wnt pathway, resulting in dynamic cytoskeleton reorganization and changes in the phenotype; as such, they are important determinants of tumor progression, serving as a suppressor of invasion and metastasis in many contexts.11–13 T-cadherin, also known as cadherin 13 (CDH13) or H-cadherin (heart), is a unique member of the cadherin superfamily since it lacks the transmembrane and cytoplasmic domains and is anchored to the cell membrane through the glycosylphosphatidylinositol (GPI) anchor.14–16 A number of studies demonstrated that T-cadherin is involved in low-density lipoproteins (LDL) hormone-like effects on Ca2+-mobilization and increased cell migration, phenotype changes, as well as insulin-dependent signaling, eNOS activation, angiogenesis, as well as in the regulation of brain network development, plasticity, and function.17–20 Abnormalities in the T-cadherin gene have been identified in several tumors, with upregulation of T-cadherin inducing cell cycle arrest, apoptosis, and inhibition of angiogenesis.21–27 The introduction of T-cadherin in human breast carcinoma cells markedly reduced their invasive potential and growth rate; it also induced the reversion of morphology from an invasive type to a normal cell-like type.28,29 T-cadherin methylation and/or gene deletion have been found to play an important role in lung alveolar differentiation regulation and epithelial tumorigenesis.30–34 Although previous studies indicated that inactivation of the T-cadherin gene is mainly induced by hypermethylation of the gene, the reported T-cadherin hypermethylation rates in NSCLC were remarkably diverse. In addition, its roles in NSCLC and clinicopathological significance have not been thoroughly investigated. There were no previous meta-analyses in the literature that covered this research question. Hence, we conducted a systematic review and meta-analysis to quantitatively evaluate the effects of T-cadherin hypermethylation on the incidence and clinical characteristics of NSCLC.
Methods
Search strategy and selection criteria
The following electronic databases were searched for relevant articles without any language restrictions: Web of Science™ (1945–2014), the Cochrane Library database, PubMed (1966–2014), Embase (1980–2014), Cumulative Index to Nursing and Allied Health Literature (CINAHL) (1982–2014), China National Knowledge Infrastructure (CNKI), Google Scholar, and the Chinese Biomedical Database (CBM) (1982–2014).
We searched articles using the search terms: “lung” and “cancer or tumor or neoplasm or carcinoma”, “methylation”, and “T-cadherin or CDH13 or cadherin 13”. We also manually searched the reference lists of the retrieved articles and reviews for additional articles.
Although our search did not have language limits initially, for the full-text reading and final evaluation, we only performed the review of the studies published in English and Chinese language. After exclusion of nonrelevant and/or redundant publications from the different databases, the remaining papers were evaluated in the full-text version for inclusion and exclusion criteria and for relevant articles in the reference lists. All searched data were retrieved. Authors’ bibliographies and references of selected studies were also searched for other relevant studies. The most complete study was chosen to avoid duplication if same patient populations were reported in several publications.
The criteria that an eligible study had to meet were as follows: 1) T-cadherin hypermethylation evaluated in the primary NSCLC tissues; 2) research revealed the relationship between T-cadherin hypermethylation and NSCLC clinicopathological parameters and prognosis; 3) T-cadherin hypermethylation examined by methylation-sensitive polymerase chain reaction; and 4) studies provided sufficient information to estimate hazard ratio (HR) for overall survival (OS) and 95% confidence interval (CI). The exclusion criteria included the following: 1) letters, reviews, case reports, conference abstracts, editorials, and expert opinion; and also, 2) all publications regarding in vitro/ex vivo studies, cell lines, and human xenograft.
Data extraction and methodological assessment
Two authors (ZW and BW) independently reviewed and extracted data from the eligible studies. Disagreements were resolved by discussion and consensus. Two authors (HG and GS) reviewed all of the articles that fit the inclusion and exclusion criteria. The following information was recorded for each study: first author name; year of publication; sample source; number of cases; clinicopathological parameter; cancer tumor/node/metastasis (TNM) stage; epidermal growth factor receptor (EGFR) mutation status; methylation detection method, methylation rate, and/or expression; and follow up. Data for study characteristics and clinical responses were summarized in table format. Heterogeneity of results was evaluated to determine whether the data of the various studies could be analyzed for a meta-analysis.
For the methodological evaluation of the studies, three investigators (ZW, BW, and XH) independently read through each publication and assessed and scored them according to the Reporting Recommendations for Tumor Marker Prognostic Studies (REMARK) guidelines and European Lung Cancer Working Party (ELCWP) quality scale.35,36 The three readers provided the quality scores and compared them, and then reached a consensus value for each item.
Statistical analysis
Analysis was conducted using the STATA 12.0 (StataCorp LP, College Station, TX, USA) and Review Manager (RevMan), Version 5.2 (The Nordic Cochrane Centre, Copenhagen, Denmark). The pooled frequency of T-cadherin hypermethylation and 95% CIs were estimated. The frequency of T-cadherin hypermethylation was compared according to different tumor characteristics. Heterogeneity among studies was evaluated with Cochran’s Q test37 and the I2 statistic.38,39 When heterogeneity was not an issue (I2 values <50%), a fixed effect model was used to calculate parameters. If there was substantial heterogeneity (I2 values ≥50%), a random effects model was used to pool data and to identify potential sources of heterogeneity, based on subgroup analyses. The pooled odds ratio (OR) was estimated for the association between T-cadherin hypermethylation and clinicopathological features. P-values tailed less than 0.05 were considered statistically significant.
Publication bias was assessed by using a method reported by Egger et al.40 We also explored reasons for statistical heterogeneity using meta-regression, subgroup analysis, and sensitivity analysis. The analysis of meta-regression and publication bias was performed using STATA 10.0 (StataCorp LP).
Results
Forty-five publications were identified by the search method as described above. Thirty-two of those were excluded as they were laboratory studies, nonoriginal articles (review), or studies irrelevant to the current analysis. Eventually, there were 15 studies included in final meta-analysis,30,33,41–53 as shown in Figure 1. Fifteen studies published from 2001 to 2012 were eligible for meta-analysis. A total of 1,172 NSCLC patients from the People’s Republic of China, Japan, South Korea, the Czech Republic, Italy, the Republic of Serbia, and the USA were enrolled. Their basic characteristics are summarized in Table 1.
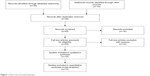  | Figure 1 Flow chart of study selection. |
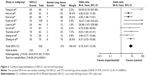
The loss of T-cadherin messenger (m)RNA and/or protein expression was strongly correlated with the promoter hypermethylation in several types of cancer, including NSCLC.32,33,54–62 T-cadherin hypermethylation was significantly higher in NSCLC than in normal lung tissue. The pooled OR from nine studies including 532 NSCLC and 372 normal lung tissue samples is shown in Figure 2 (OR=8.19, 95% CI=5.41–12.39, P<0.00001), and indicates that T-cadherin inactivation through hypermethylation plays an important role in the carcinogenesis of NSCLC.
It was previously reported that T-cadherin hypermethylation rate in female NSCLC patients was significantly higher than that in male patients.41,42 However, the pooled OR from seven studies including 388 male and 222 female NSCLC patients, as shown in Figure 3A (OR=0.87, 95% CI=0.46–1.66, P=0.68), indicates that T-cadherin hypermethylation is not strongly associated with sex status in NSCLC patients. Pesek et al reported that the T-cadherin hypermethylation rate in NSCLC patients without a smoking history was significantly higher than that in patients with a smoking history (80% vs 43%; 16/20 vs 42/98) (chi-square =0.96, P=0.002).42 The pooled OR from six studies including 447 NSCLC patients with and 237 patients without smoking history is shown in Figure 3B (OR=1.07, 95% CI=0.56–2.04, P=0.85) and indicates that T-cadherin hypermethylation is not strongly associated with the smoking status in NSCLC patients. Histology was associated with T-cadherin hypermethylation, which was observed more frequently in patients with adenocarcinoma (AD).41 The pooled OR from four studies including 111 squamous cell carcinoma patients and 106 AD patients is shown in Figure 3C (OR=0.35, 95% CI=0.19–0.66, P=0.001) and indicates that T-cadherin hypermethylation plays a more important role in the pathogenesis of AD.
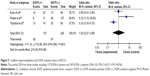
We analyzed 363 NSCLC patients pooled from four studies to assess whether the aberrant T-cadherin hypermethylation in NSCLC was associated with advanced stage. As shown in Figure 4A, aberrant T-cadherin hypermethylation was not significantly higher in advanced NSCLC (stage III and IV) than that in early stage NSCLC (stage I and II), OR=0.93, 95% CI=0.57–1.53, P=0.79. However, as shown in Figure 4B, aberrant T-cadherin hypermethylation was significantly higher in poorly differentiated NSCLC than in moderately and highly differentiated NSCLC (OR=3.61, 95% CI=1.76–7.39, P=0.0004). These results suggest that T-cadherin hypermethylation may play an important role in NSCLC progression and development. There were four studies that estimated the relationship between T-cadherin hypermethylation and OS in NSCLC. The pooled HR for OS showed that T-cadherin hypermethylation was associated with worse survival in NSCLC patients, as shown in Figure 4C (HR=3.21, 95% CI=1.41–7.31, P=0.005). In addition, there were three studies that determined the relationship between T-cadherin hypermethylation and EGFR mutation status in NSCLC. The pooled OR from three studies including 137 EGFR mutation-positive patients and 245 EGFR mutation-negative patients is shown in Figure 5 (OR=1.03, 95% CI=0.27–3.99, P=0.96) and indicates that T-cadherin hypermethylation has no significant correlation with EGFR mutation status in NSCLC.
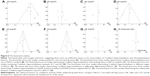
A sensitivity analysis, in which one study was removed at a time, was conducted to assess the result stability. The pooled ORs and HRs were not significantly changed, indicating the stability of our analyses. The funnel plots were largely symmetric (Figure 6), suggesting there were no publication biases in the meta-analysis of T-cadherin hypermethylation and clinicopathological features.
Discussion
Inactivation of T-cadherin by promoter hypermethylation plays an important role during normal development and in the tumorigenesis of several types of tumors, including NSCLC.41,47,56,63–69 To date, there have been some studies describing the precise expression, prognostic impact, and methylation status of T-cadherin in NSCLC; however, the role of T-cadherin inactivation by hypermethylation in NSCLC and its clinical significance have not been thoroughly investigated. Therefore, we conducted the meta-analysis to determine the correlation between T-cadherin hypermethylation and clinicopathological characteristics in NSCLC. Analysis of the pooled data showed that 1) NSCLC had higher hypermethylation than did normal lung tissue; 2) T-cadherin hypermethylation is not strongly associated with sex or smoking status in NSCLC patients; 3) T-cadherin hypermethylation plays a more important role in the pathogenesis of AD than in squamous cell carcinoma; 4) aberrant T-cadherin hypermethylation was not significantly higher in advanced NSCLC (stage III and IV) than that in early stage NSCLC (stage I and II); 5) aberrant T-cadherin hypermethylation was not significantly correlated with EGFR mutation status in NSCLC. However, aberrant T-cadherin hypermethylation was significantly higher in poorly differentiated NSCLC than in moderately and highly differentiated NSCLC; and finally, 6) NSCLC patients with T-cadherin hypermethylation had a lower OS that those without T-cadherin hypermethylation. The results from the current study demonstrated that the hypermethylation rate of the T-cadherin gene promoter in NSCLC was significantly higher than that in the normal lung tissues, indicating that T-cadherin promoter hypermethylation was common in NSCLC. Since changes in T-cadherin promoter hypermethylation are reversible, drug treatment targeting demethylation may be useful to delay carcinogenesis and progression, and to improve prognosis. Treatment of T-cadherin-negative tumor cells with the demethylating agent 5-aza-2′-deoxycytidine induced reexpression of this gene in several types of tumor cells, including colorectal cancer,54,55,60 pancreatic cancer,62 esophageal cancer,56 hepatocellular carcinoma,70,71 as well as B cell lymphoma.58 Therefore, this approach may bring new direction and hope for cancer treatment through gene-targeted therapy.
Epigenetic alterations, particularly aberrant DNA methylation, one of the best-characterized epigenetic modifications, contribute to tumor initiation and progression.5,6 T-cadherin has been reported to encode T-cadherin, an adiponectin receptor discovered after adipoR1 and adipoR2.72 T-cadherin is thought to affect cellular function and behavior, largely through its signaling properties.26 T-cadherin reexpression in most cancer cell lines inhibits cell proliferation and invasiveness, increases susceptibility to apoptosis, and reduces tumor growth in vivo models.55,73,74 Therefore, T-cadherin can be considered as tumor suppressor and its inactivation could contribute to tumor progression and poor prognosis. Although only four studies evaluated the relationship between OS and T-cadherin hypermethylation in NSCLC, they showed very similar results.30,44,47,49 Based on this meta-analysis, we may consider that T-cadherin hypermethylation in NSCLC tends to indicate a poor prognosis. Inconsistent with this finding, it was also found, using the kmplot tool (http://kmplot.com/analysis/index.php?p=service&cancer=lung), that low T-cadherin mRNA expression also indicates a poor prognosis in NSCLC patients.
Consistent results were shown in sensitivity analyses, and no evidence of publication bias was found. This study has several potential limitations. First, the possibility of information and selection biases and unidentified confounders could not be completely excluded because all of the included studies were observational. Second, the searching strategy was restricted to articles published in English or Chinese; articles with potentially high-quality data that were published in other languages were not included because of anticipated difficulties in obtaining accurate medical translation. Hence, cautions should be taken when our findings are interpreted among the general population.
In conclusion, our meta-analysis showed T-cadherin may play an important role in NSCLC initiation and progression. In addition, T-cadherin hypermethylation is associated with an increased risk and worse OS in NSCLC. Further large-scale studies, especially multicenter and well-matched cohort research, will provide more insight into the role of T-cadherin in the prognosis and clinical intervention for NSCLC patients.
Disclosure
The authors report no conflicts of interest in this work
References
Siegel R, Naishadham D, Jemal A. Cancer statistics, 2012. CA Cancer J Clin. 2012;62(1):10–29. | ||
Guo P, Huang ZL, Yu P, Li K. Trends in cancer mortality in China: an update. Ann Oncol. 2012;23(10):2755–2762. | ||
Ramalingam S, Belani C. Systemic chemotherapy for advanced non-small cell lung cancer: recent advances and future directions. Oncologist. 2008;13 Suppl 1:S5–S13. | ||
Ghavifekr Fakhr M, Farshdousti Hagh M, Shanehbandi D, Baradaran B. DNA methylation pattern as important epigenetic criterion in cancer. Genet Res Int. 2013;2013:317569. | ||
Delpu Y, Cordelier P, Cho WC, Torrisani J. DNA methylation and cancer diagnosis. Int J Mol Sci. 2013;14(7):15029–15058. | ||
Ma X, Wang YW, Zhang MQ, Gazdar AF. DNA methylation data analysis and its application to cancer research. Epigenomics. 2013;5(3):301–316. | ||
Heller G, Babinsky VN, Ziegler B, et al. Genome-wide CpG island methylation analyses in non-small cell lung cancer patients. Carcinogenesis. 2013;34(3):513–521. | ||
Sandoval J, Mendez-Gonzalez J, Nadal E, et al. A prognostic DNA methylation signature for stage I non-small-cell lung cancer. J Clin Oncol. 2013;31(32):4140–4147. | ||
Walter K, Holcomb T, Januario T, et al. DNA methylation profiling defines clinically relevant biological subsets of non-small cell lung cancer. Clin Cancer Res. 2012;18(8):2360–2373. | ||
Fleischhacker M, Dietrich D, Liebenberg V, Field JK, Schmidt B. The role of DNA methylation as biomarkers in the clinical management of lung cancer. Expert Rev Respir Med. 2013;7(4):363–383. | ||
Berx G, van Roy F. Involvement of members of the cadherin superfamily in cancer. Cold Spring Harb Perspect Biol. 2009;1(6):a003129. | ||
Jeanes A, Gottardi CJ, Yap AS. Cadherins and cancer: how does cadherin dysfunction promote tumor progression? Oncogene. 2008;27(55):6920–6929. | ||
van Roy F. Beyond E-cadherin: roles of other cadherin superfamily members in cancer. Nat Rev Cancer. 2014;14(2):121–134. | ||
Angst BD, Marcozzi C, Magee AI. The cadherin superfamily: diversity in form and function. J Cell Sci. 2001;114(Pt 4):629–641. | ||
Takeuchi T, Ohtsuki Y. Recent progress in T-cadherin (CDH13, H-cadherin) research. Histol Histopathol. 2001;16(4):1287–1293. | ||
Philippova M, Joshi MB, Kyriakakis E, Pfaff D, Erne P, Resink TJ. A guide and guard: the many faces of T-cadherin. Cell Signal. 2009;21(7):1035–1044. | ||
Philippova M, Joshi MB, Pfaff D, et al. T-cadherin attenuates insulin-dependent signalling, eNOS activation, and angiogenesis in vascular endothelial cells. Cardiovasc Res. 2012;93(3):498–507. | ||
Rubina K, Kalinina N, Potekhina A, et al. T-cadherin suppresses angiogenesis in vivo by inhibiting migration of endothelial cells. Angiogenesis. 2007;10(3):183–195. | ||
Rubina K, Talovskaya E, Cherenkov V, et al. LDL induces intracellular signalling and cell migration via atypical LDL-binding protein T-cadherin. Mol Cell Biochem. 2005;273(1–2):33–41. | ||
Rivero O, Sich S, Popp S, Schmitt A, Franke B, Lesch KP. Impact of the ADHD-susceptibility gene CDH13 on development and function of brain networks. Eur Neuropsychopharmacol. 2013;23(6):492–507. | ||
Bosserhoff AK, Ellmann L, Quast AS, Eberle J, Boyle GM, Kuphal S. Loss of T-cadherin (CDH-13) regulates AKT signaling and desensitizes cells to apoptosis in melanoma. Mol Carcinog. 2014;53(8):635–647. | ||
Ghosh S, Joshi MB, Ivanov D, et al. Use of multicellular tumor spheroids to dissect endothelial cell-tumor cell interactions: a role for T-cadherin in tumor angiogenesis. FEBS Lett. 2007;581(23):4523–4528. | ||
Wang XD, Wang BE, Soriano R, et al. Expression profiling of the mouse prostate after castration and hormone replacement: implication of H-cadherin in prostate tumorigenesis. Differentiation. 2007;75(3):219–234. | ||
Riou P, Saffroy R, Chenailler C, et al. Expression of T-cadherin in tumor cells influences invasive potential of human hepatocellular carcinoma. FASEB J. 2006;20(13):2291–2301. | ||
Duan XS, Lu J, Ge ZH, Xing EH, Lu HT, Sun LX. Effects of T-cadherin expression on B16F10 melanoma cells. Oncol Lett. 2013;5(4):1205–1210. | ||
Andreeva AV, Kutuzov MA. Cadherin 13 in cancer. Genes Chromosomes Cancer. 2010;49(9):775–790. | ||
Kim JS, Han J, Shim YM, Park J, Kim DH. Aberrant methylation of H-cadherin (CDH13) promoter is associated with tumor progression in primary nonsmall cell lung carcinoma. Cancer. 2005;104(9):1825–1833. | ||
Lee SW. H-cadherin, a novel cadherin with growth inhibitory functions and diminished expression in human breast cancer. Nat Med. 1996;2(7):776–782. | ||
Lee SW, Reimer CL, Campbell DB, Cheresh P, Duda RB, Kocher O. H-cadherin expression inhibits in vitro invasiveness and tumor formation in vivo. Carcinogenesis. 1998;19(6):1157–1159. | ||
Brock MV, Hooker CM, Ota-Machida E, et al. DNA methylation markers and early recurrence in stage I lung cancer. N Engl J Med. 2008;358(11):1118–1128. | ||
Zhong Y, Delgado Y, Gomez J, Lee SW, Perez-Soler R. Loss of H-cadherin protein expression in human non-small cell lung cancer is associated with tumorigenicity. Clin Cancer Res. 2001;7(6):1683–1687. | ||
Sato M, Mori Y, Sakurada A, Fujimura S, Horii A. The H-cadherin (CDH13) gene is inactivated in human lung cancer. Hum Genet. 1998;103(1):96–101. | ||
Toyooka KO, Toyooka S, Virmani AK, et al. Loss of expression and aberrant methylation of the CDH13 (H-cadherin) gene in breast and lung carcinomas. Cancer Res. 2001;61(11):4556–4560. | ||
Jin M, Kawakami K, Fukui Y, et al. Different histological types of non-small cell lung cancer have distinct folate and DNA methylation levels. Cancer Sci. 2009;100(12):2325–2330. | ||
McShane LM, Altman DG, Sauerbrei W, Taube SE, Gion M, Clark GM; Statistics Subcommittee of the NCI-EORTC Working Group on Cancer Diagnostics. Reporting recommendations for tumor marker prognostic studies (REMARK). J Natl Cancer Inst. 2005;97(16):1180–1184. | ||
Steels E, Paesmans M, Berghmans T, et al. Role of p53 as a prognostic factor for survival in lung cancer: a systematic review of the literature with a meta-analysis. Eur Respir J. 2001;18(4):705–719. | ||
DerSimonian R, Laird N. Meta-analysis in clinical trials. Control Clin Trials. 1986;7(3):177–188. | ||
Higgins JP, Thompson SG, Deeks JJ, Altman DG. Measuring inconsistency in meta-analyses. BMJ. 2003;327(7414):557–560. | ||
DerSimonian R. Meta-analysis in the design and monitoring of clinical trials. Stat Med. 1996;15(12):1237–1248; discussion 1249–1252. | ||
Egger M, Davey Smith G, Schneider M, Minder C. Bias in meta-analysis detected by a simple, graphical test. BMJ. 1997;315(7109):629–634. | ||
Kontic M, Stojsic J, Jovanovic D, et al. Aberrant promoter methylation of CDH13 and MGMT genes is associated with clinicopathologic characteristics of primary non-small-cell lung carcinoma. Clin Lung Cancer. 2012;13(4):297–303. | ||
Pesek M, Kopeckova M, Benesova L, et al. Clinical significance of hypermethylation status in NSCLC: evaluation of a 30-gene panel in patients with advanced disease. Anticancer Res. 2011;31(12):4647–4652. | ||
Kubo T, Yamamoto H, Ichimura K, et al. DNA methylation in small lung adenocarcinoma with bronchioloalveolar carcinoma components. Lung Cancer. 2009;65(3):328–332. | ||
Zhang Y, Wang R, Song H, et al. Methylation of multiple genes as a candidate biomarker in non-small cell lung cancer. Cancer Lett. 2011;303(1):21–28. | ||
Feng Q, Hawes SE, Stern JE, et al. DNA methylation in tumor and matched normal tissues from non-small cell lung cancer patients. Cancer Epidemiol Biomarkers Prev. 2008;17(3):645–654. | ||
Hsu HS, Chen TP, Hung CH, et al. Characterization of a multiple epigenetic marker panel for lung cancer detection and risk assessment in plasma. Cancer. 2007;110(9):2019–2026. | ||
Kim DS, Kim MJ, Lee JY, Kim YZ, Kim EJ, Park JY. Aberrant methylation of E-cadherin and H-cadherin genes in nonsmall cell lung cancer and its relation to clinicopathologic features. Cancer. 2007;110(12):2785–2792. | ||
Wang Y, Zhang D, Zheng W, Luo J, Bai Y, Lu Z. Multiple gene methylation of nonsmall cell lung cancers evaluated with 3-dimensional microarray. Cancer. 2008;112(6):1325–1336. | ||
Suzuki M, Shigematsu H, Iizasa T, et al. Exclusive mutation in epidermal growth factor receptor gene, HER-2, and KRAS, and synchronous methylation of nonsmall cell lung cancer. Cancer. 2006;106(10):2200–2207. | ||
Toyooka S, Tokumo M, Shigematsu H, et al. Mutational and epigenetic evidence for independent pathways for lung adenocarcinomas arising in smokers and never smokers. Cancer Res. 2006;66(3):1371–1375. | ||
Ulivi P, Zoli W, Calistri D, et al. p16INK4A and CDH13 hypermethylation in tumor and serum of non-small cell lung cancer patients. J Cell Physiol. 2006;206(3):611–615. | ||
Liao W, Xia L, Zeng W. Status of CDH1 promoter methylation in lung cancer patients. Journal of Chinese Modern Medicine. 2011;3:8–10. | ||
Sui X, Wang F, Huang G. The significance of CDH13 methylation in human lung cancer. Chinese Journal of Laboratory Diagnosis. 2012;5:836–983. | ||
Ren JZ, Huo JR. [5-aza-2′-deoxycytidine-induced inhibition of CDH13 expression and its inhibitory effect on methylation status in human colon cancer cells in vitro and on growth of xenograft in nude mice]. Zhonghua Zhong Liu Za Zhi. 2012;34(1):6–10. Chinese. | ||
Ren JZ, Huo JR. Correlation between T-cadherin gene expression and aberrant methylation of T-cadherin promoter in human colon carcinoma cells. Med Oncol. 2012;29(2):915–918. | ||
Jin Z, Cheng Y, Olaru A, et al. Promoter hypermethylation of CDH13 is a common, early event in human esophageal adenocarcinogenesis and correlates with clinical risk factors. Int J Cancer. 2008;123(10):2331–2336. | ||
Qian ZR, Sano T, Yoshimoto K, et al. Tumor-specific downregulation and methylation of the CDH13 (H-cadherin) and CDH1 (E-cadherin) genes correlate with aggressiveness of human pituitary adenomas. Mod Pathol. 2007;20(12):1269–1277. | ||
Ogama Y, Ouchida M, Yoshino T, et al. Prevalent hyper-methylation of the CDH13 gene promoter in malignant B cell lymphomas. Int J Oncol. 2004;25(3):685–691. | ||
Roman-Gomez J, Castillejo JA, Jimenez A, et al. Cadherin-13, a mediator of calcium-dependent cell-cell adhesion, is silenced by methylation in chronic myeloid leukemia and correlates with pretreatment risk profile and cytogenetic response to interferon alfa. J Clin Oncol. 2003;21(8):1472–1479. | ||
Toyooka S, Toyooka KO, Harada K, et al. Aberrant methylation of the CDH13 (H-cadherin) promoter region in colorectal cancers and adenomas. Cancer Res. 2002;62(12):3382–3386. | ||
Sun D, Zhang Z, Van do N, Huang G, Ernberg I, Hu L. Aberrant methylation of CDH13 gene in nasopharyngeal carcinoma could serve as a potential diagnostic biomarker. Oral Oncol. 2007;43(1):82–87. | ||
Sakai M, Hibi K, Koshikawa K, et al. Frequent promoter methylation and gene silencing of CDH13 in pancreatic cancer. Cancer Sci. 2004;95(7):588–591. | ||
Abudukadeer A, Bakry R, Goebel G, et al. Clinical relevance of CDH1 and CDH13 DNA-methylation in serum of cervical cancer patients. Int J Mol Sci. 2012;13(7):8353–8363. | ||
Lin YL, Sun G, Liu XQ, Li WP, Ma JG. Clinical significance of CDH13 promoter methylation in serum samples from patients with bladder transitional cell carcinoma. J Int Med Res. 2011;39(1):179–186. | ||
Missaoui N, Hmissa S, Trabelsi A, et al. Promoter hypermethylation of CDH13, DAPK1 and TWIST1 genes in precancerous and cancerous lesions of the uterine cervix. Pathol Res Pract. 2011;207(1):37–42. | ||
Suehiro Y, Okada T, Okada T, et al. Aneuploidy predicts outcome in patients with endometrial carcinoma and is related to lack of CDH13 hypermethylation. Clin Cancer Res. 2008;14(11):3354–3361. | ||
Hibi K, Kodera Y, Ito K, Akiyama S, Nakao A. Methylation pattern of CDH13 gene in digestive tract cancers. Br J Cancer. 2004;91(6):1139–1142. | ||
Wang L, Lin YL, Li B, Wang YZ, Li WP, Ma JG. Aberrant promoter methylation of the cadherin 13 gene in serum and its relationship with clinicopathological features of prostate cancer. J Int Med Res. 2014;42(5):1085–1092. | ||
Sacristan R, Gonzalez C, Fernández-Gómez JM, Fresno F, Escaf S, Sánchez-Carbayo M. Molecular classification of non-muscle-invasive bladder cancer (pTa low-grade, pT1 low-grade, and pT1 high-grade subgroups) using methylation of tumor-suppressor genes. J Mol Diagn. 2014;16(5):564–572. | ||
Chan DW, Lee JM, Chan PC, Ng IO. Genetic and epigenetic inactivation of T-cadherin in human hepatocellular carcinoma cells. Int J Cancer. 2008;123(5):1043–1052. | ||
Yan Q, Zhang ZF, Chen XP, et al. Reduced T-cadherin expression and promoter methylation are associated with the development and progression of hepatocellular carcinoma. Int J Oncol. 2008;32(5):1057–1063. | ||
Hug C, Wang J, Ahmad NS, Bogan JS, Tsao TS, Lodish HF. T-cadherin is a receptor for hexameric and high-molecular-weight forms of Acrp30/adiponectin. Proc Natl Acad Sci U S A. 2004;101(28):10308–10313. | ||
Ellmann L, Joshi MB, Resink TJ, Bosserhoff AK, Kuphal S. BRN2 is a transcriptional repressor of CDH13 (T-cadherin) in melanoma cells. Lab Invest. 2012;92(12):1788–1800. | ||
Kuphal S, Martyn AC, Pedley J, et al. H-cadherin expression reduces invasion of malignant melanoma. Pigment Cell Melanoma Res. 2009;22(3):296–306. |
 © 2014 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2014 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.






