Back to Journals » Drug Design, Development and Therapy » Volume 9
Adaptive evolution and elucidating the potential inhibitor against schizophrenia to target DAOA (G72) isoforms
Authors Sehgal SA, Mannan S, Kanwal S , Naveed I, Mir A
Received 12 March 2014
Accepted for publication 29 April 2014
Published 3 July 2015 Volume 2015:9 Pages 3471—3480
DOI https://doi.org/10.2147/DDDT.S63946
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Sheikh Arslan Sehgal,1,2 Shazia Mannan,2,* Sumaira Kanwal,2,* Ishrat Naveed,1 Asif Mir1
1Department of Bioinformatics and Biotechnology, International Islamic University, Islamabad, Pakistan; 2Department of Biosciences, COMSATS Institute of Information Technology, Sahiwal, Pakistan
*These authors contributed equally to this work
Abstract: Schizophrenia (SZ), a chronic mental and heritable disorder characterized by neurophysiological impairment and neuropsychological abnormalities, is strongly associated with d-amino acid oxidase activator (DAOA, G72). Research studies emphasized that overexpression of DAOA may be responsible for improper functioning of neurotransmitters, resulting in neurological disorders like SZ. In the present study, a hybrid approach of comparative modeling and molecular docking followed by inhibitor identification and structure modeling was employed. Screening was performed by two-dimensional similarity search against selected inhibitor, keeping in view the physiochemical properties of the inhibitor. Here, we report an inhibitor compound which showed maximum binding affinity against four selected isoforms of DAOA. Docking studies revealed that Glu-53, Thr-54, Lys-58, Val-85, Ser-86, Tyr-87, Leu-88, Glu-90, Leu-95, Val-98, Ser-100, Glu-112, Tyr-116, Lys-120, Asp-121, and Arg-122 are critical residues for receptor–ligand interaction. The C-terminal of selected isoforms is conserved, and binding was observed on the conserved region of isoforms. We propose that selected inhibitor might be more potent on the basis of binding energy values. Further analysis of this inhibitor through site-directed mutagenesis could be helpful for exploring the details of ligand-binding pockets. Overall, the findings of this study may be helpful in designing novel therapeutic targets to cure SZ.
Keywords: schizophrenia, bioinformatics, modeling, docking, DAOA, G72, DAO, computer-aided drug designing, phylogenetic analysis, d-amino acid oxidase activator
Introduction
Schizophrenia (SZ) affects approximately 1% of the world’s population, having similar prevalence throughout diverse ethnic groups.1 SZ is a chronic, mental, highly heritable and common disease characterized by neurophysiology impairment and neuropsychological abnormalities.1–3 The susceptibility of SZ is influenced by environment factors, polygenic components, and their interactions.4 Molecular mechanisms activating SZ remains intangible. SZ gene identification is particularly exigent and challenging due to the fact that the disorder may include limited SZ diagnostic accuracy as phenotypic definition and numerous entities have not been yet defined. Additionally, the lack of conclusive linkage from scans of the genome could also be due to the existence of several SZ vulnerability genes that are difficult to replicate and detect.5
The variations in D-amino acid oxidase activator (DAOA) (13q34) gene initially were linked with SZ.6 Furthermore, this gene has been linked with other phenotypes and psychiatric disorders such as severe bipolar disorder,7 bipolar disorder,8 and major depression.9 DAOA genetic variations have contributed to different central nervous system (CNS) disorders linked with dysfunction of glutamatergic signaling.6,10,11 The longest open reading frame of DAOA (G72) is predicted to encode a putative 153 amino acid protein isolated from testis, spinal cord, caudate nucleus, and amygdale libraries.6 DAOA (G72), a primate-specific protein, has been characterized as DAO activity modulator.6 DAOA expression in transgenic mice induced schizophrenic-related behavioral phenotypes.12,13
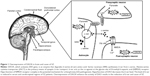
SZ patients show DAOA overexpression in the dorsolateral prefrontal cortex when compared with healthy controls.14 SZ susceptibility genes have been identified in genetic studies,15–18 but genetic interactions among SZ genes and their interplay with clinical subtypes and neurobiological abnormalities remains obscure. The product of DAO is an enzyme that degrades D-serine amino acid which acts as a co-agonist at the glycine site of the N-methyl-D-aspartic acid (NMDA) receptors.19 The product of DAOA (G72) activates the DAO enzyme.6 The biological function of DAOA and DAO are involved in the hypothesized hypofunction of NMDA receptor complex as the prospective pathogenesis of SZ (Figure 1).20
The neurotransmission of NMDA is the dominant molecular mechanism for memory, cognition, and synaptic plasticity. Numerous psychiatric and neurological disorders are linked with dysfunction of NMDA receptor–mediated neurotransmission.21 Hyperactivity and overexpression of brain DAO have been associated with SZ.22,23
DAO hyperfunction resulting from the decreased level of D-serine and NMDA receptor hypofunction has been suggested to be an underlying dysregulation in this disorder.24,25 In the cerebrospinal fluid26 and serum27 of schizophrenic patients, the level of D-serine is reduced. In addition to its role in cognition and psychosis, DAO may be involved in amyotrophic lateral sclerosis, resulting in D-serine accumulation, motoneuron death, and excitotoxicity.28 DAO activity manipulation could be an approach to regulate the level of D-serine29 (Figure 1). DAOA inhibitors may give valuable therapeutics approaches and strategies to improve schizophrenic symptoms.
DAOA has been identified in many different parts of the CNS, including spinal cord, nucleus caudatus, and amygdala.26 It has also been implicated in the regulation of dendritic branching and mitochondrial functions.30 DAOA was identified to activate DAO, while recent studies showed that DAOA modulates human DAO function as a negative effector.26,31 DAO catalyzes the oxidative deamination of D-amino acids such as D-serine to a keto-acids and D-3,4-dihydroxyphenylalanine (D-DOPA).32,33 L-DOPA enters in the basic biosynthetic pathway to homovanillic acid (HVA) and dopamine. Dopamine-β-hydroxylase converts dopamine into noradrenaline, which is degraded into 3-methoxy-4-hydroxyphenylglycol (MHPG) after entering its metabolic pathway. Kinetic data showed that the maximal velocity for the oxidative deamination of D-serine is much lower than for D-DOPA.19 D-serine is an allosteric modulator of NMDA-type glutamate receptor (NMDAR),19 having a D-serine modulatory site. D-serine occupation at this site is required for glutamate to stimulate the flow of cations.34,35 Study on the interactions between noradrenaline36 and glutamate propose that DAOA may be linked with noradrenaline via glutamatergic mechanism.
A bidirectional interaction between the dopamine system and NMDAR was also evidenced. The activation of NMDAR leads to enhance the recruitment of the dopamine D1 receptor (DRD1) to the plasma membrane.37,38 There is also a direct protein–protein coupling between NMDAR an DRD1.36,39,40 It has also been suggested that NMDAR and DRD1, early after their biosynthesis, form heteromeric complexes, which are transported as performed units to the plasma membrane.41 NMDA antagonists lead to the increase of dopamine neuron firing rates in mid brain.42 A direct association between dopamine release and D-serine has been shown, as high doses of D-serine attenuated amphetamine-induced dopamine release.31
The present work focused on the ligand analysis to reveal novel inhibitor against SZ. Experimental validation of DAOA isoforms by using X-ray crystallography and NMR (nuclear magnetic resonance) are not available yet. So as to solve structural insights, the isoforms of DAOA three-dimensional structures were predicted by applying template crystal structures from protein data bank (PDB). The inclusive in silico analysis may provide an evidence for a reliable framework that could assist in the designing and development of lead molecules that could be potential drug targets against SZ.
The significant role of DAOA on DAO regulation has been suggested;6 the specific role DAOA plays in regulating the activity of DAO in schizophrenic patients is still controversial. Sacchi et al31 demonstrated that a 1:2 molar ratio of DAOA to DAO is optimal for the inhibition of DAO. On the other hand, Chumakov et al6 revealed that DAOA enhances the activity of DAO, resulting in a reduction of D-serine concentration. DAOA is elevated in the serum of nonmediated and mediated patients of SZ,6 despite the fact that DAOA and DAOA mRNA are hardly detectable in the brain.31,43 As a result, dysfunction of NMDA and its related SZ pathological symptoms may result from DAOA overexpression that changes the activity of DAO. Sehgal et al44 explored the protein interaction between DAOA and DAO by the exercise of molecular docking. Four isoforms of DAOA (isoform 1, isoform 2, isoform 3, isoform 4) were selected from the Uniprot protein database. The longest isoform of DAOA contains 153 residues. Conserved amino acid residues are found in the selected four DAOA isoforms suggesting that the conserved region is important for DAOA folding and function.
Materials and methods
A DAOA transcribing two transcripts led to the production of four isoforms collectively, namely, DAOA-153, DAOA-126, DAOA-82, and DAOA-125. While DAOA-125 has the accession number A2T115, the rest of the three isoforms were represented by the single accession number P59103 in Uniprot.
In the current study, structure prediction, sequence analysis, phylogenetic analysis, docking studies, and comparative proteomics analysis were performed on HP Core-I-5 workstation. The amino acid sequences of three known isoforms of DAOA and one another variant of DAOA were retrieved for comparative modeling. The amino acid sequences of DAOA isoforms were retrieved from Uniprot and subjected to a protein–protein BLAST for the identification of suitable template structure against PDB.45 The automated protein modeling program MODELLER 9v1046 was employed to predict the three-dimensional structures of DAOA by satisfying spatial restraints. Five evaluation tools were used: Rampage,47 ProCheck,48 Anolea,49 and ERRAT50 were applied to assess the predicted models, and the structures were further evaluated by MolProbity.51 Finally, the poor Ramachandran outliers and rotamers were removed by utilizing WinCoot52 tool.
Sehgal et al44 utilized a ligand DAOA:O, O-diethylthiophosphoryl (Z)-2-(2-aminothiazol-4-yl)-2-trityloxyiminoacetate (C28H28N3O5PS2) to analyze the binding sites of DAOA (accession number A2T115). The ligand (C28H28N3O5PS2) was utilized for protein–ligand docking analysis using the AutoDock tools.53 The number of H-bond donors, rotatable bonds, and H-bond acceptors were obtained by employing PubChem,54 mCule, and Molinspiration. Lipinski’s rule of five55 was analyzed using the Cresset and mCule servers. mCule and Cresset were employed to estimate the mutagenesis of novel molecules, and no mutagenic risks were detected. The aim of the docking analysis was to identify the relative binding specificities and binding pattern of protein against the specific ligand (C28H28N3O5PS2). The geometrical optimization of three-dimensional structures of proteins and ligand were performed using ChemDraw Ultra and Vega ZZ. Results were visualized and analyzed by utilizing Ligplot and Chimera 1.6. ADMET refers to the absorption, distribution, metabolism, excretion, and toxicity properties of compound and was analyzed by utilizing admetSAR online server.
The evolutionary history of DAOA was constructed by protein sequence analysis. P-distance (difference between two amino acid sequences on the basis of their amino acid positions), complete-deletion, and uncorrected options were utilized for analysis. National Center for Biotechnology Information (NCBI), ENSEMBL, UCSC genome browsers, and biological databases were utilized to analyze all the mentioned species in these databases. To analyze the DAOA ancestral relationship, Molecular Evolutionary Genetic Algorithm (MEGA 5) tool was employed. A considered bootstrap value of 1,000 replications was applied and using neighbor-joining (NJ) approach to construct the phylogenetic tree. ENSEMBL, NCBI, and UCSC genome browser were utilized to search for evidence of DAOA paralogs and orthologs. The protein sequences were run through ENSEMBL BLAT and NCBI BLAST against all the species included in ENSEMBL and primates. The sequences were also run through NCBI BLAST against over 16 primate protein sequences.
Results and discussion
The objective of our study was based on the relation of DAOA isoforms with SZ and its bioinformatics analysis for analyzing the ligand. The optimally aligned templates against all the DAOA-selected proteins with maximum identity, and query coverage are mentioned in Table 1. The suitable, scrutinized templates were utilized for three-dimensional structure predictions of DAOA isoforms. Consequently, selected templates gave better evaluation results. The predicted structures were comparatively modeled using crystal structures selected from PDB.
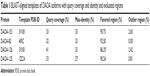  | Table 1 BLAST-aligned templates of DAOA isoforms with query coverage and identity and evaluated regions |
Evaluation tools showed the reliability and efficacy of DAOA three-dimensional predicted structures (Figure 2). Favored and outlier regions of predicted models were detected in Ramachandran plot and mentioned in Table 1. DAOA-153 and DAOA-126 showed four and three residues in outlier regions, respectively. No amino acid residue was observed in outlier region, and all the amino acids of the DAOA-82 were observed in favored and allowed region. In DAOA-125, only one residue was observed in outlier region. Poor rotamers and outliers were corrected to refine the predicted models of DAOA.
  | Figure 2 Three-dimensional structure of the four DAOA isoforms. |
However, the docking analysis of ligand revealed variations in their binding energies. Initially, the docking analysis was performed with ten poses, and 150 runs were saved, out of which the best poses with lowest binding energy were chosen from DAOA isoforms. Our findings indicated that the ligand (DAOA:O) efficiently binds to DAOA isoforms (Table 2).44
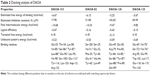  | Table 2 Docking analysis of DAOA |
Ligand docking analysis with DAOA isoforms were analyzed on the basis of their binding energy values and drug properties (Table 3). The ligand is a cyclic molecule (Figure 3) with significant biological properties. The ligand compound may be considered as potential antischizophrenic agents. Among all the selected isoforms, minimum binding energy was observed in DAOA-153.
  | Table 3 Properties of ligand |
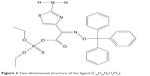  | Figure 3 Two-dimensional structure of the ligand (C28H28N3O5PS2). |
AutoDock tool was utilized for docking analysis, and top complexes of DAOA isoforms having lowest binding energies were selected for further analysis. Slight variation was observed in all the isoform complexes analyzed with lowest binding energies. Perhaps, the stability of the ligand may be due to the allowing of more conserved binding affinities.
It was revealed that the DAOA isoforms–docked complexes with analyzed ligand molecule showed reliable results. It was observed that the selected ligand binds at the conserved region of DAOA isoforms and revealed the binding pocket. It is also possible that the combination of observed binding residues leads to the lower binding energy achieved in the current study.
It was observed that Ser-100 of DAOA-153, Ser-29 of DAOA-82, and Ser-72 of DAOA-125 (Figure 4) having good binding interactions with the same docked ligand. The residues having different positions due to variation in the size of isoforms are indicated with matching superscript letters in Table 2. The conserved region has same binding residues but have different positions due to different size of isoforms (Figure 5). In an effort to comprehend the better interactions observed between ligand and amino acid residues in the active site of protein, a plot of amino acids–ligand interactions were generated employing Ligplot and Chimera 1.6 as shown in Figure 6.
  | Figure 5 Multiple sequence alignment (MSA) of all four isoforms of DAOA. |
Drug designing procedure is not only time consuming but also costly. Hence, we applied bioinformatics methodologies and approaches for drug discovery procedure. This emerging inclination has immense importance in decreasing the time required and amplifying the selected compound with better biological activity and minimal side effects for specific disease targets.
The chemical structures are evaluated to be an efficacious drug-like compound subjected to Lipinski’s rule of five55 and for their oral bioavailability. ADMET refers to the absorption, distribution, metabolism, excretion, and toxicity properties of compound and was analyzed by utilizing admetSAR online server. Eleven mathematical models (Ames toxicity, blood–brain barrier penetration, fish toxicity, aqueous solubility [LogS], honey bee toxicity, carcinogens, acute oral toxicity, Caco2 permeability, cytochrome P450 2D6 inhibition, rat acute toxicity, and human intestinal absorption) were employed to predict the ADMET properties of selected compound. Various toxicities are predicted often and are used in drug designing (Table 3). These toxicities help in evaluating intermediates, metabolites, and pollutants, along with adjusting the dose range for animal assays.
The aqueous solubility (at 25°C) prediction showed that the selected compound is soluble in water. Lipophilicity measurement (LogP) solubility is the ratio of compound in octanol compared to its solubility in water. It was observed that the compound showed less LogP value and follows Lipinki’s rule of five, thus showing better oral bioavailability. The excretion process that eliminates the compound from the body depends on LogP.56 The compound must be absorbed by human intestine. The intestinal human absorption results showed that the compound could easily be absorbed by the intestine. The selected molecule was found to be a noninhibitor of cytochrome P450 2D6, which indicated that the analyzed molecule may be well metabolized in Phase I metabolism. The cytochrome P450 2D6 is one of the significant enzymes involved in drug metabolism.57 Carcinogenicity and toxicity risk assessment were analyzed for the selected molecule, which showed it to be noncarcinogenic.
The results of docking analysis suggested that an ideal ligand must be the one that comes under the designed parameters for satisfying its drug likeness and has a less docking score. Coinciding with the statement and satisfying the least docking energy, it is suggested that C28H28N3O5PS2 is a potential drug molecule for SZ treatment that works by targeting DAOA isoforms. Our docking results revealed that binding residues mentioned in Table 2 are crucial, and mutational studies of these binding residues could be highly effective in further studies. It stands to reason that the selected ligand molecule in this study has the propensity to be a good candidate for the treatment of SZ by targeting DAOA. The proposed and used strategy led to simplification of the process of novel drug designing as biological investigation and drug analysis can be carried out on candidates with good results and without the need to waste money and time on poor activity compounds.
Protein data sets for a huge range of vertebrate and invertebrate genomes are presently available for analyzing the phylogenetic history of candidate genes of SZ. The phylogenetic NJ tree presented in current analysis reveals many interesting characteristics of the candidate gene in vertebrates (Figure 7). There are approximately 80 genomes of vertebrates that are completely sequenced, and sequences are embedded in the ENSEMBL genome browser. The BLAT alignment tool of ENSEMBL was employed for sequence alignment of DAOA canonical protein sequence against sequences of all the species present in ENSEMBL. It was observed that no orthologs of DAOA were found in any other species through NCBI and ENSEMBL. DAOA may be absent in all other sequenced species except hominoids and great apes. In silico analysis revealed that the gene was only found in humans, chimpanzees, gorillas, orangutans, and crab-eating macaques. The phylogenetic tree was reconciled with primate phylogeny.58 It is hypothesized that DAOA is a primate-specific gene according to the analysis of current available biological sequences data in biological databases (NCBI, ENSEMBL, UCSC). It is also hypothesized that may be this gene evolved approximately 35 million years ago before the divergence of the new-world monkey from the old-world monkey clade. The old-world monkeys and new-world monkeys shared derived features, and the primate’s brain is more developed than those of other species.
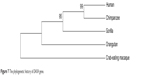  | Figure 7 The phylogenetic history of DAOA gene. |
Although the significance of DAO regulation in the function of neurophysiology has been revealed, the authenticated exact mechanism remains vague. The molecular characterization of DAOA showed that it was an endogenous modulator of the activity of DAO. This may provide a better and clearer understanding of DAO regulating activity. This study reveals the binding interactions of DAOA isoforms with ligand. By employing computational and in silico approaches, it is demonstrated that DAOA isoforms have conserved regions (Figure 8) and conserved C-terminal region. The analyzed ligand showed binding residues in conserved regions. This work also identified common binding residues that interact with selected ligand and normalize the expression of DAOA.
  | Figure 8 DAOA isoforms (DAOA-153, DAOA-126, DAOA-82) having conserved region shown in orchid color and also presented in multiple sequence alignment analysis. |
The bioinformatics analysis indicated that binding residues within C-terminal of DAOA are essential for ligand, instead of N-terminal. The manipulation in the expression and activity of DAOA can be a fruitful option for the treatment of SZ. The results of the docking analysis, by selected ligand, could be used for designing novel chemical compounds and synthetic peptides to reduce the overexpression of DAOA that leads to SZ. The level of D-serine plays a crucial role in neurotransmission of glutamate NMDA. The level of D-serine must be maintained to avoid over- or underexpression of NMDA receptor function within a relatively narrow range for avoiding CNS function impairment. In schizophrenic patients, the D-serine level in serum is 17.7% lower than normal controls, and the ratio of D-serine in serum to total serine is 28.4% lower.
Conclusion
In conclusion, the selected ligand molecule is efficacious in the treatment of SZ for targeting DAOA. The insilico analysis of DAOA has higher probability and efficacy on the basis of binding energy and other used parameters. It is also hypothesized that DAOA is a primate-specific gene, and only exists in great apes. The future analysis of this gene will be helpful in understanding the behavior of Homo sapiens. Further studies and synthesis of novel compounds considering these findings can expect similar response rates and can cure the effects of SZ.
Acknowledgments
Authors are grateful to Dr Amir Ali Abbasi, National Centre for Bioinformatics, Quaid-i-Azam University, Islamabad, Pakistan, for fruitful guidance in sequence and phylogenetic analysis. This research work is supported by the Higher Education Commission (HEC) Grant-2039.
Disclosure
The authors report no conflicts of interest in this work.
References
Bromet EJ, Fennig S. Epidemiology and natural history of schizophrenia. Biol Psychiatry. 1999;46(7):871–881. | ||
Breier A, Su T-P, Saunders R, et al. Schizophrenia is associated with elevated amphetamine-induced synaptic dopamine concentrations: evidence from a novel positron emission tomography method. Proc Natl Acad Sci U S A. 1997;94(6):2569–2574. | ||
Abi-Dargham A, Gil R, Krystal J, et al. Increased striatal dopamine transmission in schizophrenia: confirmation in a second cohort. Am J Psychiatry. 1998;155(6):761–767. | ||
Olney JW, Farber NB. Glutamate receptor dysfunction and schizophrenia. Arch Gen Psychiatry. 1995;52(12):998–1007. | ||
Pulver AE. Search for schizophrenia susceptibility genes. Biol Psychiatry. 2000;47(3):221–230. | ||
Chumakov I, Blumenfeld M, Guerassimenko O, et al. Genetic and physiological data implicating the new human gene G72 and the gene for D-amino acid oxidase in schizophrenia. Proc Natl Acad Sci U S A. 2000;99:13675–13680. | ||
Dalvie S, Horn N, Nossek C, van der Merwe L, Stein DJ, Ramesar R. Psychosis and relapse in bipolar disorder are related to GRM3, DAOA, and GRIN2B genotype. Afr J Psychiatry. 2010;13:297–301. | ||
Prata D, Breen G, Osborne S, Munro J, St Clair D, Collier D. Association of DAO and G72(DAOA)/G30 genes with bipolar affective disorder. Am J Med Genet B Neuropsychiatr Genet. 2008;147B:914–917. | ||
Rietschel M, Beckmann L, Strohmaier J, et al. G72 and its association with major depression and neuroticism in large population-based groups from Germany. Am J Psychiatry. 2008;165:753–762. | ||
Di Maria E, Bonvicini C, Bonomini C, Alberici A, Zanetti O, Gennarelli M. Genetic variation in the G720/G30 gene locus (DAOA) influences the occurrence of psychotic symptoms in patients with Alzheimer’s disease. J Alzheimers Dis. 2009;18:953–960. | ||
Drews E, Otte DM, Zimmer A. Involvement of the primate specific gene G72 in schizophrenia: from genetic studies to pathomechanisms. Neurosci Biobehav Rev. 2013;37(10):2410–2417. | ||
Cheng L, Hattori E, Nakajima A, et al. Expression of the G72/G30 gene in transgenic mice induces behavioral changes. Mol Psychiatry. 2014;19:175–183. | ||
Otte DM, Bilkei-Gorzo A, Filiou MD, et al. Behavioral changes in G72/G30 transgenic mice. Eur Neuropsychopharmacol. 2009;19:339–348. | ||
Korostishevsky M, Kaganovich M, Cholostoy A, et al. Is the G72/G30 locus associated with schizophrenia? Single nucleotide polymorphisms, haplotypes, and gene expression analysis. Biol Psychiatry. 2004;56:169–176. | ||
International Schizophrenia Consortium, Purcell SM, Wray NR, Stone JL, et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature. 2009;460:748–752. | ||
Stefansson H, Ophoff RA, Steinberg S, et al. Common variants conferring risk of schizophrenia. Nature. 2009;460:744–747. | ||
O’Donovan MC, Williams NM, Owen MJ. Recent advances in the genetics of schizophrenia. Hum Mol Genet. 2003;12(2):R125–R133. | ||
Owen MJ, Williams NM, O’Donovan MC. The molecular genetics of schizophrenia: findings promise new insights. Mol Psychiatry. 2004;9:14–27. | ||
Mothet JP, Parent AT, Wolosker H, et al. D-serine is an endogenous ligand for the glycine site of the N-methyl-D-aspartate receptor. Proc Natl Acad Sci U S A. 2000;97:4926–4931. | ||
Hashimoto K, Fukushima T, Shimizu E, et al. Decreased serum levels of D-serine in patients with schizophrenia: evidence in support of the N-methyl-D-aspartate receptor hypofunction hypothesis of schizophrenia. Arch Gen Psychiatry. 2003;60:572–576. | ||
Kemp JA, Leeson PD. The glycine site of the NMDA receptor – five years on. Trends Pharmacol Sci. 1993;14:20–25. | ||
Burnet P, Eastwood S, Bristow G, et al. D-Amino acid oxidase (DAO) activity and expression are increased in schizophrenia. Mol Psychiatry. 2008;13:658–660. | ||
Madeira C, Freitas ME, Vargas-Lopes C, Wolosker H, Panizzutti R. Increased brain D-amino acid oxidase (DAAO) activity in schizophrenia. Schizophr Res. 2008;101:76–83. | ||
Epstein FH, Lipton SA, Rosenberg PA. Excitatory amino acids as a final common pathway for neurologic disorders. N Engl J Med. 1994;330:613–622. | ||
Kalia LV, Kalia SK, Salter MW. NMDA receptors in clinical neurology: excitatory times ahead. Lancet Neurol. 2008;7:742–755. | ||
Bendikov I, Nadri C, Amar S, et al. A CSF and postmortem brain study of D-serine metabolic parameters in schizophrenia. Schizophr Res. 2007;90:41–51. | ||
Hashimoto K, Fukushima T, Shimizu E, et al. Decreased serum levels of D-serine in patients with schizophrenia: evidence in support of the N-methyl-D-aspartate receptor hypofunction hypothesis of schizophrenia. Arch Gen Psychiatry. 2003;60:572–576. | ||
Mitchell J, Paul P, Chen HJ, et al. Familial amyotrophic lateral sclerosis is associated with a mutation in D-amino acid oxidase. Proc Natl Acad Sci U S A. 2010;107:7556–7561. | ||
Hashimoto K, Fujita Y, Horio M, et al. Co-administration of a D-amino acid oxidase inhibitor potentiates the efficacy of D-serine in attenuating prepulse inhibition deficits after administration of dizocilpine. Biol Psychiatry. 2009;65:1103–1106. | ||
Kvajo M, Dhilla A, Swor DE, Karayiorgou M, Gogos JA. Evidence implicating the candidate schizophrenia/bipolar disorder susceptibility gene G72 in mitochondrial function. Mol Psychiatry. 2008;13:685–696. | ||
Sacchi S, Bernasconi M, Martineau M, et al. pLG72 modulates intracellular D-serine levels through its interaction with D-amino acid oxidase: effect on schizophrenia susceptibility. J Biol Chem. 2008;283:22244–22256. | ||
Kawazoe T, Park HK, Iwana S, Tsuge H, Fukui K. Human D-amino acid oxidase: an update and review. Chem Rec. 2007;7:305–315. | ||
Wu M, Zhou XJ, Konno R, Wang YX. D-dopa is unidirectionally converted to L-dopa by D-amino acid oxidase, followed by dopa transaminase. Clin Exp Pharmacol Physiol. 2006;33:1042–1046. | ||
Johnson JW, Ascher P. Glycine potentiates the NMDA response in cultured mouse brain neurons. Nature. 1987;325:529–531. | ||
Feldman S, Weidenfeld J. Involvement of endogeneous glutamate in the stimulatory effect of norepinephrine and serotonin on the hypothalamo-pituitary-adrenocortical axis. Neuroendocrinology. 2004;79:43–53. | ||
Pei L, Lee FJ, Moszczynska A, Vukusic B, Liu F. Regulation of dopamine D1 receptor function by physical interaction with the NMDA receptors. J Neurosci. 2004;24:1149–1158. | ||
Scott L, Kruse MS, Forssberg H, Brismar H, Greengard P, Aperia A. Selective up-regulation of dopamine D1 receptors in dendritic spines by NMDA receptor activation. Proc Natl Acad Sci U S A. 2002;99:1661–1664. | ||
Fiorentini C, Gardoni F, Spano P, Di Luca M, Missale C. Regulation of dopamine D1 receptor trafficking and desensitization by oligomerization with glutamate N-methyl-D-aspartate receptors. J Biol Chem. 2003;278:20196–20202. | ||
Lee FJ, Xue S, Pei L, et al. Dual regulation of NMDA receptor functions by direct protein–protein interactions with the dopamine D1 receptor. Cell. 2002;111:219–230. | ||
Missale C, Fiorentini C, Busi C, Collo G, Spano PF. The NMDA/D1 receptor complex as a new target in drug development. Curr Top Med Chem. 2006;6:801–808. | ||
French ED. Phencyclidine and the midbrain dopamine system: electrophysiology and behavior. Neurotoxicol Teratol. 1994;16:355–362. | ||
Smith SM, Uslaner JM, Yao L, et al. The behavioral and neurochemical effects of a novel D-amino acid oxidase inhibitor compound 8 [4H-thieno [3,2-b]pyrrole-5-carboxylic acid] and D-serine. J Pharmacol Exp Ther. 2009;328:921–930. | ||
Benzel I, Kew JN, Viknaraja R, et al. Investigation of G72 (DAOA) expression in the human brain. BMC Psychiatry. 2008;8:94. | ||
Sehgal SA, Khattak NA, Mir A. Structural, phylogenetic and docking studies of D-amino acid oxidase activator (DAOA), a candidate schizophrenia gene. Theor Biol Med Model. 2013;10(1):3. | ||
Berman HM, Westbrook J, Feng Z, et al. The protein data bank. Nucleic Acids Res. 2000;28:235–242. | ||
Eswar N, Eramian D, Webb B, Shen MY, Sali A. Protein structure modeling with MODELLER. Methods Mol Biol. 2008;426:145–159. | ||
Lovell SC, Davis IW, Arendall WB III, et al. Structure validation by Cαgeometry: φ/ψ and Cβ deviation. Proteins. 2003;50(3):437–450. | ||
Laskowski RA, MacArthur MW, Moss DS, Thornton JM. PROCHECK-a program to check the stereochemical quality of protein structures. J Appl Crystallogr. 1993;26:283–291. | ||
Melo F, Devos D, Depiereux E, Feytmans E. ANOLEA: a www server to assess protein structures. Proc Int Intell Syst Mol Biol. 1997;97:110–113. | ||
Colovos C, Yeates TO. Verification of protein structures: patterns of nonbonded atomic interactions. Protein Sci. 1993;2:1511–1519. | ||
Chen VB, Arendal WB III, Headd J, et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr D Biol Crystallogr. 2010;66:12–21. | ||
Emsley P, Lohkamp B, Scott W, Cowtan K. Features and development of coot. Acta Crystallogr D Biol Crystallogr. 2010;66;486–501. | ||
Trott O, Olson AJ. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization and multithreading. J Comput Chem. 2010;31:455–461. | ||
Bolton E, Wang Y, Thiessen PA, Bryant SH. PubChem: integrated platform of small molecules and biological activities. In: Annual Reports in Computational Chemistry, Vol 4, Washington, DC: American Chemical Society; 2008. | ||
Leeson, P. Drug discovery: Chemical beauty contest. Nature. 2012; 481(7382):455–456. | ||
Alam S, Khan F. QSAR and docking studies on xanthone derivatives for anticancer activity targeting DNA topoisomerase IIα. Drug Des Devel Ther. 2014;8:183. | ||
Lynch T, Price A. The effect of cytochrome P450 metabolism on drug response, interactions, and adverse effects. Am Fam Physician. 2007;76:391–396. | ||
Perelman P, Johnson WE, Roos C, et al. A molecular phylogeny of living primates. PLoS Genet. 2011;7(3):e1001342. |
 © 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.