Back to Journals » Hepatic Medicine: Evidence and Research » Volume 15
Analysis of SARS-CoV-2 Variant-Specific Serum Antibody Post-Vaccination Utilizing Immortalized Human Hepatocyte-Like Cells (HLC) to Assess Development of Immunity
Authors Collins DP , Steer CJ
Received 30 August 2023
Accepted for publication 21 November 2023
Published 5 December 2023 Volume 2023:15 Pages 221—231
DOI https://doi.org/10.2147/HMER.S431327
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Gerry Lake-Bakaar
Daniel P Collins,1 Clifford J Steer2,3
1CytoMedical Design Group, LLC, Saint Paul, MN, USA; 2Department of Medicine, University of Minnesota Medical School, Minneapolis, MN, USA; 3Department of Genetics, Cell Biology and Development, University of Minnesota Medical School, Minneapolis, MN, USA
Correspondence: Daniel P Collins, CytoMedical Design Group, LLC, 4280 Centerville Road, Saint Paul, MN, 55127, USA, Email [email protected]
Background: Our previous studies demonstrated that SARS-CoV-2 spike protein could bind to primary hepatocytes and immortalized Hepatocyte-like cells (HLC) via the asialoglycoprotein receptor-1 (ASGR-1). The binding of biotinylated spike protein could be inhibited by Spike-neutralizing monoclonal antibodies, anti-ASGR-1 antibodies and unlabeled spike protein. The cells were unable to bind Spike S1 and Spike S1 was incapable of blocking labeled Spike protein, suggesting that the Receptor Binding Domain (RBD) was not involved in the binding event. This study was done to investigate the utility of these cells and immortalized alveolar type 2-like (AT-2) cells in studying the development of variant-specific antibodies post-vaccination.
Methods: Serum was collected from 10 individuals pre- and post-vaccination with the J&J, Moderna or Pfizer vaccines. The serum samples were quantified for variant-specific antibodies in a flow cytometry-based immunofluorescent assay utilizing beads coated with biotinylated variant spike proteins. Inhibition of spike protein binding to HLC and AT-2 cells by donor serum was analyzed by immunofluorescent confocal analysis.
Results: All variant spike proteins bound to HLC and AT-2 cells. Post-vaccination serum samples demonstrated increases of SARS-CoV-2 antibody levels from 2 weeks to 2.5 months post-vaccination with associated increased spike-blocking capacity. It was also demonstrated that vaccination with all the available vaccines stimulated antibodies that inhibited binding of all the available variant spike proteins to both HLC and AT-2 cells.
Conclusion: HLC, along with AT-2 cells, provides a useful platform to study the development of neutralizing antibodies post-vaccination. Vaccination with the 3 available vaccines all elicited neutralizing serum antibodies that inhibited binding of each of the variant spike proteins to both AT-2 and HLC cells. This study suggests that inhibition of spike binding to target cells may be a more useful technique to assess immunity than gross quantitation of antibody.
Keywords: SARS-CoV-2, E12-HLC, E12-AT-2, spike protein variants, ASGR-1, ACE-2, TMPRSS-2
Introduction
SARS-CoV-2 is the virus responsible for the worldwide acute respiratory pandemic that has resulted in over 1 million COVID-19 deaths in the United States alone and is still contributing to morbidity and mortality worldwide as new variant strains evolve and infect susceptible individuals. Development of annual updates to vaccine boosters incorporating new variants is likely (similar to new formulations of annual flu vaccines developed to immunize against the current and predominant circulating strain), to require testing of protective immunity provided by the updated vaccines. While the main cause of morbidity and mortality associated with SAR-CoV-2 infection is respiratory in nature it has been shown that infection with SARS-CoV-2 virus also affects other organ systems.1–4 Further, it has been shown to directly infect other non-respiratory organs and tissues,5,6 potentially contributing to non-respiratory related morbidity. In our previous publication,7 we demonstrated that SARS-CoV-2 spike proteins could specifically bind to hepatocytes and immortalized hepatocyte-like cells (HLC) via the asialoglycoprotein receptor-1 (ASGR-1) providing a potential portal for attachment, internalization, and infection of hepatocytes. Hepatocytes/HLC, typically, do not express detectible ACE-2, precluding that from being a portal for viral entry. We also demonstrated that the S1 portion of the spike protein (containing the RBD) was unable to bind to hepatocytes/HLC and was unable to block the binding of complete Spike protein. From that data, we concluded that Spike S1 was not the mechanism for spike binding, but rather S2 was the portion of the spike protein responsible for the observed binding.
The expression of Transmembrane Serine Protease-2 (TMPRSS-2) on hepatocytes could provide a potential co-factor for internalization of the virus, as it cleaves the S1 and S2 portions of the spike protein and is essential to viral internalization via the ACE-2 portal pathway.8 Inhibition of spike protein binding to hepatocytes/HLC by preincubation of spike proteins with commercially available spike protein-specific monoclonal antibodies suggested that hepatocytes/HLC could provide a useful platform for assessing the effectiveness of antibodies to interfere with Spike S2 binding to cell surface receptors, including ASGR-1. To our knowledge, the development of anti-Spike S2 directed antibodies post-vaccination has not been studied or reported.
To document that immunization with the currently available vaccines elicited an antibody response against the Spike S2-mediated binding to HLC, we collected serum from 10 individuals pre- and post-vaccination and tested them for the development of serum antibodies reactive to variant spike proteins and their capacity to block the binding of spike proteins to HLC. As a positive control, E12-AT-2 cells were used as an additional target cell to follow the concurrent development of serum antibody expression against the already well-documented Spike S1 receptor-binding domain (RBD) to ACE-2. Variant-specific antibodies were detected and quantitated by a flow cytometry-based immunofluorescent assay utilizing biotinylated-spike variant proteins bound to avidin-coated paramagnetic beads. The capacity of serum antibodies to inhibit binding of spike proteins to target cells was evaluated by fluorescent confocal microscopy utilizing biotinylated variant spike proteins, pre-incubation of spike proteins with post-vaccination serum, E12-HLC (HLC) and E12-AT-2 (AT-2) target cells. This study suggests that HLC cells can provide a valuable and reproducible tool to evaluate the development of S2-blocking antibodies elicited by immunization with current and future vaccines.
Methods and Materials
Hepatocyte-Like Cells (HLC)
Human hepatocyte-like cells (HLC) were developed from the chemical fusion of immortalized E12-MLPC and primary hepatocytes to create a cell with the phenotypic and biological characteristics of fully mature hepatocytes that were immortalized but not transformed.9 These cells, like primary hepatocytes, were also demonstrated to be a target cell for the binding of SARS-CoV-2 spike proteins via binding to ASGR-1 and that this binding could be prevented by commercially available monoclonal antibodies, including antibodies to ASGR-1.7 Cells were maintained and expanded using a specific expansion medium composed of Williams Medium E supplemented with 2% fatty acid-free BSA, 1% ITS solution, 5mM hydrocortisone 21-hemisuccinate, FGF basic (20 ng/mL), FGF-4 (20 ng/mL), HFG (40 ng/mL), SCF (40 ng/mL), Oncostatin M (20 ng/mL), BMP-4 (20 ng/mL), EGF (40 ng/mL) and IL-1β (20 ng/mL). Williams Medium E supplemented with 2% fatty acid-free BSA, 1% ITS solution, 5mM hydrocortisone 21-hemisuccinate, FGF basic (20 ng/mL), FGF-4 (20 ng/mL), HFG (40 ng/mL), SCF (40 ng/mL), Oncostatin M (20 ng/mL), BMP-4 (20 ng/mL), EGF (40 ng/mL) and IL-1β (20 ng/mL).9
Alveolar Type-2-Like Cells (AT-2)
Development of AT-2-like cells from MLPC and their utility in the analysis of SARS-CoV-2 spike binding to respiratory cells has been previously reported.10 AT-2 cells developed by this technique produced cells that were phenotypically and biologically identical to AT-2 cells (marketed as Small Airway Epithelial Cells). Immortalized E12-MLPC were differentiated to AT-2-like cells by the same method previously developed for non-immortalized MLPC.11 Briefly, E12-MPLC were cultured in complete SAGM (Lonza, Walkerville, MD, cat# 3118) for 8–10 days to facilitate differentiation of AT-2-like cells, which were then expanded in the same medium for the studies.
Spike Proteins
Biotinylated variant spike proteins were used in the development of both the flow cytometry-based quantitative assay and the confocal spike binding/blocking assay. Biotinylated spike variants that were available for the study were the original isolate (RBD SPD-C822E9, ACROBiosystems) and its spike 1 region (S1N-C82E8, ACROBiosystems), the alpha (α) variant (B.1.1.7)(SPNC82E5, ACROBiosystems, Newark, DE), the beta variant (β) (B.1.351)(SPN-C82E6, ACROBiosystems), a combination of the alpha, beta and gamma (γ) variants (B.1.1.7/B.1.351/P1) (SPN-C82E3, ACROBiosystems) and the delta (δ) variant (B.1.617.2)(SPD-C82Ed, ACROBiosystems).
Quantitative Flow Cytometry Assay
Spike Protein Labeled Beads
The detection of spike protein-specific antibodies by flow cytometry was enabled by using 7 µm diameter avidin-coated paramagnetic beads (VM-60-10, Spherotech, Forest Lake, IL) with the various biotinylated bound spike proteins. Briefly, saturating concentrations of biotinylated spike proteins (8.3 µg) were reacted with 200 µL of (1% w/v) washed beads. After 2 hours of gentle mixing, the beads were washed and resuspended with 6 mL of PBS with 1% BSA for use in the assay. Final concentration of beads was approximately 5×106 beads per mL.
Quantitative Flow Cytometry
The standard curve for quantitation of spike-reactive antibodies was developed utilizing the spike-protein coated beads. The assay procedure was an initial incubation of 100 µl of bead suspension with 100 µL of known concentrations of monoclonal antibody or serum samples for one hour with mixing. Beads were washed and incubated with goat-anti-human IgG-alexa-488 (A48276, Life Technologies) for 30 minutes. Beads were again washed and resuspended for analysis on a Becton-Dickinson FACS caliber flow cytometer on the FL-1 channel. The standard curve was developed for all the variants using the anti-RBD-specific monoclonal antibody developed by ACROBioSystems (SAD-S35). All standard curves were developed using this antibody and the secondary antibody. Mean channel fluorescence of each serum sample was compared to the standard curve to determine the concentration of spike protein-reactive antibodies. While monoclonal antibodies do not reflect on the diverse polyclonal reactivity that would be seen in an immune serum, it does provide a reasonable quantification of total human antibody bound to the spike protein.
Confocal Immunofluorescent Analysis
The procedure previously used to determine the binding and blocking of spike proteins to HLC and AT-2 target cells7,10 was utilized to determine the ability of post-vaccination serum to inhibit the binding of variant spike proteins. Target cells (200 µL at 1 x 106/mL) were plated in 16 well collagen-coated chamber slides. Cells were allowed to bind to the slides overnight. After overnight incubation, medium was removed, and cells were fixed in 1% formaldehyde for one hour. Cells were then washed twice with 200 µL PrepaCyte permeabilization medium (WBP-1000, CMDG, St. Paul, MN). To determine the binding of each variant spike protein to target cells, cells in each well were incubated with 250 ng of spike proteins diluted in 100 µL of PermaCyte Medium for 30 minutes. Cells were then washed twice with PermaCyte Medium and then incubated with 100 ng of streptavidin-alexa 594 (S11227, Life Technologies, Eugene, OR) and counterstained with DAPI for 30 minutes. Cells were then washed with 200 µL of PermaCyte Medium and analyzed by confocal microscopy. All confocal images were obtained using an Olympus Fluoview 1000 confocal microscope.
To determine the degree of blocking of spike binding to target cells mediated by serum antibodies, we used a modification of the same procedure used to examine the binding of the spike protein to the target cells. Briefly, 250 ng aliquots of variant spike proteins were incubated with 100 µL of subject serum for 1 hour with mixing prior to addition to fixed and permeabilized cells. Cells were then incubated for 30 minutes, washed with PermaCyte and then incubated for another 30 minutes with streptavidin-alexa 594 and DAPI. Cells were then washed with 200 µL of Permacyte Medium. Cells were then analyzed by confocal microscopy.
Cellular Flow Cytometry
The utility of analyzing the binding of biotinylated spike proteins by flow cytometry was examined by using a similar procedure used for the analysis of spike protein binding to slide-plated cells but adapted to cells in suspension. Briefly, cells were directly subjected to similar conditions to those bound to slides (dissociation with expansion flasks by Tryp-LE, washing and resuspension). Cells (5 x 105) were fixed for one hour in 1% formaldehyde; and were then washed and permeabilized with 2 mL of PermaCyte Medium. They were then incubated for 30 minutes with 250 ng of biotinylated spike variants, washed twice with 2 mL of PermaCyte Medium and incubated with streptavidin-Alexa 594 and incubated for 30 minutes. Cells were again washed with 2 mL of Permacyte Medium and resuspended in PBS and analyzed by flow cytometry.
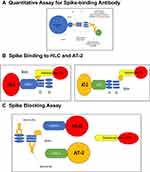
Figure 1 details all the analytical methods used in this study.
Results
Development of Quantitative Flow Cytometry for Anti-Spike Antibodies
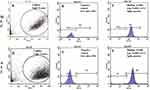
As described in the Methods and Materials section above, we utilized the biotinylated spike protein variant-coated beads to develop a standard curve utilizing the ACROBiosystems antibody as standard. While it is important to understand that comparing a monoclonal antibody to a polyclonal antibody response does not account for differences in the binding affinities of individual antibodies in a polyclonal vaccinated serum, nor their binding to different portions of the spike protein, it does provide a good estimate of antibody binding to the variant spike protein at a given concentration. Figure 2 shows a typical standard curve utilizing the ACROBiosytems antibody against spike proteins. The figure also shows the gating on beads and the fluorescent signals on FL-1 that correlate with antibody concentration. This assay, as shown, was used to analyze donor serum to determine total anti-SARS-CoV-2. The sensitivity of this assay was 0.153–2500 ng/mL.
Comparison of Anti-SARS-CoV-2 Antibody Expression Stimulated by the J&J, Moderna and Pfizer Vaccines
As determined by the quantitation flow cytometry assay, levels of spike variant-specific antibodies stimulated by the 3 vaccines were compared at 2.5 months post-vaccination. As shown in Table 1, the highest antibody levels were seen with the Moderna vaccine, while similar levels of antibody were stimulated by the J&J and Pfizer vaccines. In individual samples, there was a large variability seen between donors, and within each donor there was often a large variability seen between variants. This is shown in the table by the individual results and large standard deviations of the averages.
 |
Table 1 Quantitation of Anti-SARS-CoV-2 Serum Antibody Expressed 2.5 Months After Vaccination |
Binding of Spike Proteins to HLC and AT-2
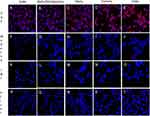
The binding of SARS-CoV-2 spike proteins to HLC and AT-2 cells was determined by fluorescent confocal microscopy. Positive binding of spike protein was demonstrated by red fluorescence. As shown in Figure 3, HLC did not bind spike 1, but was able to bind the wild type and all the variant spike proteins. AT-2 cells were able to bind each of the different spike proteins.
Analysis of Spike-Blocking Antibodies
The presence of antibodies capable of inhibiting binding of spike proteins was determined by fluorescent confocal analysis. Spike proteins were pre-incubated with subject serum prior to addition to target cells. As is seen in Figure 3, positive binding of Spike proteins was indicated by red fluorescence of cells. Inhibition of binding was shown by reduced or absent red fluorescence of the cells.
Blocking antibody against the spike variants developed from 2 weeks to 2.5 months as shown with both target HLC (Figure 4) and AT-2 cells (Figure 5). There was a significant, but not complete block of spike protein binding by serum at 2 weeks, while at 2.5 months there was complete block of detectable spike binding.
Analysis of variant-specific blocking antibodies was analyzed for serum obtained from individuals receiving the different vaccinations. As seen in Figure 6, pre-vaccination serum did not block spike protein binding to HLC, whereas each of the different vaccinations stimulated the expression of blocking antibodies to all the variants. AT-2 cells were able to bind each of the different spike proteins. Binding to AT-2 cells were also uniformly blocked by serum from individuals receiving any of the 3 vaccines (Figure 7). Adaptation of this assay for flow cytometry demonstrated that this system worked well for determining spike binding to target cells. An additional desirable quality afforded by flow cytometry is the ability to easily compare the relative fluorescence of each binding which directly relates to the amount of spike protein that is bound (Figure 8). These cellular targets were demonstrated to be useful in these analyses.
Discussion
The SARS-CoV-2 virus is spread primarily by airborne transmission. The respiratory system is the first entry point for the virus, via the ACE-2 receptor, which is the attachment point for the virus.12,13 The respiratory system is the main focal point for most of the morbidity and mortality associated with infection.14 In some cases, infection with the virus has also been associated with non-respiratory pathology including brain, kidney, cardiac, digestive system, and liver.1–4 While many organ systems contain cells that express ACE-2 and can be infected by that portal, it has been demonstrated that the SARS-CoV-2 virus is capable of infecting cells that do not possess the ACE-2 receptor.5,7 It has been shown that hepatic parenchymal cells could be infected by the virus, yet hepatocytes do not typically express ACE-2,7,8 suggesting a different mechanism for viral attachment and entry.
In a previous publication,7 we reported that hepatocytes do not express ACE-2 but still bound the viral spike protein. We also reported that hepatocytes were unable to bind the S1 portion, which contains the receptor-binding domain (RBD) that binds to ACE-2. Similarly, we showed that an immortalized HLC also did not express ACE-2, yet bound spike protein and not the Spike S1 protein. The Spike S1 protein was also unable to block the binding of biotinylated intact Spike protein. Additionally, we demonstrated that antibodies directed against the surface membrane asialoglycoprotein receptor could block the binding of spike protein to hepatocytes and HLC. Combined, these results suggested that the spike protein binds to hepatocytes and HLC via the asialoglycoprotein-1 receptor and the S2 portion of the Spike protein. The additional presence of TMPRSS-2 on hepatocytes8 potentially provides a co-factor needed for internalization of the virus. The block of spike protein binding by commercially available anti-spike monoclonal antibodies to hepatocytes and HLC suggests that HLC could serve as a target cell to determine the ability of post-vaccination serum to block the binding of spike protein, thus serving as a useful tool to assess immunity against the S2 portion of the spike protein. We have previously reported differentiation of alveolar type-2 cells (AT-2) from immortalized E12-MLPC cells as an in vitro system to study the binding of spike protein to the ACE-2 receptor.10 The model provides a cell line that can assess immunity to the S1 portion of the spike protein. The use of two different cell types from lung and liver enables the ability to assess antibodies directed towards both the S1 (extensively studied) and the S2 (rarely studied) portions of the spike protein.
This paper describes a study utilizing the HLC and AT-2 as target cells to determine if binding to the spike protein could be inhibited by post-vaccination serum. The serum of ten individuals were obtained prior to and post vaccination with the three different vaccines (J&J, Moderna, Pfizer). Serum samples were quantitated for anti-spike specific antibody by flow cytometry-based IFA (Figure 1 and Table 1) and tested for their ability to inhibit the binding of spike protein to the target cells. Samples were also tested for the ability to block spike protein binding using the same methodology previously described to study both the binding and blocking of spike proteins utilizing HLC and AT-2 cells.7,10 We observed that all variants tested bound to both HLC and AT-2 cells (we had one opportunity to test the omicron variant and found that it bound to both HLC and AT-2 cells and could be blocked by immune serum, data not shown). We also established that serum from all subjects tested after vaccination with all three vaccines stimulated the production of antibodies that completely blocked the binding of all the spike variants. When the antibody response was analyzed, we identified significant variations in antibody levels between individuals in addition to within each of the individuals themselves. Antibody levels did not necessarily reflect the degree of spike protein-binding block to target cells, as even low levels of antibody were capable of complete inhibition of binding. We believe that the spike-blocking assay provided a better assessment of protective immunity than just measuring antibody levels. While not extensively studied, we found that the binding/inhibition assays were easily adaptable to analysis by flow cytometry allowing utilization of this assay to laboratories that do not have access to confocal microscopy but do have availability of a flow cytometer.
We believe that this assay has many advantages over simply determining gross anti-SARS-CoV-2 antibody levels or the use of pseudo-viruses to test inhibition of binding. First was the development of HLC and AT-2 cell lines that are stable, immortalized, highly differentiated and express the phenotypic and biological properties of mature and fully functional primary human hepatocytes and primary small alveolar cells, respectively. The use of biotinylated spike proteins to study binding of the virus to target cells instead of live virus or pseudo-virus also provides a significant advantage in testing. The use of this combined system utilizes immortalized cell lines that can be readily grown and maintained to provide a stable and reproducible source of target cells. The use of commercially available spike protein also eliminates the need for creation of new pseudo-viruses with the evolution of each new variant or use of the actual live virus. This eliminates any dangers involved with working with live or pseudo-viruses and the need for extensive biological containment. These features allow the assay to be produced on a large-scale and safely utilized by any normally equipped analytical laboratory. We believe that our system should be widely utilized as it provides a more accurate assessment of protective immunity than simple quantitation of SARS-CoV-2-specific antibodies and is safe and easily implementable in standard analytical laboratories.
Acknowledgments
This study was funded in its entirety by CMDG LLC. Cells that were generated (MLPC, HLC and AT-2) and other tissues (cord blood and peripheral blood samples) used in this study are included as an extension of IRB approved by the Quorum Review Protocol #800, March 3, 2005, which supported previous studies (7, 9–11). Donations were collected with informed donor consent for research use only.
Disclosure
Dr. Daniel P. Collins was an owner and employee of Cytomedical Design Group LLC, during the conduct of this study. In addition, Dr. Daniel P Collins has patents 7,622,108 and 7,670,596 licensed to BioE, Inc. The authors report no other conflicts of interest in this work.
References
1. Puelles VG, Lütgehetmann M, Lindenmeyer MT, et al. Multiorgan and renal tropism of SARS-CoV-2. N Engl J Med. 2020;383(6):590–592. doi:10.1056/NEJMc2011400
2. Lin L, Jiang X, Zhang Z, et al. Gastrointestinal symptoms of 95 cases with SARS-CoV-2 infection. Gut. 2020;69(6):997–1001. doi:10.1136/gutjnl-2020-321013
3. Hamming I, Timens W, Bulthuis MLC, et al. Tissue distribution of ACE-2 protein, the functional receptor for SARS Corona virus. A first step in understanding SARS pathogenesis. J Pathol. 2004;203(2):631–637. doi:10.1002/path.1570
4. Chai X, Hu L, Zhang Y, et al. Specific ACE2 expression in cholangiocytes may cause liver damage after 2019-nCoV infection. bioRxiv. 2020. doi:10.1107/2020.02.03.931/766
5. Wang Y, Liu S, Liu H, et al. SARS-CoV-2 infection of the liver directly contributes to hepatic impairment in patients with COVID-19. J Hepatol. 2020;73(4):807–816. doi:10.1016/jhep.2020.05.002
6. Lamers MM, Beumer J, van der Vaart J, et al. SARS-CoV-2 productively infects human gut enterocytes. Science. 2020;369(6499):50–54. doi:10.1126/science.abc1669
7. Collins DP, Steer CJ. Binding of the SARS-CoV-2 spike protein to the asialoglycoprotein receptor on human primary hepatocytes and immortalized hepatocyte-like cells by confocal microscopy. Hepatic Med. 2021;13:37–44. doi:10.2147/HMER.S301979
8. X-X H, Y-X M, Lin Y-X, et al. ACE2 and TMPRSS2 expression in hepatocytes of chronic HBV infection patients. Infect Dis Immun. 2021;1(1):36–42. doi:10.1097/ID9.0000000000000007
9. Collins DP, Hapke JH, Aravalli RN, Steer CJ, Papaccio G. Development of immortalized human hepatocyte-like cells by fusion of multi-lineage progenitor cells with primary hepatocytes. PLoS One. 2020;15(6):e234002. doi:10.1371/journal.pone.0234002
10. Collins DP, Osborn MJ, Steer CJ. Differentiation of immortalized human multi-lineage progenitor to alveolar type 2-like cells: angiotensin-converting enzyme 2 expression and binding of severe acute respiratory syndrome coronavirus 2 spike and spike 1 proteins. Cytotherapy. 2021;23(12):1064–1073. doi:10.1016/j.jcyt.2021.07.017
11. Burger MJ, Adams SD, Tigges BM, et al. Differentiation of umbilical cord blood-derived multi-lineage progenitor cells into respiratory epithelial cells. Cytotherapy. 2006;8(5):480–487. doi:10.1080/14653240600941549
12. Shang J, Wan Y, Luo C, et al. Cell entry mechanisms of SARS-CoV-2. Proc Natl Acad Sci USA. 2020;117(21):11727–11734. doi:10.1073/pnas.2003138117
13. Lan J, Ge J, Yu J, et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature. 2020;581(7807):215–220. doi:10.1038/s41586-020-2180-5
14. Tang D, Comish P, Kang R, Hobman TC. The hallmarks of COVID-19 disease. PLoS Pathog. 2020;16(5):e1008536. doi:10.1371/journal.ppat.1008536
 © 2023 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2023 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.