Back to Journals » OncoTargets and Therapy » Volume 8
Upregulation of nucleostemin in colorectal cancer and its effects on cell malignancy
Authors Wei B, Huang Q, Zhong X, Zhang Z
Received 1 December 2014
Accepted for publication 24 April 2015
Published 21 July 2015 Volume 2015:8 Pages 1805—1814
DOI https://doi.org/10.2147/OTT.S78461
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 7
Editor who approved publication: Professor Daniele Santini
Bin Wei,1,* Qiaoying Huang,2,* Xiaogang Zhong3
1Department of Gastroenterology and Peripheral Vascular Surgery, The People’s Hospital of Guangxi Zhuang Autonomous Region, 2Department of Medical Molecular Biology, The First Affiliated Hospital of Guangxi University of Chinese Medicine Research, 3Department of Gastroenterology and Peripheral Vascular Surgery, The People’s Hospital of Guangxi Zhuang Autonomous Region, Nanning, Guangxi, People’s Republic of China
*These authors contributed equally to this work
Objective: Nucleostemin (NS) is a new protein localized in the nucleolus of most stem cells and tumor cells, which regulates their self-renewal and cell cycle progression. The aim of this study was to investigate the expression of NS in colorectal cancer (CRC) and the effects of NS knockdown in the Sw620 cell line to provide basis for clinical target therapy.
Methods: NS expression in 372 patients with CRC and 367 normal participants was assessed using immunohistochemistry. The expression level of NS gene was evaluated by polymerase chain reaction. Then, the relationship among NS expression, clinicopathological features, and prognosis was analyzed. Silencing of NS expression was achieved by using NS-specific small-interfering RNAs. The viability and growth rate of Sw620 cells were determined by proliferation and invasion assays. Cell cycle distribution of the cells was analyzed by flow cytometry.
Results: High NS expression was positively related with node metastasis, distant metastasis, and TNM stage. In Kaplan–Meier survival analysis, patients with low NS expression always had significantly longer survival time than those with high expression. Moreover, our results showed that knockdown of NS expression inhibited proliferation and viability of Sw620 cells in a time-dependent manner. Cell cycle studies revealed that NS depletion resulted in G1 cell cycle arrest at short times of transfection (24 hours), followed with apoptosis at longer times (48 hours and 72 hours), suggesting that post-G1 arrest apoptosis occurred in Sw620 cells.
Conclusion: Overall, these results point to the essential role of NS in Sw620 cells; thus, this gene might be considered a promising target for treatment of CRC.
Keywords: apoptosis, colorectal cancer, nucleostemin, small interfering RNA, Sw620, target therapy
Introduction
Colorectal cancer (CRC) is the third most prevalent cancer in humans worldwide and accounts for ~9% of all cancer mortalities.1,2 Early diagnosis of CRC is beneficial to guide surgical resection and improve the survival rate for CRC. However, the long-term survival rate of, as well as accurate prognosis for, patients with CRC remains poor.3 Current knowledge of molecular alterations that are important for CRC, including epigenetic and genetic changes in key tumor suppressors and oncogenes, is extensive; however, it still represents the tip of the iceberg of knowledge that needs to be resolved for a complete understanding of CRC pathogenesis.4
In 2002, Tsai and McKay discovered that a novel gene called nucleostemin (NS) is apparently expressed in stem cells of embryonic and adult rat central nervous systems.5,6 The protein coded by the NS gene was found in the nucleoli of undifferentiated cells, such as adult and embryonic stem cells, neural stem cells, and human bone marrow stem cells, but not in differentiated counterpart cells, indicating that NS gene is silenced during normal cell differentiation.7,8 Interestingly, recent reports suggest that the NS gene is also abundantly expressed in several human cancer cell lines, such as SGC-7901 (gastric), HeLa (cervical), 5637 (bladder), PC-3 (prostate), and HL-60 (acute myelocytic leukemia).9–12 Some experiments using RNA interference (RNAi) showed that inhibition of NS gene expression markedly inhibited proliferation and cell cycle progression of cancerous cells, followed with induction of differentiation and/or apoptosis.11–15 Recently, a high expression level of NS has been reported in gastric cancer patients.15 Consistent with this, RNAi-mediated NS knockdown inhibited proliferation and induced differentiation and apoptosis in gastric cancer cell lines.16 However, the importance of NS in other types of digestive cancers, especially CRCs, needs to be addressed.
This study was designed to investigate the functional importance and therapeutic potential of NS gene expression and effects of NS knockdown on cell cycle and apoptosis in CRC cell lines. Our result showed that RNAi-mediated NS silencing induced G1 cell cycle arrest, followed with apoptosis in CRC cell lines.
Materials and methods
Participants and samples
In this study, 372 patients diagnosed with CRC, who were in our hospital during 2010–2012, were recruited according to their tissue detection data (details in Table 1). Three hundred and sixty-seven patients in our department without CRC were recruited as controls. Formalin-fixed paraffin-embedded tissues were obtained from the Second and First Affiliated Hospitals of Jiangxi University of Chinese Medicine (Nangchang, People’s Republic of China). The follow-up period was defined to be the duration from the date of surgery to the date of patient death or the final follow-up time point of January 2014. Follow-up data were recorded by communicating with the patients or their relatives. This study was approved by the Ethics Committee of the Jiangxi University of Chinese Medicine. Informed consent was signed by all participants, and the study was performed in accordance with the Declaration of Helsinki.
  | Table 1 Clinicopathological characteristics and NS expression in patients with colorectal cancer |
Cell culture and transfection
LoVo, Caco2, and Sw620 cell lines were purchased from the American Type Culture Collection (Rockville, MD, USA). These cells were maintained in RPMI-1640 supplemented with 10% fetal bovine serum and incubated in 5% CO2 at 37°C. NS-specific small interfering RNAs (NS-siRNAs) and scrambled siRNA (control) were synthesized by Invitrogen Life Technologies and transfected into cells using Lipofectamine® 2000 (Invitrogen Life Technologies, Shanghai, People’s Republic of China) according to the manufacturer’s instructions.
Histopathological examination and scoring
The NS protein expression was detected by immunohistochemistry of paraffin-embedded sections of CRC samples. After deparaffinization, the sections were incubated with anti-NS antibody (Abcam, Cambridge, MA, USA), as described previously.9 Negative-control sections were incubated with preimmunized rabbit serum (Abcam). The immunostaining results were analyzed by measuring the intensity of positive regions and the percentage of positive-expression cells. The staining intensity was scored on a three-level scale (zero to three). The percentage of stained tumor cells was graded in four levels as follows: 0 (<5% positive cells),1 (5%–25% positive cells), 2 (26%–50% positive cells), 3 (51%–75% positive cells), and 4 (>75% positive cells). The final scores ranged from 0 to 12. In this study, scores between zero and four were denoted as low expression of NS, and the scores ranging from 5 to 12 were denoted as high expression. Incongruous scores would be reevaluated by two individual pathologists until a consensus score was obtained.
PCR analysis
RNA isolation and reverse transcription were performed as previously described.22 Oligonucleotide primer sequences were as follows: β-actin (264 bp): forward 5′-GAG ACC TTC AAC ACC CCA GCC-3′; reverse 5′-AAT GTC AC G CAC GATT TCC C-3′; NS (201 bp): forward 5′-TCC CCA TCG CCA TCC CC-3′; reverse 5′-CAC CAT GGC CTC GGC TGG-3′. For all the above genes, amplification was performed under the same cycling conditions (1 minute at 94°C, 50 seconds at 57°C, and 1 minute at 72°C), except for the number of cycles that were specified for each gene (32 for NS). All the experiments were repeated at least three times.
Western blot
Sw620 cells were harvested at specific time points after treatment with reagents as indicated in each experiment. Cells were mixed with loading buffer and subjected to electrophoresis. After electrophoresis, proteins were transferred to polyvinylidene difluoride membranes (Pall Filtron) using a semidry blotting apparatus (Pharmacia) and probed with mouse monoclonal antibodies, followed by incubation with peroxidase-labeled secondary antibodies. Detection was performed by the use of a chemiluminescence system (Amersham) according to the manufacturer’s instructions. Then membrane was stripped with elution buffer and reprobed with antibodies against the nonphosphorylated protein as a measure of loading control. Controls for the immunoprecipitation used the same procedure, except that agarose beads contained only mouse immunoglobulin G. All the experiments were repeated at least three times.
Migration and invasion assays
We used a Transwell insert (24-well insert, pore size 8 μm; Corning, Inc, Corning, NY, USA) to determine the effect of microRNA miR-320a on K562 migration and invasion in vitro. Briefly, the transfected cells were first starved in serum-free medium overnight, and 3×104 cells were resuspended in serum-free medium and placed in the top chambers in triplicate. The lower chamber was filled with 10% fetal bovine serum as the chemoattractant and incubated for 48 hours for the migration assay and 72 hours for the invasion assay. For the invasion assay, the inserts were previously coated with extracellular matrix gel (BD Biosciences, Bedford, MA, USA). At the end of the experiments, the cells on the upper surface of the membrane were removed, and the cells on the lower surface were fixed and stained with 0.1% crystal violet. Five visual fields of each insert were randomly chosen and counted under a light microscope.
MTT assay
To record the cell viability, the 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay was used for assessment, as previously described. After transfection, cell lines were plated in 96-well plates at a density of 104 cells per well and incubated for 24 hours, 48 hours, and 72 hours. Then, a total of 20 μL of 5 mg/mL MTT (Sigma-Aldrich) was added to the culture wells and incubated for 4 hours at 37°C. Before measurement, the supernatant was discarded, and 200 μL of dimethyl sulfoxide was added to each well to dissolve the formazan. Optical density at 490 nm was recorded by using a spectrophotometer (SpectraMax Plus384; Molecular Devices, Sunnyvale, CA, USA). All of the experiments were performed in triplicate.
Cell cycle analyses
DNA contents of cells were analyzed using flow cytometry as described previously. Control and transfected cells were harvested and washed twice with phosphate-buffered saline, fixed in 70% ethanol, and stored at −20°C until analysis. Then the cells were stained with 20 μg/mL propidium iodide containing 20 μg/mL RNase (DNase free) for 2 hours. The stained cells were analyzed by flow cytometry (Partec PAS, Münster, Germany). The populations of G0/G1, S, G2/M, and sub-G1 cells were determined using Mulicycle Cell Cycle Software. The results are expressed as percentage of the cells in each phase.
Statistical analysis
Results are expressed as mean ± standard deviation. Data were analyzed using the unpaired two-tailed Student’s t-test and the log-rank test. P-values <0.05 were considered significant.
Results
Elevated level of NS in CRC and the correlation between NS and clinicopathological variables
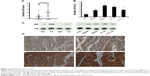
NS expression levels were found to be elevated in the CRC specimens, compared with paired normal colon tissue, by quantitative polymerase chain reaction (qPCR) (n=30, P<0.01, Figure 1A). In CRC cell lines, NS expression was also increased (P<0.05, Figure 1B). Northern blotting assay revealed that NS blot was significantly denser in CRC samples and cell lines, than in normal tissue and colon cells (Figure 1C). Moreover, the location of NS was predominantly nuclear, as shown by immunohistochemistry results (Figure 1D), and the sections showed that NS expression was significantly higher in CRC tissue than in noncancerous, normal colorectal tissue. The expression of NS protein was analyzed in 372 CRC tissue samples and 367 noncancerous colorectal tissue samples. Among the CRC tissues, 69.36% (258/372) of cases showed high NS expression (scale >5), while only 10.22% (38/367) of noncancerous colorectal tissue sections showed high NS expression (P<0.001). In addition, significant differences in NS expression were observed in the two groups in tumors with node metastasis (P=0.0017), distant metastasis (P<0.001), and different TNM stages (P=0.0016; data not shown).
Correlation between NS expression and survival rate in CRC patients
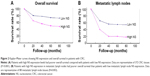
Briefly, the survival analysis revealed that patients with low NS expression (scale <5) survived significantly longer than those with high NS expression (log-rank test, P<0.001; Figure 2A). The metastatic lymph nodes in patients with low NS expression indicated a longer overall survival than in patients with high NS expression (Figure 2B, log-rank test, P=0.007). These results suggested that NS expression was closely related with the survival rate of CRC patients.
Expression of NS was efficiently inhibited by NS-siRNA in Sw620 cells
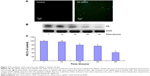
On the basis of our preliminary data on the high expression level of NS in Sw620 cell lines, we examined different RNAi techniques for silencing of this gene in Sw620 cells. One of the designed siRNAs, called NS-siRNA, could efficiently inhibit NS expression in Sw620 cells (Figure 3). As depicted in Figure 3, NS-siRNA at 200 nM was efficiently delivered into Sw620 cells (Figure 3A), and it significantly inhibited NS expression in a time-dependent manner (Figure 3B). In fact, no significant reduction in NS expression was observed after NS-siRNA transfection of Sw620 cells for 6–12 hours, whereas NS mRNA and protein levels were significantly inhibited between 16 hours and 48 hours of transfection (Figure 3B and C). The inhibition rate of NS expression in comparison with the corresponding β2-microglobulin internal control after 16 hours, 24 hours, and 48 hours were approximately 20%, 23%, and 56%, respectively (Figure 3C).
Knockdown of NS significantly inhibited CRC cell proliferation and invasion
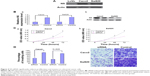
The expression of NS was analyzed in LoVo, Caco2, and Sw620 cells. Interestingly, it was expressed at higher levels in Caco2 and Sw620 cells than in LoVo cells (Figure 4A). On the basis of this observation, Caco2 and Sw620 cells were chosen for the subsequent functional analysis. Knockdown of NS by siRNA was confirmed using qPCR and Western blotting, in which obviously low expression of NS was observed after transfection (Figure 4B). The cell proliferation and invasion of Caco2 and Sw620 cells were significantly suppressed after siRNA transfection (Figure 4C and D).
Knockdown of NS leads to profound morphological and functional changes in Sw620 cells
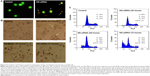
We first detected the morphology of Sw620 cells after NS-siRNA transfection (Figure 5A). Aggregation of Sw620 cells and decrease in cell confluency were typically observed in NS-depleted Sw620 cells. However, some cell death criteria, such as cell shrinking and cell debris, were observed after 48–72 hours of NS-siRNA transfection. To determine the mode of cell death in NS-siRNA-transfected cells, we studied apoptosis and necrosis by acridine orange/ethidium bromide double staining of the cells (Figure 5B). The results clearly showed that NS-siRNA-transfected cells underwent apoptosis after 48 hours. The apoptotic criteria, including nuclear fragmentation, chromatin condensation, and apoptotic bodies, were clearly observed. In these figure panels, viable cells were equally green, whereas early apoptotic cells had bright green dots or blobs in their nuclei, indicating chromatin condensation and nuclear fragmentation. Late apoptotic cells, however, stained orange and showed condensed and fragmented nuclei. Necrotic cells were uniformly orange. This means that knockdown of NS induces apoptosis in Sw620 cells.
Evidence suggests that the cell fate decision is made within the G1 phase of the cell cycle. Therefore, the cell cycle distribution of NS-siRNA-transfected Sw620 cells was also studied in this work (Figure 5C). When compared with control cells, NS-siRNA-transfected cells showed a significant increase in the G0/G1 phase of cell cycle population, with concurrent decrease in S and G2M phases after 24 hours of transfection. As might be expected, a sub-G1 peak (corresponding to apoptotic cells) was apparent after longer times of transfection. For example, after 24 hours, the G0/G1 cell cycle population of NS-siRNA-transfected cells (58%) was higher than that in control cells (47%). Moreover, the sub-G1 cell population (apoptotic cells) was increased from 21% to 38% during the 48- to 72-hour period of transfection, respectively. Results are expressed as the mean ± standard deviation. Data were analyzed using the unpaired two-tailed Student’s t-test and the log-rank test. P-values <0.05 were considered significant. All these indicated that knockdown of NS induces G0/G1 cell cycle arrest in Sw620 cells.
Discussion
CRC is a serious problem for human health.17 The incidence and mortality of CRC in the People’s Republic of China has increased rapidly in the past few decades. Early detection is essential to reduce mortality and improve survival rates, but this approach is hampered by the lack of convenient screening tools with high specificity and sensitivity for early-stage tumors.18 Therefore, novel biomarkers for detection of early-stage CRC are urgently required.
Several reports have suggested that NS is a marker of stem cells and is involved in controlling self-renewal, cell cycle progression, and proliferation in both stem cells and cancerous cells. Considering that NS plays a critical role in cell proliferation, we examined expression and function of NS in Sw620 and Caco2 cells as models of CRC cell lines. To our knowledge, the functional importance of NS in CRC has not been studied until now. Our results indicated that NS mRNA was highly expressed in Sw620 and Caco2 cells. This finding is in alignment with previous studies based on NS overexpression in several human cancer cell lines.19–24
Previous reports have indicated that NS expression was related with cancer cell metastasis, TNM stage, and mortality. Furthermore, these correlations were found to be independent of other patient characteristics.25 These findings hint that a high NS expression could be used as a prognostic marker for CRC diagnosis. An abnormally high expression of NS has been found in various human cancers, including neuroblastoma and pancreatic, lung, bladder, liver, ovarian, and breast cancers.8,13,25,26 However, the mechanism of how NS regulates CRC is not yet elucidated. In the present study, immunohistochemistry revealed that NS had a nuclear expression pattern and was upregulated in CRC tissues, compared to the expression level in noncancerous tissues.
In human breast, overexpression of NS has been proposed to contribute to malignant progression by the inactivation of wild-type p53 and p38 mitogen-activated protein kinase, as well as by decreasing p16 protein expression.27 In addition, NS is a good candidate prognostic marker in patients with lung adenocarcinoma.28 Furthermore, high NS expression has been negatively correlated with prognosis in patients with pancreatic neuroendocrine tumors and medulloblastoma.29,30 Similar to these findings, a worse outcome was observed in CRC patients with high NS expression. Furthermore, multivariate analysis indicated that high NS expression was an independent prognostic parameter for CRC patients.30 In cell lines, the transfection with NS-siRNA significantly inhibited CRC cell proliferation and invasion. Taken together, NS can be used not only as a prognostic marker, but also as a potential therapeutic target.
In our study, the roles of NS in cell cycle progress and apoptosis were determined by NS-siRNA. These oligos led to a significant decrease in the NS mRNA expression. The results showed that NS knockdown inhibited growth of Sw620 cells 24 hours after transfection. Apoptosis began after 48 hours and increased to its highest level after 72 hours. Therefore, NS depletion in Sw620 cells resulted in growth inhibition at short times and apoptosis at longer times. These results are in full agreement with cell cycle results, wherein an accumulation in G1 phase population was observed after 24 hours of NS-siRNA transfection. After this time point, however, the cell population at the G1 phase decreased and a sub-G1 peak appeared, suggesting that post-G1 arrest of apoptosis was the exact mode of NS-siRNA action in Sw620 cells. Most literature reports suggest that NS depletion inhibited proliferation and induced cell cycle arrest in cancer cell lines.31–34 For instance, NS-siRNA in bladder cancer cells led to G1 cell cycle arrest in prostate PC-3 cells and bladder cancer 5637 cells. However, NS may also induce G2/M cell cycle arrest as in the case of bladder cancer SW1710 cells. Apparently, the role of NS in the regulation of the G1 phase of the cell cycle in Sw620 cells are in full agreement with most of these literature reports.
Although several reports point to the apoptotic effects of NS depletion in different cancerous cells, induction of apoptosis following G1 cell cycle arrest is a novel finding of this study. In fact, it has been previously reported that NS depletion induced a rapid apoptosis response in HeLa cells, PC-3 cells, human bladder (5637) cells, and HL-60 cells.23,27,35 In our experiments, however, we observed a delayed apoptosis response in Sw620 and Caco2 cells. This may be related to the different levels of NS depletion and the protein contents of the cells used in distinct experiments.
In conclusion, NS may be a diagnostic biomarker for CRC and high expression of NS is closely related with poor prognosis. Further investigations are necessary to validate the findings and to elucidate the underlying mechanisms of how NS regulates the development of CRC.
Disclosure
The authors report no conflicts of interest in this work.
References
Yao Y, Li J, Wang L. Large intervening non-coding RNA HOTAIR is an indicator of poor prognosis and a therapeutic target in human cancers. Int J Mol Sci. 2014;15(10):18985–18999. | ||
Patel SA, Bhambra U, Charalambous MP, et al. Interleukin-6 mediated upregulation of CYP1B1 and CYP2E1 in colorectal cancer involves DNA methylation, miR27b and STAT3. Br J Cancer. 2014;111(12):2287–2296. | ||
Boisen MK, Dehlendorff C, Linnemann D, et al. Tissue microRNAs as predictors of outcome in patients with metastatic colorectal cancer treated with first line capecitabine and oxaliplatin with or without bevacizumab. PLoS One. 2014;9(10):e109430. | ||
Runtsch MC, Round JL, O’Connell RM. MicroRNAs and the regulation of intestinal homeostasis. Front Genet. 2014;5:347. | ||
Tsai RY, McKay RD. A nucleolar mechanism controlling cell proliferation in stem cells and cancer cells. Genes Dev. 2002;16:2991–3003. | ||
Goossens-Beumer IJ, Derr RS, Buermans HP, et al. MicroRNA classifier and nomogram for metastasis prediction in colon cancer. Cancer Epidemiol Biomarkers Prev. 2014;24(1):187–197. | ||
Peacock O, Lee AC, Cameron F, et al. Inflammation and MiR-21 pathways functionally interact to downregulate PDCD4 in colorectal cancer. PLoS One. 2014;9(10):e110267. | ||
Wang J, Paris PL, Chen J, et al. Next generation sequencing of pancreatic cyst fluid microRNAs from low grade- benign and high grade- invasive lesions. Cancer Lett. 2014;356(2 pt B):404–409. | ||
Zhang J, Fei B, Wang Q, et al. MicroRNA-638 inhibits cell proliferation, invasion and regulates cell cycle by targeting tetraspanin 1 in human colorectal carcinoma. Oncotarget. 2014;5(23):12083–12096. | ||
Jiang JX, Zhang N, Liu ZM, Wang YY. Detection of microRNA-21 expression as a potential screening biomarker for colorectal cancer: a meta-analysis. Asian Pac J Cancer Prev. 2014;15(18):7583–7588. | ||
Adams SV, Newcomb PA, Burnett-Hartman AN, et al. Rare circulating microRNAs as biomarkers of colorectal neoplasia. PLoS One. 2014;9(10):e108668. | ||
Zhao HJ, Ren LL, Wang ZH, et al. MiR-194 deregulation contributes to colorectal carcinogenesis via targeting AKT2 pathway. Theranostics. 2014;4(12):1193–1208. | ||
Xie WQ, Tan SY, Wang XF. Effect of a common genetic variant microRNA-146a rs2910164 on colorectal cancer: a meta-analysis. J Dig Dis. 2014;15(12):647–653. | ||
Li ZW, Yang YM, Du LT, et al. Overexpression of miR-223 correlates with tumor metastasis and poor prognosis in patients with colorectal cancer. Med Oncol. 2014;31(11):256. | ||
Nonaka R, Nishimura J, Kagawa Y, et al. Circulating miR-199a-3p as a novel serum biomarker for colorectal cancer. Oncol Rep. 2014;32(6):2354–2358. | ||
Ress AL, Stiegelbauer V, Winter E, et al. MiR-96-5p influences cellular growth and is associated with poor survival incolorectal cancer patients. Mol Carcinog. Epub 2014 Sep 25. | ||
Xu L, Li M, Wang M, Yan D, Feng G, An G. The expression of microRNA-375 in plasma and tissue is matched in humancolorectal cancer. BMC Cancer. 2014;14:714. | ||
Hara T, Jones MF, Subramanian M, et al. Selective targeting of KRAS-mutant cells by miR-126 through repression of multiple genes essential for the survival of KRAS-mutant cells. Oncotarget. 2014;5(17):7635–7650. | ||
Du M, Liu S, Gu D, et al. Clinical potential role of circulating microRNAs in early diagnosis of colorectal cancer patients. Carcinogenesis. 2014;35(12):2723–2730. | ||
Saadatian Z, Masotti A, Nariman Saleh Fam Z, Alipoor B, Bastami M, Ghaedi H. Single-nucleotide polymorphisms within microRNAs sequences and their 3′ utr target sites may regulate gene expression in gastrointestinal tract cancers. Iran Red Crescent Med J. 2014;16(7):e16659. | ||
Zheng G, Du L, Yang X, et al. Serum microRNA panel as biomarkers for early diagnosis of colorectaladenocarcinoma. Br J Cancer. 2014;111(10):1985–1992. | ||
Yau TO, Wu CW, Dong Y, et al. microRNA-221 and microRNA-18a identification in stool as potential biomarkers for the non-invasive diagnosis of colorectal carcinoma. Br J Cancer. 2014;111(9):1765–1767. | ||
Feng J, Yang Y, Zhang P, et al. miR-150 functions as a tumour suppressor in human colorectal cancer by targeting c-Myb. J Cell Mol Med. 2014;18(10):2125–2134. | ||
Lin Z, Chen L, Song M, Shi KQ, Tang KF. Association between a polymorphism in miR-34b/c and susceptibility to cancer – a meta-analysis. Asian Pac J Cancer Prev. 2014;15(17):7251–7255. | ||
Orang AV, Barzegari A. MicroRNAs in colorectal cancer: from diagnosis to targeted therapy. Asian Pac J Cancer Prev. 2014;15(17):6989–6999. | ||
Zhou JJ, Zheng S, Sun LF, Zheng L. MicroRNA regulation network in colorectal cancer metastasis. World J Biol Chem. 2014;5(3):301–307. | ||
Cao Y, Hu J, Fang Y, Chen Q, Li H. Association between a functional variant in microRNA-27a and susceptibility tocolorectal cancer in a Chinese Han population. Genet Mol Res. 2014;13(3):7420–7427. | ||
Yamada N, Tsujimura N, Kumazaki M, et al. Colorectal cancer cell-derived microvesicles containing microRNA-1246 promote angiogenesis by activating Smad 1/5/8 signaling elicited by PML down-regulation in endothelial cells. Biochim Biophys Acta. 2014;1839(11):1256–1272. | ||
Zhang H, Jia R, Wang C, Hu T, Wang F. Piceatannol promotes apoptosis via up-regulation of microRNA-129 expression incolorectal cancer cell lines. Biochem Biophys Res Commun. 2014;452(3):775–781. | ||
Li L, Sheng Y, Lv L, Gao J. The association between two microRNA variants (miR-499, miR-149) and gastrointestinal cancer risk: a meta-analysis. PLoS One. 2013;8(11):e81967. | ||
Zhou Y, Feng X, Liu YL, et al. Down-regulation of miR-126 is associated with colorectal cancer cells proliferation, migration and invasion by targeting IRS-1 via the AKT and ERK1/2 signaling pathways. PLoS One. 2013;8(11):e81203. | ||
Kamata I, Ishikawa Y, Akishima-Fukasawa Y, et al. Significance of lymphatic invasion and cancer invasion-related proteins on lymph node metastasis in gastric cancer. J Gastroenterol Hepatol. 2009;24(9):1527–1533. | ||
Peng TL, Chen J, Mao W, Song X, Chen MH. Aryl hydrocarbon receptor pathway activation enhances gastric cancer cell invasiveness likely through a c-Jun-dependent induction of matrix metalloproteinase-9. BMC Cell Biol. 2009;10:27. | ||
Nomura T, Kamio Y, Takasu N, et al. Kimura intrahepatic micrometastases around liver metastases from gastric cancer. J Hepatobiliary Pancreat Surg. 2009;16(4):493–501. | ||
Achyut BR, Ghoshal UC, Moorchung N, Mittal B. Transforming growth factor-B1 and matrix metalloproteinase-7 promoter variants induce risk for Helicobacter pylori-associated gastric precancerous lesions. DNA Cell Biol. 2009;28(6):295–301. |
 © 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.