Back to Journals » Cancer Management and Research » Volume 13
Upregulation of Linc-ROR Promotes the Proliferation, Migration, and Invasion of Gastric Cancer Cells Through miR-212-3p/FGF7 Axis
Authors Mi Y, Li Y, He Z, Chen D, Hong Q, You J
Received 20 October 2020
Accepted for publication 24 December 2020
Published 2 February 2021 Volume 2021:13 Pages 899—912
DOI https://doi.org/10.2147/CMAR.S287775
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Eileen O'Reilly
Yanjun Mi,1,* Yongwen Li,2,* Zhuo He,3,* Donghan Chen,2 Qingqi Hong,2 Jun You2
1Department of Medical Oncology, The First Affiliated Hospital of Xiamen University, Teaching Hospital of Fujian Medical University, Xiamen, Fujian Province, 361003, People’s Republic of China; 2Department of Gastrointestinal Surgery, The First Affiliated Hospital of Xiamen University, Teaching Hospital of Fujian Medical University, Xiamen, Fujian Province, 361003, People’s Republic of China; 3Department of Gastropancreatoduodenal Surgery, Hunan Cancer Hospital, Changsha, Hunan Province, 410013, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Jun You
Department of Gastrointestinal Surgery, The First Affiliated Hospital of Xiamen University, Teaching Hospital of Fujian Medical University, No. 55 Zhenhai Road, Si Ming District, Xiamen, Fujian Province, 361003, People’s Republic of China
Tel +86 13906051681
Email [email protected]
Background: Linc-ROR is a long non-coding RNA, that is found aberrantly expressed in various human cancers. We aim here to unveil the role of Linc-ROR in gastric cancer (GC) progression.
Methods: qPCR was used to determine gene expression. Cell viability was measured by CCK-8 assay. Transwell assays were performed to evaluate the GC cells’ migratory and invasive abilities. Xenograft mouse model was conducted to measure tumor growth.
Results: We found that Linc-ROR were overexpressed in GC tissues compared to the adjacent tissues. High Linc-ROR predicts poor prognosis of GC patients. The prediction of bioinformatics online revealed that Linc-ROR could bind to miR-212-3p. Further, dual-luciferase reporter assay confirmed a direct interaction between Linc-ROR and miR-212-3p. Overexpression of miR-212-3p facilitated GC cells’ migration and invasion, while the silencing of miR-212-3p attenuated GC cell migratory and invasive abilities. Moreover, Linc-ROR knockdown significantly suppressed the proliferation, migration, and invasion of GC cells, whereas miR-212-3p antagomir partially reversed Linc-ROR knockdown-induced phenotypes. Fibroblast growth factor 7 (FGF7), a downstream molecule of miR-212-3p, was overexpressed in GC cells. The recovery of FGF7 expression partially reversed the phenotypes caused by Linc-ROR silencing. Mechanistically, silencing of Linc-ROR contributed to the downregulation of CDK4, CDK6, Cyclin D1, N-Cadherin, Vimentin, MMP-9, MMP-2, but caused the upregulation of P21, P27, E-Cadherin, CK-19 in MGC-803 cells; however, FGF7 treatment could reverse the results induced by Linc-ROR silencing. Results in vivo further suggested that Linc-ROR knockdown repressed GC tumor growth, where the expression of miR-212-3p was up-regulated and FGF7 expression was downregulated in tumor tissues of mice.
Conclusion: These findings indicated that Linc-ROR/miR-212-3p/FGF7 axis played an important role in gastric cancer progression. Linc-ROR expression level was associated with the prognosis of GC patients.
Keywords: Linc-ROR, miR-212-3p, FGF7, gastric cancer
Introduction
Gastric cancer (GC) causes hundreds of thousands of deaths every year and is ranked as the fifth most common cancer globally.1 With the development of the gastrointestinal endoscopy, the survival rate of GC patients has been significantly improved.2 However, because the symptoms are nonspecific, most patients are diagnosed at an advanced stage of gastric cancer, losing the best time window for surgery.3 Besides, the therapeutic efficacy of chemotherapy and/or radiotherapy in metastatic gastric cancer is not satisfactory.4,5 Furthermore, the metastasis of gastric cancer is responsible for high mortality in patients.6,7 Although many risk factors, such as poor diet patterns, contribute to gastric cancer development,3,8–10 the underlying mechanisms of gastric cancer progression require further investigation.
In recent years, an explosion of interest in long non-coding RNA has revealed a multitude of regulatory functions for them in various cancers.11,12 Linc-ROR, a long non-coding RNA, modulates the behavior of human-induced pluripotent stem cells13 and has been implicated in tumorigenesis. Previous studies have suggested that Linc-ROR facilitates the occurrence and development of breast cancer by modulating the Hippo/YAP signaling pathway.14 Additionally, Linc-ROR increases the stability of c-Myc mRNA, thereby up-regulating the expression of downstream gene c-Myc, and resulting in cell proliferation and tumorigenesis.15 Preliminarily, we predicted a relationship between Linc-ROR and miR-212-3p; however, the role of the Linc-ROR/miR-212-3p axis in the development of gastric cancer remains unclear.
microRNAs (miRNAs), a small non-coding RNA containing 20–25 nucleotides, is closely associated with a variety of pathological or physiological processes.16,17 Mature miRNAs recognize the 3ʹ-untranslated region (3ʹ UTR) of targeted mRNAs (messenger RNAs) via complementary base pairing, and degrade the mRNAs or inhibit the translation of mRNAs by loading onto the RNA-induced silencing complex (RISC).18,19 miR-212-3p is significantly down-regulated in a variety of human cancers, including glioblastoma,20 gastric cancer,21 lung cancer,22 and bladder cancer.23 Besides, miR-212-3p induces apoptosis in a variety of tumor cells,22,23 thereby inhibiting tumor development. Previous research has reported that the dysregulation of miR-212-3p in gastric cancer tissues is significantly related to the patient’s prognosis, and affects the proliferative and migratory abilities of gastric cancer cells.24
In the present study, the findings suggested that Linc-ROR and miR-212-3p were abnormally expressed in gastric cancer tissues compared with the adjacent normal tissues. Besides, the high expression of Linc-ROR and low expression of miR-212-3p predicted a favorable prognosis of GC patients. Moreover, Linc-ROR/miR-212-3p axis modulated GC cell proliferation, migration, and invasion in vitro. Further, fibroblast growth factor 7 (FGF7) belonging to the fibroblast growth factor (FGF) family was identified as a downstream signaling molecule of the Linc-ROR/miR-212-3p axis. Overall, these findings reveal that the Linc-ROR/miR-212-3p/FGF7 axis is implicated in the progression of gastric cancer.
Materials and Methods
Human Gastric Cancer Tissues
Tumor tissues and paired normal samples were collected from 72 patients with gastric adenocarcinoma from May 2012 to April 2014 at the Department of Pathology, The First Affiliated Hospital of Xiamen University. Written informed consent was obtained from all the patients enrolled in this study. The clinicopathological factors of GC patients are listed in Table 1. All GC patients without other diseases were pathologically confirmed by two pathologists, and they had completed a 5-year follow-up after resection. The study was conducted in accordance with the Declaration of Helsinki, and was approved by the ethics committee of The First Affiliated Hospital of Xiamen University.
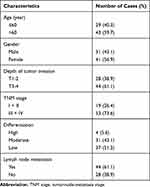 |
Table 1 The Clinicopathological Factors of Patients with Gastric Cancer |
Cell Culture, Virus Infection, and miR-212-3p Mimics or Antagomir Transfection
Human gastric adenocarcinoma cell lines AGS and MGC-803 were purchased from the Cell Bank of Shanghai Institutes for Biological Sciences, CAS (China). The cells were cultured in 89% DMEM (Gibco) (supplemented with 10% FBS (Gibco) and 1% streptomycin-penicillin (Gibco)) and incubated at 37°C with 5% CO2. Linc-ROR silencing vector possessing puromycin resistance was constructed by Genechem (Shanghai, China). Lentiviral particles were synthesized by co-transfection with lentiviral packaging plasmids and Linc-ROR silencing vector into 293T cells. 40 nM of miR-212-3p mimics or antagomir (Sangon Biotechnology Company, Shanghai, China) were transfected into the human gastric adenocarcinoma cells (5×105 cells/well in six-well plate) using Lipofectamine 2000 (Beijing Solarbio Science and Technology Co., Ltd., Beijing, China). The cells transfected with nonsense miRNA served as the control group.
RNA Isolation and Quantitative Real-Time PCR
TRIzol reagent (Life Technologies) was used to extract total RNA (including miRNA) according to the manufacturer’s instructions. Reverse transcription of mRNA was performed using EasyScript® First-Strand cDNA Synthesis SuperMix, while reverse transcription of miRNA was performed using TransScript® miRNA First-Strand cDNA Synthesis SuperMix (Transgen Biotechnology Inc., Beijing, China), respectively. The reaction procedures were as follows: 37°C for 5min; 42°C for 15min; 85°C for 5 seconds, and stored at 4°C. Subsequently, TransScript® One-Step RT-PCR SuperMix (Transgen Biotechnology) was used for qPCR. Sangon Biotechnology Company provided all the primer sequences. The PCR reaction procedures were as follows: 95°C for 50 s; 95°C for 15 s; 58.5°C for 35 s; and 40 cycles were performed. β-actin and U6 were used as internal controls for Linc-ROR and miR-212-3p expression, respectively. The experiment was repeated three times.
Transwell Migration and Invasion Assays
Cells at the logarithmic growth phase were prepared for transfection experiments. After transfection with miR-212-3p mimics or antagomir for 24 hours, cell migratory and invasive capacities were analyzed using transwell assay (BD Sciences) as described previously.25 Cell suspensions (5×104 cells/mL) were prepared using a serum-free medium. Into each well of the upper chamber, 100 μL of the cell suspension was added; the lower chamber contained 600 μL of the culture medium supplemented with 10% FBS. For transwell invasion assay, 100μL Matrigel (1mg/mL; BD Sciences) was added into the upper chamber and incubated at 37°C for 30 min. The cells were washed, seeded, and incubated at 37°C with 5% CO2 for 24 hours and the uninvaded cells swabbed. The cells in the lower chamber were fixed with 75% ethanol for 10 min and stained using 0.5% crystal violet (Solarbio) at room temperature for 20 minutes and observed under an optical microscope (Leica, Mannheim, Germany). Each experiment was repeated three times.
Wound Healing Assay
Cells at the logarithmic growth phase were digested and seeded into 6-well plates (1×106 cells/well). When the cells reached 90% confluence, they were subjected to 1 μg/mL of mitomycin for 1 hour to prevent the cells from proliferating. The culture medium was changed and a 200 μL pipette was used to evenly scratch the center of the 6-well plate to create a “wound”, and the images were captured at the beginning and after 24 h. The wound closure rate was quantified. The experiment was repeated three times.
Dual-Luciferase Reporter Assay
Dual-luciferase reporter assay was performed to determine the interaction between LincROR and miR-212-3p, miR-212-3p, and the 3ʹUTR of FGF7 mRNA. pmirGLO -Linc-ROR wild type (WT) plasmid (including the binding sites with miR-212-3p), pmirGLO -Linc-ROR mutant plasmid (including the mutated binding sites with miR-212-3p), pmirGLO -FGF7 3ʹUTR WT plasmid (including the binding sites with miR-212-3p) and pmirGLO-FGF7 3ʹUTR mutant plasmid (including the mutated binding sites with miR-212-3p) were constructed. The plasmids and microRNA mimics were co-transfected into 239T cells. The luciferase activity was monitored after 48 h. The pmirGLO plasmid encoding Renilla luciferase was used as the control. The experiment was repeated three times.
Cell Proliferation CCK-8 Assay
Cell proliferative ability was measured using the CCK-8 kit (Beijing Zoman Biotechnology Co., Ltd., Beijing, China) following the manufacturer’s instruction, 24 h after transfection. The cells were seeded into 96-well plates (1×104 cells/well) as described previously.26 After overnight acclimatization, 10% CCK-8 reagent was added into the cells and incubated for 2 h at 37°C. The absorbance at 450 nm was measured at various time points: 0-day, 1-day, 2 days, 3 days, 4 days, and 5 days. The experiment was repeated three times.
Western Blotting Analysis
Linc-ROR-silenced GC cells or xenograft tumor tissues were collected and lysed using RIPA lysis buffer (KenGen Biotech, China). Protein lysates were separated on 10% SDS-polyacrylamide gels, transferred onto PVDF membranes (Millipore), and blocked in 5% non-fat milk (Sangon Biotech, China), followed by primary antibodies (anti-KGF/FGF7: 1:2000, ab131162, Abcam; anti-GAPDH: 1:5000, ab9485, Abcam) incubation overnight at 4°C. Afterwards, the membranes were incubated with anti-rabbit HRP-conjugated secondary (1:5000, ab6721, Abcam) for 2 h at room temperature. Western blots were visualized using enhanced chemiluminescent reagent (Millipore, Billerica, MA) and quantified by Image J software (National Institutes of Health; http://imagej.nih.gov).
Animal Experiments
All animal experiments were performed under the Chinese animal welfare guidances and were approved by the Animal Research Ethics Committee of The First Affiliated Hospital of Xiamen University. Six-week-old female BALB/c nude mice weighing about 20 g were purchased from Shanghai Laboratory Animal Center of Chinese Academy Sciences (Shanghai, China), and maintained under specific pathogen-free condition. A total of 12 mice were used in this experiment (N=6 mice per group). To establish the tumor xenograft model, MGC-803 cells with or without Linc-ROR knockdown (2×106 tumor cells mixed with Matrigel) were injected subcutaneously into the flank of the nude mice. The growth of xenograft tumors in nude mice was monitored using caliper measurements every week for six weeks. The nude mice were sacrificed at 42 days and the tumor tissues excised immediately, measured, photographed, and stored at −80°C for subsequent use. Tumor graft volumes were calculated using the formula: Volume = (width2 × length)/2.
Statistical Analysis
Statistical analysis was performed using SPSS version 21.0. Kaplan-Meier method was used to draw survival curves for survival estimation. The correlation between the expression levels of Linc-ROR and miR-212-3p was analyzed by Pearson correlation. A comparison among the groups was analyzed using One-way ANOVA with LSD post hoc test. A comparison between groups was performed using the Student’s t-test. All experiments were performed in triplicate. Data were represented as mean ± standard deviation (S.D). P < 0.05, was considered statistically significant.
Results
Linc-ROR is Significantly Upregulated in Human Gastric Cancer Tissues and Linc-ROR Silencing Attenuated the Malignant Phenotypes of Gastric Cancer Cells in vitro and in vivo
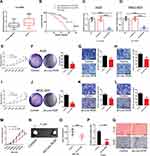
The expression level of Linc-ROR in GC and paired normal tissues was evaluated by qPCR. Linc-ROR was highly expressed in GC tissues compared with normal tissues (Figure 1A). And patients were sorted into a high Linc-ROR expression group and a low Linc-ROR expression group, according to the median Linc-ROR expression level (Median relative expression). The overall survival (OS) was significantly increased in patients with low expression of Linc-ROR (Figure 1B). Then, AGS and MGC-803 cell lines were infected with sh-Linc-ROR lentiviral vectors for 24 h and selected using puromycin (2 μg/mL). As shown in Figure 1C and D, the expression of Linc-ROR was significantly reduced in GC cell lines (AGS, MGS-803) infected with sh-Linc-ROR lentiviral vectors compared with the matched control cells. To further uncover the biological functions of Linc-ROR, cell proliferation in gastric cancer AGS and MGC-803 cells were analyzed using CCK-8 and colony formation assay. The migratory and invasive abilities of gastric cancer AGS and MGC-803 cells were detected by transwell assays. We found that Linc-ROR silencing suppressed the proliferative, migratory and invasive capabilities of gastric cancer AGS (Figure 1E–H) and MGC-803 (Figure 1I–L) cells compared with the matched control cells. The in vivo results (N=6 mice per group) further suggested that Linc-ROR knockdown significantly repressed GC tumor growth (Figure 1M–Q).
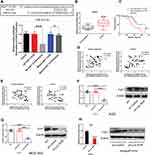
The Direct Interaction Between Linc-ROR and miR-212-3p is Determined and Linc-ROR Down-Regulates miR-212-3p Expression
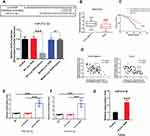
A search on LncBase (https://omictools.com/diana-lncbase-tool), the binding sites between Linc-ROR and miR-212-3p was predicted (Figure 2A). Further, dual-luciferase reporter assay revealed that the luciferase activity did not change significantly in 293T cells transfected with WT-Linc-ROR plasmids containing the miRNA binding sites compared to the cells transfected with control plasmids alone. However, the fluorescence activity of cells co-transfected with the WT-Linc-ROR plasmid containing the miRNA binding sites and miR-212-3p mimics was significantly attenuated compared to 293T cells infected with the WT-Linc-ROR plasmid. Meanwhile, after transfection with Mut-Linc-ROR plasmids containing mutant ROR (Mut-Linc ROR) sequences of the predicted miR-212-3p binding sites, firefly luciferase gene expression was not affected, which was consistent with the results in 293T cells transfected with WT-Linc-ROR plasmids. Moreover, co-transfection with miR-212-3p mimics and Mut-Linc ROR plasmids has no effect on firefly luciferase activity in 293T cells in comparison to the group transfected with WT-Linc-ROR plasmids. These findings suggested that miR-212-3p could directly bind to the WT Linc-ROR not the Mut Linc-ROR (Figure 2A). Additionally, miR-212-3p level was significantly lower in human GC tumor tissues than in adjacent normal tissues (Figure 2B). And the overall survival was greatly increased in GC patients with high expression of miR-212-3p (Figure 2C). Correlation analysis between Linc-ROR and miR-212-3p showed that miR-212-3p expression was negatively correlated with Linc-ROR expression levels in tumor tissues and adjacent normal tissues (Figure 2D). To further validate whether the Linc-ROR/miR-212-3p expression is the independent risk factor leading to poor overall survival in patients of GC, univariate and multivariate Cox proportional hazards analysis were performed (Table 2). Data showed that Linc-ROR and miR-212-3p expression levels significantly affected the overall survival in patients of GC. Besides, functional in vitro experiments demonstrated that Linc-ROR silencing led to a significant increase in miR-212-3p expression in AGS (Figure 2E) and MGC-803 (Figure 2F) cells compared to the matched control cells. In the tumor-bearing mouse model (N=6 mice per group), the expression of miR-212-3p was significantly up-regulated in Linc-ROR knockdown tumor tissues compared with the control group (Figure 2G).
 |
Table 2 Univariate and Multivariate Analysis of Clinic Pathologic Factors for Overall Survival in 72 GC Patients |
Linc-ROR Promotes Malignant Phenotypes of Gastric Cancer Cells by Sponging miR-212-3p
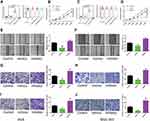
To further unveil how Linc-ROR exerts its tumor-promoting functions by acting as miR-212-3p sponge in GC cells, we performed a series of functional experiments and analysis. The in vitro results indicated that miR-212-3p mimics or inhibitors could significantly increase or decrease the level of miR-212-3p in GC cells. And miR-212-3p overexpression/silencing showed no effect on the expression of Linc-ROR in AGS (Figure 3A; right) and MGC-803 cells (Figure 3C; right). In addition, the overexpression of miR-212-3p weakened cell viability, whereas the silencing of miR-212-3p had the opposite effects on AGS (Figure 3B) and MGC-803 (Figure 3D) cells. The results further confirmed that miR-212-3p overexpression inhibited GC cell migration and invasion, while miR-212-3p silencing promoted GC cell migratory (Figure 3E–H) and invasive abilities (Figure 3I–J). Moreover, Linc-ROR silencing contributed to lower GC cell activity (Figure 4A and C), evidently attenuated ability of cell proliferation (Figure 4B and D), migration (Figure 4E–H) and invasion (Figure 4I–J) compared with the control cells. Notably, miR-212-3p mimics combined with Linc-ROR silencing synergistically inhibited the proliferation, migration, and invasion of GC cells; however, the miR-212-3p inhibitor could rescue the phenotypes caused by Linc-ROR silencing. These findings indicated that Linc-ROR promotes the proliferation, migration, and invasion of gastric cancer cells by acting as miR-212-3p sponge.
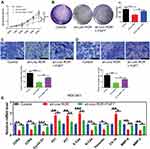
Fibroblast Growth Factor 7 (FGF7), a Downstream Molecule of miR-212-3p, is Overexpressed in Human GC Tissues
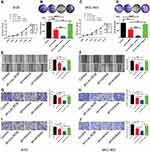
Fibroblast growth factor 7 (FGF7) was predicted to be a downstream target gene of miR-212-3p via microRNA database (http://mirdb.org/). Dual-luciferase reporter assay system showed that the fluorescence activity of cells co-transfected with the WT-3ʹUTR of FGF7 mRNA plasmid and miR-212-3p mimics was significantly attenuated compared to 293T cells infected with the control plasmid. Moreover, due to mutation of the binding sites, the fluorescence activity of cells co-transfected with Mutant-3ʹUTR of FGF7 mRNA and miR-212-3p mimics showed no significant differences, indicating a direct interaction between miR-212-3p and FGF7 (Figure 5A). Furthermore, we found that FGF7 was highly expressed in GC tissues compared with normal tissues (Figure 5B). High expression of FGF7 was associated with poor prognosis in patients with gastric cancer (Figure 5C). Correlation analysis showed that FGF7 expression was positively correlated with the expression level of Linc-ROR (Figure 5D); however, FGF7 expression was negatively correlated with miR-212-3p expression level in tumor tissues and adjacent normal tissues (Figure 5E). Linc-ROR silencing led to a significant decrease in FGF expression in AGS (Figure 5F) and MGC-803 (Figure 5G) cells compared to the matched control cells. In MGC-803 cell-bearing nude mouse tissues, FGF7 was significantly down-regulated in Linc-ROR knockdown tumor tissues compared with the control group (Figure 5H).
Additionally, Linc-ROR knockdown caused significant inhibition of GC cell activity (Figure 6A), proliferation (Figure 6B), migration (Figure 6C) and invasion (Figure 6D) compared with the control cells. In rescue experiments, the restored expression of FGF7 in Linc-ROR-silenced GC cells partially reversed Linc-ROR knockdown-induced phenotypes (Figure 6A–D). Consistent with the results observed in phenotypes, the expression levels of cell cycle-related genes (CDK4, CDK6, CyclinD1, P21, P27) and epithelial-mesenchymal transition (EMT) process-associated genes (E-Cadherin, N-Cadherin, Vimentin, CK-19, MMP-9, MMP-2) were changed significantly as well (Figure 6G). Collectively, our data uncovered that Linc-ROR promoted the malignant phenotypes of gastric cancer cells by sponging miR-212-3p, thereby promoting FGF7 expression.
Discussion
Mounting evidence has highlighted Linc-ROR as a potential oncogene in various types of cancer.14,15,27 In this study, the up-regulation of Linc-ROR was associated with the poor prognosis of GC patients. Silencing of Linc-ROR significantly suppressed the malignant phenotypes of gastric cancer cells by directly binding to miR-212-3p, which is a well-researched tumor suppressor miRNA in different types of human malignant tumors.20,23 Furthermore, fibroblast growth factor 7 (FGF7), a fibroblast growth factor (FGF) family member, was found to be a downstream signaling molecule of the Linc-ROR/miR-212-3p axis. Besides, Linc-ROR/miR-212-3p/FGF7 regulatory signaling was associated with the proliferation, migration, and invasion of GC cells.
Aberrant expression of Linc-ROR promotes tumor cell invasion.14,15,27 Besides, the aberrant expression of Linc-ROR in esophageal cancer contributes to cancer progression.27,28 In this study, high expression of Linc-ROR in tumor tissues from gastric cancer patients was reported compared with the normal tissues. The high expression of Linc-ROR was significantly associated with poor prognosis of GC patients, suggesting that Linc-ROR may be a potential predictive biomarker for gastric cancer. In vitro, Linc-ROR promoted the proliferation, migration, and invasion of gastric cancer cells, indicating that Linc-ROR is a multifunctional lncRNA in gastric cancer.
The miR-212-3p expression is significantly reduced in human cancers,29 and its down-regulation is associated with not only short-term treatment outcomes but also long-term survival.30,31 Studies have reported that miR-212-3p expression is a potential diagnostic and prognostic biomarker for gastric cancer.32 Consistently, the results in this study demonstrated that the down-regulation of miR-212-3p predicted poor survival of GC patients. Moreover, consistent with previous studies,23,29 the results in this study showed a decrease in the migratory and invasive abilities of gastric cancer cells overexpressing miR-212-3p. These findings confirm the antitumor effect of miR-212-3p in gastric cancer.
Previous studies have reported FGF7 as an oncogenic factor which enhances the malignant phenotype of tumors by binding FGF receptor 2 (FGFR2).33,34 It has been reported that FGF signaling could regulate epithelial-to-mesenchymal transition (EMT) process and cell invasion through PI3K-AKT pathway.35,36 In addition, work in GC cells has indicated that lncRNA AFAP1-AS1 promotes proliferation and metastasis via miR-155-5p/FGF7 regulatory axis.37 In this study, Linc-ROR was negatively correlated with miR-212-3p, while Linc-ROR was positively correlated with FGF7 expression in gastric tissues. Further, the dual-luciferase reporter assay confirmed that the direct interaction between Linc-ROR and miR-212-3p, miR-212-3p and FGF7 3ʹUTR mRNA. And Linc-ROR/miR-212-3p/FGF7 axis was implicated in regulating the proliferation, migration, and invasion of GC cells. However, the detailed mechanism needs further clarification, including the reason for the significant up-regulation of Linc-ROR in GC tissues.
Conclusions
In summary, Linc-ROR and FGF7 are significantly up-regulated, while miR-212-3p is down-regulated in gastric cancer tissues. High expression of Linc-ROR or FGF7 and low expression of miR-212-3p predicts poor prognosis of GC patients. Linc-ROR has a direct interaction with miR-212-3p, while miR-212-3p has direct interaction with FGF7. Moreover, Linc-ROR/miR-212-3p/FGF7 axis promotes the proliferative, migratory, and invasive capabilities of GC cells.
Abbreviations
GC, gastric cancer; FGF7, fibroblast growth factor 7; miRNAs, microRNAs; 3ʹ UTR, 3ʹ-untranslated region; RISC, RNA-induced silencing complex; qPCR, quantitative real-time polymerase chain reaction.
Data Sharing Statement
All data generated or analysed during this study are available from the corresponding author on reasonable request.
Ethics Approval and Informed Consent
Written informed consents were obtained from patients and the study was approved by the ethics committee of The First Affiliated Hospital of Xiamen University. All animal experiments were performed under the Chinese animal welfare guidances and were approved by the Animal Research Ethics Committee of The First Affiliated Hospital of Xiamen University.
Acknowledgments
We thank Dr Li Qinyuan (National Institute of Data Science in Health and Medicine, School of Medicine, Xiamen University) for his help with this study.
Author Contributions
All authors made a significant contribution to the work reported, whether that is in the conception, study design, execution, acquisition of data, analysis and interpretation, or in all these areas; took part in drafting, revising or critically reviewing the article; gave final approval of the version to be published; have agreed on the journal to which the article has been submitted; and agree to be accountable for all aspects of the work.
Funding
This work is financed by Wu Jieping Medical Foundation (Grant Number 320.6750.17509) and Youth Foundation of Xiamen Cancer Center (Grant Number ZLYYA201703).
Disclosure
The authors declare that there are no conflicts of interest.
References
1. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2019. CA Cancer J Clin. 2019;69(1):7–34. doi:10.3322/caac.21551
2. Miller KD, Nogueira L, Mariotto AB, et al. Cancer treatment and survivorship statistics, 2019. CA Cancer J Clin. 2019;69(5):363–385. doi:10.3322/caac.21565
3. Karimi P, Islami F, Anandasabapathy S, Freedman ND, Kamangar F. Gastric cancer: descriptive epidemiology, risk factors, screening, and prevention. Cancer Epidemiol Biomarkers Prev. 2014;23(5):700–713. doi:10.1158/1055-9965.EPI-13-1057
4. Obermannova R, Lordick F. Management of metastatic gastric cancer. Hematol Oncol Clin North Am. 2017;31(3):469–483. doi:10.1016/j.hoc.2017.01.006
5. Yan Y, Wang LF, Wang RF. Role of cancer-associated fibroblasts in invasion and metastasis of gastric cancer. World J Gastroenterol. 2015;21(33):9717–9726. doi:10.3748/wjg.v21.i33.9717
6. Abbas M, Habib M, Naveed M, et al. The relevance of gastric cancer biomarkers in prognosis and pre- and post-chemotherapy in clinical practice. Biomed Pharmacother. 2017;95:1082–1090. doi:10.1016/j.biopha.2017.09.032
7. Chen LT, Oh DY, Ryu MH, et al. Anti-angiogenic therapy in patients with advanced gastric and gastroesophageal junction cancer: a systematic review. Cancer Res Treat. 2017;49(4):851–868. doi:10.4143/crt.2016.176
8. Cavatorta O, Scida S, Miraglia C, et al. Epidemiology of gastric cancer and risk factors. Acta bio-medica: Atenei Parmensis. 2018;89(8–s):82–87. doi:10.23750/abm.v89i8-S.7966
9. Kashihara H, Shimada M, Yoshikawa K, et al. Risk factors for recurrence of gastric cancer after curative laparoscopic gastrectomy. J Med Invest. 2017;64(1.2):79–84. doi:10.2152/jmi.64.79
10. Choi YJ, Kim N. Gastric cancer and family history. Korean J Intern Med. 2016;31(6):1042–1053. doi:10.3904/kjim.2016.147
11. Li W, Xu C, Guo J, et al. Cis- and trans-acting expression quantitative trait loci of long non-coding RNA in 2,549 cancers with potential clinical and therapeutic implications. Front Oncol. 2020;10:602104. doi:10.3389/fonc.2020.602104
12. Bhan A, Soleimani M, Mandal SS. Long noncoding RNA and cancer: a new paradigm. Cancer Res. 2017;77(15):3965–3981. doi:10.1158/0008-5472.CAN-16-2634
13. Wang Y, Xu Z, Jiang J, et al. Endogenous miRNA sponge lincRNA-RoR regulates Oct4, Nanog, and Sox2 in human embryonic stem cell self-renewal. Dev Cell. 2013;25(1):69–80. doi:10.1016/j.devcel.2013.03.002
14. Chen W, Wang H, Liu Y, et al. Linc-RoR promotes proliferation, migration, and invasion via the Hippo/YAP pathway in pancreatic cancer cells. J Cell Biochem. 2020;121(1):632–641. doi:10.1002/jcb.29308
15. Huang J, Zhang A, Ho TT, et al. Linc-RoR promotes c-Myc expression through hnRNP I and AUF1. Nucleic Acids Res. 2016;44(7):3059–3069. doi:10.1093/nar/gkv1353
16. Deng K, Wang H, Guo X, Xia J. The cross talk between long, non-coding RNAs and microRNAs in gastric cancer. Acta Biochim Biophys Sin (Shanghai). 2016;48(2):111–116. doi:10.1093/abbs/gmv120
17. Hayes J, Peruzzi PP, Lawler S. MicroRNAs in cancer: biomarkers, functions and therapy. Trends Mol Med. 2014;20(8):460–469. doi:10.1016/j.molmed.2014.06.005
18. Jonas S, Izaurralde E. Towards a molecular understanding of microRNA-mediated gene silencing. Nat Rev Genet. 2015;16(7):421–433. doi:10.1038/nrg3965
19. Huntzinger E, Izaurralde E. Gene silencing by microRNAs: contributions of translational repression and mRNA decay. Nat Rev Genet. 2011;12(2):99–110. doi:10.1038/nrg2936
20. Wang D, Zheng J, Liu X, et al. Knockdown of USF1 inhibits the vasculogenic mimicry of glioma cells via stimulating SNHG16/miR-212-3p and linc00667/miR-429 axis. Mol Ther Nucleic Acids. 2019;14:465–482. doi:10.1016/j.omtn.2018.12.017
21. Wu R, Li F, Zhu J, et al. A functional variant at miR-132-3p, miR-212-3p, and miR-361-5p binding site in CD80 gene alters susceptibility to gastric cancer in a Chinese Han population. Med Oncol. 2014;31(8):60. doi:10.1007/s12032-014-0060-2
22. Qiu HB, Yang K, Yu HY, Liu M. Downregulation of long non-coding RNA XIST inhibits cell proliferation, migration, invasion and EMT by regulating miR-212-3p/CBLL1 axis in non-small cell lung cancer cells. Eur Rev Med Pharmacol Sci. 2019;23(19):8391–8402. doi:10.26355/eurrev_201910_19150
23. Wu X, Chen H, Zhang G, et al. MiR-212-3p inhibits cell proliferation and promotes apoptosis by targeting nuclear factor IA in bladder cancer. J Biosci. 2019;44:4. doi:10.1007/s12038-019-9903-5
24. Dang Y, Liu T, Yan J, et al. Gastric cancer proliferation and invasion is reduced by macrocalyxin C via activation of the miR-212-3p/Sox6 pathway. Cell Signal. 2019;66:109430. doi:10.1016/j.cellsig.2019.109430
25. Wang X, Ye T, Xue B, et al. Mitochondrial GRIM-19 deficiency facilitates gastric cancer metastasis through oncogenic ROS-NRF2-HO-1 axis via a NRF2-HO-1 loop. Gastric Cancer. 2020.
26. Ye T, Yang M, Huang D, et al. MicroRNA-7 as a potential therapeutic target for aberrant NF-κB-driven distant metastasis of gastric cancer. J Exp Clin Cancer Res. 2019;38(1):55. doi:10.1186/s13046-019-1074-6
27. Wang L, Yu X, Zhang Z, et al. Linc-ROR promotes esophageal squamous cell carcinoma progression through the derepression of SOX9. J Exp Clin Cancer Res. 2017;36(1):182. doi:10.1186/s13046-017-0658-2
28. Sahebi R, Malakootian M, Balalaee B, et al. Linc-ROR and its spliced variants 2 and 4 are significantly up-regulated in esophageal squamous cell carcinoma. Iran J Basic Med Sci. 2016;19(10):1131–1135.
29. Liu H, Li C, Shen C, et al. MiR-212-3p inhibits glioblastoma cell proliferation by targeting SGK3. J Neurooncol. 2015;122(3):431–439. doi:10.1007/s11060-015-1736-y
30. Hou P, Kang Y, Luo J. Hypoxia-mediated miR-212-3p downregulation enhances progression of intrahepatic cholangiocarcinoma through upregulation of Rab1a. Cancer Biol Ther. 2018;19(11):984–993. doi:10.1080/15384047.2018.1456608
31. Lu X, Wang F, Fu M, Li Y, Wang L. Long non-coding RNA KCNQ1OT1 accelerates the progression of ovarian cancer via microRNA-212-3/LCN2 axis. Oncol Res. 2019. doi:10.3727/096504019X15719983040135
32. He L, Qu L, Wei L, Chen Y, Suo J. Reduction of miR1323p contributes to gastric cancer proliferation by targeting MUC13. Mol Med Rep. 2017;15(5):3055–3061. doi:10.3892/mmr.2017.6347
33. Fan EW, Li CC, Wu WJ, et al. FGF7 over expression is an independent prognosticator in patients with urothelial carcinoma of the upper urinary tract and bladder. J Urol. 2015;194(1):223–229. doi:10.1016/j.juro.2015.01.073
34. Huang T, Wang L, Liu D, et al. FGF7/FGFR2 signal promotes invasion and migration in human gastric cancer through upregulation of thrombospondin-1. Int J Oncol. 2017;50(5):1501–1512. doi:10.3892/ijo.2017.3927
35. Katoh M, Nakagama H. FGF receptors: cancer biology and therapeutics. Med Res Rev. 2014;34(2):280–300. doi:10.1002/med.21288
36. Inan S, Hayran M. Cell signaling pathways related to epithelial mesenchymal transition in cancer metastasis. Crit Rev Oncog. 2019;24(1):47–54. doi:10.1615/CritRevOncog.2018029509
37. Ma HW, Xi DY, Ma JZ, et al. Long noncoding RNA AFAP1-AS1 promotes cell proliferation and metastasis via the miR-155-5p/FGF7 axis and predicts poor prognosis in gastric cancer. Dis Markers. 2020;2020:8140989. doi:10.1155/2020/8140989
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.