Back to Journals » Pharmacogenomics and Personalized Medicine » Volume 12
Toxicogenomic analysis of publicly available transcriptomic data can predict food, drugs, and chemical-induced asthma
Authors Hachim MY , Hachim IY, Elemam NM , Hamoudi RA
Received 29 May 2019
Accepted for publication 30 July 2019
Published 26 August 2019 Volume 2019:12 Pages 181—199
DOI https://doi.org/10.2147/PGPM.S217535
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Martin H Bluth
Mahmood Yaseen Hachim,1 Ibrahim Yaseen Hachim,2 Noha M Elemam,1 Rifat A Hamoudi1–3
1Sharjah Institute for Medical Research, University of Sharjah, Sharjah 27272, United Arab Emirates; 2Department of Clinical Sciences, College of Medicine, University of Sharjah, Sharjah 27272, United Arab Emirates; 3Division of Surgery and Interventional Science, University College London, London, UK
Correspondence: Mahmood Yaseen Hachim
Sharjah Institute for Medical Research, University of Sharjah, Sharjah 27272, United Arab Emirates
Tel +971 50 945 2674
Email [email protected]
Rifat A Hamoudi
Department of Clinical Sciences, College of Medicine, University of Sharjah, Sharjah 27272, United Arab Emirates
Tel +971 56 715 4756
Email [email protected]
Background: With the increasing incidence of asthma, more attention is focused on the diverse and complex nutritional and environmental triggers of asthma exacerbations. Currently, there are no established risk assessment tools to evaluate asthma triggering potentials of most of the nutritional and environmental triggers encountered by asthmatic patients.
Purpose: The objective of this study is to devise a reliable workflow, capable of estimating the toxicogenomic effect of such factors on key player genes in asthma pathogenesis.
Methods: Gene expression extracted from publicly available datasets of asthmatic bronchial epithelium were subjected to a comprehensive analysis of differential gene expression to identify significant genes involved in asthma development and progression. The identified genes were subjected to Gene Set Enrichment Analysis using a total of 31,826 gene sets related to chemical, toxins, and drugs to identify common agents that share similar asthma-related targets genes and signaling pathways.
Results: Our analysis identified 225 differentially expressed genes between severe asthmatic and healthy bronchial epithelium. Gene Set Enrichment Analysis of the identified genes showed that they are involved in response to toxic substances and organic cyclic compounds and are targeted by 41 specific diets, plants products, and plants related toxins (eg adenine, arachidonic acid, baicalein, caffeic acid, corilagin, curcumin, ellagic acid, luteolin, microcystin-RR, phytoestrogens, protoporphyrin IX, purpurogallin, rottlerin, and salazinic acid). Moreover, the identified chemicals share interesting inflammation-related pathways like NF-κB.
Conclusion: Our analysis was able to explain and predict the toxicity in terms of stimulating the differentially expressed genes between severe asthmatic and healthy epithelium. Such an approach can pave the way to generate a cost-effective and reliable source for asthma-specific toxigenic reports thus allowing the asthmatic patients, physicians, and medical researchers to be aware of the potential triggering factors with fatal consequences.
Keywords: toxicogenomic, transcriptomic, GSEA, chemical-induced asthma
Introduction
It is widely accepted that asthma has a multifactorial etiology, where genetic predisposition plays an essential role in disease susceptibility,1 while environmental factors play a critical role in disease development and progression.2 Due to the rise in the incidence of asthma, there is a growing concern over the environmental exposures that may trigger asthma exacerbations.3 Although many theories were suggested regarding how exposure to drugs, toxins, chemicals, and infections can participate in asthma development and/or exacerbation, the exact mechanism is still not fully understood.4
Asthma was linked to food allergy as children with food allergies have a higher risk of developing food-induced episodes of asthma that can end up with anaphylaxis; nevertheless, this link is not fully understood yet.5 Changes in dietary habits were suggested as a possible cause of increased asthma prevalence6 in developed and developing countries. Besides food, air pollution can adversely influence lung function in asthmatic individuals,7 but which particles in the air can precisely trigger such an effect is still a matter of debate between researchers except for few well-studied examples.8 Exposure to chemicals at work is a significant risk factor for occupational asthma and should be brought to the attention and awareness of every asthmatic patient.9 Occupational asthma should be distinguished from the non-immunologic asthma-like syndrome10 called Reactive Airways Dysfunction Syndrome (RADS), which develops after a single high-level exposure to a pulmonary irritant.11 Many substances used in consumer products are associated with occupational asthma or asthma-like syndromes.12 Besides occupation-induced asthma, common household chemicals can be another uncountable trigger for asthma in adults.13 Drug-induced asthma, especially aspirin-induced asthma, is well-defined, relatively common, and often an underdiagnosed asthma phenotype.14,15
Currently, there are no ideal asthma risk assessment tools for food, drugs, occupational, and household chemicals. Moreover, there is no means of prediction of potential respiratory sensitization for all possible food or environmental items that we encounter in our daily life.16 Only a few of these tools are available in the clinical setting, with a limited list of items.17 Recently, toxicogenomic investigation of different toxic agents’ interaction with the cellular genome improved our understanding of the effect of different chemicals, hazardous agents, drugs, and environmental stressors on different cellular and biological systems. Through multi-omics analysis, the response of all genes to chemical exposure can be examined in order to gain a more comprehensive insight into the potential hazards of that toxicant.18 Although toxicogenomics was proposed to be a useful tool in health risk assessment,18 this approach has not been tried yet for asthma triggers’ assessment. Since the bronchial epithelium is the key player in asthma initiation and progression that orchestrates airway inflammation and remodeling, toxicogenomic analysis of bronchial epithelium in asthma is mandated.19
In this study, we used an in-house bioinformatics pipeline that has shown a remarkable performance in clustering complex diseases previously using publicly available omics data.20 We aimed at identifying the effect of dietary, environmental, and occupational influences on genes that are differentially expressed between healthy and severe asthmatic bronchial epithelium. Therefore, this appraoch can facilitate the development of a comprehensive toxicogenomic database that can link and predict asthma susceptibility or progression in response to a given chemical.
Materials and methods
Bioinformatics approach: microarray analysis
To identify differentially expressed genes in asthmatic patients’ bronchial epithelium (in both small and large airways) compared to healthy controls, publicly available transcriptomic datasets from Gene Expression Omnibus (GEO) (https://www.ncbi.nlm.nih.gov/geo/) were extracted. We decided to use dataset (GSE64913) due to its appropriate design, a complete characterization and proper categorization of patients as well as being a representative of the two extremes of the disease (healthy versus severe asthma). The study was done using Affymetrix Human Genome U133 Plus 2.0 Array, which has the advantages of complete coverage of over 53,000 transcripts for analysis. Additionally, this dataset shows the effect of sampling of the bronchial tree as it included central and peripheral airway samples from each participant. Accordingly, we hypothesized that genes that are differentially expressed between severe asthmatic and healthy bronchial epithelium in both central and peripheral airways must have a role in the initiation or progression of the disease.
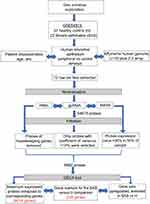
We used a novel in-house R Bioconductor based pipeline as described previously by Hamoudi et al.20 The pipeline is composed of 5 steps: (1) preprocessing and QC assessment of the downloaded raw microarray image files, (2) normalization to remove background noise and (3) filtration of nonvariant probes between severe asthmatics and healthy controls to (4) precisely identify differentially expressed genes (DEG). Finally, the DEG between the two groups will be used for (5) Gene Set Enrichment Analysis (GSEA) to identify top pathways where the identified genes are enriched. Such an approach will give us a clear list of genes that may participate in the pathogenesis of severe asthma. Figures 1 and 2 outline the pipeline steps used in this study.
 |
Figure 2 The flowchart of the bioinformatics approach to identify gene sets related to chemical, toxins, and drugs. |
Raw microarray image processing and normalization
Raw CEL files (n=70) that stores the results of the intensity calculations on the pixel values were extracted, then the dataset underwent pre-processing and normalization separately. Affy, Robust Multiarray Averaging (RMA), GeneChip RMA (gcRMA), Affymetrix Microarray Suite 5 (MAS5) packages of R Bioconductor statistical software version 3.0.2 were applied to normalize and remove the background noise. gcRMA and MAS5 expression values were used for the next non-specific filtering based on the coefficient of variation (CV). The CV was calculated as the mean/standard deviation of each probe across all cases.
Non-specific filtration
To filter out non-variant genes, only probes with a MAS5 value of 50 or more and CV value of 10–100% in the gcRMA across all cases, were passed and intersected to obtain a common set of variant probes. Out of the 54,675 probes present in the chip, only 9682 probes passed the filtration process. These filtered probes were annotated, collapsed to their corresponding genes using GSEA software (http://software.broadinstitute.org/gsea/downloads.jsp) by choosing probes with the maximum expression for each gene.21 The housekeeping probes, along with those that are not assigned to a gene, were excluded. Hence the resultant filtered probes were the only variant probes as per the GSEA manual.
Limma package to identify DEG
R Bioconductor Limma package was used to identify DEG between severe asthma and healthy controls. Out of the 6014 filtered genes, 225 genes with an adjusted p-value less than 0.05 were identified to be differentially expressed between severe asthma and healthy controls. To visualize top pathways and biological processes shared by the DEG gene list, a simplified and customizable web portal (http://www.metascape.org) was used.22 The gene list enrichment analysis was carried out with the following ontology sources: KEGG Pathway, GO Biological Processes, Reactome Gene Sets.
GSEA
The resultant 6014 filtered gene list was used as input for the GSEA to identify the significantly enriched pathways among gene sets related to chemical, toxins, and drugs, as shown in Figure 2. 31,826 gene sets, were downloaded from two major resources : DSigDB (http://tanlab.ucdenver.edu/DSigDB/DSigDBv1.0/) and DrugMatrix (ftp://anonftp.niehs.nih.gov/ntp-cebs/datatype/Drug_Matrix/) databases. DSigDB organizes drugs and small molecules-related gene sets into four collections based on quantitative inhibition, and drug-induced gene expression changes data.23 The DrugMatrix database is one of the world’s most massive toxicogenomic reference resources. Table 1 shows the details of the gene sets, the gene coverage, and the number of sets included in each set. The results of GSEA were ranked according to the nominal p-value (<0.05) and false discovery rate (≤0.25) as described previously.24
 |
Table 1 Details of datasets extracted from DSigDB and DrugMatrix and used for GSEA |
Cell of origin
ARCHS4 is a web resource that makes the majority of published RNA-sequencing data from human and mouse available at the gene and transcript levels. This resource was used to determine which cell type or tissue can express the genes that are differentially expressed between severe asthmatic and healthy bronchial epithelium and are enriched in a given gene set.
Finding a common pathway between identified chemicals
In order to identify common pathways targeted by most of the identified chemicals in the GSEA step, we used the Comparative Toxicogenomics Database (CTD) batch query webtool (http://ctdbase.org/tools/batchQuery).25 All the earlier identified drugs and chemicals were uploaded to the query tool to search for genes and pathways that were reported to be affected by the queried chemicals. The tool will generate a list of pathways where the given chemical affects genes related to that pathway significantly (adjusted p-value <0.05). Only pathways that are shared by at least 50 percent and above of the identified chemicals are selected. As illustrated in Figure 3, a schematic flowchart of this step is outlined.
 |
Figure 3 The flowchart outline using the Comparative Toxicogenomics Database (CTD) batch query tool (http://ctdbase.org/tools/batchQuery) to identify common pathways targeted by most of the GSEA-identified chemicals. (A) All the earlier identified drugs and chemicals were uploaded to the query tool to search for genes and (B) pathways that were documented to be affected by queried chemicals. The tool will generate (C) a list of pathways where the given chemical affects genes related to that pathway significantly (adjusted p-value <0.05). (D) Only pathways that are shared by at least 50 percent and above of the identified chemicals are selected. |
Results & discussion
Transcriptomic analysis reveals significant enrichment of genes related to cell division between asthmatic and healthy bronchial epithelial cells
Our analysis identified 225 differentially expressed genes between severe asthmatic bronchial epithelium and healthy bronchial epithelium, as shown in Figure 4A and B. Furthermore, the identified genes shared common pathways related to epithelial cell differentiation, response to growth factors, extracellular stimulus, mechanical stimulus, and wounding (Figure 4C). Interestingly, pathways related to the response to toxic substances and organic cyclic compounds were among the top enriched pathways. These findings indicate that genes altered by environmental substances might play a significant role in asthma development and/or progression to severe asthma.
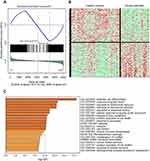 |
Figure 4 Gene Set Enrichment Analysis (GSEA) of the differentially expressed genes between severe asthmatic bronchial epithelium (n=22) and healthy bronchial epithelium (n=37) in GSE64913. (A) Distribution of the identified genes ranked according to their position (B) Heatmap image generated from the 2952 DEG between severe asthma and healthy controls which were later filtered into 225 genes (C) the top enriched pathways whether upregulated or downregulated in severe asthma compared to healthy controls using metascape (http://metascape.org): a gene annotation and analysis online resource that generates a graphical representation. |
Genes that are differentially expressed in the asthmatic bronchial epithelium are targeted by specific diets, plants products, and plants related toxins
Our further analysis revealed that the significant differentially expressed genes in asthmatic epithelium compared to healthy controls are targets for many substances that have not been previously associated or documented to trigger asthma, as shown in Table 2. Additionally as shown in Table 3, these substances can be categorized into three subgroups: (1) Occupational hazards, (2) Drugs, (3) Dietary factors: plant, plant toxins and food. This is substantial as most of the asthmatic individuals are not explicitly aware that such factors might have a potential effect on their disease status.
 |
Table 3 The top enriched chemicals by GSEA categorized into different subgroups |
The identified chemicals share exciting immune/inflammation-related pathways
In order to examine which pathways are associated with the largest number of the identified 41 chemicals, we used the Comparative Toxicogenomics Database (CTD) batch query webtool (http://ctdbase.org/tools/batchQuery). The tool can generate a report listing the pathways that show significant association with the given chemical, thus having a potentially significant effect on a proportion of the genes of that pathway. More than 70% of the 41 identified chemicals are associated with common pathways mainly involved in the immune response, as shown in Table 4. Those pathways are: Immune system, Cytokine signaling in immune system, IL-17 signaling pathway, Pathways in cancer, Signaling by interleukins, Innate immune system, Apoptosis, TNF signaling pathway, Cellular responses to stress, Toll-like receptor signaling pathway, Influenza A, Adaptive immune system, Downstream signaling events of B Cell Receptor (BCR), Senescence-Associated Secretory Phenotype (SASP), Fc epsilon receptor (FCERI) signaling, Signaling by EGFR, Th17 cell differentiation, Toll-Like receptors cascades, Cellular senescence, Interleukin-10 signaling, Activated TLR4 signaling.
 |
Table 4 List of pathways significantly associated with the largest number of the identified 41 chemicals using Comparative Toxicogenomics Database (CTD) batch query webtool (http://ctdbase.org/tools/batchQuery). Only pathways that are shared by more than 50% of the identified chemicals were listed |
 |
Table 4 (Continued). |
 |
Table 4 (Continued). |
22 of the identified chemicals are enriched for NF-kB pathways
Notably, 22 out of the total 41 identified chemicals showed significant enrichment for the NF-κB pathway and those were ellagic acid, baicalein, curcumin, SB 202190, acrylamide, myricetin, arachidonic acid, luteolin, ammonium hexachloroplatinate(iv), bisindolylmaleimide I, 67526-95-8, vinblastine, thimerosal, MG-132, thalidomide, adenine, caffeic acid, dronabinol, rottlerin, rapamycin, gefitinib and acrolein.
The transcription factor NF-κB regulates innate and adaptive immune functions through upregulation of pro-inflammatory genes, and when deregulated, it can contribute to the pathogenic processes of various inflammatory diseases.25 NF-κB was shown to be activated predominantly in the epithelial cells of the conducting airways, which have been reported to be the main source of NF-κB-dependent mediators that play a role in asthma.26 Any inhalational stimuli can activate bronchial epithelial NF-κB pathway sufficiently to promote allergic sensitization to innocuous inhaled antigens.27 The mainstay of therapy for asthma is the anti-inflammatory glucocorticoids that act mainly by inhibiting NF-κB induced gene transcription.28 These reports indicate the central role of the NF-κB pathway in asthma pathogenesis and hence propose it as an important therapeutic target.29
Plants, plant toxins and food-related asthma triggers
In this study, we focused on the identified plants, plant toxins, and food-related chemicals. This is due to the fact that there is no specific risk assessment for their potential effect on asthma development or exacerbation, even though asthmatic patients are in contact with one or more of these triggers in their close environment. Drugs, chemotherapy, chemical compounds, and occupational hazards are usually associated with a specific warning and awareness regarding asthma, although the exact underlying mechanisms are not fully understood.
Food
DNA, present in food, can survive harsh processing and be absorbed to circulate through the blood to other tissues of human and animals.30 Dietary purines like adenine are found in virtually all foods.31 Adenine and guanine comprise more than 60% of total purine-rich foods (such as cereals, beans, soybean products, and seaweeds),31 with greater bioavailability of adenine than guanine.32 There are many circumstantial pieces of evidence that purine and its metabolites might have a role in asthma with no conclusive findings. Allergic asthmatic plasma metabolomics showed aberrant purine metabolism that may change the consequence of having a more purine-rich diet in such patients.33 It has been previously reported that allergy to purine-rich wheat flour is the leading cause of serious occupational asthma among bakery workers called baker’s asthma.34 Another possible indirect link between purine-rich diet and asthma is through gout, sleep apnea, and circadian rhythm. A purine-rich diet is associated with a high risk of gout,35 which in turn is linked to sleep apnea.36 Unrecognized obstructive sleep apnea (OSA) can potentiate poor asthma control despite optimal therapy.37 The endogenous circadian system prolongs respiratory events across the night and can modulate sleep apnea.38 Circadian regulation of de novo purine synthesis is an important mechanism conferring circadian rhythmicity on the cell cycle.39 Intriguingly, our results showed that genes affected by dietary adenine were differentially expressed in severe asthmatic bronchial epithelium. Most of these genes are lung epithelial cell tissue-specific genes (BLM; CHAF1A; GDF15; KIF20A; CDC25A). Of interest, one of these genes (PRKAB1) is involved in circadian rhythmicity.40 Furthermore, one of the characteristics of asthma is worsening of symptoms overnight that has been linked to circadian variations controlled by clock genes.41 Therefore, activation of circadian rhythm genes by purine-rich food can be the link between gout, sleep apnea, and asthma.38
Another interesting food-related chemical identified by our method is Arachidonic acid(ARA), an omega-6 polyunsaturated fatty acid found in the phospholipids of the cell membranes and is abundant in the brain, muscles, and liver.42 Arachidonic acid occurs in the animal source diet such as eggs, poultry, and meat.43 This could explain and suggest a possible contributing factor to the increased incidence of asthma in western societies due to their consumption of such a pro-inflammatory diet and thus promoting the release of pro-inflammatory arachidonic acid metabolites (leukotrienes and prostanoids).44 Prostaglandins and leukotrienes are arachidonic acid-derived lipid mediators converted via cyclooxygenase and lipoxygenase, respectively and play a major role in asthma.45 However, there are insufficient studies to draw any firm conclusions about the relationship between ARA and asthma risk.46 Surprisingly, our results showed that genes targeted by arachidonic acid are specific to alveolar macrophages (ABCA1; JUN; SERPINB2; HPGD; IL1B; PLA2G4A; PPARG; FOS; PTGS2; PHLDA1; PLA2G7; ATF3). Alveolar macrophages serve as the first line of defense against foreign invaders to the lung tissue and have a critical role in asthma.47 Unlike blood-borne monocytes, resident alveolar macrophages have a suppressive role to inflammation but could gain pathogenic functions after repeated exposures.48
Strawberries are considered as functional food and nutraceutical source, mainly because of their high concentration of ellagic acid (EA) and its precursors.49 EA is derived from ellagitannins (ETs) and is found in some nuts, seeds, and fruits, especially berries and fruit juices.50 Dietary ETs are partially hydrolyzed in the gut to EA then to urolithin A (UA) by colonic microflora to enter the circulation.51 ETs are natural polyphenolic compounds that show potent anti-inflammatory properties in various diseases such as that observed in OVA-induced asthma mouse model, possibly through inhibition of NF-κB activation.52 Furthermore, EA has an anti-eosinophilic activity in a murine model of asthma53 and was suggested as a potential therapeutic agent for accelerating the resolution of allergic airways inflammation.54
Another identified food component is the flavone luteolin, which is found in several plant products, including broccoli, pepper, thyme, and celery. Due to its anti-inflammatory and neuroprotective function, plants rich in luteolin have been used in Chinese traditional medicine for treating various diseases such as hypertension, inflammatory disorders, and cancer.55 Through intrinsic and extrinsic signaling pathways, luteolin as an active compound showed anti-oxidant, anti-tumor, anti-inflammatory, and anti-apoptotic activities.56 Our results showed that targeted genes by luteolin in asthmatic epithelium are related to inflammation pathways like TNF signaling pathway (NFKBIA; JUN; IL1B; FOS; PTGS2; JUNB), Th17 cell differentiation and IL-17 signaling pathway (NFKBIA; JUN; IL1B; FOS, FOSB; PTGS2), Arachidonic acid metabolism (PTGS2; CBR3), Toll-like receptor signaling pathway (NFKBIA;JUN; IL1B; FOS) and NF-κB signaling pathway (NFKBIA; GADD45B; IL1B; PTGS2).
Caffeic acid is an active anti-oxidative component, that has been shown to have beneficial effects on several respiratory disorders, such as chronic obstructive pulmonary disease and lung cancer.57 Caffeic acid has powerful antimicrobial, antioxidant activities, and can influence collagen production and block premature aging.58 It was shown previously that caffeic acid has immunoregulatory effects by inhibition of cytokine and chemokine production as well as enhancement of transforming growth factor-beta 1 production in asthmatics.59,60 Of interest, another enriched pathway between severe asthmatic and healthy epithelium and related to coffee is Pyrogallol which is converted under alkaline conditions into purpurogallin,61 generating reactive oxygen species. Another source for purpurogallin is Galls, the abnormal growth in plants. Galls are induced by viruses, bacteria, fungi, nematodes, arthropods, or even other plants, which are similar to cancers in fauna and used in folkloric medicine.62 Purpurogallin was shown to exert antiplatelet, antithrombotic63 and anti‑inflammatory effects by inhibiting NF‑κB and MAPK signaling pathways in lipopolysaccharide‑stimulated cells.64 Due to these anti-inflammatory activities, it was suggested to be a therapeutic target for various systemic inflammatory diseases.65 Our results showed that genes affected by purpurogallin and upregulated in severe asthmatic epithelium (TNNI3, HPGD, UHRF1, TNNC1, SELL, BLM, GFER, PLA2G7, TDP1, CDK5, SENP8, MCL1, GAPDH, RNASEH1, GSK3A, RUNX1, PABPC1, BCL2L1) are related to positive regulation of apoptotic process, leukocyte cell-cell adhesion and DNA repair.
Protoporphyrin IX (PPIX) is a heterocyclic organic compound, which consists of four pyrrole rings, and is the final intermediate in the heme biosynthetic pathway. It is ubiquitously present in all living cells in small amounts.66 PPIX is a naturally occurring pigment in meat products that is increased by higher pH conditions in the context of nitrite reduction.67 Also, PPIX is the main pigment resulting in the brown coloration of eggshells.68 PPIX can induce heme oxygenase which was shown to inhibit Th17 cell-mediated immune response and prevent ovalbumin-induced neutrophilic airway inflammation.69
Plants
Baicalein (5,6,7 trihydroxyflavone) is a famous phenolic flavonoid present in the dry roots of Scutellaria baicalensis plant and is a component of the traditional herbal remedy known as Chinese skullcap (or Huang Qin).70 It was shown to attenuate inflammatory responses by suppressing TLR4 mediated NF-κB and MAPK signaling pathways.71 Furthermore, baicalein protects cells from hydrogen peroxide by inhibiting 12-lipoxygenase thus blocking the increase in ROS levels.72 Our results have shown that genes (HPGD, SELL, BLM, CYP2D6, PLA2G7, TDP1, GAPDH) that are affected by baicalein are significantly enriched in asthmatic bronchial epithelium. HPGD (15-Hydroxyprostaglandin Dehydrogenase) gene contributes to the regulation of events that are under the control of prostaglandin levels, and its expression is affected by aspirin.73 This can be part of the Aspirin-Exacerbated Respiratory Disease (AERD) which is a syndrome that includes asthma, recurrent nasal polyps, and pathognomonic reactions to aspirin and other nonselective cyclooxygenase inhibitors.74 On the other side, SELL (selectin S) gene was previously shown to be upregulated in different lung inflammatory diseases.75
Corilagin is one of the major active components of many ethnopharmacological plants isolated from Caesalpinia and was reported to exhibit anti-tumor and anti-inflammatory activities.76 Corilagin was shown to inhibit the release of cytokines such as TNF-α, IL-1β, and IL-6 as well as the production of nitric oxide.77 More specifically, corilagin possess anti-anaphylactic and anti-allergic activities by inhibiting the release of mediators from mast cells and by decreasing the serum concentration of immunoglobulin E (IgE).78 Furthermore, the potent inhibition of the Corilagin on the phagocytic activity of neutrophils makes it an interesting herbal asthma remedy.79 Our results showed that the upregulated genes in the asthmatic epithelium and are part of Corilagin targets include TNNI3, HPGD, TNNC1, BLM, GFER, PLA2G7, SQLE, MCL1, PPARG. Four of them are alveolar macrophage-specific genes (HPGD; PPARG; PLA2G7; MCL1).
Phytoestrogens are plant-derived compounds found in a wide variety of foods and plants, being most abundant in soy,80 which is known to induce allergy, affecting approximately 0.4% of children.81 This could be the reason why it is discouraged to use soya protein in children in the first six months of life to avoid sensitization and exposure to phytoestrogens.82 Soy sauce is a traditional fermented seasoning of Japan, that is made from soybeans and wheat, both of which are established food allergens.80 On the other hand, phytoestrogens were reported to have a protective role against heart disease, breast cancer, and menopausal symptoms of osteoporosis.83 Phytoestrogens have structural similarities to estrogen and hence can bind to its receptor causing (anti)oestrogenic effects84 and could cause potential adverse health effects as well.83 With regards to asthma, it was found that increased consumption of phytoestrogens may help prevent or treat asthma and allergic disease.85 Furthermore, phytoestrogens can reduce antigen-induced eosinophilia in the lung.86 It was not surprising that the genes affected by phytoestrogens and upregulated in the severe asthmatic epithelium in our analysis (TOP2A, ANLN, MKI67, UHRF1, BIRC5, TFF1, MND1, RRM2, VWF, NUSAP1), are related to cell nuclear division, mitotic nuclear division, and cell cycle. TOP2A, ANLN, RRM2, UHRF1, NUSAP1, BIRC5, MND1, and MKI67 are enriched specifically in bronchial epithelial cells.
Rottlerin, also called mallotoxin, is the principal phloroglucinol constituent of the Mallotus Philippinensis (known as Kamala Tree). Previous studies have shown that rottlerin induces apoptosis, autophagy, and suppresses NF-κB and PKCδ in cancer cells, such as lung cancer.87,88 Rottlerin was shown to inhibit microvascular endothelial cells tube formation, block cell senescence, and intracellular ROS generation in psoriasis.89 In the lung, rottlerin was shown to be anti-inflammatory, airways smooth muscles relaxant90 and suppressant of airway hyperreactivity in mouse models of experimental asthma.91 Additionally, rottlerin induces apoptosis of human blood eosinophils , hence, can attenuate allergic reactions.92 On the other hand, rottlerin increases barrier dysfunction in pulmonary endothelial cell monolayers and causes pulmonary edema in rats.93
Salazinic acid can be isolated from Xanthoparmelia camtschadalis, Rimelia cetrata, and Parmelia caperata. It can be used as an antioxidant agent which plays an important role in macrophage killing of bacteria and tumors.94 Beside the anti-oxidative effect, salazinic acid has immunostimulatory, antimicrobial and antiproliferative potentials.94,95 Lichens contain large amounts of salazinic acid and have been used since ancient times as a therapeutic agent for the treatment of bronchitis, asthma, and inflammation.96 In our analysis, genes targeted by salazinic acid and upregulated in severe asthmatic bronchial epithelium (TNNI3, TNNC1, GFER, PLA2G7, TDP1, MCL1, PIN1, RUNX1, PABPC1, BCL2L1, RGS12) are related to the regulation of neuron apoptotic process and cardiac muscle tissue development.
Plant toxins
Microcystins (MCs) are hepatotoxins, produced by various species of cyanobacteria, whose occurrence is increasing worldwide owing to climate change and anthropogenic activities.97 Various edible aquatic organisms, plants, and food supplements based on algae can bioaccumulate these toxins.98 This occurs at times when blooms form and accumulate as scum on the water surface after which the death and decay of cells release large amounts of cyanotoxins which become toxic to eukaryotic organisms, including humans.99 Contact dermatitis, asthma-like symptoms, and symptoms resembling hay fever have been attributed to microcystins chemical sensitivity.100 Hence, water-based recreational activities can expose people to very low concentrations of aerosol-borne microcystins101 or even aerosolized cyanotoxins, making inhalation a potential route of exposure.102 Exposure to such aerosolized toxins in asthmatic subjects can have adverse effects.3 Our results showed that targeted genes by Microcystin in the asthmatic epithelium are specific to the trachea (ZNF57, LMNA, SERPINB5) and that the gene GSTT1 showed significant association with asthma risk.103
Conclusion
Our analysis using the publicly available gene expression data and linking it to toxicological omics’ data was able to explain and predict the toxicity in terms of affwcting the differentially expressed genes between severe asthmatic and normal epithelium. Many of the identified chemicals using this approach have no special warnings or precautions to avoid them by asthma patients. Even if some of the identified genes were reported earlier and linked to asthma, the exact mechanism is still poorly understood. The enriched pathways shared by most of the chemicals identified were related to significant players in the signaling pathways that are associated with triggering or exacerbation of asthma development. Such an approach can pave the way to generate a cost-effective and reliable source for asthma-specific toxigenic reports thus allowing the asthmatic patients, physicians, and medical researchers to be aware of the potential triggering factors with fatal consequences.
Acknowledgment
R.H. is funded by the Sharjah Research Academy (Grant code: MED001), University of Sharjah (Grant code: 1901090254) and Al-Jalila Foundation (Grant code: AJF201741).
Disclosure
The authors report no conflicts of interest in this work.
References
1. Ober C, Yao T-C. The genetics of asthma and allergic disease: a 21st century perspective. Immunol Rev. 2011;242(1):10–30. doi:10.1111/j.1600-065X.2011.01029.x
2. Thomsen SF. Genetics of asthma: an introduction for the clinician. Eur Clin Respir J. 2015;2. doi:10.3402/ecrj.v3402.24643
3. Fleming LE, Kirkpatrick B, Backer LC, et al. Aerosolized red-tide toxins (brevetoxins) and asthma. Chest. 2007;131(1):187–194. doi:10.1378/chest.06-1830
4. Elina T, KD W. Asthma risk factors. Int Forum Allergy Rhinol. 2015;5(S1):S11–S16. doi:10.1002/alr.21557
5. Foong R-X, Du Toit G, Fox AT. Asthma, food allergy, and how they relate to each other. Fronti Pediatr. 2017;5:89. doi:10.3389/fped.2017.00089
6. Litonjua AA. Dietary factors and the development of asthma. Immunol Allergy Clin North Am. 2008;28(3):603–ix. doi:10.1016/j.iac.2008.03.005
7. Ierodiakonou D, Zanobetti A, Coull BA, et al. Ambient air pollution, lung function, and airway responsiveness in asthmatic children. J Allergy Clin Immunol. 2016;137(2):390–399. doi:10.1016/j.jaci.2015.05.028
8. Liu S, Xia T, Zhu Y, Mu L, Zhang Z-F. Pulmonary diseases induced by ambient ultrafine and engineered nanoparticles in twenty-first century. Natl Sci Rev. 2016;3(4):416–429. doi:10.1093/nsr/nww064
9. Torén K, Blanc PD. Asthma caused by occupational exposures is common – A systematic analysis of estimates of the population-attributable fraction. BMC Pulm Med. 2009;9(1):7. doi:10.1186/1471-2466-9-7
10. Brooks SM, Weiss MA, Bernstein IL. Reactive airways dysfunction syndrome (RADS). Persistent asthma syndrome after high level irritant exposures. Chest. 1985;88(3):376–384. doi:10.1378/chest.88.3.376
11. Hewitt DJ. Can reactive airways dysfunction syndrome (RADS) be iatrogenic? Respir Care. 2011;56(8):1188–1194. doi:10.4187/respcare.01104
12. Maier A, Vincent MJ, Gadagbui B, et al. Integrating asthma hazard characterization methods for consumer products. Regul Toxicol Pharmacol. 2014;70(1):37–45. doi:10.1016/j.yrtph.2014.06.009
13. Zock J-P, Plana E, Jarvis D, et al. The use of household cleaning sprays and adult asthma: an international longitudinal study. Am J Respir Crit Care Med. 2007;176(8):735–741. doi:10.1164/rccm.200612-1793OC
14. Khan DA, Solensky R. Drug allergy. J Allergy Clin Immunol. 2010;125(2,Supplement 2):S126–S137.e121. doi:10.1016/j.jaci.2009.10.028
15. Varghese M, Lockey RF. Aspirin-exacerbated asthma. Allergy Asthma Clin Immunol. 2008;4(2):75. doi:10.1186/1710-1492-4-2-75
16. Vincent MJ, Bernstein JA, Basketter D, LaKind JS, Dotson GS, Maier A. Chemical-induced asthma and the role of clinical, toxicological, exposure and epidemiological research in regulatory and hazard characterization approaches. Regul Toxicol Pharmacol. 2017;90:126–132. doi:10.1016/j.yrtph.2017.08.018
17. Gomes ER, Brockow K, Kuyucu S, et al. Drug hypersensitivity in children: report from the pediatric task force of the EAACI Drug Allergy Interest Group. Allergy. 2016;71(2):149–161. doi:10.1111/all.12774
18. Moffat I, Chepelev N, Labib S, et al. Comparison of toxicogenomics and traditional approaches to inform mode of action and points of departure in human health risk assessment of benzo[a]pyrene in drinking water. Crit Rev Toxicol. 2015;45(1):1–43. doi:10.3109/10408444.2014.973934
19. Carsin A, Mazenq J, Ilstad A, Dubus J-C, Chanez P, Gras D. Bronchial epithelium in children: a key player in asthma. Eur Respir Rev. 2016;25(140):158–169. doi:10.1183/16000617.0101-2015
20. Hamoudi RA, Appert A, Ye H, et al. Differential expression of NF-kappaB target genes in MALT lymphoma with and without chromosome translocation: insights into molecular mechanism. Leukemia. 2010;24(8):1487–1497. doi:10.1038/leu.2010.118
21. Hung J-H, Yang T-H, Hu Z, Weng Z, DeLisi C. Gene set enrichment analysis: performance evaluation and usage guidelines. Brief Bioinform. 2012;13(3):281–291. doi:10.1093/bib/bbr049
22. Tripathi S, Pohl MO, Zhou Y, et al. Meta- and orthogonal integration of influenza “OMICs” data defines a role for UBR4 in virus budding. Cell Host Microbe. 2015;18(6):723–735. doi:10.1016/j.chom.2015.11.002
23. Kuo CS, Pavlidis S, Loza M, et al. A transcriptome-driven analysis of epithelial brushings and bronchial biopsies to define asthma phenotypes in U-BIOPRED. Am J Respir Crit Care Med. 2017;195(4):443–455. doi:10.1164/rccm.201512-2452OC
24. Subramanian A, Tamayo P, Mootha VK, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005;102(43):15545–15550. doi:10.1073/pnas.0506580102
25. Liu T, Zhang L, Joo D, Sun S-C. NF-κB signaling in inflammation. Signal Transd Targeted Ther. 2017;2:17023. doi:10.1038/sigtrans.2017.23
26. YMW J-H, Poynter ME, Aesif SW, et al. Nuclear factor kappaB, airway epithelium, and asthma: avenues for redox control. Proc Am Thorac Soc. 2009;6(3):249–255. doi:10.1513/pats.200806-054RM
27. Ather JL, Hodgkins SR, YMW J-H, Poynter ME. Airway epithelial NF-κB activation promotes allergic sensitization to an innocuous inhaled antigen. Am J Respir Cell Mol Biol. 2011;44(5):631–638. doi:10.1165/rcmb.2010-0106OC
28. Schuliga M. NF-kappaB signaling in chronic inflammatory airway disease. Biomolecules. 2015;5(3):1266–1283. doi:10.3390/biom5031266
29. Rico-Rosillo G, Vega-Robledo GB. The involvement of NF-κB transcription factor in asthma. Rev Alerg Méx. 2011;58(2):107–111.
30. Nawaz MA, Mesnage R, Tsatsakis AM, et al. Addressing concerns over the fate of DNA derived from genetically modified food in the human body: a review. Food Chem Toxicol. 2019;124:423–430. doi:10.1016/j.fct.2018.12.030
31. Kaneko K, Aoyagi Y, Fukuuchi T, Inazawa K, Yamaoka N. Total purine and purine base content of common foodstuffs for facilitating nutritional therapy for gout and hyperuricemia. Biol Pharm Bull. 2014;37(5):709–721. doi:10.1248/bpb.b13-00967
32. Clifford A, Riumallo J, Young V, Scrimshaw N. Effect of oral purines on serum and urinary uric acid of normal, hyperuricemic and gouty humans. J Nutr. 1976;106(3):428–434. doi:10.1093/jn/106.3.428
33. Yu M, Cui FX, Jia HM, et al. Aberrant purine metabolism in allergic asthma revealed by plasma metabolomics. J Pharm Biomed Anal. 2016;120:181–189. doi:10.1016/j.jpba.2015.12.018
34. Bittner C, Peters U, Frenzel K, Müsken H, Brettschneider R. New wheat allergens related to baker’s asthma. J Allergy Clin Immunol. 2015;136(5):1416–1418.e1412. doi:10.1016/j.jaci.2015.05.010
35. Zhang Y, Chen C, Choi H, et al. Purine-rich foods intake and recurrent gout attacks. Ann Rheum Dis. 2012;71(9):1448–1453. doi:10.1136/annrheumdis-2011-201215
36. Zhang Y, Peloquin CE, Dubreuil M, et al. Sleep apnea and the risk of incident gout: a population-based, body mass index–matched cohort study. Arthritis Rheum. 2015;67(12):3298–3302. doi:10.1002/art.39330
37. Teodorescu M, Polomis DA, Hall SV, et al. Association of obstructive sleep apnea risk with asthma control in adults. Chest. 2010;138(3):543–550. doi:10.1378/chest.09-3066
38. Butler MP, Smales C, Wu H, et al. The circadian system contributes to apnea lengthening across the night in obstructive sleep apnea. Sleep. 2015;38(11):1793–1801. doi:10.5665/sleep.5166
39. Li Y, Li G, Görling B, Luy B, Du J, Yan J. Integrative analysis of circadian transcriptome and metabolic network reveals the role of de novo purine synthesis in circadian control of cell cycle. PLoS Comput Biol. 2015;11(2):e1004086–e1004086. doi:10.1371/journal.pcbi.1004086
40. Vecsey CG, Peixoto L, Choi JHK, et al. Genomic analysis of sleep deprivation reveals translational regulation in the hippocampus. Physiol Genomics. 2012;44(20):981–991. doi:10.1152/physiolgenomics.00084.2012
41. Durrington HJ, Farrow SN, Loudon AS, Ray DW. The circadian clock and asthma. Thorax. 2014;69(1):90–92. doi:10.1136/thoraxjnl-2013-203482
42. Maples R. Arachidonic acid food sources and recommendation for the vegetarian. 2013;21–32.
43. Friesen RW, Innis SM. Dietary arachidonic acid to EPA and DHA balance is increased among canadian pregnant women with low fish intake. J Nutr. 2009;139(12):2344–2350. doi:10.3945/jn.109.112565
44. Mickleborough TD, Rundell KW. Dietary polyunsaturated fatty acids in asthma- and exercise-induced bronchoconstriction. Eur J Clin Nutr. 2005;59(12):1335–1346. doi:10.1038/sj.ejcn.1602250
45. Miyata J, Arita M. Role of omega-3 fatty acids and their metabolites in asthma and allergic diseases. Allergol Int. 2015;64(1):27–34. doi:10.1016/j.alit.2014.08.003
46. Kakutani S, Egawa K, Saito K, et al. Arachidonic acid intake and asthma risk in children and adults: a systematic review of observational studies. J Nutr Sci. 2014;3:e12–e12. doi:10.1017/jns.2014.9
47. Balhara J, Gounni AS. The alveolar macrophages in asthma: a double-edged sword. Mucosal Immunol. 2012;5(6):605–609. doi:10.1038/mi.2012.74
48. Draijer C, Peters-Golden M. Alveolar macrophages in allergic asthma: the forgotten cell awakes. Curr Allergy Asthma Rep. 2017;17(2):12. doi:10.1007/s11882-017-0681-6
49. Muthukumaran S, Tranchant C, Shi J, Ye X, Xue SJ. Ellagic acid in strawberry (Fragaria spp.): biological, technological, stability, and human health aspects. Food Qual Saf. 2017;1(4):227–252. doi:10.1093/fqsafe/fyx023
50. Ismail T, Calcabrini C, Diaz AR, et al. Ellagitannins in cancer chemoprevention and therapy. Toxins. 2016;8(5):151. doi:10.3390/toxins8050151
51. Vicinanza R, Zhang Y, Henning SM, Heber D. pomegranate juice metabolites, ellagic acid and urolithin A, synergistically inhibit androgen-independent prostate cancer cell growth via distinct effects on cell cycle control and apoptosis. Evid-based Compl Alt Med. 2013;2013:247504. doi:10.1155/2013/247504
52. Zhou E, Fu Y, Wei Z, Yang Z. Inhibition of allergic airway inflammation through the blockage of NF-kappaB activation by ellagic acid in an ovalbumin-induced mouse asthma model. Food Funct. 2014;5(9):2106–2112. doi:10.1039/C4FO00384E
53. Rogerio AP, Fontanari C, Borducchi E, et al. Anti-inflammatory effects of Lafoensia pacari and ellagic acid in a murine model of asthma. Eur J Pharmacol. 2008;580(1–2):262–270. doi:10.1016/j.ejphar.2007.10.034
54. de Freitas Alves C, Angeli GN, Favarin DC, et al. The effects of proresolution of ellagic acid in an experimental model of allergic airway inflammation. Mediators Inflamm. 2013;2013:9. doi:10.1155/2013/863198
55. Nabavi SF, Braidy N, Gortzi O, et al. Luteolin as an anti-inflammatory and neuroprotective agent: a brief review. Brain Res Bull. 2015;119:1–11. doi:10.1016/j.brainresbull.2015.09.002
56. Luo Y, Shang P, Luteolin: LD. A flavonoid that has multiple cardio-protective effects and its molecular mechanisms. Front Pharmacol. 2017;8:692. doi:10.3389/fphar.2017.00692
57. Ma Y, Zhang JX, Liu YN, et al. Caffeic acid phenethyl ester alleviates asthma by regulating the airway microenvironment via the ROS-responsive MAPK/Akt pathway. Free Radic Biol Med. 2016;101:163–175. doi:10.1016/j.freeradbiomed.2016.09.012
58. Magnani C, Isaac V, Corrêa M, Salgado H. Caffeic acid: a review of its potential use in medications and cosmetics. Anal Methods. 2014;6:3203.
59. Wang L-C, Lin Y-L, Liang Y-C, et al. The effect of caffeic acid phenethyl ester on the functions of human monocyte-derived dendritic cells. BMC Immunol. 2009;10(1):39. doi:10.1186/1471-2172-10-39
60. Sy LB, Yang L-K, Chiu C-J, Wu W-M. The Immunoregulatory effects of caffeic acid phenethyl ester on the cytokine secretion of peripheral blood mononuclear cells from asthmatic children. Pediatr Neonatol. 2011;52(6):327–331. doi:10.1016/j.pedneo.2011.08.005
61. Honda S, Fukuyama Y, Nishiwaki H, Masuda A, Masuda T. Conversion to purpurogallin, a key step in the mechanism of the potent xanthine oxidase inhibitory activity of pyrogallol. Free Radical Biol Med. 2017;106:228–235. doi:10.1016/j.freeradbiomed.2017.02.037
62. Patel S, Rauf A, Khan H. The relevance of folkloric usage of plant galls as medicines: finding the scientific rationale. Biomed Pharmacother. 2018;97:240–247. doi:10.1016/j.biopha.2017.10.111
63. Ku S-K, Bae J-S. Antiplatelet and antithrombotic activities of purpurogallin in vitro and in vivo. BMB Rep. 2014;47(7):376–381. doi:10.5483/bmbrep.2014.47.7.195
64. Park HY, Kim TH, Kim CG, et al. Purpurogallin exerts anti‑inflammatory effects in lipopolysaccharide‑stimulated BV2 microglial cells through the inactivation of the NF‑κB and MAPK signaling pathways. Int J Mol Med. 2013;32(5):1171–1178. doi:10.3892/ijmm.2013.1478
65. Hoon Kim T, Ku S-K, Lee I-C, Bae J-S. Anti-inflammatory functions of purpurogallin in LPS-activated human endothelial cells. BMP Rep. 2012;45:200–205.
66. Sachar M, Anderson KE, Ma X. Protoporphyrin IX: the Good, the Bad, and the Ugly. J Pharmacol Exp Ther. 2016;356(2):267–275. doi:10.1124/jpet.115.228130
67. De Maere H, Chollet S, De Brabanter J, Michiels C, Paelinck H, Fraeye I. Influence of meat source, pH and production time on zinc protoporphyrin IX formation as natural colouring agent in nitrite-free dry fermented sausages. Meat Sci. 2018;135:46–53. doi:10.1016/j.meatsci.2017.08.024
68. Li G, Chen S, Duan Z, Qu L, Xu G, Yang N. Comparison of protoporphyrin IX content and related gene expression in the tissues of chickens laying brown-shelled eggs. Poult Sci. 2013;92(12):3120–3124. doi:10.3382/ps.2013-03484
69. Zhang Y, Zhang L, Wu J, Di C, Xia Z. Heme oxygenase-1 exerts a protective role in ovalbumin-induced neutrophilic airway inflammation by inhibiting Th17 cell-mediated immune response. J Biol Chem. 2013;288(48):34612–34626. doi:10.1074/jbc.M113.494369
70. Donald G, Hertzer K, Eibl G. Baicalein–an intriguing therapeutic phytochemical in pancreatic cancer. Curr Drug Targets. 2012;13(14):1772–1776.
71. He X, Wei Z, Zhou E, et al. Baicalein attenuates inflammatory responses by suppressing TLR4 mediated NF-kappaB and MAPK signaling pathways in LPS-induced mastitis in mice. Int Immunopharmacol. 2015;28(1):470–476. doi:10.1016/j.intimp.2015.07.012
72. Yeh C-H, Ma K-H, Liu P-S, Kuo J-K, Chueh S-H. baicalein decreases hydrogen peroxide-induced damage to NG108-15 cells via upregulation of Nrf2. J Cell Physiol. 2015;230(8):1840–1851. doi:10.1002/jcp.24900
73. Fink SP, Yamauchi M, Nishihara R, et al. Aspirin and the risk of colorectal cancer in relation to the expression of 15-hydroxyprostaglandin dehydrogenase (HPGD). Sci Transl Med. 2014;6(233):233re232–233re232. doi:10.1126/scitranslmed.3008481
74. Laidlaw TM. Clinical updates in aspirin-exacerbated respiratory disease. Allergy and Asthma Proc. 2019;40(1):4–6. doi:10.2500/aap.2019.40.4188
75. Pennings JLA, Kimman TG, Janssen R. Identification of a common gene expression response in different lung inflammatory diseases in rodents and macaques. PLoS One. 2008;3(7):e2596. doi:10.1371/journal.pone.0002596
76. Li X, Deng Y, Zheng Z, et al. Corilagin, a promising medicinal herbal agent. Biomed Pharmacother. 2018;99:43–50. doi:10.1016/j.biopha.2018.01.030
77. Okabe S, Suganuma M, Imayoshi Y, Taniguchi S, Yoshida T, Fujiki H. New TNF-α releasing inhibitors, geraniin and corilagin, in leaves of Acer nikoense, Megusurino-ki. Biol Pharm Bull. 2001;24(10):1145–1148. doi:10.1248/bpb.24.1145
78. Long X, Zhou W, Wang Y, Liu S. Prognostic significance of ANLN in lung adenocarcinoma. Oncol Lett. 2018;16(2):1835–1840. doi:10.3892/ol.2018.8858
79. Jantan I, Ilangkovan M, Yuandani MHF. Correlation between the major components of Phyllanthus amarus and Phyllanthus urinaria and their inhibitory effects on phagocytic activity of human neutrophils. BMC Complement Altern Med. 2014;14(1):429. doi:10.1186/1472-6882-14-429
80. Jargin SV. Soy and phytoestrogens: possible side effects. German Med Sci. 2014;12:Doc18–Doc18.
81. Savage JH, Kaeding AJ, Matsui EC, Wood RA. The natural history of soy allergy. J Allergy Clin Immunol. 2010;125(3):683–686. doi:10.1016/j.jaci.2009.12.994
82. Paediatric group position statement on the use of soya protein for infants. J Fam Health Care. 2003;13(4):93.
83. Patisaul HB, Jefferson W. The pros and cons of phytoestrogens. Front Neuroendocrinol. 2010;31(4):400–419. doi:10.1016/j.yfrne.2010.03.003
84. Rietjens IMCM, Louisse J, Beekmann K. The potential health effects of dietary phytoestrogens. Br J Pharmacol. 2017;174(11):1263–1280. doi:10.1111/bph.13622
85. Cardet J-C, Johns C, Savage JR. Urinary levels of phytoestrogens are inversely associated with wheezing, asthma, and atopy. J Allergy Clin Immunol. 2014;133(2):AB163. doi:10.1016/j.jaci.2013.09.002
86. Regal JF, Fraser DG, Weeks CE, Greenberg NA. Dietary phytoestrogens have anti-inflammatory activity in a guinea pig model of asthma. Proc Soc Exp Biol Med. 2000;223(4):372–378. doi:10.1046/j.1525-1373.2000.22353.x
87. Maioli E, Torricelli C, Valacchi G. Rottlerin and cancer: novel evidence and mechanisms. TheScientificWorldJournal. 2012;2012:350826. doi:10.1100/2012/350826
88. Wang L, Hou Y, Yin X, et al. Rottlerin inhibits cell growth and invasion via down-regulation of Cdc20 in glioma cells. Oncotarget. 2016;7(43):69770–69782. doi:10.18632/oncotarget.11974
89. Min M, Yan B-X, Wang P, et al. Rottlerin as a therapeutic approach in psoriasis: evidence from in vitro and in vivo studies. PLoS One. 2017;12(12):e0190051. doi:10.1371/journal.pone.0190051
90. Goldklang MP, Perez-Zoghbi JF, Trischler J, et al. Treatment of experimental asthma using a single small molecule with anti-inflammatory and BK channel-activating properties. FASEB J. 2013;27(12):4975–4986. doi:10.1096/fj.13-235176
91. Chesne J, Braza F, Mahay G, Brouard S, Aronica M, Magnan A. IL-17 in severe asthma. Where do we stand? Am J Respir Crit Care Med. 2014;190(10):1094–1101. doi:10.1164/rccm.201405-0859PP
92. Berro AI, Bharadwaj A, Agrawal DK. Rottlerin induces apoptosis of human blood eosinophils: a possible role for PKC-δ in mediating eosinophil survival. J Allergy Clin Immunol. 2005;115(2):S193. doi:10.1016/j.jaci.2004.10.029
93. Klinger JR, Murray JD, Casserly B, et al. Rottlerin causes pulmonary edema in vivo: a possible role for PKCδ. J Appl Physiol. 2007;103(6):2084–2094. doi:10.1152/japplphysiol.00695.2007
94. White P, Oliveira R, Oliveira A, et al. Antioxidant activity and mechanisms of action of natural compounds isolated from lichens: a systematic review. Molecules. 2014;19:14496–14527.
95. Mitrović T, Stamenković S, Cvetković V, et al. Antioxidant, antimicrobial and antiproliferative activities of five lichen species. Int J Mol Sci. 2011;12(8):5428–5448. doi:10.3390/ijms12085428
96. Salgado F, Albornoz L, Cortez C, et al. Secondary metabolite profiling of species of the genus usnea by UHPLC-ESI-OT-MS-MS. Molecules. 2017;23(1). doi:10.3390/molecules23010054
97. Díez-Quijada L, Puerto M, Gutiérrez-Praena D, Llana-Ruiz-Cabello M, Jos A, Cameán AM. Microcystin-RR: occurrence, content in water and food and toxicological studies. A review. Environ Res. 2019;168:467–489. doi:10.1016/j.envres.2018.07.019
98. Gutiérrez-Praena D, Á J, Pichardo S, Moreno IM, Cameán AM. Presence and bioaccumulation of microcystins and cylindrospermopsin in food and the effectiveness of some cooking techniques at decreasing their concentrations: a review. Food Chem Toxicol. 2013;53:139–152. doi:10.1016/j.fct.2012.10.062
99. Rai AK, Chaturvedi R, Kumar A. Proteomic evidences for microcystin-RR-induced toxicological alterations in mice liver. Sci Rep. 2018;8(1):1310. doi:10.1038/s41598-018-19299-w
100. Torokne A, Palovics A, Bankine M. Allergenic (sensitization, skin and eye irritation) effects of freshwater cyanobacteria - Experimental evidence. Environ Toxicol. 2001;16:512–516.
101. Backer LC, Carmichael W, Kirkpatrick B, et al. Recreational exposure to low concentrations of microcystins during an algal bloom in a small lake. Mar Drugs. 2008;6(2):389–406. doi:10.3390/md20080018
102. Backer L, McNeel SV, Barber T, et al. Recreational exposure to microcystins during algal blooms in two California lakes. Toxicon. 2009;55:909–921.
103. Liang S, Wei X, Gong C, et al. Significant association between asthma risk and the GSTM1 and GSTT1 deletion polymorphisms: an updated meta-analysis of case-control studies. Respirology. 2013;18(5):774–783. doi:10.1111/resp.12097
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.