Back to Journals » International Journal of Nanomedicine » Volume 13
The therapeutic effect in gliomas of nanobubbles carrying siRNA combined with ultrasound-targeted destruction
Authors Cai W, Lv W, Feng Y, Yang H , Zhang Y, Yang G, Duan Y , Wang J
Received 6 February 2018
Accepted for publication 5 June 2018
Published 24 October 2018 Volume 2018:13 Pages 6791—6807
DOI https://doi.org/10.2147/IJN.S164760
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Professor Lei Yang
Wenbin Cai,1,2,* Wei Lv,1,3,* Yang Feng,4 Hengli Yang,1 Yajun Zhang,1 Guodong Yang,5 Yunyou Duan,1 Jia Wang1
1Department of Ultrasound Diagnosis, Tang Du Hospital, Fourth Military Medical University, Xi’an 710038, China; 2General Hospital of Tibet Military Command, Lhasa, Tibet Autonomous Region, 850007, China; 3Department of Radiology, 305 Hospital of Chinese People’s Liberation Army, Xicheng District, Beijing, 100017, China; 4Xijing Hospital, Traditional Chinese Medicine, Xi’an 710032, China; 5Department of Biochemistry and Molecular Biology, The Fourth Military Medical University, Xi’an 710032, China
*These authors contributed equally to this work
Background: Nanobubbles (NBs) combined with ultrasound-targeted destruction (UTD) have become promising potential carriers for drug or siRNA delivery. Due to their nano-size, NBs could penetrate tumor blood vessels and accumulate in intercellular spaces so that “sonoporation” induced by UTD would act directly on the tumor cells to increase cell membrane permeability.
Methods: Based on the successful the fabrication of NBs, we synthesized NBs carrying siRNA (NBs-siRNA) by using a biotin-streptavidin system. We then utilized ultrasound irradiation (UI)-targeted NBs-siRNA to improve siRNA transfection and achieve the inhibition of glioma growth.
Results: NBs as carriers combined with UI effectively enhanced siRNA transfection and the effect of silencing targeted genes in vitro. Additionally, a better therapeutic effect was shown in the NBs-siRNA with UI group in vivo compared with that of microbubbles (MBs) with UI or NBs-siRNA without UI.
Conclusion: These results indicated that NBs combined with UTD might be an ideal delivery vector for siRNA to achieve the noninvasive treatment of glioma.
Keywords: nanobubbles, ultrasound, siRNA, glioma
Introduction
Glioma – one of the most invasive and infiltrative tumors – has an exceedingly poor prognosis with a 5-year survival rate of <10% and a 12- to 15-month median overall survival duration.1–4 The current treatments for gliomas are mainly surgical resection, irradiation, and chemotherapy. However, due to special pathological and physiological characteristics, some problems, such as surgical injury and side effects, remain.5 Therefore, noninvasive treatment of glioma with low side effects is still a huge challenge and task.
With the development of genetic technology, the gene silencing represented by small interference RNAs (siRNAs) plays an increasingly important role in noninvasive cancer treatment.6–9 SiRNAs are a class of double-stranded RNA molecules of 20–25 base pairs in length that can recognize and degrade complementary messenger RNAs (mRNAs).10 Therefore, designing siRNAs could silence multiple mutational genes that result in tumor formation and affect tumor growth.11,12 Recently, siRNA molecules have entered the human trial phase and are considered an efficient and promising treatment for cancer.13–15 However, the in vivo application of siRNA still faces great challenges. For example, physiological conditions could affect the stability of siRNA, as this molecule has to pass through the tumor blood vessel and enter the intercellular space. Thus, the tumor-targeted delivery of siRNA needs to permeate through the membrane into the cell.16,17 Therefore, delivery systems have become a key point for the application of siRNA. Although many delivery systems, such as inorganic nanoparticles,18,19 polymers,20–24 lipids,25–27 and virus vectors,28 have been developed to address the above challenges, these systems have some imperfections. The toxicity of cationic lipids, primarily resulting from their cationic characteristics, is a big problem for the application of these molecules in siRNA delivery;29 liposomes are limited by poor entrapment efficiency;30 and viral vectors have high transfection efficiencies but show immunogenicity, genotoxicity, and mutagenicity.31,32 Therefore, an effective, tumor-targeting, and safe siRNA delivery strategy is significant.
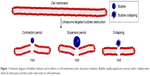
Microbubbles (MBs) – which have been widely used in the clinic as ultrasound contrast agents (USA) – play a critical role in the ultrasound diagnosis of many diseases. The revolution brought by MBs – that is, spherical core shell structures filled with gases such as perfluorocarbons – is not confined only to the diagnosis33–35 but also addresses the treatment of diseases.36–40 Previous studies have shown that MBs rapidly expand and contract and/or collapse in response to ultrasound exposure,41,42 referred to as ultrasound-targeted destruction (UTD). The localized significant shear stress, fluid flow, and other mechanical impacts generated by MBs combined with UTD would change cells and biological structures to produce “sonoporation”, which generates voids on cell membranes of approximately 300 nm, with a half-life of 20–50 ms (Figure 1).43–50 This sonoporation could improve the permeability of the cell membrane, increase the transport of extracellular impermeable compounds into cells, and permit cells to more easily take up drugs and genes.44,51–53 Due to their minimal inflammatory and immunological responses, MBs combined with UTD represent a potential strategy for gene delivery to noninvasively treat tumors.54,55 However, the size of MBs limits their application for the treatment of tumors, as MBs cannot pass through the endothelial gaps (380–780 nm)56,57 of tumor blood vessels and enter tissues.58 Therefore, to overcome this limitation, nanobubbles (NBs) with diameters of 300–700 nm have attracted considerable research attention.
NBs combined with UTD represent a potential ideal vector for gene delivery. Previous studies have shown that, due to their nano size, NBs could penetrate tumor blood vessels and achieve tumor-targeted imaging59,60 by utilizing the endothelial gaps of tumor blood vessels, which are 7 nm in normal tissue.61 First, UTD is limited to the ultrasound irradiation area, which would reduce its impact on normal tissue and, thus, effectively increase the siRNA concentration in the targeted tissue, and reduce systemic toxic side effects. Second, the sonoporation driven by NBs combined with UTD would increase cell membrane permeability and gene transfection efficiency (Figure 2). Therefore, in this study, we developed NBs carrying siRNA by using a biotin and streptavidin system and demonstrated that NBs carrying siRNA combined with UTD could improve siRNA transfection efficiency, providing experimental support for NBs carrying siRNA combined with UTD as a noninvasive treatment of glioma.
Material and methods
Materials
The phospholipids 1,2-dipalmitoyl-snglycero-3-phosphocholine (DPPC; MW, 734.039 g/mol) and 1,2-distearoyl-sn-glycero-3-phosphoethanolamine-N-[biotinylated(polyethyleneglycol)–2000] (biotinylated DSPE-PEG[2000]; MW, 3,016.781 g/mol), which were used in the fabrication of NBs, were purchased from Avanti Polar Lipids Inc. in powder form and used without further purification. Octafluoropropane (C3F8) gas was purchased from the R&D Center for Specialty Gases at the Research Institute of Physical and Chemical Engineering of Nuclear Industry (Beijing, China). The dynamic light scattering (DLS) analyzer was purchased from Beckman Coulter (USA). The U87 MG cell line was purchased from the American Type Culture Collection. ImmunoPure® streptavidin was purchased from Pierce (Rockford, IL, USA). The siRNA duplexes designed to target the SIRT2 gene, scrambled siRNA (SCR), and fluorescein amidite (FAM)-labeled scrambled siRNA (FAM-SCR) were purchased from Genpharm (Shanghai, China). The sequences are shown in Table S1.
Fabrication and characteristics of NBs
The NBs were fabricated by a thin-film hydration method as previously reported. Briefly, 14 mg biotinylated DSPE-PEG (2000) and DPPC, in a fixed ratio, were weighed and added to a 25-mL rotary evaporation bottle. Then, 2 mL chloroform was added to the bottle to dissolve the phospholipid. Subsequently, 8 μL red fluorescent probe DiI solution (1.2 mg/mL, Beyotime, Haimen, China) was added for the NB fluorescence experiments. Then, the bottle was placed on a rotary evaporator (New Brunswick Scientific, Enfield, CT, USA), and rotary evaporation was undertaken at 55°C and 120 rpm/min. After ten minutes, the chloroform was vaporized, and a milky white phospholipid thin film formed on the rotary evaporation bottle wall. Next, 1 mL of hydration liquid comprising 10% glycerol and 90% 1×PBS (V/V) was added to the bottle. The bottle was placed in an incubator shaker (New Brunswick Scientific), followed by shaking for 60 minutes at 37°C and 130 rpm to form a liposomal film suspension. The suspension was equally divided into two tubes with plastic caps. The air in the sealed tubes was replaced with C3F8 gas using a syringe. Finally, every tube was oscillated for 45 seconds in an amalgamator. The bubbles in each tube were separately diluted to 8 mL with PBS. The MBs used in the present study were fabricated by the same method, except that 28 mg phospholipid was used, on the basis of a previous report. MBs were used to compare the ability of MBs and NBs to penetrate tumor blood vessels. The NBs and MBs were sterilized via CO60 irradiation. DLS with a laser wavelength of 660 nm at an angle of 90° was used to measure the particle sizes of the NBs and MBs. The experiments were repeated three times. To visualize the NBs, a drop of NB suspension was loaded onto dust-free foil, placed in a desiccator, gold sputter-coated for five minutes, and then examined by scanning electron microscopy (SEM; HITACHI S-4800, Japan). A Cell Counting Kit-8 (CCK-8; Dojindo Laboratories, Kumamoto, Japan) was used to assess the cytotoxicity of NBs. Briefly, U87 cells were seeded onto 96-well plates at 4,000 cells/well with 100 μL/well of RPMI-1640 medium containing 10% fetal bovine serum (FBS). After 24 hours, the medium was replaced with the same volume of fresh medium containing various concentrations of phospholipids (2.5–640 μg/mL). After 24 hours, 10 μL CCK-8 reagent was added to each well. After incubation at 37°C for one hour, the absorbance at 450 nm in each well was measured by an Infinite F200 Multimode plate reader (Tecan, Männedorf, Switzerland). The curve was drawn, and a statistical analysis was conducted.
Synthesis and detection of NB-siRNA conjugates
A 100-μL aliquot of NBs suspension was diluted with RNase-free PBS to 850 μL, and 100 μL streptavidin (concentration 10 mg/mL) was added to the tube. The tube was placed in Mini-Tube Rotators (Fisher Scientific™ USA) and subjected to oscillation for 30 minutes at 18 rpm. Then, 50 μL of 20 μM siRNA suspension was added, and the mixture was oscillated again. The air in the tube was replaced with C3F8 gas, and the tube was maintained in an ice box at 4°C for 90 minutes. After delamination appeared in the suspension (Figure 4A), the lower layer of clear suspension was withdrawn by a syringe, and equal amounts of PBS were injected. This step was repeated two or three times to wash the excess streptavidin and siRNA. Finally, the NBs-siRNA were formed. Similarly, NBs-SCR and NBs-FAM-SCR were fabricated using the same method. The particle size and stability of NBs-siRNA were measured by DLS and hemocytometer count. A 1-mL aliquot of NBs-siRNA was immediately measured after the completion of fabrication at 25°C, and the remaining NBs-siRNAs were maintained at 25°C. The particle size of the remaining NBs-siRNAs was then measured after 15, 30, 45, and 60 minutes. At each time point, the NBs-siRNAs were concurrently transferred to a hemocytometer for counting. A drop of the NBs-FAM-SCR diluted sample was placed onto slide glasses and observed by fluorescence microscopy with a 100× oil-immersion objective lens (microscope: Zeiss Axioskop, WEL Instrument Co., LLC; Camera: Zeiss Axiocam MRc5). Bare NBs were used as controls.
Measurements of transfection efficiency
Cell transfection
A total of 5×104 glioma U87 cells were seeded onto confocal 12-well culture dishes containing RPMI-1640 medium supplemented with 10% FBS and incubated at 37°C with 5% CO2 for 24 hours. The medium was replaced with 900 μL medium and 100 μL sample suspension. Gentle shaking was used to evenly distribute the samples in the culture medium. To ensure good penetration of the ultrasound, after disinfection, the probe was placed in medium without contact with the cells. The ultrasound exposure parameters were 1 MHz frequency, 0.88 W/cm2 intensity, and 45 seconds exposure time.
Confocal laser scanning microscopy
The transfection efficiency of FAM-SCR was detected by confocal laser scanning microscopy (CLSM). There were three groups: NBs carrying FAM-SCR with ultrasound irradiation (NBs-FAM-SCR-UI), FAM-SCR with ultrasound irradiation (FAM-SCR-UI), and bare FAM-SCR (control). The cells were cultured in confocal culture dishes and transfected as described earlier. At 24 hours after transfection, the cells were washed three times with PBS, and the cell nuclei were stained with DAPI to identify the location of the cells. The cells were observed on CLSM (Olympus, Fluoview FV10i, Tokyo, Japan).
Target gene mRNA expression level
The mRNA expression of target genes was detected by real-time PCR assay. The experiment groups were similar to those described earlier: NBs carrying siRNA with ultrasound irradiation (NBs-siRNA-UI), siRNA with ultrasound irradiation (siRNA-UI), and NBs carrying SCR with UI (control). RNA was isolated from the cells at 48 hours after transfection using the TRIzol Kit (BioTeke, Beijing, China) and converted into cDNA. Real-time PCR was conducted by the StepOnePlus Real-Time PCR System (Applied Biosystems, American). The GAPDH gene was used as an internal control. Primers used in real-time PCR are shown in Table S2.
Target gene protein expression level
The protein expression of the target gene was detected by Western blot analysis. The experimental groups were similar to those described above. Total protein was extracted from all groups. The 30-mg protein samples were separated on 12% SDS-PAGE gels and transferred to polyvinylidene fluoride (PVDF) membranes, which were initially incubated with the primary antibody, and then washed and incubated with the secondary antibody. The proteins were detected by a chemiluminescence system.
Detection of cell apoptosis
Annexin-V was used to detect cell apoptosis. The experimental groups were similar to those described above. At 48 hours after transfection, the cells were trypsinized, collected, and washed with PBS. The cells were dyed according to the Annexin V-FITC Apoptosis Kit (Abnova). Briefly, the cells were resuspended in 500 μL Binding Buffer, and 5 μL of Annexin V-FITC and 5 μL of propidium iodide were added. The cells were incubated at room temperature for five minutes in the dark. Flow cytometry (Ex=488 nm; Em=530 nm) was done to analyze Annexin-V.
Cell viability test
A CCK-8 assay was applied to evaluate the cell viability after transfection. The U87 cells were seeded onto 96-well plates at 3,000 cells per well and cultured for 24 hours. The experimental groups were similar to those described above. Transfection was conducted with 90 μL medium and 10 μL sample suspension by using the same process described earlier. Each group contained five wells. At 48 hours after transfection, 10 μL CCK-8 solution was added to each well of the plate. Then, the plate was incubated for four hours. The absorbance at 450 nm in each well was measured by the Infinite F200 Multimode plate reader. The cell viability = (A1−A2)/(A3−A2)×100%, where A1 was the absorbance of wells in that experimental group, A2 was the absorbance of wells containing medium and CCK-8 solution without cells and samples, and A3 was the absorbance of wells containing medium, CCK-8 solution, and cells without samples.
Animal models
The glioma in vivo model was established by a subcutaneous injection of 5×106 U87 cells into the right back area of athymic nude mice (five weeks). All animal experiments were approved by and conducted according to the guidelines of the Tangdu Hospital committee on Animal Care and Use in accordance with the National Institutes of Health (NIH) guidelines.
NBs-siRNA distribution in vivo
The Esaote MyLab Twice ultrasound diagnostic apparatus with a broad bandwidth (3–9 MHz) routine clinical linear array transducer (Esaote, LA522) was used to compare the NBs-siRNA-enhanced tumor images before and after ultrasound-targeted destruction. After isoflurane anesthesia, the mice were injected with NBs-siRNA suspension (200 μL) through the tail vein. At 15 seconds after the enhancement appeared, targeted destruction ultrasound – at 1 MHz frequency, with 1.5 W/cm2 intensity, and 1-minute exposure time – was implemented. The enhanced tumor images before and after ultrasound irradiation were recorded (UI [+] group). The enhanced tumor images without ultrasound irradiation at the same time point were used as a control (UI [−] group). The gray scale intensity of the digital clips and images was obtained by WCIF ImageJ software (v1.37; National Institutes of Health, Bethesda, MA, USA), and phase-intensity curves were created by GraphPad Prism (v5.01; GraphPad Software, Inc.) and statistically analyzed using SPSS software (v19.0; SPSS Inc., an IBM Company).
DiI-labeled NBs-siRNA and MBs-siRNA (200 μL) were separately injected into nude mice bearing glioma xenograft tumors. After ultrasound-targeted destruction, the nude mice were sacrificed, and the tumors were immediately separated to prepare frozen sections. The frozen sections were incubated with Isolectin-b4 (1:1,000, Beyotime, Haimen, China) at 25°C for eight hours and then washed three times with PBS. Subsequently, the sections were incubated for two hours with anti-biotin as a secondary antibody (1:1,000, Beyotime, Haimen, China) and washed three times with 1×PBS. The nucleus was stained with DAPI. Finally, CLSM was used to observe the tumor sections.
The NBs-siRNA distribution and metabolism in the tumors were measured by tissue fluorescence imaging by using the IVIS Spectrum Imaging System. At ten minutes, 30 minutes, one hour, six hours, 12 hours, and 24 hours after the injection of DiI-NBs-siRNA solution, the mice were sacrificed, and the liver, spleen, kidney, heart, and lungs were harvested. The fluorescence images of the tumors and organs were obtained. Based on the fluorescent signal, the liver and spleen were selected as positive and negative controls. The DiI-NBs-siRNA solution was injected into tumor-carrying nude mice, and the mice were simultaneously sacrificed. The tumor, liver, and spleen were harvested and subsequently observed and analyzed.
Effect of in vivo treatment
After the cells were injected, additional trials were conducted on animals until the tumor volume was approximately 100–300 mm3. For in vivo experiments, the mice were randomly separated into four groups: NBs carrying siRNA with ultrasound irradiation (NBs-siRNA-UI), MBs carrying siRNA with ultrasound irradiation (MBs-siRNA-UI), NBs carrying siRNA without ultrasound irradiation (NBs-siRNA), and NBs carrying SCR with ultrasound irradiation (NBs-SCR-UI). An aliquot of 200 μL of NBs carrying siRNA or SCR suspensions was injected through the tail vein. Then, the tumors were exposed to UI under the same parameters as described earlier. This operation was repeated two times per week for four weeks. The tumor volume was calculated according to the formula for the spherical segment volume: V=(A×B2)/2. A and B represent the length and the width of the tumor, respectively. The volume of the tumor was measured once every four days. The measurement was repeated 40 days later when the mouse began to die. The survival rate of mice in the four groups was analyzed by Kaplan–Meier survival analysis.
After the mice were sacrificed, the tumors were separated to prepare paraffinized sections, and hematoxylin and eosin (H&E) staining was done after deparaffinization. A TUNEL assay was conducted according to the TdT In Situ Apoptosis Detection Kit (Roche, Indianapolis, IN, USA). Briefly, the sections were incubated with dimethylbenzene and ethanol and washed with PBS. Then, proteinase K (0.3 mg/mL) was used to process the sections at room temperature for 25 minutes. Next, 3% H2O2 aqueous solution was added to quench the endogenous peroxidase. After TdT Enzyme and digoxigenin-labeled dUTP were added, the sections were incubated in equilibration buffer for one hour at 37°C.
Results
Characteristics of NBs
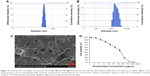
The average particle sizes of the bubbles were measured by DLS. As shown in Figure 3A and B, the average particle size of the NBs is 625.4±63.8 nm (n=3), and that of MBs is 2,355.8±512.8 nm (n=3). In the SEM micrographs (Figure 3C), the NBs were observed as un-aggregated hollows, similar to the findings of previous studies,59,60 with particle sizes of approximately 400–800 nm, corresponding to the size distributions measured by DLS. Figure 3D shows the cytotoxicity curve for the phospholipid concentration. Statistical analysis revealed obvious cytotoxicity when the concentration increased to 10 μg/mL.
NBs carrying siRNA
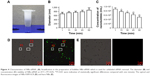
The stability of NBs-siRNA at room temperature was evaluated by particle size and concentration. The average particle sizes of NBs carrying siRNA that were stored separately at 25°C for 1, 15, 30, 45, and 60 minutes were 688.23±61.4 nm (n=3), 740.33±54.88 nm (n=3), 756.37±51.20 nm (n=3), 789.13±62.22 nm (n=3), and 870.17±93.09 nm (n=3), respectively (Figures 4 and S1). Compared with the average particle size of NBs carrying siRNA stored for one minute, there was no significant difference at each time point. The concentrations of NBs carrying siRNA were 88.34×105±6.59×105/mL (n=5), 77.02×105±6.62×105/mL (n=5), 70.76×105±4.29×105/mL (n=5), 62.00×105±7.53×105/mL (n=5), and 50.72×105±6.91×105/mL (n=5) at 1, 15, 30, 45, and 60 minutes, respectively (Figure 4C). The statistical analysis showed a significant difference between the concentrations at each time point and that at one minute. Figure 4D and E shows the optical and fluorescence images of the NBs-FAM-SCR group and the bare NBs control group. In the NBs-FAM-SCR group, green fluorescence was observed, and the conformity with NBs under optical and fluorescence microscopy was notable. No fluorescence was observed on the bare NBs. Additionally, an irregular fluorescence region appeared at some positions without bubbles, which are indicated by a red frame in Figure 4D.
siRNA transfection efficiency of NBs carrying siRNA with UI
CLSM was used to evaluate the cell uptake of FAW-SCR. As shown in Figure 5, the cell nucleus was blue in all groups, and green florescence in the perinuclear and cytoplasm was observed in the NBs-FAM-SCR-UI and FAM-SCR-UI groups, but not in the control group. However, compared to the FAM-SCR-UI group, the green florescence was higher and stronger in the NBs-FAM-SCR-UI group.
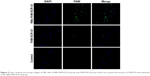  | Figure 5 Laser confocal microscopic images of U87 cells of NBs-FAM-SCR-UI group and FAM-SCR-UI group. Much more green fluorescence of FAM-SCR was observed in the NBs-FAM-SCR-UI group. |
Transfection efficiency assay by target gene expression level
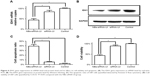
The transfection efficiency was assessed on the basis of the mRNA level by real-time PCR assay and the protein level was evaluated by Western blotting analysis. Compared with the control group, the IDH1 mRNA in both NBs-siRNA-UI and siRNA-UI groups decreased, as shown in Figure 6A, and statistical analysis showed that the IDH1 mRNA in the NBs-siRNA-UI group (44.1% ± 6.7%) is lower than that in the siRNA-UI group (84.0% ± 6.9%). Similar to the results of real-time PCR, Western blotting analysis (Figure 6B) revealed that IDH1 expression in the NBs-siRNA-UI group at the protein level was much lower than that in the siRNA-UI group, although the latter was lower as compared with that in the control group.
Cell apoptosis and viability
The cell apoptosis analysis by flow cytometry indicated that the NBs-siRNA-UI group showed a 39.4% ± 3.7% apoptosis rate, which was the highest. Additionally, the siRNA-UI and control groups only showed 5.4% ± 1.3% and 2.1% ± 0.6% apoptosis rates, respectively. Compared with the other two groups, the NBs-siRNA-UI group had the highest rate of apoptosis (Figures 6C and S2). Cell viability was measured by CCK-8 assay. The U87 glioma cells viability in the NBs-siRNA-UI group was 43.6% ± 3.6%, which was lower than that in the siRNA-UI group (89.1% ± 4.6%) and control group (100.2% ± 1.5%), as shown in Figure 6D.
NBs-siRNA distribution in vivo
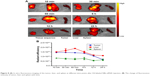
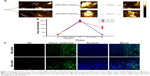
Figure 7A shows the enhanced tumor images of NBs-siRNA with or without ultrasound irradiation. Imaging contrast enhancement appeared after NBs-siRNA injection in both the UI (+) and UI (−) groups. However, the contrast enhancement in the UI (−) group was stronger than that in the control UI (+) group after ultrasound irradiation. Figure 7B shows the phase-intensity histogram, indicating that the gray scale of the UI (+) and UI (−) groups changed in different phases. A significant difference between the two groups was observed after UI (P<0.01).
The CLSM examination of the frozen tumor sections was obtained to locate DiI-labeled NBs-siRNA. The blood vessels were dyed green, whereas the nuclei were dyed blue. DiI-labeled MBs-siRNA was not detected, and NBs-siRNA was observed in the extravascular intercellular space, as shown in Figure 7C.
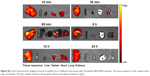
To further confirm the NBs-siRNA distribution and metabolism in tumors, a tissue fluorescence imaging experiment was conducted. The autofluorescence was low or even lack so that the fluorescence signal intensity inside the organs could reflect the NBs-siRNA content.62 Fluorescent images of the liver, spleen, kidney, heart, and lungs of healthy mice were initially observed, and the liver initially showed a strong signal that gradually decreased, whereas the spleen showed no signal (Figure S3). Based on this result, the liver and spleen were used as positive and negative controls, respectively. The fluorescence signals in the tumor, liver, and spleen were analyzed at different time points after injection. As shown in Figure 8A, the fluorescence in the tumor obviously reached a level similar to that in the liver after the injection of DiI-labeled NBs-siRNA, then gradually decreased to the level of fluorescence in the spleen after 24 hours. A region of interest (ROI) analysis was conducted to semiquantitatively assess the uptake by the tissue. Figure 8B shows the change in fluorescence intensity with time. Similar to Figure 8A, the fluorescence intensity of the tumor showed a level comparable to that in the liver at ten minutes after injection and it lasted for six hours. Then, the intensity began to decline and decreased to a level similar to that in the spleen after 24 hours.
Evaluation of therapeutic effect in vivo
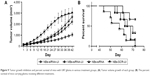
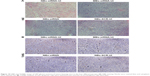
To evaluate the therapeutic effect of NBs-siRNA with UI, tumor growth was measured, and the results are shown in Figure 9A. Compared to the other three groups, the tumor volume in the NBs-siRNA-UI group was smaller (P<0.05). The cumulative survival (Figure 9B) showed that nude mice in the MBs-siRNA-UI, NBs-siRNA, and NBs-SCR-UI groups began to die on days 54, 56, and 44 days, respectively. In comparison, the NBs-siRNA-UI group showed a better therapeutic effect. The histological changes in each group were observed by H&E staining (Figure 10A). The NBs-siRNA-UI group showed fewer cells and lower levels of polymorphism, in contrast to the other three groups. The TUNEL assays were conducted to evaluate tumor apoptosis. The cell apoptosis in the NBs-siRNA-UI group was obviously higher than that in the other three groups (Figure 10B).
Discussion
As a potential carrier for drug and gene delivery, MB application in therapy has received increasing attention because the sonoporation phenomenon caused by MBs combined with UTD would improve cell membrane permeability, resulting in the increased uptake of drugs and genes.44,51–53 However, the large particle size of MBs limits their application in tumors. MBs cannot penetrate the tumor blood vessel, which has 380- to 780-nm gaps in the vessel endothelium. Therefore, based on previous studies,59,60 we combined NBs with IDH1-siRNA to form NBs carrying siRNA by a biotin–streptavidin system, which was combined with UTD to increase siRNA uptake, inhibit tumor growth, and achieve glioma treatment.
We first fabricated NBs and detected their characteristics. The particle size of NBs tested by DLS confirmed that the bubbles fabricated by using an optimized film hydration method showed a nanoscale particle size similar to that obtained in a previous study. The NBs observed by SEM were not spherical, as described in other reports,63 due to the rupture of NBs in the vacuum environment during SEM scanning. Calculations revealed that the phospholipid concentration of NBs in all experiments was less than 10 μg/mL; therefore, the NBs would not lead to cell apoptosis or decrease viability. Due to their hypocytotoxicity, DPPC and DSPE-PEG have been used as materials for MBs or NBs in many studies.59,60,63,64
The NBs-siRNA showed a similar particle size to NBs after fabrication. The stability of this complex indicated that the particle size of NBs-siRNA is more stable than the concentration. However, the particle size does not affect the application of these molecules because NBs-siRNA only needs to coexist with cells for a few minutes at room temperature. The gas core of NBs is C3F8, which is stable, with low dispersion and blood solubility, resulting in the relative stability of NBs in the blood.65
Fluorescence microscope analysis (Figure 4D and E) of the control (bare NBs) and NBs-FAM-SCR groups verified that the green fluorescence of the NBs-FAM-SCR group resulted from FAM-SCR, and not from the autofluorescence of NBs, which illustrated that the NBs were connected with siRNA by biotin–streptavidin. Additionally, fluorescence microscopy provided direct observations of NBs-siRNAs and the particle sizes based on a scale similar to the measurement of DLS. The irregular fluorescence region observed at some positions without bubbles might be a small amount of liposome residue, resulting from broken bubbles.
We assumed that the siRNA-NBs combined with UTD would improve the transfection efficiency of siRNA, which had been confirmed by CLSM, real-time PCR assay, Western blotting analysis, and cell apoptosis and viability experiments. UI could facilitate the transport of some molecules into living cells by the sonoporation phenomenon.66–69 The presence of bubbles can enhance this effect. Thus, CLSM showed that the NBs-FAM-SCR-UI group had higher and stronger green florescence compared with that of the FAM-SCR-UI group. The real-time PCR assay and Western blotting analysis confirmed the function of NBs and UI in siRNA transfection based on mRNA and protein levels, respectively. Cell apoptosis and viability were analyzed by flow cytometry and CCK-8 assay. The reduction in IDH1 expression in wild-type IDH1 cells would inhibit tumor cell growth70 by lipid production, redox homeostasis, and the regulation of cellular differentiation.
The comparison of enhanced tumor images with or without the use of UI illustrated that UI would destruct bubbles in vivo. Upon caudal vein injection, the enhancement in the tumor immediately increased, as previously reported.59,60 After ultrasound-targeted destruction, the enhanced image was significantly lower than that in the group without UI, and this effect was limited to the area of UI. The CLSM examination of tumor frozen sections confirmed the ability of NBs to enter the tumor through the vessel wall, which MBs are unable to achieve. We observed the DiI-labeled NBs-siRNA signal in the tumor, liver, and spleen at different time points after injection. The tumor signal initially corresponded to that in the liver and was similar to that in the spleen after 24 hours. NBs-siRNA accumulate in the tumor for a period of time, as NBs possess an ability to passively target the tumor, resulting from the enhanced permeability and retention (EPR) effect.71
In the in vivo experiments, the MBs-siRNA-UI and NBs-siRNA groups demonstrated that, without NBs or UI, the inhibition efficacy was reduced. A significant inhibitory effect was observed in the NBs-siRNA-UI groups, which indicated that UI could effectively promote siRNA transfection by the targeted destructed of NBs. The mechanism involves the penetration of NBs into blood vessels, followed by accumulation in the tumor and production of acoustic cavitation with UI – an effect that could enhance the permeability of cell membranes and increase siRNA intake for improved transfection and therapy.72 Additionally, targeted UI could enable siRNA to locally accumulate to a higher concentration in tumor tissues.73
Conclusion
In summary, we fabricated nanosized bubbles by a thin film hydration method, and the applied concentration showed no obvious cytotoxicity – consistent with the requirement of biosafety for siRNA carriers. NBs could be associated with siRNA by avidin–biotin to form NBs-siRNA complexes with good particle size stability. The combination of NBs-siRNA and UTD could achieve the improvement of siRNA transfection and interference efficiency in vitro. Furthermore, due to its small size, NBs-siRNA would penetrate the tumor blood vessel into the intercellular space in vivo and realize the suppression of tumor growth in vivo. NBs combined with UTD could be used as a vector for siRNA delivery for the noninvasive treatment of glioma.
Acknowledgments
The present study was financially supported by a grant from the National Natural Science Foundation of China (grant no 81301227). The authors thank Dr Gang Zhu (Neurosurgery Department, Tangdu Hospital) for assistance with the U87 cell culture.
Disclosure
The authors report no conflicts of interest in this work
References
Biddlestone-Thorpe L, Sajjad M, Rosenberg E, et al. ATM kinase inhibition preferentially sensitizes p53-mutant glioma to ionizing radiation. Clin Cancer Res. 2013;19(12):3189–3200. | ||
Chirio D, Gallarate M, Peira E, et al. Positive-charged solid lipid nanoparticles as paclitaxel drug delivery system in glioblastoma treatment. Eur J Pharm Biopharm. 2014;88(3):746–758. | ||
Ohgaki H, Kleihues P. Epidemiology and etiology of gliomas. Acta Neuropathol. 2005;109(1):93–108. | ||
Wang SC, Wu CC, Wei YY, Hong JH, Chiang CS. Inactivation of ataxia telangiectasia mutated gene can increase intracellular reactive oxygen species levels and alter radiation-induced cell death pathways in human glioma cells. Int J Radiat Biol. 2011;87(4):432–442. | ||
Dy GK, Adjei AA. Understanding, recognizing, and managing toxicities of targeted anticancer therapies. CA Cancer J Clin. 2013;63(4):249–279. | ||
Bumcrot D, Manoharan M, Koteliansky V, Sah DW. RNAi therapeutics: a potential new class of pharmaceutical drugs. Nat Chem Biol. 2006;2(12):711–719. | ||
Carstea ED, Hough S, Wiederholt K, Welch PJ. State-of-the-art modified RNAi compounds for therapeutics. IDrugs. 2005;8(8):642–647. | ||
Guo J, Bourre L, Soden DM, O’Sullivan GC, O’Driscoll C. Can non-viral technologies knockdown the barriers to siRNA delivery and achieve the next generation of cancer therapeutics? Biotechnol Adv. 2011;29(4):402–417. | ||
Tatiparti K, Sau S, Kashaw SK, Iyer AK. siRNA Delivery Strategies: A Comprehensive Review of Recent Developments. Nanomaterials. 2017;7(4):E77:77. | ||
Agrawal N, Dasaradhi PVN, Mohmmed A, et al. Mechanism, and Applications. Microbiology and Molecular Biology Reviews. 2003;67(4):657–685. | ||
van de Veire S, Stalmans I, Heindryckx F, et al. Further pharmacological and genetic evidence for the efficacy of PlGF inhibition in cancer and eye disease. Cell. 2010;141(1):178–190. | ||
Zhao M, Sun J, Zhao Z. Synergetic regulatory networks mediated by oncogene-driven microRNAs and transcription factors in serous ovarian cancer. Mol Biosyst. 2013;9(12):3187–3198. | ||
Davis ME, Zuckerman JE, Choi CH, et al. Evidence of RNAi in humans from systemically administered siRNA via targeted nanoparticles. Nature. 2010;464(7291):1067–1070. | ||
Vaishnaw AK, Gollob J, Gamba-Vitalo C, et al. A status report on RNAi therapeutics. Silence. 2010;1(1):14. | ||
Wang J, Mi P, Lin G, Wáng YX, Liu G, Chen X. Imaging-guided delivery of RNAi for anticancer treatment. Adv Drug Deliv Rev. 2016;104:44–60. | ||
Kim HJ, Kim A, Miyata K, Kataoka K. Recent progress in development of siRNA delivery vehicles for cancer therapy. Adv Drug Deliv Rev. 2016;104:61–77. | ||
Whitehead KA, Langer R, Anderson DG. Knocking down barriers: advances in siRNA delivery. Nat Rev Drug Discov. 2009;8(2):129–138. | ||
Kim T, Hyeon T. Applications of inorganic nanoparticles as therapeutic agents. Nanotechnology. 2014;25(1):012001. | ||
Mody VV, Cox A, Shah S, Singh A, Bevins W, Parihar H. Magnetic nanoparticle drug delivery systems for targeting tumor. Appl Nanosci. 2014;4(4):385–392. | ||
Cheng D, Cao N, Chen J, Yu X, Shuai X. Multifunctional nanocarrier mediated co-delivery of doxorubicin and siRNA for synergistic enhancement of glioma apoptosis in rat. Biomaterials. 2012;33(4):1170–1179. | ||
Kim JS, Oh MH, Park JY, Park TG, Nam YS. Protein-resistant, reductively dissociable polyplexes for in vivo systemic delivery and tumor-targeting of siRNA. Biomaterials. 2013;34(9):2370–2379. | ||
Kwok A, Hart SL. Comparative structural and functional studies of nanoparticle formulations for DNA and siRNA delivery. Nanomedicine. 2011;7(2):210–219. | ||
Truong NP, Gu W, Prasadam I, et al. An influenza virus-inspired polymer system for the timed release of siRNA. Nat Commun. 2013;4:4. | ||
Yin L, Song Z, Kim KH, Zheng N, Gabrielson NP, Cheng J. Non-viral gene delivery via membrane-penetrating, mannose-targeting supramolecular self-assembled nanocomplexes. Adv Mater. 2013;25(22):3063–3070. | ||
Huang L, Liu Y. In vivo delivery of RNAi with lipid-based nanoparticles. Annu Rev Biomed Eng. 2011;13:507–530. | ||
Prakash TP, Lima WF, Murray HM, et al. Lipid nanoparticles improve activity of single-stranded siRNA and gapmer antisense oligonucleotides in animals. ACS Chem Biol. 2013;8(7):1402–1406. | ||
Rahman MA, Amin AR, Wang X, et al. Systemic delivery of siRNA nanoparticles targeting RRM2 suppresses head and neck tumor growth. J Control Release. 2012;159(3):384–392. | ||
Hutson TH, Foster E, Moon LD, Yáñez-Muñoz RJ. Lentiviral vector-mediated RNA silencing in the central nervous system. Hum Gene Ther Methods. 2014;25(1):14–32. | ||
Ballarín-González B, Howard KA. Polycation-based nanoparticle delivery of RNAi therapeutics: adverse effects and solutions. Adv Drug Deliv Rev. 2012;64(15):1717–1729. | ||
Li W, Hu ZF, Chen B, Ni GX, Zf H, Gx N. Response of C2C12 myoblasts to hypoxia: the relative roles of glucose and oxygen in adaptive cellular metabolism. Biomed Res Int. 2013;2013:326346–10. | ||
Aronovich EL, Mcivor RS, Hackett PB. The Sleeping Beauty transposon system: a non-viral vector for gene therapy. Hum Mol Genet. 2011;20(R1):R14–20. | ||
Huang S, Kamihira M. Development of hybrid viral vectors for gene therapy. Biotechnol Adv. 2013;31(2):208–223. | ||
Fokong S, Theek B, Wu Z, et al. Image-guided, targeted and triggered drug delivery to tumors using polymer-based microbubbles. J Control Release. 2012;163(1):75–81. | ||
Marxer EE, Brüssler J, Becker A, et al. Development and characterization of new nanoscaled ultrasound active lipid dispersions as contrast agents. Eur J Pharm Biopharm. 2011;77(3):430–437. | ||
Wilson SR, Burns PN. Microbubble-enhanced US in body imaging: what role? Radiology. 2010;257(1):24–39. | ||
Banche G, Prato M, Magnetto C, et al. Antimicrobial chitosan nanodroplets: new insights for ultrasound-mediated adjuvant treatment of skin infection. Future Microbiol. 2015;10(6):929–939. | ||
Basilico N, Magnetto C, D’Alessandro S, et al. Dextran-shelled oxygen-loaded nanodroplets reestablish a normoxia-like pro-angiogenic phenotype and behavior in hypoxic human dermal microvascular endothelium. Toxicol Appl Pharmacol. 2015;288(3):330–338. | ||
Gulino GR, Magnetto C, Khadjavi A, et al. Oxygen-Loaded Nanodroplets Effectively Abrogate Hypoxia Dysregulating Effects on Secretion of MMP-9 and TIMP-1 by Human Monocytes. Mediators Inflamm. 2015;2015:964838. | ||
Khadjavi A, Magnetto C, Panariti A, et al. Chitosan-shelled oxygen-loaded nanodroplets abrogate hypoxia dysregulation of human keratinocyte gelatinases and inhibitors: New insights for chronic wound healing. Toxicol Appl Pharmacol. 2015;286(3):198–206. | ||
Prato M, Khadjavi A, Magnetto C, et al. Effects of oxygen tension and dextran-shelled/2H,3H-decafluoropentane-cored oxygen-loaded nanodroplets on secretion of gelatinases and their inhibitors in term human placenta. Biosci Biotechnol Biochem. 2016;80(3):466–472. | ||
Cosgrove D. Ultrasound contrast agents: an overview. Eur J Radiol. 2006;60(3):324–330. | ||
O’Brien WD. Ultrasound-biophysics mechanisms. Prog Biophys Mol Biol. 2007;93(1–3):212–255. | ||
Marmottant P, Hilgenfeldt S. Controlled vesicle deformation and lysis by single oscillating bubbles. Nature. 2003;423(6936):153–156. | ||
Mccreery TP, Sweitzer RH, Unger EC, Sullivan S. DNA delivery to cells in vivo by ultrasound. Methods Mol Biol. 2004;245:293–298. | ||
Miller DL, Pislaru SV, Greenleaf JE. Sonoporation: mechanical DNA delivery by ultrasonic cavitation. Somat Cell Mol Genet. 2002;27(1–6):115–134. | ||
Ohl CD, Arora M, Ikink R, et al. Sonoporation from jetting cavitation bubbles. Biophys J. 2006;91(11):4285–4295. | ||
Tachibana K, Uchida T, Ogawa K, Yamashita N, Tamura K. Induction of cell-membrane porosity by ultrasound. Lancet. 1999;353(9162):1409. | ||
Wu G, Mikhailovsky A, Khant HA, Fu C, Chiu W, Zasadzinski JA. Remotely triggered liposome release by near-infrared light absorption via hollow gold nanoshells. J Am Chem Soc. 2008;130(26):8175–8177. | ||
Wu J. Shear stress in cells generated by ultrasound. Prog Biophys Mol Biol. 2007;93(1–3):363–373. | ||
Zarnitsyn V, Rostad CA, Prausnitz MR. Modeling transmembrane transport through cell membrane wounds created by acoustic cavitation. Biophys J. 2008;95(9):4124–4138. | ||
Korpanty G, Chen S, Shohet RV, et al. Targeting of VEGF-mediated angiogenesis to rat myocardium using ultrasonic destruction of microbubbles. Gene Ther. 2005;12(17):1305–1312. | ||
Newman CM, Bettinger T. Gene therapy progress and prospects: ultrasound for gene transfer. Gene Ther. 2007;14(6):465–475. | ||
Unger EC, Hersh E, Vannan M, Matsunaga TO, Mccreery T. Local drug and gene delivery through microbubbles. Prog Cardiovasc Dis. 2001;44(1):45–54. | ||
Hauff P, Seemann S, Reszka R, et al. Evaluation of gas-filled microparticles and sonoporation as gene delivery system: feasibility study in rodent tumor models. Radiology. 2005;236(2):572–578. | ||
Suzuki R, Maruyama K. Effective in vitro and in vivo gene delivery by the combination of liposomal bubbles (bubble liposomes) and ultrasound exposure. Methods Mol Biol. 2010;605:473–486. | ||
Brigger I, Dubernet C, Couvreur P. Nanoparticles in cancer therapy and diagnosis. Adv Drug Deliv Rev. 2002;54(5):631–651. | ||
Hobbs SK, Monsky WL, Yuan F, et al. Regulation of transport pathways in tumor vessels: role of tumor type and microenvironment. Proc Natl Acad Sci U S A. 1998;95(8):4607–4612. | ||
Ferrara K, Pollard R, Borden M. Ultrasound microbubble contrast agents: fundamentals and application to gene and drug delivery. Annu Rev Biomed Eng. 2007;9:415–447. | ||
Cai WB, Yang HL, Zhang J, et al. The Optimized Fabrication of Nanobubbles as Ultrasound Contrast Agents for Tumor Imaging. Sci Rep. 2015;5:13725. | ||
Yang H, Cai W, Xu L, et al. Nanobubble-Affibody: Novel ultrasound contrast agents for targeted molecular ultrasound imaging of tumor. Biomaterials. 2015;37:279–288. | ||
Rapoport NY, Nam KH, Gao Z, Kennedy A. Application of Ultrasound for Targeted Nanotherapy of Malignant Tumors. Acoust Phys. 2009;55(4–5):594–601. | ||
Gao J, Chen K, Xie R, et al. In vivo tumor-targeted fluorescence imaging using near-infrared non-cadmium quantum dots. Bioconjug Chem. 2010;21(4):604–609. | ||
Yin T, Wang P, Zheng R, et al. Nanobubbles for enhanced ultrasound imaging of tumors. Int J Nanomedicine. 2012;7:895–904. | ||
Yin T, Wang P, Li J, et al. Tumor-penetrating codelivery of siRNA and paclitaxel with ultrasound-responsive nanobubbles hetero-assembled from polymeric micelles and liposomes. Biomaterials. 2014;35(22):5932–5943. | ||
Abou-Elkacem L, Bachawal SV, Willmann JK. Ultrasound molecular imaging: Moving toward clinical translation. Eur J Radiol. 2015;84(9):1685–1693. | ||
de Temmerman ML, Dewitte H, Vandenbroucke RE, et al. mRNA-Lipoplex loaded microbubble contrast agents for ultrasound-assisted transfection of dendritic cells. Biomaterials. 2011;32(34):9128–9135. | ||
Frenkel V. Ultrasound mediated delivery of drugs and genes to solid tumors. Adv Drug Deliv Rev. 2008;60(10):1193–1208. | ||
Karshafian R, Samac S, Bevan PD, Burns PN. Microbubble mediated sonoporation of cells in suspension: clonogenic viability and influence of molecular size on uptake. Ultrasonics. 2010;50(7):691–697. | ||
Lee JL, Lo CW, Ka SM, Chen A, Chen WS. Prolonging the expression duration of ultrasound-mediated gene transfection using PEI nanoparticles. J Control Release. 2012;160(1):64–71. | ||
Calvert AE, Chalastanis A, Wu Y, et al. Cancer-Associated IDH1 Promotes Growth and Resistance to Targeted Therapies in the Absence of Mutation. Cell Rep. 2017;19(9):1858–1873. | ||
Iyer AK, Khaled G, Fang J, Maeda H. Exploiting the enhanced permeability and retention effect for tumor targeting. Drug Discov Today. 2006;11(17–18):812–818. | ||
Oeffinger BE, Wheatley MA. Development and characterization of a nano-scale contrast agent. Ultrasonics. 2004;42(1–9):343–347. | ||
Wang L, Zhang M, Tan K, et al. Preparation of nanobubbles carrying androgen receptor siRNA and their inhibitory effects on androgen-independent prostate cancer when combined with ultrasonic irradiation. PLoS One. 2014;9(5):e96586. |
Supplementary materials
  | Table S1 IDH1 siRNA and scrambled siRNA sequence |
  | Table S2 Primers used in real-time PCR for IDH1 and GAPDH |
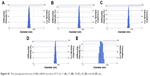  | Figure S1 The average particle size of NBs-siRNA stored at 25°C for 1 (A), 15 (B), 30 (C), 45 (D), and 60 (E) min. |
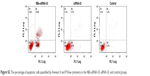  | Figure S2 The percentage of apoptotic cells quantified by Annexin V and PI flow cytometry in the NBs-siRNA-UI, siRNA-UI, and control groups. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.