Back to Journals » Cancer Management and Research » Volume 12
The Role of Prefoldin and Its Subunits in Tumors and Their Application Prospects in Nanomedicine
Authors Mo S, Zhao H , Tian Y, Zhao HL
Received 1 July 2020
Accepted for publication 24 August 2020
Published 23 September 2020 Volume 2020:12 Pages 8847—8856
DOI https://doi.org/10.2147/CMAR.S270237
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Chien-Feng Li
Shao-jian Mo,1 Hai-Chao Zhao,1 Yan-zhang Tian,2 Hao-Liang Zhao2
1Department of General Surgery, The Affiliated Bethune Hospital of Shanxi Medical University, Taiyuan 030032, People’s Republic of China; 2Department of General Surgery, Shanxi Bethune Hospital, Shanxi Academy of Medical Sciences, Taiyuan 030032, People’s Republic of China
Correspondence: Yan-zhang Tian
Department of General Surgery, Shanxi Bethune Hospital, Shanxi Academy of Medical Sciences, Taiyuan 030032, People’s Republic of China
Email [email protected]
Abstract: Prefoldin (PFDN) is a hexameric chaperone complex that is widely found in eukaryotes and archaea and consists of six different subunits (PFDN1-6). Its main function is to transfer actin and tubulin monomers to the eukaryotic cell cytoplasmic chaperone protein (c-CPN) specific binding during the assembly of the cytoskeleton, to stabilize the newly synthesized peptides so that they can be folded correctly. The current study found that each subunit of PFDN has different functions, which are closely related to the occurrence, development and prognosis of tumors. However, the best characteristics of each subunit have not been fully affirmed. The connection between research and tumors can change the understanding of PFDN and further extend its potential prognostic role and structural function to cancer research and clinical practice. This article mainly reviews the role of canonical PFDN and its subunits in tumors and other diseases, and discusses the potential prospects of the unique structure and function of PFDN in nanomedicine.
Keywords: prefoldin, disease, tumor, nanomedicine
Introduction
Prefoldin, a hexameric chaperone complex, is widely found in eukaryotes and archaea. Its main function is to stabilize newly synthesized peptides, promote the correct folding of cytoskeletal proteins, and prevent protein misfolding and aggregation.1 This function is particularly important for maintaining protein homeostasis, and its abnormality can lead to protein aggregation and induce various diseases. Each subunit of prefoldin has different functions, and it is abnormally expressed in many tumors, which is closely related to the occurrence and development of tumors. This abnormal expression can also be used as a biomarker to indicate the prognosis of the disease, which is of great significance in assisting the decision-making of disease treatment.2,3 In terms of structure, prefoldin has a special and flexible hydrophobic protein binding region, which can capture proteins. Therefore, it can be used as a nanobiological template and designed as a nanoactuator to capture nanocargoes with different charges and various sizes. It can be used to treat diseases in the field of nanomedicine.4
Therefore, prefoldin and prefoldin-subunits are essential for life activities and exploration of the occurrence and development of tumors. In addition, its special structure and function are an excellent choice for designing nano-actuators (drug delivery), opening up a new path for the hot emerging field of bio-nanorobots. Based on these unique advantages of prefoldin, it is necessary to have a detailed understanding of prefoldin and prefoldin-subunits.
The Prefoldin Complex
The Composition and Structure of Prefoldin
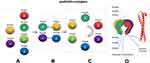
In 1998, Irina E Vainberg1 discovered a novel chaperone protein Prefoldin (PFDN, originally called GimC), which is a hexameric chaperone complex that can assist the transfer of actin and tubulin to eukaryotic cytoplasmic chaperone proteins (named TRiC/CCT, also called c-CPN), and then promote protein folding. In eukaryotes, canonical PFDN hexamer is composed of two different α subunits (PFDN3 and PFDN5) and four different β subunits (PFDN6, PFDN4, PFDN2 and PFDN1).5 Each subunit interacts with each other at the protein level to form a PFDN complex. First, PFDN2-PFDN3 and PFDN5-PFDN6 form sub-complexes and assemble together, then PFDN1 and PFDN4 are recruited to these sub-complexes, and finally form a clockwise order of PFDN3, PFDN2, PFDN1, PFDN5, PFDN6 and PFDN4 hexamer complex.6 The hexamer complex has a unique quaternary structure, similar in shape to jellyfish (Figure 1). In the PFDN complex, a large central cavity is surrounded by six long tentacles, which are formed by rods of an α spiral coil. The coiled coil tentacles point to a platform that is composed of two eight-strand β-barrels. Each PFDN subunits forms a coiled coil and the coil is firmly fixed on the platform through its proximal end, the distal regions of the coiled coils expose hydrophobic sites, which is a multivalent binding site for nonnative proteins. However, there is virtually no interaction between the coiled coils of each subunit.7 Interestingly, the expression levels of the PFDN subunits are mutually regulated. This regulation is carried out at the protein level by the ubiquitin-proteasome system. The PFDN subunits escape the degradation of the ubiquitin-proteasome system through the coiled coil structure between subunits.6 When a subunit is knocked down, the protein level of other subunits is reduced, and there are specific combinations that reduce the expression level between these subunits, which implies the mutual regulation of protein levels between PFDN subunits.
The Function of Prefoldin to Maintain Protein Homeostasis and Facilitate the Folding of Cytoskeletal Proteins
In eukaryotic cells, the homeostasis of proteins depends on chaperones which it is essential for many proteins to perform their life activities in cells.8,9 As a chaperone, PFDN can mediate the folding of nascent proteins or misfolded proteins. During protein synthesis or denaturation under cellular stress, it selectively interacts with unfolded hydrophobic peptides, folds and stabilizes them. The function of PFDN is mainly to interact with c-CPN to promote the folding.1,10–13 The c-CPN complex is involved in protein folding in about 10% of the cytoplasm, and the PFDN is necessary to transport unfolded proteins to the complex.8
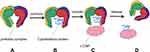
Unlike other chaperones (such as heat shock proteins), the PFDN promotes protein folding without regulating the binding and release of substrates by ATP binding and hydrolysis.1,5 In participating the assembly process of the cytoskeleton, the PFDN tentacles have great flexibility, which enables the hexamer to adapt to the different space requirements of the substrate molecule. The combined substrate extends into the central cavity formed by the coiled tentacles7 to form a binary complex, when PFDN binds to c-CPN, it quickly release the substrate to transfer to c-CPN, stabilize the newly synthesized peptides, so that they can fold correctly (Figure 2). Actin and tubulin are the main substrates of c-CPN and PFDN1,10 in eukaryotic organisms. In the PFDN-c-CPN system, the specific interaction between the two partners may help provide a suitable environment in which nonnative peptides can be effectively folded without misfolding and aggregation.1,11 If disrupting the PFDN-c-CPN interaction in the body, it will be very harmful and will lead to the accumulation of amyloid aggregates, so this system is essential to prevent toxic conformations and ensure effective cellular protein stability.14
PFDN can stabilize many nonnative proteins and prevent aggregation. When a subunit of the complex is deleted, it can cause cytoskeletal defects and protein aggregation.1,5 In addition, abnormal expression of PFDN can cause many diseases, so eliminating misfolded proteins to mitigate the harmful effects, thereby inhibiting protein toxicity and disease development, it is necessary to conduct in-depth research on PFDN. In recent years, studies have found that the function of the prefoldin-subunits differs from that of prefoldin complex, they can cause many diseases (Table 1). The following sections will introduce the role of the PFDN complex and prefoldin-subunits in disease.
 |
Table 1 Summary Table of the Correlation Between Prefoldin and disease |
Function of Prefoldin Subunits in Diseases
PFDN1 and EMT, Cell Cycle, and Cell Metabolism
PFDN1 is a 122 amino acid protein with a cytogenetic band of 5q31.3 and a molecular mass of 14,210 Dalton. Study has found that PFDN1 is associated with invasion, cell cycle and cell metabolism.15 As we all know, epithelial–mesenchymal transition (EMT) exists in many tumors, which play an important role in tumor invasion and metastasis. Existing reports show that PFDN1 can activate EMT to promote the migration, invasion and metastasis gastric cancer,3 lung cancer,16 breast cancer.17 PFDN1 expression is significantly upregulated in gastric cancer tissues, and this upregulation is associated with tumor metastasis and poor prognosis. Study on its mechanism found that PFDN1 promotes the migration, invasion and metastasis of gastric cancer cells by activating WMT/β-catenin signaling-mediated EMT.3 During the progression of lung cancer, PFDN1 acts as a tumor promoter and directly interacts with the cyclin A promoter at the transcription initiation site to inhibit the expression of cyclin A, thereby inducing EMT and increasing the ability of cell migration and invasion.16 Recent studies have shown that overexpression of PFDN is associated with mortality and metastatic risk of non-small cell lung cancer, especially when PFDN1 is overexpressed, it is more prone to distant metastasis.18 Another in vitro cell study found19 that PFDN1 promotes EMT transformation of breast cancer BT474 cells and affects their ability to invade and migrate, but its mechanism has not been elucidated. Other studies have shown that depletion of cyclin A2 can lead to increase cytoskeletal rearrangement and cell migration in normal mammary epithelial cells.17 On the other hand, knockdown of cyclin A2 can induce mammary epithelial cells EMT.20 From this point, the mechanism may be the same as in activating lung cancer EMT. Since cytoskeletal rearrangement occurs during the EMT process, and PFDN1 can promote EMT, so the mechanism of PFDN1 promotes EMT may be through affecting cytoskeletal reorganization.
In colorectal cancer, silence of PFDN1 inhibits the G2/M cell cycle and results in dysfunction of the cytoskeleton system, thereby inhibiting tumor cell growth and movement. It is also found that PFDN1 expression is positively correlated with colorectal tumor size and invasion that is indicator of poor prognosis.21 Cyclin A2 is known to be associated with cyclin-dependent kinases CDK2 and CDK1, and is a regulator of S-phase progression and G2/M transition.22 Whether PFDN1 regulates colorectal cancer through cyclin A2 has not been determined yet.
In addition, the lack of PFDN1 will cause the phenotype of cytoskeletal function defects, which can lead to ciliary dyskinesia, neuronal loss, and the development and function of B and T cells, indicating that it is also necessary for lymphocyte development and function.23
PFDN2 Can Be Used as a Tumor Biomarker
PFDN2 is a 154 amino acid protein with a cytogenetic band of 1q23.3 and a molecular mass of 16,648 Dalton. It was found that PFDN2 is also closely related to tumors and can be used as a biological indicator of some diseases. For example, López Virginia24 and Markus Riester25 have successively found that the 1q23.3 genome has changed in bladder tumors, and determined that the PFDN2 is a prognostic biomarker for bladder tumor stratification and clinical outcome. The expression of PFDN2 in various subtypes of gastric cancer shows poor overall survival and is a biological indicator of poor prognosis.26 In type 2 diabetes, anti-PFDN2 antibodies may be useful biomarkers for the disease.27
PFDN2 has also undergone genetic changes in breast cancer.28 This change may make taxane chemotherapeutics resistant. Because PFDN2 is responsible for the folding of tubulin, its mutation or abnormal function may cause the misfolding of tubulin. Taxane is a type of chemotherapeutic agent, mainly by stabilizing microtubules to destroy the mitotic spindle, thereby inhibiting cell division to play an anticancer effect, so abnormal PFDN2 can make breast cancer resistant to taxane treatment.
Chang KW29 discovered a κ-actin, which is expressed in hepatocellular carcinoma tissues, replaces β-actin and becomes the main component of the actin cytoskeleton, resulting in a greatly reduced interaction between actin and PFDN2, and the reduction in interaction may cause poor cytoskeletal organization and change the biological characteristics of liver cancer, thereby promoting the growth, proliferation or metastasis of abnormal cells. It has been reported30 that the interaction of hepatitis C virus (HCV) F protein with PFDN2 disrupts the normal function of the PFDN complex, leading to abnormal of the tubulin cytoskeleton, and viruses in HCV infection persist. It has also been reported that PFDN2 plays an important role in the development of chronic HCV infection and liver disease.31 The long-term hepatitis can cause liver cancer, so there may be some interaction between PFDN2 and liver cancer to affect tumors. Recent studies have found32 that various subunits of PFDN have a high affinity with HDAC1 in hepatocellular carcinoma HepG2 cells, which further provides a basis for this hypothesis. PFDN2 has an effect on liver cancer, but the underlying mechanism still needs to be determined.
PFDN3 and Protein Degradation
PFDN3 is a 197 amino acid protein with a cytogenetic band of Xq 28 and a molecular mass of 22,626 Dalton. PFDN3 is necessary to maintain the stability of Von Hippel-Lindau (VHL).33 Known as Von Hippel-Lindau binding protein 1 (VBP1),34 PFDN3 interacts with VHL protein to form a complex that can promote the stability of microtubules and regulate protein stability.35 VHL is a tumor suppressor protein, which can interact with c-CPN to mediate the folding and assembly of tumor suppressor complexes.36–38 In addition, PFDN3 requires c-CPN to assist the function of protein folding. Therefore, c-CPN, VHL, and PFDN may have a certain relationship to regulate tumor formation.
The product of the VHL tumor suppressor gene, pVHL, is the active component of E3 ubiquitin protein ligase, and there is a direct connection with the ubiquitination process.39 Therefore, in addition to interacting with c-CPN and promoting the folding of cytoskeletal proteins, PFDN3 is also suggested to participate in the ubiquitination process. Study has demonstrated that PFDN3 can degrade hMSH4 in the human embryonic kidney cell line HEK293T through ubiquitination.40 Another study found41 that during the integration-transcriptional transition of the HIV replication cycle, integrase degrades through the PFDN3-vhl-proteasome pathway, affecting HIV-1 gene expression. It is believed that PFDN3 is the binding partner of HIV-1 viral integrase and can serve as a bridge connecting the interaction between cullin2 and VHL ubiquitin ligase, so that HIV-1 integrase integrates the provirus into the host genome and viral gene transcription. On the other hand, VHL mediates the degradation of HIF-1α by the ubiquitin-proteasome pathway,42 and PFDN3 can inhibit tumor metastasis by enhancing pVHL-induced degradation of HIF-1α,43 so PFDN3 ubiquitin-degraded proteins are also involved in tumors Progress. In summary, PFDN3 is involved in protein degradation.
Function of PFDN4 and PFDN6 in Tumors and Transcriptional Regulation
PFDN4 is a 134 amino acid protein with a cytogenetic band of 20q13.2 and a molecular mass of 15,314 Dalton. PFDN4, also known as PFD4 or C1, is a transcription factor that regulates the cell cycle.44 PFDN4 may be up-regulated as an activated transcription factor in B-cell non-Hodgkin’s lymphoma.45 Sequence analysis of breast cancer genomes shows that46 PFDN4 is up-regulated in breast cancer and may promote cancer progression. At the same time, it was found to be abnormally expressed in colorectal cancer and hepatocellular carcinoma. When its low expression indicates a poor prognosis, and may be related to metastasis, it can be considered as a prognostic indicator.47,48
PFDN6 is a 129 amino acid protein with a cytogenetic band of 6p21.32 and a molecular mass of 14,583 Dalton. HKE2 is a gene encoding PFDN6,1 and some studies believe that49 It plays a role in adaptive immunity and cancer, which means that PFDN6 is also closely related to tumors. However, there is little research about PFDN6 and the only study about it showed that50 it is related to cancer, suggesting that it may be a prognostic biomarker for acute lymphoblastic leukemia in children. In addition, studies have found51 that PFDN6 can mediate the effect of thermal shock factor 1 on FOXO transcriptional activity and increase the lifespan of C. elegans, indicating that PFDN6 may also play a role in transcriptional regulation.
Taken together, these studies show that PFDN4 and PFDN6 are involved in transcriptional regulation and tumors, but the underlying mechanisms are still unclear, and it would be interesting to investigate in the future.
PFDN5 and Diseases
PFDN5 and Neurological Diseases
PFDN5 (also called MM-1) is a 154 amino acid protein with a cytogenetic band of 12q13.13 and a molecular mass of 17,328 Dalton. Abnormal PFDN can lead to protein misfolding and diseases, of which the most studied is in the nervous system diseases, which are characterized by the accumulation of misfolded proteins. PFDN plays a role in controlling protein aggregation, and dysfunction of PFDN is one of the causes of neurodegenerative diseases, which can protect cells from protein aggregation-induced cell death.52 For example, in Parkinson’s disease53 and Huntington’s disease,54 the knockdown of PFDN2 and PFDN5 destroys the formation of PFDN in cells, leading to protein accumulation and inducing cell death, indicating PFDN plays a protective role in cell death. Other data show that genetic variation in the PFDN family is related to the risk of Alzheimer’s disease.55 PFDN1 is an Alzheimer’s disease-associated gene,56 and the lack of PFDN in Drosophila causes the polarity of neuroblasts to be destroyed, causing the neuroblasts in the brain to overgrow.57 PFDN3 can promote the development of neuroblastoma.58 Recent studies have pointed out that PFDN5 is the main pathogenic gene of Alzheimer’s disease.59 Although each subunit of PFDN may affect the nervous system, PFDN5 may be the most important one. First, normal sensory and neuronal development require PFDN5,60 and secondly, the level of PFDN5 protein in mouse hippocampal neurons can be changed according to synaptic plasticity, which is considered to be one of the important components of synaptic plasticity.61 Synaptic plasticity is considered to be the basis of plasticity changes related to learning and memory. Degenerative diseases of the nervous system (such as Alzheimer’s disease) are mainly based on progressive memory impairment, so PFDN5 may be the most important gene in the nervous system diseases.
PFDN5 as a Tumor Suppressor Gene
In addition to its important role in neurological diseases, whether PFDN5, like other subunits, plays a role in tumors? It is said that PFDN5 gene is up-regulated in pancreatic cancer.62 Studies have found that PFDN5 has a potential carcinogenic effect in canine breast tumors and may be a tumor driver gene.63 However, its specific mechanism of action is still poorly understood. The c-MYC is a classic proto-oncogene product, and PFDN5 was previously identified as a c-MYC binding protein by Mori K.64 PFDN5 can be used as a connection factor of c-Myc transcription repression pathway.65 Therefore, it can be considered that PFDN5 has an inhibitory effect on c-Myc and acts as a potential tumor suppressor gene. Existing studies have found the inhibitory mechanism of PFDN5 and c-Myc on tumors. For example, PFDN5 negatively regulates the Wnt signaling pathway (which is a non-classical Wnt pathway) by suppressing the expression of wnt4 gene and also through the classic Wnt-β-catenin pathway. This dual mechanism inhibits c-Myc, thereby inhibiting tumors;66 on the other hand, it combines with the N-terminal reverse transcription region of c-Myc to play an inhibitory role.67 Studies have shown that the role of MM-1 in controlling the transcriptional activity of c-Myc and inhibiting tumors can be used as a drug candidate for cancer suppressors in leukemia/lymphoma and tongue cancer.68
Prefoldin and Nanotechnology
Prefoldin Can Be Used as a Nano-Actuator to Capture and Load Cargo
Nanotechnology uses protein as a template to construct advanced biomaterials to achieve precise molecular positioning, which has great potential. PFDN does not require ATP to capture and transfer proteins. Its substrate-binding cavity is surrounded by six long tentacles to form a hydrophobic area. These six tentacles have high flexibility and can swing to change the size of the cavity to adapt to the size of the substrate.1,5,7 This unique structure is an excellent choice for designing nanotechnology protein templates. With further research,69–72 it was discovered that PFDN has the ability to act as a nano-component in bio-nano-robots, and was gradually designed as a prefoldin nano-actuator (PNA) to act as a drug delivery tool and carrier.4,69,73,74
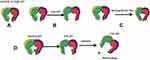
The six tentacles of the PNA make it very flexible. When the temperature and pH of the environmental medium are changed, the overall structure and space size of the PNA are significantly affected, and the combination and release of nano-cargo are controlled. In alkaline and neutral pH environments, the PNA cavity size is basically the same. When the pH decreases, the cavity area increases, and as the temperature increases, the PNA cavity area decreases. This change belongs to the change of conformation, which is caused by the interaction of Prefoldin subunits with each other and with the solvent, and it has no effect on the performance of PNA. The conformational change controls the opening and closing of the substrate-binding cavity and determines the size of the nano-cargo that PNA can carry (Figure 3). Combining these two control parameters can provide a suitable environment to capture nano-cargo with specific dimensions for drug delivery in vivo.4,73
PNA captures nano-cargo, mainly relying on the hydrophobic interaction with the cargo to attract the cargo and maintain the established stability, and secondly rely on the hydrogen bond formed between them.75 Molecular dynamics method found that mutant type PNA tentacles (namely, PFDN hydrophobic amino acids are replaced by hydrophilic amino acids to enhance the role of tentacles in PNA) can better capture cargo and have better stability.76 Wild-type PNA (unmutated) has only two tentacles that can capture cargo, while four of the six tentacles of the mutated PFDN can effectively attract cargo. This mutant PNA can be used as a potential tool to suppress pathogenic factors and transfer hydrophobic cargo.75 In addition, the mutant PNA changes the hydrophobic interaction with the cargo and the number of hydrogen bonds, so that it also captures the positively charged cargo in a neutral environment and releases it in an acidic environment.74
The Possible Application of PNA in Tumor Treatment
Recent studies have assembled PFDN and connexin into a nanostructure of macromolecules with a defined shape, which can be used as therapeutic drug delivery and inhibit pathogenic factors.71,75,76 Masoud et al77 used PNA as an inhibitor of Aβ oligomers to treat Alzheimer’s disease, they found that it has a higher efficacy. Therefore, PFDN nano-drug delivery system can achieve disease treatment. Does this also mean that it brings new opportunities for tumor treatment?
Nano-actuators play a role in delivering drugs to treat diseases in nanomedicine.78 It can transfer the drug to the target area without damaging the drug and other cells, minimize the adverse toxicity during the treatment process, improve the utilization rate of the drug, and provide a new strategy for tumor chemotherapy, radiotherapy, and immunotherapy. Therefore, it has a better curative effect in overcoming the multi-drug resistance and systemic toxicity of anti-tumor79–82
The PNA nano drug delivery system is a stimulus-responsive drug delivery system that controls the binding and release of drugs under the influence of the temperature and pH of the environment medium (especially at low pH).4 Studies have pointed out that83 the difference in pH between normal tissues and tumor tissues can become specific targets for cancer treatment. The endogenous stimuli of tumors, including low pH and temperature rise caused by inflammation, are different from normal tissues.84,85 This abnormality can become a specific target for cancer treatment and cause the release of drugs. By regulating the temperature and pH parameters of PNA binding and releasing cargo, and controlling the binding of drugs and the release of specific locations, precise treatment of tumors can be achieved. Since mutant PNA can regulate these two parameters, the future application of PNA to tumor therapy has great prospects. The current PNA research76,86,87 is mainly to understand the mechanical properties of PNA and the treatment of neurological diseases, paving the way for the application of PNA in intelligent drug delivery and capturing pathogenic cargo. In the future, a large number of researchers are still needed to commit to PNA research to promote the progress of medical care.
Summary
PFDN complex and its subunits play an important role in maintaining protein homeostasis, preventing protein misfolding diseases, and tumor progression. The unique structure and function of PFDN make it as a nanomaterial designed as a PNA for intelligent drug delivery. Based on the different pH and temperature of the tumor site from the normal tissue, PNA can accurately control the release of nano-drugs at the tumor, achieving precise tumor treatment and bringing new hope for the treatment of diseases. To open up a new path for precision treatment of nanomedicine. However, in order to achieve precise treatment of tumors or other diseases, in-depth research and development on PNA properties such as the interaction with the cargo, how to better regulate the combination and release of the cargo, are required. Such as re-mutation or interaction with other proteins to form better nano-components, expand the function of PNA, expand the scope of PNA in disease treatment, and achieve precise treatment of diseases. Due to the limited research and understanding of them, their research prospects are still huge. In the future, whether they become tumor markers or the application of nanotechnology to treat diseases will be of great significance. In general, the research of PFDN and its subunits in tumors and other diseases and their potential as nanotechnology applications can provide new directions for future clinical research and treatment.
Disclosure
The authors declare no conflicts of interest for this work.
References
1. Vainberg IE, Lewis SA, Rommelaere H. Prefoldin, a chaperone that delivers unfolded proteins to cytosolic chaperonin. Cell. 1998;93(5):863–873. doi:10.1016/s0092-8674(00)81446-4
2. Yesseyeva G, Aikemu B, Hong H, et al. Prefoldin subunits (PFDN1-6) serve as poor prognostic markers in gastric cancer. Biosci Rep. 2020;40(2). doi:10.1042/BSR20192712
3. Zhou C, Guo Z, Xu L, et al. PFND1 Predicts Poor Prognosis of Gastric Cancer and Promotes Cell Metastasis by Activating the Wnt/beta-Catenin Pathway. Onco Targets Ther. 2020;13:3177–3186. doi:10.2147/OTT.S236929
4. Shokuhfar A, Ghaffari A, Ghasemi RH. Cavity Control of Prefoldin Nano Actuator (PNA) by Temperature and pH. Nano Micro Let. 2012;4(2):110–117. doi:10.1007/BF03353701
5. Leroux MR. MtGimC, a novel archaeal chaperone related to the eukaryotic chaperonin cofactor GimC/prefoldin. EMBO J. 1999;18(23):6730–6743. doi:10.1093/emboj/18.23.6730
6. Miyazawa M, Tashiro E, Kitaura H. Prefoldin subunits are protected from ubiquitin-proteasome system-mediated degradation by forming complex with other constituent subunits. J Biol Chem. 2011;286(22):19191–19203. doi:10.1074/jbc.M110.216259
7. Siegert R, Leroux MR, Scheufler C, Hartl FU, Moarefi I. Structure of the molecular chaperone prefoldin: unique interaction of multiple coiled coil tentacles with unfolded proteins. Cell. 2000;103(4):621–632. doi:10.1016/s0092-8674(00)00165-3
8. Hartl FU. Molecular chaperones in the cytosol: from nascent chain to folded protein. Science. 2002;295(5561):1852–1858. doi:10.1126/science.1068408
9. Balchin D, Hayer-Hartl M, Hartl FU. In vivo aspects of protein folding and quality control. Science. 2016;353(6294):aac4354. doi:10.1126/science.aac4354
10. Geissler S. A novel protein complex promoting formation of functional alpha- and gamma-tubulin. EMBO J. 1998;17(4):952–966. doi:10.1093/emboj/17.4.952
11. Siegers K. Compartmentation of protein folding in vivo: sequestration of non-native polypeptide by the chaperonin-GimC system. EMBO J. 1999;18(1):75–84. doi:10.1093/emboj/18.1.75
12. Hansen WJ, Cowan NJ, Welch WJ. Prefoldin-nascent chain complexes in the folding of cytoskeletal proteins. J Cell Biol. 1999;145(2):265–277. doi:10.1083/jcb.145.2.265
13. Zako T, Murase Y, Iizuka R. Localization of prefoldin interaction sites in the hyperthermophilic group II chaperonin and correlations between binding rate and protein transfer rate. J Mol Biol. 2006;364(1):110–120. doi:10.1016/j.jmb.2006.08.088
14. Gestaut D, Roh SH, Ma B. The Chaperonin TRiC/CCT Associates with Prefoldin through a Conserved Electrostatic Interface Essential for Cellular Proteostasis. Cell. 2019;177(3):751–65.e15. doi:10.1016/j.cell.2019.03.012
15. Liu H, Liu X, Jin H, Yang F, Gu W, Li X. Proteomic analysis of knock-down HLA-G in invasion of human trophoblast cell line JEG-3. Int J Clin Exp Pathol. 2013;6(11):2451–2459.
16. Wang D, Shi W, Tang Y. Prefoldin 1 promotes EMT and lung cancer progression by suppressing cyclin A expression. Oncogene. 2017;36(7):885–898. doi:10.1038/onc.2016.257
17. Bendris N, Cheung CT, Leong HS. Cyclin A2, a novel regulator of EMT. Cell Mol Life Sci. 2014;71(24):4881–4894. doi:10.1007/s00018-014-1654-8
18. Peñate Salas X. Overexpression of Canonical Prefoldin Associates with the Risk of Mortality and Metastasis in Non-Small Cell Lung Cancer. Cancers. 2020;12(4). doi:10.3390/cancers12041052
19. Effect of Prefoldin 1 on epithelial–mesenchymal transition in breast cancer. J Pract Med. 2019;35(01):47–50. doi:10.3969/j.issn.1006-5725.2019.01.010
20. Cheung CT, Bendris N, Paul C. Cyclin A2 modulates EMT via β-catenin and phospholipase C pathways. Carcinogenesis. 2015;36(8):914–924. doi:10.1093/carcin/bgv069
21. Wang P, Zhao J, Yang X. PFDN1, an indicator for colorectal cancer prognosis, enhances tumor cell proliferation and motility through cytoskeletal reorganization. Med Oncol. 2015;32(12):264. doi:10.1007/s12032-015-0710-z
22. Pagano M, Pepperkok R, Verde F, Ansorge W, Draetta G. Cyclin A is required at two points in the human cell cycle. EMBO J. 1992;11(3):961–971. doi:10.1002/j.1460-2075.1992.tb05135.x
23. Cao S, Carlesso G, Osipovich AB. Subunit 1 of the prefoldin chaperone complex is required for lymphocyte development and function. J Immunol. 2008;181(1):476–484. doi:10.4049/jimmunol.181.1.476
24. López V, González-Peramato P, Suela J. Identification of prefoldin amplification (1q23.3-q24.1) in bladder cancer using comparative genomic hybridization (CGH) arrays of urinary DNA. J Transl Med. 2013;11(1):182. doi:10.1186/1479-5876-11-182
25. Riester M, Werner L, Bellmunt J. Integrative analysis of 1q23.3 copy-number gain in metastatic urothelial carcinoma. Clin Cancer Res. 2014;20(7):1873–1883. doi:10.1158/1078-0432.Ccr-13-0759
26. Aljahani AH, Alarjani KM, Hassan ZK. Prefoldin subunits (PFDN1-6) serve as poor prognostic markers in gastric cancer. Biosci Rep. 2020;40(2). doi:10.1042/bsr20192712
27. Chang DC, Piaggi P, Hanson RL. Autoantibodies against PFDN2 are associated with an increased risk of type 2 diabetes: A case-control study. Diabetes Metab Res Rev. 2017;33(8). doi:10.1002/dmrr.2922
28. Hübner M, Hinske CL, Effinger D. Genetics and Expression Profile of the Tubulin Gene Superfamily in Breast Cancer Subtypes and Its Relation to Taxane Resistance. Cancers. 2018;10(8). doi:10.3390/cancers10080274
29. Chang K, Yang P, Lai H, Yeh T, Chen T, Yeh C. Identification of a novel actin isoform in hepatocellular carcinoma. Hepatology Res. 2006;36(1):33–39. doi:10.1016/j.hepres.2006.05.003
30. Tsao M-L, Chao C-H, Yeh C-T. Interaction of hepatitis C virus F protein with prefoldin 2 perturbs tubulin cytoskeleton organization. Biochem Biophys Res Commun. 2006;348(1):271–277. doi:10.1016/j.bbrc.2006.07.062
31. Musavi Z, Hashempour T, Moayedi J, et al. Antibody Development to HCV Alternate Reading Frame Protein in Liver Transplant Candidate and its Computational Analysis. Curr Proteomics. 2020;17(2):154–170. doi:10.2174/1570164617666190822103329
32. Farooq M, Hozzein WN, Elsayed EA, Taha NA, Wadaan MAM. Identification of histone deacetylase 1 protein complexes in liver cancer cells. Asian Pacific j Cancer Prevention. 2013;14(2):915–921. doi:10.7314/apjcp.2013.14.2.915
33. Le Goff X, Chesnel F, Delalande O. Aggregation dynamics and identification of aggregation-prone mutants of the von Hippel-Lindau tumor suppressor protein. J Cell Sci. 2016;129(13):2638–2650. doi:10.1242/jcs.184846
34. Tsuchiya H, Iseda T, Hino O. Identification of a novel protein (VBP-1) binding to the von Hippel-Lindau (VHL) tumor suppressor gene product. Cancer Res. 1996;56(13):2881–2885.
35. Delgehyr N, Wieland U, Rangone H. Drosophila Mgr, a Prefoldin subunit cooperating with von Hippel Lindau to regulate tubulin stability. Proc Natl Acad Sci U S A. 2012;109(15):5729–5734. doi:10.1073/pnas.1108537109
36. Feldman DE, Thulasiraman V, Ferreyra RG, Frydman J. Formation of the VHL–elongin BC tumor suppressor complex is mediated by the chaperonin TRiC. Mol Cell. 1999;4(6):1051–1061. doi:10.1016/s1097-2765(00)80233-6
37. Melville MW, McClellan AJ, Meyer AS, Darveau A, Frydman J. The Hsp70 and TRiC/CCT chaperone systems cooperate in vivo to assemble the von Hippel-Lindau tumor suppressor complex. Mol Cell Biol. 2003;23(9):3141–3151. doi:10.1128/mcb.23.9.3141-3151.2003
38. Feldman DE, Spiess C, Howard DE, Frydman J. Tumorigenic mutations in VHL disrupt folding in vivo by interfering with chaperonin binding. Mol Cell. 2003;12(5):1213–1224. doi:10.1016/s1097-2765(03)00423-4
39. Lisztwan J, Imbert G, Wirbelauer C, Gstaiger M, Krek W. The von Hippel-Lindau tumor suppressor protein is a component of an E3 ubiquitin-protein ligase activity. Genes Dev. 1999;13(14):1822–1833. doi:10.1101/gad.13.14.1822
40. Xu Y, Her C. facilitates proteasome and autophagy-mediated degradation of MutS homologue hMSH4. FASEB j. 2013;27(12):4799–4810. doi:10.1096/fj.13-235127
41. Mousnier A, Kubat N, Massias-Simon A. von Hippel-Lindau binding protein 1-mediated degradation of integrase affects HIV-1 gene expression at a postintegration step. Proc Natl Acad Sci U S A. 2007;104(34):13615–13620. doi:10.1073/pnas.0705162104
42. Salceda S, Caro J. Hypoxia-inducible factor 1alpha (HIF-1alpha) protein is rapidly degraded by the ubiquitin-proteasome system under normoxic conditions. Its stabilization by hypoxia depends on redox-induced changes. J Biol Chem. 1997;272(36):22642–22647. doi:10.1074/jbc.272.36.22642
43. Kim JA, Choi DK, Min JS. VBP 1 represses cancer metastasis by enhancing HIF −1α degradation induced by pVHL. FEBS J. 2018;285(1):115–126. doi:10.1111/febs.14322
44. Iijima M, Kano Y, Nohno T, Namba M. Cloning of cDNA with possible transcription factor activity at the G1-S phase transition in human fibroblast cell lines. Acta Med Okayama. 1996;50(2):73–77. doi:10.18926/amo/30489
45. Kurata M, Maesako Y, Ueda C. Characterization of t(3;6)(q27;p21) breakpoints in B-cell non-Hodgkin’s lymphoma and construction of the histone H4/BCL6 fusion gene, leading to altered expression of Bcl-6. Cancer Res. 2002;62(21):6224–6230.
46. Collins C. Comprehensive genome sequence analysis of a breast cancer amplicon. Genome Res. 2001;11(6):1034–1042. doi:10.1101/gr.gr1743r
47. Miyoshi N, Ishii H, Mimori K. Abnormal expression of PFDN4 in colorectal cancer: a novel marker for prognosis. Ann Surg Oncol. 2010;17(11):3030–3036. doi:10.1245/s10434-010-1138-5
48. Wang D, Zhu Z, Jiang H, et al. Multiple genes identified as targets for 20q13.12-13.33 gain contributing to unfavorable clinical outcomes in patients with hepatocellular carcinoma. Hepatol Int. 2015;9(3):438–446. doi:10.1007/s12072-015-9642-0
49. Ostrov DA, Barnes CL, Smith LE. Characterization of HKE2: an ancient antigen encoded in the major histocompatibility complex. Tissue Antigens. 2007;69(2):181–188. doi:10.1111/j.1399-0039.2006.00730.x
50. Dehghan-Nayeri N, Rezaei-Tavirani M, Omrani MD, Gharehbaghian A, Goudarzi Pour K, Eshghi P. Identification of potential predictive markers of dexamethasone resistance in childhood acute lymphoblastic leukemia. J Cell Commun Signal. 2017;11(2):137–145. doi:10.1007/s12079-016-0357-3
51. Son HG, Seo K, Seo M. Prefoldin 6 mediates longevity response from heat shock factor 1 to FOXO in C. elegans. Genes Dev. 2018;32(23–24):1562–1575. doi:10.1101/gad.317362.118
52. Abe A, Takahashi-Niki K, Takekoshi Y. Prefoldin plays a role as a clearance factor in preventing proteasome inhibitor-induced protein aggregation. J Biol Chem. 2013;288(39):27764–27776. doi:10.1074/jbc.M113.476358
53. Takano M, Tashiro E, Kitamura A. Prefoldin prevents aggregation of α-synuclein. Brain Res. 2014;1542:186–194. doi:10.1016/j.brainres.2013.10.034
54. Tashiro E, Zako T, Muto H. Prefoldin protects neuronal cells from polyglutamine toxicity by preventing aggregation formation. J Biol Chem. 2013;288(27):19958–19972. doi:10.1074/jbc.M113.477984
55. Broer L, Ikram MA, Schuur M. Association of HSP70 and its co-chaperones with Alzheimer’s disease. J Alzheimer’s Dis. 2011;25(1):93–102. doi:10.3233/jad-2011-101560
56. Jun GR, Chung J, Mez J. Transethnic genome-wide scan identifies novel Alzheimer’s disease loci. Alzheimer’s Dementia. 2017;13(7):727–738. doi:10.1016/j.jalz.2016.12.012
57. Zhang Y, Rai M, Wang C, Gonzalez C, Wang H. Prefoldin and Pins synergistically regulate asymmetric division and suppress dedifferentiation. Sci Rep. 2016;6(1):23735. doi:10.1038/srep23735
58. Cimmino F, Spano D, Capasso M. Comparative Proteomic Expression Profile in All-Retinoic Acid Differentiated Neuroblastoma Cell Line. J Proteome Res. 2007;6(7):2550–2564. doi:10.1021/pr060701g
59. Tao Y, Han Y, Yu L, Wang Q, Leng SX, Zhang H. The Predicted Key Molecules, Functions, and Pathways That Bridge Mild Cognitive Impairment (MCI) and Alzheimer’s Disease (AD). Front Neurol. 2020;11:233. doi:10.3389/fneur.2020.00233
60. Lee Y. Prefoldin 5 is required for normal sensory and neuronal development in a murine model. J Biol Chem. 2011;286(1):726–736. doi:10.1074/jbc.M110.177352
61. Kadoyama K, Matsuura K, Takano M, Maekura K, Inoue Y, Matsuyama S. Changes in the expression of prefoldin subunit 5 depending on synaptic plasticity in the mouse hippocampus. Neurosci Lett. 2019;712:134484. doi:10.1016/j.neulet.2019.134484
62. Alldinger I, Dittert D, Peiper M. Gene expression analysis of pancreatic cell lines reveals genes overexpressed in pancreatic cancer. Pancreatology. 2005;5(4–5):370–379. doi:10.1159/000086537
63. Hennecke S, Beck J, Bornemann-Kolatzki K. Prevalence of the Prefoldin Subunit 5 Gene Deletion in Canine Mammary Tumors. PLoS One. 2015;10(7):e0131280. doi:10.1371/journal.pone.0131280
64. Mori K, Maeda Y, Kitaura H. MM-1, a novel c-Myc-associating protein that represses transcriptional activity of c-Myc. J Biol Chem. 1998;273(45):29794–29800. doi:10.1074/jbc.273.45.29794
65. Satou A, Taira T, Iguchi-Ariga SMM, Ariga H. A novel transrepression pathway of c-Myc. Recruitment of a transcriptional corepressor complex to c-Myc by MM-1, a c-Myc-binding protein. J Biol Chem. 2001;276(49):46562–46567. doi:10.1074/jbc.M104937200
66. Yoshida T, Kitaura H, Hagio Y, Sato T, Iguchi-Ariga SMM, Ariga H. Negative regulation of the Wnt signal by MM-1 through inhibiting expression of the wnt4 gene. Exp Cell Res. 2008;314(6):1217–1228. doi:10.1016/j.yexcr.2008.01.002
67. Satou A, Hagio Y, Taira T, Iguchi-Ariga SMM, Ariga H. Repression of the c-fms gene in fibroblast cells by c-Myc-MM-1-TIF1β complex. FEBS Lett. 2004;572(1–3):211–215. doi:10.1016/j.febslet.2004.07.034
68. Fujioka Y, Taira T, Maeda Y. MM-1, a c-Myc-binding protein, is a candidate for a tumor suppressor in leukemia/lymphoma and tongue cancer. J Biol Chem. 2001;276(48):45137–45144. doi:10.1074/jbc.M106127200
69. Ghaffari A. Design and simulation of a novel bio nano actuator by prefoldin. Nanotechnology. 2010.
70. Whitehead T. Rational Shape Engineering of the Filamentous Protein gamma Prefoldin Through Incremental Gene Truncation. Biopolymers. 2009;6(91). doi:10.1002/bip.21157
71. Clark DS, Glover DJ. Controlled Assembly of the Filamentous Chaperone Gamma-Prefoldin into Defined Nanostructures. Methods Mol Biol. 2018;1798:293–306. doi:10.1007/978-1-4939-7893-9_22
72. Askarian M, Moavenian M, Ghasemi RH. Prefoldin β 1: A New Bio-Nanorobots Component. Adv Sci Eng Med. 2013. doi:10.1166/asem.2013.1363
73. Ghaffari A, Shokuhfar A, Ghasemi RH. Prefoldin: A Nano Actuator for Carrying the Various Size Nano Drugs. J Comput Theor Nanosci. 2011;8(10):2078–2086. doi:10.1166/jctn.2011.1929
74. Ghasemi SRH. Capturing and releasing a nano cargo by Prefoldin nano actuator. Sens Actuators B Chem. 2012. doi:10.1016/j.snb.2012.06.077
75. Ghasemi RH, Keramati M. Design a protein gripper to capture a hydrophobic cargo. Iet Nanobiotechnol. 2019;13(5):546–552. doi:10.1049/iet-nbt.2018.5208
76. Ghasemi RH, Keramati M, Mojarrad MHS. The effect of structure on improvement of the PNA Young modulus: A study of steered molecular dynamics. Comput Biol Chem. 2019;83:107133. doi:10.1016/j.compbiolchem.2019.107133
77. Keramati M, Ghasemi RH. A molecular dynamics investigation of the effects of mutation on prefoldin nano actuator in inhibiting amyloid beta 42-dimer. Sensors Actuators B Chem. 2017;248:536–544. doi:10.1016/j.snb.2017.03.165
78. Dubey A, Sharma G, Mavroidis C, Tomassone M, Nikitczuk K, Yarmush ML. Computational Studies of Viral Protein Nano-Actuators. J Comput Theor Nanosci. 2004;1(1):18–28. doi:10.1166/jctn.2003.003
79. Cheng X, He L, Xu J, et al. Oxygen-producing catalase-based prodrug nanoparticles overcoming resistance in hypoxia-mediated chemo-photodynamic therapy. Acta Biomater. 2020;112:234–249. doi:10.1016/j.actbio.2020.05.035
80. Shuai G, Liang Q, Yafei C, et al. TMEM16A-inhibitor loaded pH-responsive nanoparticles: a novel dual-targeting antitumor therapy for lung adenocarcinoma. Biochem Pharmacol. 2020. doi:10.1016/j.bcp.2020.114062
81. Lan J, Ze-Ming G, Lin Y, Ai-Ying Z, Zeng-Guo F. A pH-sensitive nano drug delivery system of doxorubicin-conjugated amphiphilic polyrotaxane-based block copolymers. Biomater Sci. 2013;1(12). doi:10.1039/c3bm60112a
82. Jinbing X, Ying L, Yi C, et al. Photo synthesis of protein-based drug-delivery nanoparticles for active tumor targeting. Biomater Sci. 2013;1(12). doi:10.2142/biophys.53.s162_6
83. Gerweck LE. The pH difference between tumor and normal tissue offers a tumor specific target for the treatment of cancer. Drug Resistance Updates. 2000;3(1):49–50. doi:10.1054/drup.2000.0122
84. Gerweck LE, Seetharaman K. Cellular pH gradient in tumor versus normal tissue: potential exploitation for the treatment of cancer. Cancer Res. 1996;6(56).
85. Colotta F. Cancer-related inflammation, the seventh hallmark of cancer: links to genetic instability. Carcinogenesis. 2009;7(30). doi:10.1093/carcin/bgp127
86. Salehi Mojarrad M, Hasanzadeh Ghasemi R, Keramati M. Nano-mechanical Study of a Bio Nano Actuator Under External Forces. 2017;17.
87. Salehi Mojarrad MH, Goodarzi V, Taheri RA. Force Sensing in Nanoscale: integration of Virtual Spring for Sensing the Interactions of β-Amyloid Grabbed by Prefoldin. Appl Biochem Biotechnol. 2019;188(4):1108–1116. doi:10.1007/s12010-019-02975-6
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.