Back to Journals » Cancer Management and Research » Volume 12
The Role and Mechanism of S1PR5 in Colon Cancer
Authors Zhou H, Yin X, Bai F, Liu W, Jiang S, Zhao J
Received 18 November 2019
Accepted for publication 12 May 2020
Published 19 June 2020 Volume 2020:12 Pages 4759—4775
DOI https://doi.org/10.2147/CMAR.S239118
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Beicheng Sun
This paper has been retracted.
Huijun Zhou, 1 Xianli Yin, 2 Fei Bai, 3 Wu Liu, 2 Shaofeng Jiang, 2 Jinfeng Zhao 1
1Key Laboratory of Nanobiological Technology of National Health Commission of China, Xiangya Hospital, Central South University, Changsha, Hunan, People’s Republic of China; 2Department of Gastroenterology and Urology, Hunan Cancer Hospital & the Affiliated Hospital of Xiangya School of Medicine, Central South University, Changsha 410013, Hunan, People’s Republic of China; 3Department of Gastroduodeno Pancreatic Surgery, Hunan Cancer Hospital & the Affiliated Hospital of Xiangya School of Medicine, Central South University, Changsha 410013, Hunan, People’s Republic of China
Correspondence: Jinfeng Zhao
Key Laboratory of Nanobiological Technology of National Health Commission of China, Xiangya Hospital, Central South University, No. 87 Xiangya Road, Kaifu District, Changsha, Hunan 410008, People’s Republic of China
Tel +86-15874856567
Email [email protected]
Purpose: To investigate the role and mechanism of S1PR5 in colon cancer.
Materials and Methods: Lentiviral infection and drug screening helped to establish colon cancer cell lines with stable overexpression and knockdown of S1PR5. Effects of S1PR5 expression on cell growth, proliferation, migration, and invasion were analyzed using a subcutaneous xenograft model in nude mice. Western blot (WB) was used to detect the effects of S1PR5 expression on p-AKT, STAT3, NF-κB, and p-JNK. The distribution of p65 was evaluated in nuclear and cytoplasmic fractions using WB. CCK-8, Transwell migration, and Transwell invasion assays analyzed cell growth, proliferation, migration, and invasion. qRT-PCR analysis revealed that S1PR5 expression was associated with altered expression levels of NF-κB downstream target genes, such as IL-6, TNF-α, and indoleamine 2, 3-dioxygenase 1 (IDO1).
Results: qRT-PCR and WB analysis showed that the S1PR5 level in colon cancer cell lines—SW480, SW620, HCT116, and LoVo—was significantly higher than in NCM460, a healthy colonic epithelial cell line. SW620 and SW480, with high and low expression of S1PR5, respectively, were selected as model cell lines. S1PR5 knockdown in SW620 caused the growth rate, proliferation, migration, invasion, and subcutaneous tumor formation rate to decrease in mice, whereas S1PR5 overexpression in SW480 caused all of these parameters to increase. WB analysis showed an increase in phospho-p65 and its nuclear translocation. S1PR5 knockdown caused a decrease in phospho-p65 levels and its nuclear import, thereby inhibiting its activity. In S1PR5 knockdown and overexpressing cells, p65 was overexpressed and knocked down, respectively. qRT-PCR and WB showed that S1PR5 over-expression up-regulates IDO1, and S1PR5 knockdown inhibits IDO1. CCK-8 and Transwell assays showed that p65 and IDO1 overexpression antagonizes the antitumor effect of S1PR5 knockdown, and that p65 and IDO1 knockdown antagonizes the tumorigenic effect of S1PR5 overexpression.
Conclusion: S1PR5 overexpression promotes the growth, migration, and invasion of cancer by activating the NF-κB/IDO1 signaling pathway.
Keywords: S1PR5, NF-κB, IDO1, colon cancer
Introduction
Colon cancer is a high-incidence malignant tumor of the digestive tract. It ranks third in the world amongst malignant tumors, and fourth in terms of mortality. The incidence of colon cancer is higher in developed western countries; however, with the rapid economic growth that developing countries are experiencing, which is leading to improved standards of living, westernized diet structures, and schedule prevalence, the incidence of colon cancer in developing countries is rising rapidly as well.1
Epidemiological studies show that genetic factors, inflammatory bowel disease, eating habits, consumption of alcohol, and smoking are risk factors for colon cancer.2 From a mechanical point of view, the high-risk factors for colon cancer and the imbalance of intestinal homeostasis contribute to the formation of inflammatory and immunosuppressive microenvironments that encourage the malignant transformation of cells. For example, exposure to long-term risk factors can change the composition and distribution of the intestinal microbiome and promote the survival of pro-inflammatory microorganisms, thereby forming an immunosuppressive microenvironment in which small molecules, such as inflammatory factors, can act as ligands. On interaction with the cell surface receptors of intestinal epithelial cells, the regulatory signals are altered, reshaping cellular gene expression and metabolism, and eventually leading to malignant transformation of the cells. Cell surface receptors are key mediating factors for the interaction between the microenvironment and the cell. Changes in the composition and distribution of cell surface receptors are required for malignant changes to occur and for microenvironmental information to adapt to the microenvironment. Numerous studies have shown that targeted therapies and immunotherapeutic techniques based on surface receptors, such as EGFR, PD-1, and CART, play an important role in the treatment of malignant tumors, including colon cancer.3 Therefore, it is important to identify the receptors that impact the development of colon cancer, develop new therapeutic targets, and reduce the risk of resistance to individual drugs.
During the course of treatment, the side-effects caused by anti-cancer drugs limit their use; at the same time, tumors are prone to drug resistance. Currently, there is no effective solution to this problem;4 however, the discovery of S1PR regulators brings new ideas for potential solutions. S1PR1 is the first cloned S1PR gene; it was discovered and cloned in 1990 while researchers were screening for key genes involved in the early differentiation of endothelial cells.5 In the following decade, S1PR2, S1PR3, S1PR4, and S1PR5 were successively discovered and cloned. The distribution of S1PRs in different tissues is different; however, they are the most highly expressed in cells with immune functionality.6,7 This discovery first revealed the role of S1PRs in immune regulation. Inhibitors against all S1PRs or particular S1PRs have been developed; some have been used as immunomodulators in clinical applications, such as Fingolimod, which combines with S1PR1, 3, 4, and 5. Fingolimod has been approved by the US FDA to treat multiple sclerosis.8,9 As the key role of S1P in the regulation of tumors has been revealed, the role of S1PRs in tumors is beginning to be understood. RNA interference and gene knockout studies in cell lines and mouse models have revealed the roles of S1PRs in tumor growth, invasion, and angiogenesis-related metastasis.10–12 Studies have shown that S1PRs exert their effects on tumors in a tissue-specific manner. Thus, the specific roles and mechanisms of S1PRs in different tissue types can be utilized for the development of new therapeutic techniques. Although studies have confirmed that S1P is closely related to colon cancer development, the expression level, function, and mechanism of S1PRs in colon cancer have not been reported.
The comprehensive role of S1P in inflammatory and immune microenvironments and the existing research on S1PRs indicate that S1PRs may play an important role in cancers, especially those associated with inflammation, such as colon cancer. Therefore, S1PR5, which is significantly elevated in colon cancer, was either over-expressed or knocked down in colon cancer cell lines, and its function and mechanism were studied at the cellular and animal levels. This study will help to reveal the pathogenesis of colon cancer and provide the basis for using S1PR5 as a target for diagnosing and treating colon cancer.
Materials and Methods
Material and Instrument
Reagents
Reagents for this study included: TRIzol reagent (Thermo Fisher Scientific, USA), citrate buffer (pH=6.0) (Wuhan Boshide Corporation, China), qSYBR Green PCR kit (Shanghai GenePharma, Shanghai, China), reverse reaction kit (Promega, USA), 10×RT buffer (Dalian Bao Biotech Company, China), DEPC water (Sigma-Aldrich, USA), MMLV reverse transcriptase (Dalian Bao Biotech Company, China), 2.5 mM dNTP mixture (Nanjing Kaiji Biotech., China), 10× PCR buffer (Promega, USA), fetal bovine serum (Thermo Fisher Scientific, USA), trypsin/EDTA (Thermo Fisher Scientific, USA), Transwell chamber (Merck, USA), Lipofectamine TM 2000 (Thermo Fisher Scientific, USA), CCK-8 (Tongren Chemical Research Institute, Japan), Annexin V-FITC Apoptosis Detection Kit (Promega, USA), and RPMI-1640 medium (Life Technologies, Gaithersburg, MD).
Instruments
Instruments for this study included: a thermostatic magnetic stirrer (Jiangsu Jintan Medical Instrument Co., Ltd., China), an oven (Jiangsu Jintan Medical Instrument Co., Ltd., China), a low temperature high speed centrifuge (Sigma-Aldrich, USA), an incubator (Sigma-Aldrich, USA), an Olympus optical microscope (Olympus Corporation, Japan), a carbon dioxide incubator (Sanyo Co., Japan), an FE-20 pH meter (Mitler, Switzerland), a DK-2000-IIIL constant temperature water bath (Tianjin Test Instrument Factory, China), an SOP 62–0025 Pure water meter (Millipore, USA), a 7500 fluorescence quantitative PCR instrument (ABI, USA), a flow cytometer (BD Biosciences, USA), pipettes (Eppendorf, Germany), a micro high-speed benchtop cryogenic centrifuge (Sigma-Aldrich, USA), and BCLB/C nude mice (Slyke Jingda company, Hunan, China).
Experimental Method
Specimen Sources
All samples in this study were derived from cancer tissues and the corresponding paracancer tissues (no less than 2 cm from the tumor edge) of 98 colon cancer patients who received surgical treatment in the affiliated cancer hospital, Xiangya Medical College of Central South University, from January 2017 to December 2017. The human tissue samples collected and used for testing were approved by the informed consent of the patients and the ethics committee of the affiliated cancer hospital, Xiangya Medical College of Central South University.
Cell Experiment
Four colon cancer cell lines, SW480, SW620, HCT-116, and LoVo, and a healthy colonic epithelial cell line, NCM460 (Shanghai institute of cell biology, Chinese academy of sciences), were selected for the study. The cells were cultured in RPMI 1640 medium (Life Technologies, Gaithersburg, MD), containing 10% fetal bovine serum (Thermo Fisher Scientific, USA) and 100 U/mL streptomycin, in an incubator (Sigma-Aldrich, USA) at 37 °C and 5% CO2.
Establishment of the Subcutaneous Xenograft Model
SW480/NC, SW480/S1PR5, SW620/NC, and SW620/shS1PR5 were cultured to logarithmic growth phase, trypsinized, and counted. Fetal bovine serum (Thermo Fisher Scientific, USA), trypsin/EDTA (Thermo Fisher Scientific, USA), RPMI-1640 medium (Life Technologies, Gaithersburg, MD), and 2×106 cells were injected into approximately 5-week-old female nude mice (BCLB/C nude mice, Slyke Jingda company, Hunan, China). Tumor growth was monitored every third day, and the long and short diameters of the tumor were recorded. After 21 days, the nude mice were sacrificed using the routine procedure, the tumors were photographed and weighed, and the subcutaneous tumors were fixed with paraformaldehyde after being photographed. Log phase cells were collected and resuspended in the medium after centrifugation, and the concentration of the cell suspension was adjusted by adding 100 μL of complete medium to each well. Thereafter, the density of the cells to be tested was adjusted to 4000 cells per well, sterile PBS was added to the wells, and the cells were incubated in a 37 °C incubator with 5% CO2. Subsequently, 100 μL of fresh medium was added to each well at 0, 12, 24, 48, and 72 h. Three wells with 100 μL cell culture medium alone were used as blank controls. After adding 10 μL of CCK-8 (Tongren Chemical Research Institute, Japan) solution to each well, incubation was continued for 0.5–4 h in a cell culture incubator. Initially, detection with a microplate reader was conducted at 0.5, 1, 2, and 4 h. Thereafter, suitable time points were selected for measuring the absorbance range for the next four experiments. The absorbance (OD value) of each well was determined at 450 nm on the microplate reader, and the average value of each group was calculated by repeating the experiment thrice.
Invasion Experiment
The matrigel was removed from the −20 °C refrigerator and placed at 4 °C before being transferred to a clean bench. The matrigel was diluted with serum-free medium (1:8 = matrigel:medium) and added to the upper chamber of a 24-well Transwell chamber, which was placed at 4 °C overnight and then kept at room temperature for 30 min the next day. The residual liquid in the culture plate was aspirated, 50 μL of serum-free medium containing 10 g/LBSA was added to each well, and the cells were serum-starved for 12–24 h at 37 °C for 30 min to remove the effect of serum before the cell suspension was prepared. The transfected experimental and negative control group SW480 and SW620 cells were trypsinized, resuspended, and counted. A total of 200 x liters of cell suspension containing 2 x 105 cells was added to the upper chamber of the Transwell unit, and 600 μL of DMEM medium, containing 20% FBS, was added to the lower chamber. Following incubation for 2 h at 37 °C in a CO2 incubator, the Transwell chamber was rinsed thrice with PBS, and 4% paraformaldehyde was used to fix cells at room temperature for 30 min. Thereafter, the matrigel and the tumor cells in the upper chamber were gently wiped off with a cotton swab and stained with 0.1% crystal violet for 20 min. Excess crystal violet was washed off with water, and the Transwell chamber was oven dried. Photographs were taken using an upright light microscope, and nine fields were randomly chosen for cell counts. The experiment was repeated three times, independently.
Western Blot
The cells cultured to logarithmic growth phase were scraped off, and 1 mL lysate was used following homogenization. After 30 min, the supernatant was collected by centrifugation (Sigma-Aldrich, USA) at 12,000 rpm for 10 min. The protein concentration of each tube was measured by the BCA method and adjusted with the loading buffer. A total of 20 μg of protein was mixed with SDS buffer, denatured at 95 °C for 5 min, and electrophoresed on a 10% SDS-PAGE gel. The protein sample and the gel loading buffer were mixed at a volume ratio of 4:1, boiled, and denatured on an electric furnace. After cooling to room temperature, 40 μg of protein sample was loaded onto each lane. The electrophoresis buffer was added to the electrophoresis tank, and the samples were electrophoresed. When the bromophenol blue dye indicator reached the bottom of the separation gel, the power supply was disconnected. The gel was removed from the electrophoresis tank, immersed in the transfer buffer for half an hour to equilibrate, and electrophoresed for about 60 min. The membrane was removed, stained with Ponceau for 5 min, and rinsed with TBST. Thereafter, the membrane was placed in a hybridization bag with 5% skim milk powder solution as the blocking agent and then incubated at room temperature for 1 h with gentle shaking. After removing the blocking solution, appropriately diluted primary antibody (S1PR5 1:500) was added to the membrane, which was sealed after removing air bubbles and incubated at 4 °C overnight. Subsequently, the filter was rinsed thrice with BST for 10 min and placed in a hybridization bag, and the corresponding secondary antibody labeled with horseradish peroxidase (anti-rabbit IgG 1:500) was added at 0.1 mL/cm2. After removing the bubbles, the membrane was incubated at room temperature for 1 h and rinsed thrice with TBST for 10 min. Chemiluminescence was detected with a gel imaging system.
The Expression of Interstitial Phenotype Related Factor S1PR5 mRNA Was Detected by qPCR
Colonic cancer and normal colon epithelial cell lines were collected, total RNA was extracted by the TRIzol method (TRIzol reagent, Thermo Fisher Scientific, USA;qSYBR Green PCR kit, Shanghai GenePharma, Shanghai, China), RNA concentration was determined with an ultraviolet spectrophotometer, and purity was determined. Total RNA reverse transcription was synthesized in cDNA using a reverse transcription kit. The reaction conditions were 25 °C, 5 min, 42 °C, 30 min, 85 °C, 5 min and 4 °C. QPCR was amplified and detected by fluorescence quantitative PCR. QPCR reaction conditions were as follows: pre-denaturation at 95 °C for 5 min, 95 °C for 5 s, 60 °C for 30 s, 72 °C for 20 s, and 65 °C for 5 s, for a total of 40 cycles. The relative mRNA expression level of S1PR5 was calculated by 2-ΔΔCt.
The Expression of S1PR5 Was Detected by Immunohistochemistry
The expression of S1PR5 protein was detected in paraffin pathological sections according to the SP immunohistochemical kit instructions. Rabbit anti-human S1PR5 monoclonal antibody was diluted to a 1:200 ratio with biotinylated goat anti-rabbit secondary antibody and horseradish peroxidase labeled chain enzyme oviparin (triantibody) as the original solution in the kit. After adding DAB reagent to develop color, the rabbit anti-rabbit secondary antibody was stained with hematosin and sealed with neutral resin. Immunohistochemical staining showed that brown-yellow granules were positive in the cytoplasm and in vivo. The double-blind method was used to observe the whole field of view in tissue sections. The semi-quantitative integration method was used to determine the results and the positive location, as follows: (1) number of positive cells: 0 points for no positive cells, <10% is 1 point, 10% ~ 50% is 2 points,>50%50% is 3; (2) the positive intensity of staining, colorless or similar to the background color was 0 points, light yellow was 1 point, tan was 2 points, tan was 3 points. The integrals of (1) and (2) were multiplied; the integral <4 was divided into low expression, and the integral ≥4 was divided into high expression. (3) Ki67 index (labeling index, LI). Ten fields of each section were randomly selected. Ki67 positive cells were counted, and the percentage was calculated under a low-power microscope (×200). The average value was Ki67LI. As there were multiple fields with different scores in all tablets, the mean values of the largest and smallest were selected for immunohistochemical scoring. Strong staining of the nucleus was positive, weak staining of the nucleus or cytoplasm was negative. Among them, if the number of positive cells was ≥10%, it was referred to as high Ki67LI, and if the number of positive cells was <10%, it was referred to as low Ki67LI.
Data Processing
SPSS20.0 was used for statistical analysis. Continuous variables were marked with mean ± standard deviation, the t-test and Mann–Whitney test were used for comparison, and the rate or percentage of categorization variables were marked. The Chi-square test and Fisher’s exact test were mainly used for comparison. P<0.05 was considered statistically significant.
Results and Discussion
Expression of S1PR5 in Colon Cancer Cell Lines
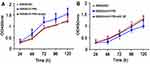
The expression levels of S1PR5 mRNA in four colon cancer cell lines, SW480, SW620, hct-116, and Lovo, and in normal colonic epithelial cells from the NCM460 cell line were detected using qRT-PCR. The results showed that the expression of S1PR5 mRNA in all four colon cancer cell lines was significantly higher than that of normal colonic epithelial cell lines (NCM460) (p<0.05). Among the four colon cancer cell lines, S1PR5 mRNA expression was the lowest in SW480 and the highest in SW620 (Figure 1A). Western blot also confirmed that the expression of S1PR5 protein in colon cancer cells was lower than that in normal colonic mucosal tissues (Figure 1B). Therefore, the role and mechanism of S1PR5 in colon cancer were investigated by down-regulating the expression of S1PR5 in SW620 cells and up-regulating the expression of S1PR5 in SW480 cells.
Tumor Weight of Each Group of Nude Mice
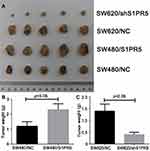
The nude mouse model was prepared by subcutaneous injection. Tumors were removed and weighed. Tumors of SW480/NC, SW480/S1PR5, SW620/NC, and SW620/shS1PR5 groups weighed 1.54 ± 0.1, 2.8 ± 0.4, 1.8 ± 0.2, and 0.5 ± 0.1 g, respectively (Figure 2). Based on these tumor volume results, it was found that S1PR5 promotes the subcutaneous tumorigenic ability of colon cancer cells.
Molecular Role of S1PR5
To further analyze the potential role of S1PR5 in colon cancer, two cell lines were established: the SW480 colon cancer cell line with stable overexpression of S1PR5, and the SW620 colon cancer cell line knocked out by S1PR5 expression, which is commonly detected using Western blot in colon cancer. Western blot analysis of the signaling molecules of common signaling pathways, such as AKT, STAT3, JNK, and NF-кB, showed that the change in S1PR5 expression only affected the NF-кB signaling pathway (Figure 3). These results suggest that S1PR5 may play a role in regulating the NF-кB signaling pathway. The hallmark event of the NF-кB signaling pathway is the translocation of p65 into the nucleus. Therefore, the relationship between S1PR5 and the NF-кB signaling pathway was investigated through evaluation of the distribution of p65 in the cytoplasm and the nucleus.
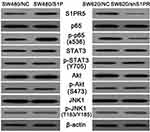 |
Figure 3 Western blot analysis of the effects of S1PR5 expression on common signaling molecules. |
S1PR5 Regulates P65 Activity in Colon Cancer Cells
Nucleolar protein is used to support the nuclear envelope and participate in the disintegration and re-formation of the nuclear envelope in the cell cycle. The effects of S1PR5 expression on p65 expression in the cytoplasm and nucleus were detected by WB. In this study, the results showed that the expression of S1PR5 was correlated with the presence of p65 in the nucleus. In S1PR5 knockout cells, p65 levels decreased in the nucleus. However, in S1PR5 overexpressing cells, p65 levels in the nucleus increased, with the difference being statistically significant (P < 0.05) (Figure 4).
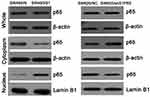 |
Figure 4 Effects of S1PR5 on cytoplasmic and nuclear p65 levels. |
Effect of P65 Intervention on S1PR5-Related Cell Biological Function
Furthermore, the relationship between NF-κB and S1PR5-related biological functions was confirmed. The SW480 cell line, which overexpresses S1PR5, was transfected with shRNA p65, and the S1PR5 knockdown SW620 cell line was transfected with p65 eukaryotic expression plasmid. The effect of p65 on the biological function of S1PR5 was evaluated.
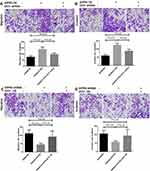
The CCK-8 experiment showed that SW480 cells proliferated significantly after S1PR5 overexpression, while SW480 cells proliferated after transfection of shRNA p65, which did not differ from SW480 cells in the control group (Figure 5A). After S1PR5 expression was knocked down, SW620 cell proliferation was significantly inhibited, and after transfection with p65 eukaryotic expression plasmid, SW620 cell proliferation ability was restored, with no difference between SW620 cells and the control group (Figure 5B). This suggests that p65 is important for S1PR5 as a target.
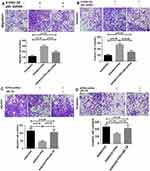
Transwell experiments showed that after S1PR5 overexpression, the migration and invasion ability of SW480 cells was significantly enhanced, and after transfection with shRNA p65, the proliferation and migration of SW480 cells decreased. There was no difference between SW480 cells and the control group. After S1PR5 expression was knocked down, the migration and invasion abilities of SW620 cells were significantly inhibited, and after transfection with p65 eukaryotic expression plasmid, SW620 cells migrated and invaded with no difference between the SW620 cells and the control group, suggesting that p65 is important for S1PR5 as a target (Figures 6–9).
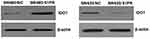 |
Figure 8 Effect of S1PR5 expression on IDO1 protein expression. |
Screening for Downstream Target Genes and Validation of NF-κB
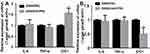
NF-κB is an important transcription factor that exhibits increased activity in most tumor cells. A variety of carcinogenic factors may promote cell growth by activating the NF-κB pathway, thereby causing malignant transformation of the cells and promoting tumor cell metastasis.13 As a transcription factor, NF-κB plays a biological role mainly by regulating downstream gene expression. Since colon cancer is mostly accompanied by an inflammatory response, it was speculated that NF-κB promotes the development of colon cancer by regulating the expression of inflammatory factors. Biosignal analysis and literature reviews were used to identify IL-6, TNF-α, and indoleamine 2, 3-dioxygenase 1 (IDO1) as downstream effectors of the NF-κB pathway,14,15 which are involved in the inflammatory response. These three genes were speculated to be potential target genes for NF-κB in colon cancer. qRT-PCR was first used to detect the expression of common NF-κB downstream target genes, including IL family members and IDO1, in SW480/S1PR5 and SW480/shS1PR5 cells (Figures 6 and 7). The results showed that IDO1 expression was consistent with the trend of NF-κB expression, suggesting that S1PR5 regulates the expression IDO1 through NF-κB, thereby promoting the proliferation and metastasis of colon cancer. It has been shown previously that IDO1, a cancer gene, is abnormally elevated in various tumors.16
IDO1 Is an Important Target for S1PR5 in Colon Cancer
The expression S1PR5 was found to correspond with the expression of IDO1, and this expression trend was consistent with the change in NF-κB expression. IDO1 is a downstream target gene of NF-κB, which lends the speculation that S1PR5 is involved in colon cancer. Since IDO1 is regulated by NF-κB, it could play a role in cancer cell proliferation, migration, and invasion. To confirm this hypothesis, the expression of IDO1 was inhibited in SW480/S1PR5 cells and up-regulated in SW620/shS1PR5 cells, and changes in cell proliferation, migration, and invasion were evaluated.
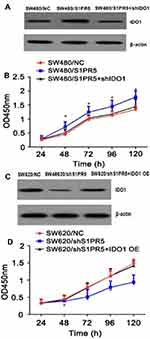
Overexpression of S1PR5 resulted in an increase in IDO1 levels, as revealed by Western blot, while the CCK-8 assay showed that SW480 cell growth increased significantly. Overexpression of S1PR5 in SW480 cells transfected with IDO1 shRNA resulted in a decrease in IDO1 expression levels, as revealed by Western blot, while the CCK-8 assay results showed that SW480 cell growth decreased. There was no difference between the SW480 cells and the control group (P > 0.05) (Figure 12). Knockdown of S1PR5 expression gave rise to a decrease in IDO1 expression, as revealed using Western blot, while the CCK-8 assay results showed that SW620 cell proliferation was significantly inhibited. S1PR5 expressing SW1 cells transfected with IDO1 overexpression plasmid showed a restoration in SW620 cell proliferation ability. No statistically significant (P > 0.05) difference between SW620 and control group cells was observed (Figure 13).
Transwell experiments showed that S1PR5 overexpression caused a significant enhancement in the migration and invasion abilities of SW480 cells, while transfection with IDO1 shRNA caused a decrease in the proliferation and migration abilities of SW480 cells, with no difference between SW480 cells and the control group (P > 0.05). After S1PR5 expression was knocked down, the migration and invasion abilities of SW620 cells were significantly inhibited. Further, after transfection with the IDO1 eukaryotic expression plasmid, SW620 cells migrated and invaded, and no difference was found between SW620 cells and the control group (P > 0.05) (Figures 9 and 10).
IDO1 and p-p65 are Highly Expressed in Colon Cancer Clinical Samples, and S1PR5 is Positively correlated with IDO1 and p-p65 Expression
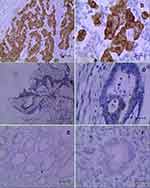
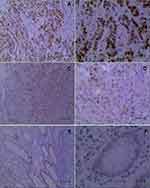
Immunohistochemistry was used to detect the expression IDO1 and p-p65 in clinical samples, and their correlation with S1PR5 expression was analyzed. The results showed that IDO1 and p-p65 were highly expressed in the colon cancer samples and correlated with S1PR5 expression (Tables 1–4). IDO1 was mainly expressed in the cytoplasm and could be seen as brown granules. Positive expression was judged under the microscope, using the following criteria: the cell membrane or cytoplasm had brownish-yellow particles, and the staining was significantly higher than the background. p-p65 is mainly expressed in the nucleus, and brown granules indicate positive expression (Figures 11 and 12).
 |
Table 1 Expression of IDO1 in Colon Cancer and Adjacent Tissues (n=98) |
 |
Table 2 Expression of p-p65 in Colon Cancer and Adjacent Tissues (n=98) |
 |
Table 3 Correlation Analysis Between S1PR5 and IDO1 in Colon Cancer |
 |
Table 4 Correlation Analysis Between S1PR5 and p-p65 in Colon Cancer |
Expression of S1PR5, IDO1, and p-p65 in Subcutaneous Tumors of Colon Cancer in Nude Mice
After the subcutaneous tumor-bearing nude mice were sacrificed, the tumors were removed, and Western blot was used to detect the expression levels of S1PR5, IDO1, and p-p65 (Figure 13). Consistent with the results from clinical samples, the expression trends of p-p65 and IDO1 aligned with the expression trend of S1PR5.
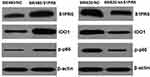 |
Figure 13 Expression of S1PR5, IDO1, and p-p65 in subcutaneous tumor model of colon cancer in nude mice. |
Discussion
S1PRs belong to the G protein-coupled receptor family and are involved in the regulation of almost all important cellular signaling pathways.17 Under S1P stimulation or constitutive activation, S1PRs may activate pathways, such as ERK2, JNK, PI3K/p38 MAPK, and PI3K/AKT/mTOR, through different subunits of the coupled G protein, which further activate YAP, AP-1, and transcription factors, such as STAT3 and NF-κB, in order to promote the expression of effector molecules and exert their biological effects. Consistent with the results of functional studies of S1PRs in tumors, information on the specific mechanism by which S1PRs regulate tumors is also mainly obtained from related studies of S1PR-3. S1PR1 can promote cell proliferation, survival, and migration by regulating multiple signaling pathways, such as Ras/ERK, PI3K/AKT, PI3K/Rac, STAT3, and PLC. Among them, the positive feedback regulation between S1PR1 and STAT3 is an important mechanism for S1PR1-mediated promotion of tumor progression. Overexpression of S1PR1 can activate STAT3, which can induce the transcription of S1PR1, thereby maintaining the activity of STAT3, sustained expression and secretion of IL-6, and activation of IL6-JAK-STAT3 in tumors. The positive feedback regulation between S1PR1 and STAT3 plays important roles in implantation at the distal end of myeloma cells, gemcitabine resistance in pancreatic cancer, angiogenesis and metastasis of hepatocellular carcinoma, activation of diffuse large B-cell lymphoma, and regulation of T-cells in tumor-related processes, such as infiltration.18,19 S1PR2 may regulate PI3K/AKT, mTOR, and other signaling pathways to promote tumor progression. It can also act as a signaling molecule downstream of the TGF-β/TGF-βR2/SMAD1 signaling pathways, inhibiting the progression of diffuse large B-cell lymphoma. Typically, S1PR2 promotes survival and invasive proliferation of hepatoma cells and esophageal cancer stem cells by activating the YAP/Hippo signaling pathway. S1PR3 can also activate pathways such as YAP and PI3K/AKT in order to affect tumorigenesis. S1PR3 is more representative, specifically owing to its regulation of the Notch signaling pathway. In triple-negative breast cancer cells, S1PR3 mediates the role of SphK1/S1P in promoting malignant progression of tumor cell proliferation, invasion, and chemotherapy resistance. In breast cancer stem cells, S1PR3 also mediates the activation of Notch SphK1/S1P, where activation of Notch is independent of the presence of ligands for Notch.20
There is little research on S1PR5 in tumors. Studies in HeLa cells have shown that S1PR5 can mediate the activation of the PI3K/AKT pathway by S1P, thereby promoting mitosis. In addition, studies in a mouse model of Huntington’s disease have shown that S1PR5 may be involved in the activation of signals, such as BDNF, AKT, and ERK. Therefore, the molecular mechanism by which S1PR5 plays a role in tumors remains to be elucidated. In the present study, the effects of S1PR5 expression on the activity of signaling molecules known to be regulated by S1PRs, such as AKT, ERK, NF-kB, and STAT3, were examined in order to reveal the molecular mechanism by which S1PR5 plays a role in colon cancer. The results showed that activation of NF-kB was the most significant, especially after a change in S1PR5 expression, which suggests that NF-κB may be a key mediating factor for the tumorigenic function of S1PR5 in colon cancer. Further, functional experiments showed that inhibition of NF-κB activity may significantly antagonize the cancer-promoting function of S1PR5 in colon cancer. Therefore, our study confirmed that S1PR5 promotes colon cancer progression mainly by activating NF-κB.
The NF-κB signaling pathway plays a key role in numerous physiological and pathological processes. It is responsible for regulating the immune system, inflammation, infection, and other stress responses.21–25 An abnormal NF-κB signaling pathway is closely related to tumors, atherosclerosis, diabetes, chronic inflammatory diseases, and other major diseases that threaten human life.1,26 In most tumors, abnormal NF-κB signaling is one of the main features, and it helps promote tumorigenesis and drug resistance by regulating the expression of genes involved in apoptosis, invasion, tumor development, treatment resistance, and recurrence. In this study, we demonstrated that S1PR5 can activate NF-κB activity in a classical manner, and inhibition of NF-κB activity is resistant to the effects of S1PR5 on the proliferation, migration, and invasion of colon cancer cells, indicating that NF-κB mediates the role of S1PR5 in promoting the malignant progression of colon cancer. This study not only reveals the molecular mechanism by which S1PR5 promotes tumorigenesis in colon cancer but also demonstrates that NF-κB plays an important role in colon cancer; thus, targeting the combination of S1PR5 and the NF-κB pathway may prove to be an effective treatment modality for the inhibition of colon cancer.
IDO1 is one of the rate-limiting enzymes that catalyzes tryptophan catabolism by the kynurenic acid pathway.27–29 Abnormal metabolism of tryptophan or IDO1 may cause severe inhibition of the immune system and have serious consequences. Therefore, enhanced tryptophan metabolism is an important mechanism for tumor cell immune escape. In immune and inflammatory tumor microenvironments of colon cancer,30–33 IDO1 can be regulated by NF-κB transcription.
Conclusions
It was speculated that IDO1 may be affected by S1PR5 regulation, since NF-κB mediates the function S1PR5. The results confirmed that S1PR5 may regulate the expression level of IDO1 and be NF-κB dependent. Knockdown or overexpression of IDO1 may antagonize or restore, respectively, the effects of S1PR5 overexpression/knockdown on the growth, migration, and invasion of colon cancer cells. Additionally, in colon cancer tissues, high expression levels of S1PR5, NF-κB, and IDO1 showed a significant positive correlation with each other. Nucleolar protein supports the nuclear envelope and participates in the disintegration and re-formation of the nuclear envelope in the cell cycle. In summary, this study demonstrated that S1PR5 induces the expression of IDO1 in an NF-κB-dependent manner, thereby promoting the malignant progression of colon cancer, and that targeting S1PR5/NF-κB/IDO1 may help inhibit colon cancer.
Ethical Review
All of the procedures were carried out in accordance with the Chinese Guidelines for Animal Welfare and Experimental Protocol and were approved by the Animal Care and Use Committee of the Xiangya Hospital, Central South University.
Acknowledgments
This work was supported by the Natural Science Foundation of Hunan Province (Grant No. 2018JJ2239).
Disclosure
The authors report no conflicts of interest in this work.
References
1. Zhong L, Luo Y, Huang C, et al. Effect of NF-κB decoy on insulin resistance of adipocytes from patients with type 2 diabetes mellitus. Diabetes Metab. 2011;37(6):520–526. doi:10.1016/j.diabet.2011.04.004
2. Pham TT, Talukder AM, Walsh NJ, et al. Clinical and epidemiological factors associated with suicide in colorectal cancer. Supportive Care Cancer. 2018;27(3):617–621.
3. Gu W, Zhu Y, Ye D. Beyond chemotherapy for advanced disease-the role of EGFR and PD-1 inhibitors. Transl Androl Urol. 2017;6(5):848–854. doi:10.21037/tau.2017.03.92
4. Tabung FK, Steck SE, Ma Y, et al. Changes in the inflammatory potential of diet over time and risk of colorectal cancer in postmenopausal women. Am J Epidemiol. 2017;186(5):21–27. doi:10.1093/aje/kwx038
5. Dashti SG, Buchanan DD, Jayasekara H, et al. Alcohol consumption and the risk of colorectal cancer in mismatch repair gene mutation carriers. Cancer Epidemiol Biomark Prev. 2017;26(3):366–373. doi:10.1158/1055-9965.EPI-16-0496
6. Patmanathan SN, Wang W, Yap LF, et al. Mechanisms of sphingosine 1-phosphate receptor signalling in cancer. Cell Signal. 2017;34:66–75. doi:10.1016/j.cellsig.2017.03.002
7. Xie Z, Liu H, Geng M. Targeting sphingosine-1-phosphate signaling for cancer therapy. Sci China Life Sci. 2017;60(6):585–595. doi:10.1007/s11427-017-9046-6
8. Park SJ, Im DS. Sphingosine 1-phosphate receptor modulators and drug discovery. Biomol Ther (Seoul). 2017;25(1):80–90. doi:10.4062/biomolther.2016.160
9. Sukocheva OA. Expansion of sphingosine kinase and sphingosine- 1-phosphate receptor function in normal and cancer cells: from membrane restructuring to mediation of estrogen signaling and stem cell programming. Int J Mol Sci. 2018;19(2):420–430. doi:10.3390/ijms19020420
10. Delgado A, Martínezcartró M. Therapeutic potential of the modulation of sphingosine 1 phosphate receptors. Curr Med Chem. 2016;23(3):1–5. doi:10.2174/0929867323666151207111509
11. Pyne S, Adams DR, Pyne NJ. Sphingosine 1-phosphate and sphingosine kinases in health and disease: recent advances. Prog Lipid Res. 2016;62:93–106. doi:10.1016/j.plipres.2016.03.001
12. Huwiler A, Zangemeister-Wittke U. The sphingosine 1-phosphate receptor modulator fingolimod as a therapeutic agent: recent findings and new perspectives. Pharmacol Ther. 2018;185:34–49. doi:10.1016/j.pharmthera.2017.11.001
13. Pires BR, Silva RC, Ferreira GM, et al. NF-kappaB: two sides of the same coin. Genes. 2018;9(1):24–31. doi:10.3390/genes9010024
14. Wang L, Ming L, Mei Y, et al. Caffeic acid attenuates the autocrine IL-6 in hepatocellular carcinoma via the epigenetic silencing of the NF-kB-IL-6-STAT-3 feedback loop. RSC Adv. 2015;5(65):52952–52957. doi:10.1039/C5RA05878C
15. Subedi L, Venkatesan R, Kim SY. Neuroprotective and anti-inflammatory activities of allyl isothiocyanate through attenuation of JNK/NF-κB/TNF-α signaling. Int J Mol Sci. 2017;18(7):1423–1427.
16. Cesario A, Rocca B, Rutella S. The interplay between indoleamine 2,3-dioxygenase 1 (IDO1) and cyclooxygenase (COX)-2 in chronic inflammation and cancer. Curr Med Chem. 2010;18(15):2263–2271. doi:10.2174/092986711795656063
17. Tang HB, Jiang XJ, Wang C, et al. S1P/S1PR3 signaling mediated proliferation of pericytes via Ras/pERK pathway and CAY10444 had beneficial effects on spinal cord injury. Biochem Biophys Res Commun. 2018;498(4):830–836. doi:10.1016/j.bbrc.2018.03.065
18. Lankadasari MB, Aparna JS, Mohammed S, et al. Targeting S1PR1/STAT3 loop abrogates desmoplasia and chemosensitizes pancreatic cancer to gemcitabine. Theranostics. 2018;8(14):3824–3840. doi:10.7150/thno.25308
19. Silva VR, Katashima CK, Bueno Silva CG, et al. Hypothalamic S1P/S1PR1 axis controls energy homeostasis in middle-aged rodents: the reversal effects of physical exercise. Aging. 2017;9(1):142–155.
20. Wang YC, Tsai CF, Chuang HL, et al. Benzyl butyl phthalate promotes breast cancer stem cell expansion via SPHK1/S1P/S1PR3 signaling. Oncotarget. 2016;7(20):29563–29576. doi:10.18632/oncotarget.9007
21. Schmitz M, Shaban M, Albert B, et al. The crosstalk of endoplasmic reticulum (ER) stress pathways with NF-κB: complex mechanisms relevant for cancer, inflammation and infection. Biomedicines. 2018;6(2):58–69. doi:10.3390/biomedicines6020058
22. Hop HT, Arayan LT, Reyes A, et al. Heat-stress-modulated induction of NF-κB leads to brucellacidal pro-inflammatory defense against Brucella abortus infection in murine macrophages and in a mouse model. BMC Microbiol. 2018;18(1):44–57. doi:10.1186/s12866-018-1185-9
23. Maracle CX, Agca R, Helder B, et al. Noncanonical NF-κB signaling in microvessels of atherosclerotic lesions is associated with inflammation, atheromatous plaque morphology and myocardial infarction. Atherosclerosis. 2018;270:33–41. doi:10.1016/j.atherosclerosis.2018.01.032
24. Sahanfirat S, Temizresitoglu M, Guden DS, et al. Protection by mTOR inhibition on zymosan-induced systemic inflammatory response and oxidative/nitrosative stress: contribution of mTOR/MEK1/ERK1/2/IKKβ/IκB-α/NF-κB signalling pathway. Inflammation. 2018;41(1):276–298. doi:10.1007/s10753-017-0686-2
25. Akanda MR, Nam HH, Tian W, et al. Regulation of JAK2/STAT3 and NF-κB signal transduction pathways; Veronica polita alleviates dextran sulfate sodium-induced murine colitis. Biomed Pharmacother. 2018;100(2):296–303. doi:10.1016/j.biopha.2018.01.168
26. Maracle CX, Agca R, Helder B, et al. AB0265 non-canonical NF-κB signaling in microvessels of atherosclerotic lesions in coronary arteries is associated with inflammatory cell infiltration and myocardial infarction. Ann Rheum Dis. 2016;75(2):982–990.
27. Zhang G, Xing J, Wang Y, et al. Discovery of novel inhibitors of indoleamine 2,3-dioxygenase 1 through structure-based virtual screening. Front Pharmacol. 2018;9(2):277–287. doi:10.3389/fphar.2018.00277
28. Zhang SN, Liu X, Yang Q, et al. The study of indoleamine 2,3- dioxygenase 1 and its inhibitors. Fudan Univ J Med Sci. 2017;44(1):1–7.
29. Zhai L, Spranger S, Binder DC, et al. Molecular pathways: targeting IDO1 and other tryptophan dioxygenases for cancer immunotherapy. Clin Cancer Res off J Am Assoc Cancer Res. 2015;21(24):5427–5434. doi:10.1158/1078-0432.CCR-15-0420
30. Muller AJ, Smith C, Chang MY, et al. Abstract 3665: IDO1 is an integrative determinant of tumor-promoting, pathogenic inflammation. Cancer Res. 2014;74(19):3665–3673.
31. Shinde R, Shimoda M, Chaudhary K, et al. B Cell-intrinsic IDO1 regulates humoral immunity to T cell-independent antigens. J Immunol. 2015;195(5):2374–2382. doi:10.4049/jimmunol.1402854
32. Wang XF, Wang HS, Wang H, et al. The role of indoleamine 2,3- dioxygenase (IDO1) in immune tolerance: focus on macrophage polarization of THP-1 cells. Cell Immunol. 2014;289(2):42–48. doi:10.1016/j.cellimm.2014.02.005
33. Wang C, Mao J, Redfield S, et al. Systemic distribution, subcellular localization and differential expression of sphingosine-1-phosphate receptors in benign and malignant human tissues. Exp Mol Pathol. 2014;97(2):259–265. doi:10.1016/j.yexmp.2014.07.013
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.