Back to Journals » Cancer Management and Research » Volume 13
The Prevalence and Concurrent Pathogenic Mutations of KRASG12C in Northeast Chinese Non-small-cell Lung Cancer Patients
Authors Liu Y, Li H, Zhu J, Zhang Y, Liu X, Li R, Zhang Q, Cheng Y
Received 23 September 2020
Accepted for publication 17 February 2021
Published 15 March 2021 Volume 2021:13 Pages 2447—2454
DOI https://doi.org/10.2147/CMAR.S282617
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Beicheng Sun
Yan Liu,1 Hui Li,1 Jing Zhu,2 Yang Zhang,2 Xianhong Liu,2 Rixin Li,1 Qiang Zhang,3 Ying Cheng1,2
1Medical Oncology Translational Research Lab, Jilin Provincial Key Laboratory of Molecular Diagnostics for Lung Cancer, Jilin Cancer Hospital, Changchun, 130012, People’s Republic of China; 2Department of Medical Thoracic Oncology, Jilin Cancer Hospital, Changchun, 130012, People’s Republic of China; 3Department of Bioinformatics, Burning Rock Biotech, Guangzhou, People’s Republic of China
Correspondence: Ying Cheng
Jilin Cancer Hospital, No. 1066 Jinhu Road, Chaoyang District, Changchun, Jilin Province, 130012, People’s Republic of China
Tel +86 43185879901
Email [email protected]
Objective: KRAS mutation is one of important driver genes in non-small-cell lung cancer (NSCLC) and the patients with KRASG12C mutations benefit from the inhibitor AMG510. However, the frequency, concurrent pathogenic mutations, and clinical characteristic of KRASG12C is unknown in the NSCLC population of Northeast China.
Methods: The retrospective analysis was derived from 431 NSCLC patients in Jilin Cancer Hospital between January 2018 and June 2019. The mutation frequency and concurrent mutations of KRASG12C in tumor or peripheral blood was detected by next-generation sequencing (NGS).
Results: The RAS mutant rate was observed in 10.7% (46/431) of this cohort. All RAS-driver cancers are caused by mutations in the KRAS isoform, while the NRAS and HRAS isoforms were not detected. Among KRAS-mutant patients, 42 (91.3%) showed exon 2 mutation in 12 codon and 13 codon. KRASG12C showed a 4.6% (20/431) mutation rate in this cohort and the highest frequency (43.5%, 20/46) in KRAS-mutant-positive patients. There was no difference between tumor tissue and plasma in terms of either KRAS or KRASG12C mutation. The most frequent co-occurrence mutations with KRASG12C were TP53, followed by PTEN. Furthermore, KRASG12C was exclusive with STK11 mutation. KRASG12C mutation was associated with age, disease stage, and smoking status (P=0.024; P=0.02; P=0.006), smoking remained an independent factor for KRASG12C mutation (P=0.037), and higher mutation frequency in patients older than 60, stage I–III, or smoking in NSCLC (P=0.0151, P=0.0343, P=0.0046, respectively).
Conclusion: KRAS mutation was the only isoforms of RAS family, of these 43.5% harbored the KRASG12C subtype in northeastern Chinese NSCLC patients. KRASG12C is associated with age, pathological stage and smoking status, more commonly harbored TP53/PTEN mutations, and providing more genome profile for targeted therapy in local clinical practice.
Keywords: next-generation sequencing, non-small-cell lung cancer, KRASG12C, tissue, plasma, mutations
Introduction
Non-small-cell lung cancer (NSCLC) is the most common histological type of lung cancer, accounting for 80–85% of lung cancers and has become the most fatal cancer in the world.1 Recently, targeted therapy based on various driver oncogene variants (EGFR, ALK and ROS1, KRAS, MET, PIK3CA, RET, BRAF) has shown great antitumor activity; unfortunately, KRAS mutations had a more complicated mechanism in comparison with other driver genes such as EGFR, with poor prognosis and high risk of tumor recurrence.2 Although prevalent, no specific treatment has been successfully developed for these NSCLCs.
KRAS mutations are some of the most prevalent alterations, approximately 10% of Asian NSCLC patients and 7.5% of Chinese NSCLC patients harbor the KRAS mutation, with codon 12 and 13 mutations being the most frequent and the most common subtypes are G12C, G12V and G12D.3,4 KRASG12C is a mutant type of KRAS guanosine triphosphatase (GTPase), and an inhibitor targeting KRASG12C is a promising novel tumor-specific therapy for tumors driven by mutant proteins.5 Current studies on KRASG12C inhibitors and the mechanism of drug resistance have confirmed that patients with KRASG12C mutations benefit from the inhibitor AMG510,6 which has also been approved by the FDA as an orphan drug for NSCLC and colon cancer with KRASG12C mutation. KRASG12C can induce allosteric switch II pocket (s-iip) and take cys-12 as the specific covalent target of alleles, which were considered as potential drug targets.2 Now, KRASG12C mutation was verified by the NGS, various clinical parameters and genetic mutation have been proposed to predict the relevance with KRASG12C (such as sex, age, smoking, co-mutation gene). In the current study we aim to discover a more precise delineation of candidate target populations and distinctive KRASG12C co-mutation subtypes in the northeast Chinese population. We retrospectively investigated and evaluated the KRASG12C mutation in northeast Chinese NSCLC, and the association between clinical factors and KRASG12C mutation status.
Materials and Methods
Patients and Samples
Four hundred and thirty-one samples were collected from Jilin Cancer Hospital between January 2018 and June 2019, 268 cases were tested through eight gene panel, 81 cases by 168 gene panel and 82 matched cases using 520 gene panel, respectively (Figure 1). Clinic pathological data were collected from the electronic medical records in Jilin Cancer Hospital, and the factors included age, sex, and clinical stage, smoking history, brain metastasis, PS score and histology. All participants signed the informed consent agreement before participating in the study, the data were anonymized, the study was approved by the Clinical Research Ethics Committee of Jilin Cancer Hospital and was conducted in accordance with the Declaration of Helsinki.
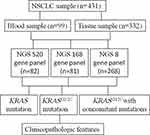 |
Figure 1 Study flowchart. Abbreviations: NSCLC, non-small-cell lung cancer; NGS, next-generation sequencing. |
DNA Extraction
DNA was extracted by DNA FFPE tissue kit (AmoyDx, China) and ctDNA extraction kit (QIAGEN, Germany) according to the manufacturer’s instructions. DNA concentration was quantified by Nanodrop 3000C and Qubit 4.0 (Thermo Fisher Scientific, Waltham, MA, USA).
Next-generation Sequencing Analysis
Library preparation was performed following manufacturer’s protocol (Burning Rock Biotech, Guangzhou, China). DNA Fragments (range: 200–400 bp) were purified by AMPure beads (Beckman Coulter, CA, USA), and captured with probe baits, hybrid selection with magnetic beads by RT-PCR amplification. Subsequently, DNA quality and size were assessed by high-sensitivity DNA assay. Indexed samples were sequenced on a MiSeq system (Beckman Coulter) with paired-end reads. The input of extracted DNA should be in the range of (30−200 ng). Sequencing platform was used by Illumina NextSeq 500 Sequencing Platform with tissue DNA (1000X) and cfDNA (20000X). All samples were analyzed by NGS targeted panel (Burning Rock Dx, China), which eight-gene panel covers well-known lung adenocarcinoma driver genes, 168 genes covers known lung cancer-related genes and 520 genes covers solid tumor-related genes. (Supplemental Table 1).
Statistical Analysis
All data was performed by SPSS Statistics 19.0 software (IBM Corporation, Armonk, NY, USA). Fisher’s exact test was used to evaluate mutation differences and clinical factor between KRASG12C and KRASwt. Logistic regression analysis was used to identify as independent factors for KRASG12C mutations. A P-value of <0.05 was considered statistically significant.
Results
Patient Population
Among 431 samples were those from tumor tissue 332 (77.04%), 99 (22.96%) plasma; 198 women (54.07%) and 233 men (45.93%), with a median age of 63 years (range: 34–86 years), respectively. Of the 431 patients, 263 (61.02%) were smokers, and 168 were nonsmokers. The histological characterization of tumors revealed that 370 samples were adenocarcinoma (85.85%), 61 were squamous cell carcinoma (14.15%). Of the 431 patients, characterization of the pathological stage showed 115 samples in stage I–III (26.68%), and 316 samples in stage IV (73.32%) (Table 1).
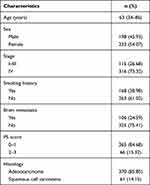 |
Table 1 Patient Characteristics |
KRASG12C is the Most Common Mutation Type of KRAS in NSCLC
The RAS mutation rate was 10.7% (46/431), and KRAS was the only mutation subtype of RAS (NRAS, KRAS, HRAS). 42 (91.3%) indicated KRAS gene exon 2 mutation, 12 and 13 codon of KRAS gene mutations were detected, and KRASG12C showed the highest frequency, the total mutation rate of KRASG12C in NSCLC was 4.6% (20/431) and 43.5% (20/46) of KRAS mutant subtypes, followed by 17.4% (8/46) of KRASG12D, 8.7% (4/46) of KRASG12V, and 8.7% (4/46) of KRASG12A. The mutation frequency of other KRAS types was lower (Figure 2).
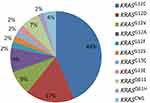 |
Figure 2 Mutation frequencies of KRAS subtypes. |
KRASG12C Mutation Between Tumor Tissue and Plasma
We compared the KRAS mutation spectrums between tumor tissue and ctDNA derived from peripheral blood in this study. Collectively, 37 (11.14%) and 16 (4.81%) patients had KRAS and KRASG12C mutation spectrum in tumor tissue, nine (9.09%) and four (4.04%) patients in ctDNA, but no significant difference was found in the two sample types (P=0.711, P=1.000, Table 2), respectively.
 |
Table 2 Mutation Frequencies of KRAS Subtypes Between Tumor Tissue and Plasma |
Co-occurring Genomic Alterations Between KRASG12C and Lung Cancer Pathogenic Gene
Lung cancer driver genes (include EGFR, RAS, ALK, ROS1, MET, RET BRAF, and HER-2) mutation samples were observed in 332 (77.3%) of 431 patients. Eight (40%) of 20 patients harbored only KRASG12C mutations, and 12 (60%) had multiple KRASG12C mutations, including eight (40%) KRASG12C patients had co-occurring driver oncogenes, was higher trend than KRASother with driver oncogenes mutations (6/26,23%), but no statistical significance (P=0.33), the most commonly co-occurring genomic alterations with KRASG12C were EGFR (10%, 2/20), ROS1 (10%, 2/20), MET (10%), HER2 (5%, 1/20), ALK (5%, 1/20), BRAF (5%, 1/20), and RET (0%), respectively (Figure 3, Supplemental Table 2). One hundred and sixty-three patients from 168 gene panel or 520 gene panel found that the KRASG12C gene is often accompanied by TP53 and PTEN mutation, the mutation rates were 50% (3/6) and 16.7% (1/6), respectively, but STK11 (0.0%, 0/6).
Age, Smoking History and Pathological Stage Associated with KRASG12C Mutation
The mutation rate of KRASG12C gene in smokers was higher than that in nonsmokers, 8.33% (14/168) vs 2.28% (6/263), P=0.0046). KRASG12C has a higher mutation rate in age (≥60 years) 15.2% (18/274) vs 1.27% (2/157); P=0.0151). KRASG12C mutation was associated with the pathological staging of the patients, 8.69% (10/115) vs 3.16% (10/316), P=0.0343), but was not associated with gender, brain metastasis, PS score, and histology (P=0.2515, P=0.4282, P=0.5266 and P=0.7526) (Table 3), to further identify the values of clinical factor on KRASG12C mutations, logistic regression analysis was included. In the univariate logistic analysis, age, smoker, clinical stage were identified as independent factors for KRASG12C mutations (OR=0.551, P=0.024; OR=5.449, P=0.02; OR=0.343, P=0.006). In the multivariate logistic model, smoker (OR=0.306, P=0.037) remained independent factors for KRASG12C (Table 4). Furthermore, we found that KRASG12C was dominant in male smokers (100%, 4/4)
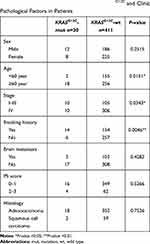 |
Table 3 431 Correlation Analysis Between KRASG12C and Clinic Pathological Factors in Patients |
 |
Table 4 Univariate and Multivariate Analysis of KRASG12C and Clinical Factor |
Discussion
Previously reported RAS was detected in about 25–30% of tumors, several studies consistently reported that Westerners have a higher mutation rate than Asians (26% vs 11%).7 Another report similarly indicated 30% of RAS mutations in Western patients and 5–15% in the Asian population,8 which accounts for about 86% KRAS, 11% NRAS and 3% HRAS mutation of RAS-induced NSCLC, KRAS accounts for 90% of RAS gene mutations in lung adenocarcinoma and is the most common oncogene in NSCLC.9 Our data are consistent with recent studies, our results might indicate the current view that KRAS was the only RAS-mutant isoform, the mutation rate was 10.7% in 431 NSCLC patients, similar to the rates reported by Jia’s group and Liu’s group.10,11 Further studies showed that the KRASG12C mutation rate is 4.6% in lung cancer, and 43.5% in KRAS mutation for our study. It was similar to several studies in that the KRASG12C mutation frequency range is from 35% to 45% followed by KRASG12V and KRASG12D in KRAS mutant lung cancer, but a lower frequency reported by Liu’s group.9,11–15 One key finding of our study was that KRAS, including KRASG12C mutation of NSCLC reflected no difference in tissue and blood. Furthermore, this study also reveals the widespread existence of concomitant mutations in patients with KRASG12C mutant advanced NSCLC, especially driver gene mutations. The three predominant KRAS co-mutations were detected including EGFR-KRASG12C (10%), equal to ROS1-KRASG12C (10%) and MET-KRASG12C (10%). We found the four cases with EGFR-KRAS concomitant mutations in our cohort were all tested before EGFR-TKI treatment, thus partly ruling out the possibility that EGFR-KRAS co-mutation was related to EGFR-TKI resistance.16 Unfortunately, neither were the four cases derived from two separate tumor tissue. The incidence rate of EGFR-KRAS in the Chinese cohort might be likely ethnic-unique, based on the knowledge that the prevalence of EGFR mutation is higher in the Asian population.17 The co-occurrence of EGFR and KRAS was 0.92% (4/431) in our study, which was supported by Scheffler et al13 (1.2%). The four concomitant mutations were KRASG12C (n=2) co-occurring with either EGFR V1097I (n=1) or EGFR amplification (n=1) and KRASother (n=2) co-occurring with EGFR 19del (n=2). Although previous studies had reported that KRAS are mutually exclusive with mutations in EGFR and ALK in NSCLC,18,19 but coexisting EGFR and KRAS mutations have also been reported..20,21 (Zhu et al reported that three patients with coexisting EGFR and KRAS mutations were found in 206 patients (1.4%).22 We infer that genetic mutation status could be related with different races, sample numbers, as well as test methodology. Nevertheless, current data about KRAS co-occurring mutations in lung cancer is insufficient. Co-occurrence with TP53 or STK11 mutations is common in KRAS mutations.23,24 KRAS and TP53 co-mutations indicated that tumors harboring those mutations couldbe more responsive to immune checkpoint inhibition in lung cancer.25 Conversely, tumors harboring concurrent KRAS and STK11 mutations could be associated with an immunosuppressive microenvironment.26,27 Furthermore, the absence of PTEN promotes resistance to T cell-mediated immunotherapy.28 So we evaluated the mutation status of TP53, STK11 or PTEN in KRASG12C mutant patients, and it indicated that in the landscape of concurrent genetic alterations in patients with KRASG12C, the co-mutation rates were 50% and 16.7%, but KRASG12C was exclusive with STK11 mutation.
KRASG12C (c.34G>T) alteration is a transversion and KRAS transversion mutations (G→T or G→C) were more common in smokers, in contrast, transition mutations (G→A) were more common in never-smokers in lung adenocarcinomas (n=500).29 Our data showed that smokers more commonly harbored KRASG12C mutations than KRASwt (70% vs 37.5%), which is consistent with reports by Liu et al and Dogan et al.11,30 Data showed that KRAS-mutant NSCLC is genetically complex, with a higher frequency of co-occurring mutations with TP53, STK11, MET and ERBB2 amplifications,29 however, no conclusions implied that the co-occurrence mutations were related to the transversion. In comparison to KRASother, KRASG12C showed higher mutation frequency in patients older than 60 years, and stage I–III. Our findings were supported by other studies.11,13,31
In summary, our study indicated that KRASG12C mutations were the most frequent mutant subtype of KRAS in northeast Chinese NSCLC patients and might be involved in the smoking, age, and clinical stage, especially we demonstrated a high frequency of KRASG12C concomitant TP53/PTEN/EGFR. In addition, no difference was observed between tissue and plasma in the KRASG12C subgroup of the northeast Chinese NSCLC patients. Our findings might contribute to distinct therapeutic guidance in NSCLC. More data should be collected and explored to address predictive and prognostic value of KRASG12C in future studies.
Acknowledgments
This research was supported by Scientific Research Project of Jilin Provincial Health and Family Planning Commission (grant numbers 2018Q007, 2019J077); Science and Technology Agency of Jilin Provincial Project (grant numbers 20200201518JC, 202002063JC); Special Project for Significant New Drug Research and Development in the Major National Science and Technology Projects of China (grant numbers 2020ZX09201-024).
Disclosure
Qiang Zhang is an employee of Burning Rock Biotech. The authors report no other potential conflicts of interests in this work.
References
1. Chen W, Zheng R, Baade PD, et al. Cancer statistics in China, 2015. CA Cancer J Clin. 2016;66(2):115–132. doi:10.3322/caac.21338
2. Janes MR, Zhang J, Li LS, et al. Targeting KRAS Mutant Cancers with a Covalent G12C-Specific Inhibitor. Cell. 2018;172:578–589.e17. doi:10.1016/j.cell.2018.01.006
3. Cancer Genome Atlas Research Network. Comprehensive molecular profiling of lung adenocarcinoma. Nature. 2014;511(7511):543–550. doi:10.1038/nature13385
4. Zhuang X, Zhao C, Li J, et al. Clinical features and therapeutic options in non-small cell lung cancer patients with concomitant mutations of EGFR, ALK, ROS1, KRAS or BRAF. Cancer Med. 2019;8(6):2858–2866. doi:10.1002/cam4.2183
5. Lou K, Steri V, Ge AY, et al. KRAS G12C inhibition produces a driver-limited state revealing collateral dependencies. Sci Signal. 2019;12(583):eaaw9450. doi:10.1126/scisignal.aaw9450
6. Lanman BA, Allen JR, Allen JG, et al. Discovery of a Covalent Inhibitor of KRASG12C (AMG 510) for the Treatment of Solid Tumors. J Med Chem. 2020;63:52–65. doi:10.1021/acs.jmedchem.9b01180
7. Ricciuti B, Leonardi GC, Metro G, et al. Targeting the KRAS variant for treatment of non-small cell lung cancer: potential therapeutic applications. Expert Rev Respir Med. 2016;10(1):53–68. doi:10.1586/17476348.2016.1115349
8. Pylayeva-Gupta Y, Grabocka E, Bar-Sagi D. RAS oncogenes: weaving a tumorigenic web. Nat Rev Cancer. 2011;11:761–774.
9. Ni D, Li X, He X, Zhang H, Zhang J, Lu S. Drugging K-RasG12C through covalent inhibitors: mission possible? Pharmacol Ther. 2019;202:1–17. doi:10.1016/j.pharmthera.2019.06.007
10. Jia Y, Jiang T, Li X, et al. Characterization of distinct types of KRAS mutation and its impact on first-line platinum-based chemotherapy in Chinese patients with advanced non-small cell lung cancer. Oncol Lett. 2017;14:6525–6532. doi:10.3892/ol.2017.7016
11. Liu SY, Sun H, Zhou JY, et al. Clinical characteristics and prognostic value of the KRAS G12C mutation in Chinese non-small cell lung cancer patients. Biomark Res. 2020;8:22. doi:10.1186/s40364-020-00199-z
12. Aredo JV, Padda SK, Kunder CA, et al. Impact of KRAS mutation subtype and concurrent pathogenic mutations on non-small cell lung cancer outcomes. Lung Cancer. 2019;133:144–150. doi:10.1016/j.lungcan.2019.05.015
13. Scheffler M, Ihle MA, Hein R, et al. K-ras mutation subtypes in NSCLC and associated co-occuring mutations in other oncogenic pathways. J Thorac Oncol. 2019;14(4):606–616. doi:10.1016/j.jtho.2018.12.013
14. Izar B, Zhou H, Heist RS, et al. The prognostic impact of KRAS, its codon and amino acid specific mutations, on survival in resected stage I lung adenocarcinoma. J Thorac Oncol. 2014;9(9):1363–1369. doi:10.1097/JTO.0000000000000266
15. Nadal E, Chen G, Prensner JR, et al. KRAS-G12C mutation is associated with poor outcome in surgically resected lung adenocarcinoma. J Thorac Oncol. 2014;9(10):1513–1522. doi:10.1097/JTO.0000000000000305
16. Ortiz-Cuaran S, Scheffler M, Plenker D, et al. Heterogeneous mechanisms of primary and acquired resistance to third-generation EGFR inhibitors. Clin Cancer Res. 2016;22(19):4837–4847. doi:10.1158/1078-0432.CCR-15-1915
17. Han SW, Kim TY, Hwang PG, et al. Predictive and prognostic impact of epidermal growth factor receptor mutation in non-small-cell lung cancer patients treated with gefitinib. J Clin Oncol. 2005;23(11):2493–2501. doi:10.1200/JCO.2005.01.388
18. Gainor JF, Varghese AM, Ou SH, et al. ALK rearrangements are mutually exclusive with mutations in EGFR and KRAS in non-small cell lung cancer. Clin Cancer Res. 2013;19(15):4273–4281. doi:10.1158/1078-0432.CCR-13-0318
19. Unni AM, Lockwood WW, Zejnullahu K, et al. that synthetic lethality underlies the mutual exclusivity of oncogenic KRAS and EGFR mutations in lung adenocarcinoma. Elife. 2015;4(4):e06907. doi:10.7554/eLife.06907
20. Gumerlock PH, Holland WS, Chen H, et al. Mutational analysis of K-RAS and EGFR implicates K-RAS as a resistance marker in the Southwest Oncology Group (SWOG) trial S0126 of bronchioalveolar carcinoma (BAC) patients (pts) treated with gefitinib. J Clin Oncol. 2005;23:623s. doi:10.1200/jco.2005.23.16_suppl.7008
21. Han SW, Kim TY, Jeon YK, et al. Optimization of patient selection for gefitinib in non-small cell lung cancer by combined analysis of epidermal growth factor receptor mutation, K-ras mutation, and Akt phosphorylation. Clin Cancer Res. 2006;12:2538–2544. doi:10.1158/1078-0432.CCR-05-2845
22. Zhu CQ, Sants GC, Ding K, et al. Role of KRAS and EGFR as biomarkers of response to erlotinib in National Cancer institute of Canada Clinical Trial Group study BR.21. J Clin Oncol. 2008;26(26):
23. Pao W, Girard N. New driver mutations in non-small-cell lung cancer. Lancet Oncol. 2011;12:175e180.
24. Ding L, Getz G, Wheeler DA, et al. Somatic mutations affect key pathways in lung adenocarcinoma. Nature. 2008;455(7216):1069e1075. doi:10.1038/nature07423
25. Dong ZY, Zhong WZ, Zhang XC, et al. Potential predictive value of TP53 and KRAS mutation status for response to PD-1 Blockade Immunotherapy in Lung Adenocarcinoma. Clin Cancer Res. 2017;23(12):3012–3024. doi:10.1158/1078-0432.CCR-16-2554
26. Skoulidis F, Goldberg ME, Greenawalt DM, et al. STK11/LKB1 Mutations and PD-1 Inhibitor Resistance in KRAS-Mutant Lung Adenocarcinoma. Cancer Discov. 2018;8:822–835. doi:10.1158/2159-8290.CD-18-0099
27. Schabath MB, Welsh EA, Fulp WJ, et al. Differential association of STK11 and TP53 with KRAS mutation-associated gene expression, proliferation and immune surveillance in lung adenocarcinoma. Oncogene. 2016;35(24):3209–3216. doi:10.1038/onc.2015.375
28. Peng W, Chen JQ, Liu C, et al. Loss of PTEN Promotes Resistance to T Cell-Mediated Immunotherapy. Cancer Discov. 2016;6:202–216. doi:10.1158/2159-8290.CD-15-0283
29. El Osta B, Behera M, Kim S, et al. Characteristics and outcomes of patients with metastatic KRAS-mutant lung adenocarcinomas: the lung cancer mutation consortium experience. J Thorac Oncol. 2019;14(5):876–889. doi:10.1016/j.jtho.2019.01.020
30. Dogan S, Shen R, Ang DC, et al. Molecular epidemiology of EGFR and KRAS mutations in 3026 lung adenocarcinomas: higher susceptibility of women to smoking-related KRAS-mutant cancers. Clin Cancer Res. 2012;18:6169–6177. doi:10.1158/1078-0432.CCR-11-3265
31. Arbour KC, Jordan E, Kim HR, et al. Effects of co-occurring genomic alterations on outcomes in patients with KRAS-mutant non-small cell lung cancer. Clin Cancer Res. 2018;24:334–340. doi:10.1158/1078-0432.CCR-17-1841
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.

